Abstract
Rice grain quality is a multifarious attribute mainly governed by multiple nutritional factors. Grain protein is the central component of rice grain nutrition dominantly affecting eating–cooking qualities. Grain protein content is quantitatively influenced by its protein fractions. Genetic quantification of five protein fractions—albumins, globulins, prolamins, glutelin, and grain protein content—were evaluated by exploiting two BC3F2 mapping populations, derived from Kongyu131/TKM9 (population-I) and Kongyu131/Bg94-1 (population-II), which were grown in a single environment. Correlation studies among protein fractions and grain protein content were thoroughly investigated. A genetic linkage map was developed by using 146 single sequence repeat (SSR) markers in population-I and 167 markers in population-II. In total, 40 QTLs were delineated for five traits in both populations. Approximately 22 QTLs were dissected in population-I, derived from Kongyu131/TKM9, seven QTLs for albumin content, four QTLs for globulin content, three QTLs for prolamin content, four QTLs for glutelin content, and four QTLs for grain protein content. In total, 18 QTLs were detected in population-II, derived from Kongyu131/Bg94-1, five QTLs for albumin content, three QTLs for globulin content, four QTLs for prolamin content, two QTLs for glutelin content, and four QTLs for grain protein content. Three QTLs, qAlb7.1, Alb7.2, and qGPC7.2, derived from population-II (Kongyu131/Bg94-1) for albumin and grain protein content were successfully validated in the near isogenic line (NIL) populations. The localized chromosomal locus of the validated QTLs could be helpful for fine mapping via map-based cloning to discover underlying candidate genes. The functional insights of the underlying candidate gene would furnish novel perceptivity for the foundation of rice grain protein content and trigger the development of nutritionally important rice cultivars by combining marker-assisted selection (MAS) breeding.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Rice (Oryza sativa L.) is regarded as a staple food crop for Asian people and feeds more than 3.5 billion of the biosphere’s populace. A total of 90% of its production and consumption occurs in Asiatic countries; China and India alone account for approximately 55% of total production (Kong et al. 2015). Among cereal crops, rice contributes 20% of the prescribed calorie intake and provides the primary diet for the growing world population. To ensure food nutritional security, the estimated rice grain production must be elevated up to 852 million tons by 2035 (Brar and Khush 2018). Currently, rice geneticists and breeders are extensively engaged in the evolution of high-yielding rice varieties. Miracles in high-yield have almost reached a plateau and are saturated, raising concern for another balanced (yield–quality) green revolution by considering current growing population demands (Chen et al. 2019; Wu et al. 2020a). Nevertheless, the revolution in rice breeding for enhancing grain quality characteristics remains in the background. Elevated human living standards demand a high quality rice grain. Therefore, an advanced breeding strategy for the demanded grain quality traits has become the current market challenge. High nutritional grain quality is the hotspot of grain superiority. A high protein content of rice food is fundamentally known to enhance the individual nutritional factor for impoverished families, particularly in high rice-consuming families. However, enhancing fractions of proteins and grain protein content (GPC) is the major bailiwick of the current breeding strategy to ameliorate highly nutritional rice grain varieties.
Rice endosperm constitutes an abundant amount of proteins after starch. Rice protein content is widely varied throughout the distinct rice varieties and is recorded as about 5% to 16%. Additionally, Indica rice cultivar restores higher protein content (2% to 3%) compared to the Japonica rice cultivar (Lin et al. 1993; Zhou et al. 2009; Yang et al. 2019). Rice comprises fractions of proteins such as albumin, globulin, prolamin, and glutelin. Nevertheless, glutelin constitutes an abundant quantity of protein and synthesizes the maximum proportion of nutritional importance, which is essential for high lysine content and digestibility (He et al. 2021). A significant manipulation within the glutelin protein content may be likely to modulate rice nutritional superiority. Grain protein becomes the central element to determine rice nutritional superiority and covalently affects eating–cooking superiority (Long et al. 2023).
The sequence similarities of amino acids indicate that glutelin is further categorized into four subunits: Glu A, Glu B, Glu C, and Glu D (Chen et al. 2018). Comprehensively, cloning and functional characterization of genes associated with protein fractions are mostly obtained from mutants mapping (Ren et al. 2014). Rough endoplasmic reticulum produces a 57 kDa precursor for glutelin, such as 57H mutants, assimilating a high level of 57 kDa pro-glutelin and constitutes milky endosperm (Wang et al. 2009; Chen et al. 2018). In 57H mutants, only gpa3, Osvpe1, and OsRab5a were completely cloned (Wang et al. 2009). Prolamin was recognized by multiple gene families, involving 34 gene replicates. Prolamin was structurally delineated into three sub sections; 10 kDa (RP10), 13 kDa (RM1, RM2, RM4, & RM9), and 16 kDa (Kawakatsu et al. 2008). Albumins and globulins are jointly synthesized in the rice bran, and proportionally removed during the milling process (Shewry 2007). Globulins retains tranquil digestion (Zhang et al. 2008) and is restricted for gene identification, as explained by (Bhullar and Gruissem 2013). Albumin proteins, RA16 and RA17, were preliminarily designated as allergenic proteins, but current reports emphasized that they play a vital role in reducing blood sugar and plasma insulin (Adachi et al. 1993; Swamy et al. 2016). Preliminarily research indicated that high grain protein content has a negative correlation with eating–cooking superiority; contrarily, lowered protein content has immense significance in improving eating–cooking superiority (Li et al. 2019; Zhang et al. 2021; Xiong et al. 2023).
Grain protein fractions are quantitatively inherited and extensively affected by environmental factors (Chattopadhyay et al. 2019; Pradhan et al. 2019). Genetic elucidation of the protein content was initiated during the creation of QTL mapping, genetic markers, and linkage maps (Peng et al. 2014). Countless efforts have been antecedently made to investigate the genetic architecture of grain protein content. Numerous QTLs for rice grain protein contents have been previously summarized (Yang et al. 2015; Tan et al. 2001; Wang et al. 2008; Lou et al. 2009; Ye et al. 2010; Zheng et al. 2011, 2012; Cheng et al. 2013; Kashiwagi and Munakata 2018; Chattopadhyay et al. 2019). However, protein fractions and GPC are extremely sensitive to environmental factors; therefore, nearly all investigations of protein fractions and GPC are based on a single environment condition. Hence, QTLs dissected for protein fractions and GPC are frequently varied based on a given population’s structure or environmental conditions. To date, only two imperative QTLs, qPC1 and qGPC-10, were successfully cloned and functionally elucidated under a natural environmental condition, resulting in controlling rice grain protein (Peng et al. 2014; Yang et al. 2019). The qPC1 functionally encodes an amino acid carrier like OsAAP6, increasing grain protein content by synthesizing and accumulating albumin, globulin, prolamin, glutelin, and starch content. The qPC1 gene positively regulates rice grain protein content via an elevated gene expression level of OsAAP6 (Peng et al. 2014). The qGPC-10 functionally encodes an OsGluA2 precursor and positively regulates rice grain protein content by a pleiotropic effect. The qPC10 is functionally elevating GPC via increasing glutelin content and ameliorating nutritional properties of rice grains (Yang et al. 2019). Reduced genetic expression of OsAPP6 and OsGluA2 were predominantly allocated for low GPC breeding by combining MAS. In addition, countless efforts have been previously made to upgrade rice grain protein content and eating–cooking superiority by utilizing various genetic resources. Ordinarily, high grain protein content is considered for high nutritional superiority, but high GPC is negatively correlated with taste quality in rice, resulting in a compact rice endosperm structure, which negatively impacts its palatability (Hamaker and Griffin 1993; Martin and Fitzgerald 2002). Thereafter, it is important to investigate the genetic mechanism of rice grain protein to balance the nutritional and eating–cooking superiority.
Huge quantities of QTLs have been antecedently mapped for rice nutritional quality traits, and most of them were concentrated on a common character of protein content. At this time, very limited QTLs have been mapped, cloned and characterized in protein fractions. In this investigation, we have executed QTL mapping for protein fractions (albumins, globulins, prolamins, & glutelins) and grain protein content by utilizing two mapping populations of BC3F2 generations—population-I (Kongyu131/TKM9) and population-II (Kongyu131/Bg94-1), derived from crossing Kongyu131 × TKM9 and Kongyu131 × Bg94-1, respectively.
Materials and methods
Development of plant materials
Genetic materials of two mapping populations were originally developed by a crossing between three parents, Kongyu131, TKM9, and Bg94-1, respectively. In this mating design, Kongyu131 was used as a common recipient parent, while TKM9 and Bg94-1 were used as donor parents, respectively. Kongyu131 is a short-bold grain type and early maturing variety, TKM9 is a red-slender grain type, drought and flood resistant variety, while Bg94-1 is a long-medium grain type and highly disease susceptible variety. Population-1 was derived from a crossing between Kongyu131 × TKM9, while population-II was originated from a crossing between Kongyu131 × Bg94-1, respectively. The genetic mapping populations utilized in this investigation were derived from BC3F2 generations and obtained by a multiple backcrossing-selfing process. The development process of genetic material was provided in supplementary Fig. 1. Population-I was constituted from 271 individual lines and population-II from 372 individual lines. Among them, each line constituted 12 individual plants; mixed samples of 12 individual plants were thoroughly genotyped and phenotyped. In addition, both populations were planted under natural environment conditions at an experimental station of Huazhong Agricultural University, Hubei province (N 30.49°, E 114.36°), Wuhan, during 2018. The progenies of each recombinant line were grown at Lingshui (N 18.51°, E110.04°), Hainan province, during 2018. Twelve individual plants of 30-day-old seedlings derived from each line were transplanted at a plant-to-plant distance of 16.5 cm and row-to-row distance of 26.4 cm apart. Field management followed local practices.
Experimental field management
The field experiment was conducted at Huazhong Agricultural University, Hubei province, Wuhan, China, during 2018. Preliminarily, seeds were poured into clean water for 72 h and shifted for pre-sprouting (24 h) at room temperature. Pre-sprouted seeds were dispersed on the seed bed during 25 May 2018 and around 21–25-day-old seedlings were finally transplanted. Each line contained 12 individual plants and were planted in three replications. Randomized block design was adopted by keeping plant-to-plant distance at 16.5 cm and row-to-row distance 26.4 cm, respectively. All agronomic and intercultural practices were adopted according to locally standardized protocols. Chemical fertilizers were applied as nitrogen @ 48.7 kg/ha, phosphorus @ 58.5 kg/ha, and potassium @ 93.75 kg/ha followed by two consecutive topdressing with nitrogen fertilizer @ 86.25 kg/ha during tillering and @ 26.6 kg/ha during the booting stage of rice.
Quantitative estimation of albumin, globulin, prolamin, glutelin, and grain protein content
Fully mature rice seeds were individually harvested, threshed, dried, and stocked at room temperature for a minimum of 3 months to optimize the threshold level of moisture content. Approximately, 50 g of paddy seeds were de-hulled via TR 200 Sheller (Kett, Tokyo, Japan), and de-hulled brown rice grains were subjected to Pearle’s polish-mill (Kett, Tokyo, Japan). Rice grain flour made from CT410 grinding-mill (FOSS, Hillerod, Denmark) was sieved with 80, stocked at –20 °C for short-term storage, and –80 °C for long-term storage (Chen et al. 2018). Components of protein fractions, albumin (Alb), globulin (Gol), prolamin (Pro), and glutelin (Glu) content were quantified using the method described by Kumamaru et al. (1988), while grain protein content (GPC) was measured using a near infrared spectrum instrument system (NIRS), as previously described by (Chen et al. 2018). For albumin extraction, weigh a rice flour sample of 0.1 g and transfer into a centrifuge tube; add 1 ml of 10 mM Tris–HCl stock solution (pH 7.5); for globulin extraction, 0.1 g rice grain flour sample, add 1 ml of 1 M NaCl solution; for prolamin extraction, weigh 0.1 g of rice grain flour, add 1 ml of 60% n-propanol-1 mM EDTA-2Na stock solution; for glutelin extraction, weight 0.1 g of rice grain flour, add 1 ml of 0.05 M NaOH stock solution. The mixed samples and solutions were thoroughly mixed for at least 2 h at room temperature, centrifuged at 12,000 rpm at 4 °C for 15 min, and extracts were isolated into new tubes. This protocol was consecutively replicated three times, and accumulated extracts were stocked at –20 °C until the next step. All four proteins were extracted from a single sample and were examined using G-250 Coomassie brilliant blue dye, as explained by (Bradford 1976). Bovine serum was used as standard solution. Quantitative analysis was attempted using an infinite M200 (Tecan Group, Männedorf, Switzerland) (Peng et al. 2014; Chen et al. 2018). The phenotypic values of each protein fraction were computed by combining three replications.
QTL mapping, genetic linkage map, and validation
Individual plants of two BC3F2 populations were thoroughly genotyped, including involved parents by extracting high-quality genomic DNA from rice leaves. The DNA extraction was achieved using the CTAB-DNA extraction method, as described by (Paterson et al. 1993). Approximately 834 SSR markers were executed in three parents to distinguish 146 and 167 SSR markers. The size of SSR markers was kept between 100–300 bp and GC percentage between 40–60%. An amplicon size was determined by the available Nipponbare genome sequence (http://redb.ncpgr.cn/accessedon2018). Approximately 146 and 167 single sequence repeat (SSR) markers were applied in genotyping for population-I and population-II, respectively (markers were provided in supplementary Table 1). The phenotypic and genotypic data were aligned together to develop a genetic linkage map. The genetic linkage map and genetic effect analysis were employed using QTL cartographer 2.5, as described by (Wang 2006), and Map-Maker/QTL1.1, as explained by (Lincoln et al. 1992). The Kosambi mapping technique was employed to determine the recombination distance (Kosambi 2016), and QTLs additive effect was detected by composite interval mapping. The logarithm of odds (LOD) score was kept above threshold level 2.5 to distinguish QTLs reliability. QTLs peaks for five traits above threshold levels were detected and highly significant QTLs were recorded. Descriptive analysis of both populations for five traits were determined using Microsoft Office Excel 2010. Correlation analysis of two populations was performed using R program version R4.2.3. The Student’s t test was utilized to distinguish the individual average mean of the two populations.
Results
Phenotypic variations of nutritional grain quality traits
The phenotypic performances of two segregating populations of the BC3F2 generation, including recipient and donor parents, are portrayed in Fig. 1. The phenotypic mean of parent Kongyu131 for albumin, prolamin, glutelin, and grain protein content was found dominant over the parent TKM9 and Bg94-1, except for globulin content. The transgressive segregations and phenotypic variations were determined in both BC3F2 populations (Fig. 1 and Table 1). The mean performance of albumin, globulin, prolamin, glutelin, and grain protein content for population-I (Kongyu131/TKM9) was determined as 7.06 mg/g, 7.05 mg/g, 5.30 mg/g, 87.16 mg/g, and 111.60 mg/g, respectively (Fig. 1 and Table 1), whereas for population-II (Kongyu131/Bg94-1), it was estimated as 7.43 mg/g, 10.49 mg/g, 2.99 mg/g, 97.82 mg/g, and 113.90 mg/g, respectively (Fig. 1 and Table 1). The frequency distribution for all five traits reveals a normal distribution pattern, exemplifying that these nutritional quality traits were quantitatively inherited and dominantly controlled by polygenes.
Blue bar represents population-I and red bar designates population-II. Black, green, and blue arrows designate mean values of Kongyu131, TKM9, and Bg94-I, respectively. Histograms A–E depict the albumin (Alb), globulin (Gol), prolamin (Pro), glutelin (Glu), and grain protein content (Gpc) evaluated in 2018. Data were derived from three biological replicates.
Correlation studies in two populations
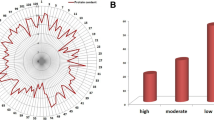
The correlation analyses for five traits were extensively executed. The correlation studies among albumin, globulin, prolamin, glutelin, and grain protein content are demonstrated in Fig. 2. For population-I, the correlation coefficient between glutelin content and prolamin content is the highest at 0.44, followed by the correlation coefficient between globulin content and glutelin content at 0.37. Similarly, the correlation coefficient between albumin content and globulin content is 0.36. In addition, weak correlations were found between globulin content and prolamin content, prolamin content and grain protein content, and glutelin content and grain protein content. For population-II, there was a clear positive correlation between albumin content and the other three protein contents except for grain protein content. A similar situation was also observed between globulin and the other three protein contents.
A represents population-I and B represents population-II, respectively. Alb—albumin; Gol—golbulin; Pro—prolamin; Glu—glutelin; Gpc—grain protein content in the year 2018. *** Significant at P < 0.001.
QTL dissection of albumin, globulin, prolamin, glutelin, and grain protein content
Through the application of a comprehensive composite interval mapping method, a total of 40 QTLs for albumin, globulin, prolamin, glutelin, and grain protein content traits were dissected in two BC3F2 populations (Fig. 3, Table 2 and supplementary Fig. 2). Among those, 22 QTLs delineated in population-I (Kongyu131/TKM9) and 18 QTLs in population-II (Kongyu131/Bg94-1), respectively. All the QTLs identified in population-I were distributed on chromosomes 1, 2, 3, 4, 6, 7, 10, and 11 for all five traits, explaining 4.1–23.63% of phenotypic variation. Among them, seven QTLs for albumin, qAlb1.1, qAlb2.1, qAlb2.2, qAlb4.1, qAlb6.1, qAlb10.1, and qAlb11.1, were characterized on chromosomes 1, 2, 4, 6, 10, and 11, explaining 7.23–21.13% of phenotypic variation; qAlb2.1 and qAlb6.1 exhibit maximum additive effect with 19.92% of phenotypic variation. Four QTLs for globulin, qGol2.1, qGol3.1, qGol6.1, and qGol11.1, were noticed on chromosomes 2, 3, 6, and 11, determining 11.34–18.35% of phenotypic variation, and qGol3.1 covers the maximum additive effect by 12.61% of phenotypic variation. Three QTLs for prolamin qPro2.1, qPro6.1, and qPro11.1, were distributed on the chromosomes 2, 6, and 11, contributing 11.61–17.50% of phenotypic variation, and qPro11.1 produces a dominating additive effect with 11.61% of phenotypic variation. Four QTLs for glutelin, qGlu2.1, qGlu3.1, qGlu10.1, and qGlu11.1, were identified on chromosomes 2, 3, 10, and 11, demonstrating 14.17–21.12% of phenotypic variation, and qGlu3.1 scored a maximum additive effect with 14.17% of phenotypic variation. Four QTLs for grain protein qGpc1.1, qGpc6.1, qGpc7.1, and qGpc10.1, were identified on chromosomes 1, 6, 7, and 10, and explained 4.10–23.63% of phenotypic variation, and qGpc7.1 constitutes the highest additive effect with 23.63% of phenotypic variation (Fig. 3 and Table 2).
Likewise, for population-II (Kongyu131/Bg94-1), 18 QTLs were reported for albumin, globulin, prolamin, glutelin, and grain protein content (Fig. 3 and Table 2). All the QTLs were extensively distributed on chromosomes 1, 2, 3, 4, 5, 7, 8, and 10, explaining 3.13–51.49% of phenotypic variation. Out of those, five QTLs for albumin qAlb2.3, qAlb2.4, qAlb7.1, qAlb7.2, and qAlb8.1, were confined on chromosomes 2, 7, and 8, determining 9.24–18.75% of phenotypic variation, qAlb7.2 and qAlb8.1 contain the maximum additive effect with 18.75% and 14.24% of phenotypic variation, respectively. Three QTLs for globulin, qGol2.2, qGol2.3, and qGol10.1, were identified on chromosomes 2 and 10, describing 50.00–51.49% of phenotypic variation, and qGol10.1 contributes the largest additive effect with 50.18% of phenotypic variation. Four QTLs for prolamin, qPro1.1, qPro2.2, qPro5.1, and qPro8.1, were dissected on chromosomes 1, 2, 5, and 8, elucidating 5.40–16.33% of phenotypic variation for each QTL, and qPro5.1 donates the largest additive effect with 16.3% of phenotypic variation. Two QTLs for glutelin, qGlu1.1 and qGlu7.1, were delineated on chromosomes 1 and 7, explicating 5.62–5.89% of phenotypic variation, and qGlu1.1 contributes the largest additive effect with 5.89% of phenotypic variation. Four QTLs for grain protein, qGpc3.1, qGpc4.1, qGpc7.2, and qGpc10.2, were identified on chromosomes 3, 4, 7, and 10, describing 3.13–23.17% of phenotypic variations, and qGpc7.2 contains the largest additive effect with 23.17% of phenotypic variation (Fig. 3 and Table 2).
Co-localizations of linked loci with antecedently reported QTLs/genes associated with nutritional grain quality characteristics
Many QTLs for grain nutritional quality traits have been previously dissected and cloned in rice crops. To distinguish QTLs reliability, localization of linked positions was compared with antecedently conveyed QTLs/genes in cultivated rice varieties for grain superiority traits from publicly available studies of previous QTL mappings. Among 40 QTLs, 37 QTLs exhibit co-localizations with earlier identified QTLs/interval/genes associated with grain quality traits (Table 2). For albumin content, 12 QTLs derived from both the populations, qAlb1.1, qAlb2.1, qAlb2.2, qAlb2.3, qAlb2.4, qAlb4.1, qAlb6.1, qAlb7.1, qAlb7.2, qAlb8.1 qAlb10.1, and qAlb11.1, exhibit co-localization with corresponding genomic interval of previously identified QTLs qAlb4, qGLB2, qPro8, qPC7, and qAAC7 on chromosomes 1, 2, 4, 6,7, 8, 10, and 11, respectively (Table 2). For globulin content, seven QTLs derived from populations, qGol2.1, qGol2.2, qGol2.3, qGol3.1, qGol6.1, qGol10.1, and qGol11.1, determine co-localization with previously reported QTLs qGLB2, qGLT2, qCph2, and qPC6, on chromosomes 2, 3, 6, 10, and 11 respectively, except for qGol3.1 (Table 2). For prolamin content, seven QTLs derived from both populations, qPro1.1, qPro2.1, qPro2.2, qPro5.1, qPro6.1, qPro8.1, and qPro11.1, expressing co-localized with previously recorded QTLs qPro6, qPR1, qALB1, and qPC2 on chromosomes 1, 2, 5, 6, 8, and 11, except for qPro5.1 (Table 2). For glutelin content, six QTLs derived from both populations, qGlu1.1, qGlu2.1, qGlu3.1, qGlu7.1, qGlu10.1, and qGlu11.1, explaining co-localizations with previously identified QTLs GluA-2, qGLB-1, qRPC-1, qAAC1.6, and qPC7.1 on chromosomes 1, 2, 3, 7, 10, and 11 (Table 2). For grain protein content, eight QTLs derived from both populations, qGpc1.1, qGpc3.1, qGpc4.1, qGpc6.1, qGpc7.1, qGpc7.2, qGpc10.1, and qGpc10.2, describing co-localization with previously detected QTLs qPC-1a, qALB-1, Gpc3-1, qPC-4, qPC-6, qPC7.1, qAAC7.1, and qPC10, on chromosomes 1, 3, 4, 6, 7, and 10 (Table 2).
Validation of the QTLs qAlb7.1, qAlb7.2, and qGpc7.2
Identified QTLs were validated in a segregating population of BC3F3 generation, derived from a BC3F2 individual plant with heterozygous target region. Two albumin QTLs, qAlb7.1, qAlb7.2, and one grain protein content QTL, qGpc7.2, were successfully validated through two near isogeneic lines (NIL). In the NIL population of qAlb7.1, the grain albumin content in the homozygous line of qAlb7.1 alleles is significantly increased by 2.47 mg/g and encodes 34.13% of change (Fig. 4 and Table 3). For qAlb7.2, compared to the control, the grain albumin content in the homozygous line of qAlb7.2 alleles significantly increased by 1.17 mg/g and encodes 24.81% of change (Fig. 4 and Table 3). Similarly, the grain protein content of Kongyu131 type homozygous lines is significantly improved by 2.60 mg/g and encodes 2.64% of change when compare to that of Bg94-1 type homozygous lines (Fig. 4 and Table 3).
The P values are calculated based on two-tailed t tests. Indicated error bars showed standard deviations for each genotype. Black and gray bars (A and B) represent Kongyu131 and Bg94-1 isogenic lines, respectively.
Discussion
Phenotypic variations of nutritional grain quality attributes
Kongyu131 is a short grain type of Japonica rice variety and commercially cultivated in temperate regions of Heilongjiang Province, China (Nan et al. 2018). Large studies have previously reported on QTL mapping in inter-specific crosses between Indica and Japonica rice (Harushima et al. 2001; Chin et al. 2011). Genetic improvement by an introgression line has become a popular strategy to develop transgressive segregants in the elite genetic background of rice cultivar. This is gained either through de novo raised genetic variations or wild allelic introgressions. In this study, all nutritional grain quality characteristics have produced transgressive segregants in both the populations. QTL mapping on rice grain quality characteristics with transgressive segregants has been previously reported by (Tan et al. 2001; Aluko et al. 2004; Kepiro et al. 2008; Yu et al. 2009; Liu et al. 2011; Bruno et al. 2017; Chen et al. 2018; Chattopadhyay et al. 2019; Jang et al. 2020; Zhao et al. 2022).
In this investigation, all nutritional grain quality characteristics were normally distributed in both populations and potentially affected by environmental factors (Fig. 1). Both populations have demonstrated wider variations for all five characteristics, which signifies that there would be diverse genetic interactions and complex genetic regulation systems among the five traits. The phenotypic variations described in this investigation for all five traits concur with antecedently reported literature by (Chen et al. 2018; Yang et al. 2019). Out of all protein fractions, glutelin constitutes the maximum proportions, and produces a higher degree of essential amino acids that are vital for human nutrition. It is emphasized that grain protein content would be quantitatively improved by enhancing the quantity of protein fractions and may establish a multifactorial bonding relationship between protein fractions and grain protein contents in rice (Hillerislambers et al. 1973). Interestingly, in this study, significant positive correlations were delineated for albumin, globulin, prolamin, glutelin, and grain protein content in both populations. These results concur with the previous report of (Kawakatsu et al. 2010; Chen et al. 2018). Phenotypic results suggest that protein fractions rarely share a common genetic pool and regulation mechanism. Individual protein fractions either have dependent or independent genetic complexity and regulation patterns.
QTL localizations and pleiotropic associations of the Alb, Gol, Pro, Glu, and Gpc
Albumin protein is an integral part of protein fractions and widely distributed in the rice seed endosperm. Confined studies on albumin content in relation to human health were previously conducted and emphasized that albumin protein constitutes an allergenic protein (Zhang et al. 2008). In addition, a recent investigation demonstrated that albumin is an important indicator for diabetes (blood sugar or glucose level) and plasma insulin reduction in the human body (Ina et al. 2016). Limited genetic studies on albumin content have been done, namely qALB-1, qALB-2, and qAlb4, QTLs identified on chromosomes 1, 2, and 4, but none of the genes have been cloned yet (Zhang et al. 2008; Chen et al. 2018). It was previously assumed that the Wx gene may have regulatory functions for albumin content; fortunately, the Wx gene does regulate rice grain protein content (Chen et al. 2018).
Generally, globulin protein is synthesized in the rice bran and mostly removed during the milling process. However, globulin and glutelin proteins were synthesized and co-located in the protein body type II (PB-II) compartment, playing significant roles in higher digestibility (Yamagata et al. 1982). It has been previously reported that globulin and glutelin proteins jointly regulate rice grain protein. Limited QTLs were mapped for globulin content on chromosomes 1, 2, and 5 (Zhang et al. 2008). Therefore, QTLs/genes mining for globulin and glutelin protein would furnish a new revolution in rice grain quality (Kumamaru et al. 1988). Prolamin roughly constitutes 20–30% of SSPs, and an abundant quantity of prolamin is thought to deteriorate rice nutritional properties, hence deducing prolamin content will enhance desirable nutritional quality (Aluko et al. 2004). Generally, Indica rice has a high prolamin content and developing an elite breeding strategy for reducing prolamin quantity would enhance the nutritional property of Indica rice (Chen et al. 2018). QTL mining on prolamin content was initiated a decade earlier, but, to date, confined QTLs/genes have been discovered on chromosomes 1, 3, 9, and 10 (Zhang et al. 2008; Park et al. 2019). Glutelin is the major nutritional component among all protein fractions, extensively manipulating eating–cooking quality, nutritional quality, palatability, and constitutes the highest nutritional substances compared to the rest of the protein fractions (Kawakatsu and Takaiwa 2010; Kawakatsu et al. 2008). A limited number of QTLs/genes for glutelin content have been discovered on chromosomes 2, 10, 11, and 12 (Zhang et al. 2008). Notably, one glutelin content gene, OsGluA2, enhances grain protein content in the natural population (Yang et al. 2019). Therefore, dissecting novel QTLs/genes for glutelin content would be an asset to develop rice varieties with accelerated palatability, eating–cooking quality, and high nutritional quality. Many QTLs associated with grain protein content were anciently reported, and thought to play a major role in regulating grain protein, and numerous QTLs have been successfully cloned and functionally studied (Cheng et al. 2013). Notwithstanding, grain protein content is extremely susceptible to environmental factors, particularly by application of a nitrogen fertilizers dose during the dough stage of rice. Hence, the detected QTLs with diverse genetic materials were enormously fluctuating, resulting in a lower detection rate. However, genetic characterization of grain protein content is extensively undone, possibly due to the complexity of QTLs/gene identification and cloning (Yang et al. 2019). To date, only two QTLs, qPC1 and qPC10, have been successfully characterized on chromosome 1 and 10, which positively regulate grain protein content in rice (Peng et al. 2014; Yang et al. 2019).
In this investigation, a total of 40 QTLs for albumin, globulin, prolamin, glutelin, and grain protein content were determined in both populations, extensively scattered on all the chromosomes of rice, except on chromosomes 9 and 12 (Fig. 3 and Table 2). Out of all QTLs, 12 QTLs for albumin were distributed on all chromosomes of rice, except on chromosomes 3, 5, 9, and 12; seven QTLs for globulin were scattered on chromosomes 2, 3, 6, and 10 of rice; seven QTLs for prolamin were distributed on chromosomes 1, 2, 5, 6, 8, and 11 of rice; six QTLs for glutelin were localized on chromosomes 1, 2, 3, 7, 10, and 11 of rice; eight QTLs for grain protein were distributed on chromosomes 1, 3, 4, 6, 7, and 10 of rice. The identified QTLs for all nutritional grain quality traits demonstrated co-localization with previously discovered QTLs in rice, except for qGol3.1, qPro5.1, and qGpc7.1, which were identified for the first time. The co-localization of the above QTLs concur with (Tan et al. 2001; Aluko et al. 2004; Hu et al. 2004; Kepiro et al. 2008; Wang et al. 2008, 2017; Zhang et al. 2008; Yu et al. 2009; Liu et al. 2011; Zheng et al. 2011; Zhong et al. 2011; Yun et al. 2014; Bruno et al. 2017; Kinoshita et al. 2017; Chen et al. 2018; Lou et al. 2009; Chattopadhyay et al. 2019; Jang et al. 2020; Zhao et al. 2022). Potentially, qAlb7.1, qAlb7.2, and qGpc7.2 have a significant effect on albumin and grain protein content. This research provides an outstanding resource of QTLs for albumin, globulin, prolamin, glutelin, and grain protein content that has potential to ameliorate rice grain nutritional quality. In summary, these QTLs may be used for further improvement of rice eating–cooking quality, palatability, and grain nutritional quality.
Genetic linkage Map of the QTLs dissected in Population-I (Kongyu131/TKM9) and population-II (Kongyu131/Bg94-1). Chr—chromosome, numerical digits—chromosome numbers, q—quantitative trait loci, Alb—albumin, Gol—globulin, Pro—prolamin, Glu—glutelin, Gpc—grain protein content. QTLs are presented at the right side of each chromosome in italic fonts, QTLs presented in orange color identified in population-I, and blue color in population-II, first numeric digit in each QTL indicates chromosome number and second digit indicates number of QTLs. Physical distances (Mb) of individual QTLs are located on the left side of each chromosome, pentagonal shape at the right side of the QTLs determines co-localization with previously identified QTLs, and triangle shape indicates validated QTLs. The scale bar is located on the left side of the chromosomes
Conclusion
Highly nutritious rice grain has become a promising requirement of current growing populations. High nutritive value of rice grains can be achieved by enhancing the quantitative content of protein fractions such as albumin, globulin, prolamin, glutelin, and grain protein content. This study authenticates that germplasm Kongyu131, TKM9, and Bg94-1 can be used to obtain major/minor QTLs/genes to proliferate protein fractions and grain protein content in rice. Together, the outlined consequences significantly validated that included parents have enough allelic potential to furnish protein fractions and grain protein content throughout the distinct genomic regions, and offspring populations have potential alleles which were derived from appropriate parents. QTL localization in mapping populations and validation of QTLs such as qAlb7.1, qAlb7.2, and qGpc7.2 will drive genetic improvement of rice albumin and grain protein content. Furthermore, this exploration has delivered a broad range for protein fractions and grain protein content. Detected near isogenic lines can be productively exploited for new breeding schemes, utilized as new sources of genetic materials to improve rice protein fractions and grain protein content, especially in South Asian and South East Asian countries, where rice intake is the primary source of food and nutrition. Additionally, these consequences could potentially imply desirable allelic pyramiding via inter-specific crosses, which would be an essential tool for expanding high nutritional rice grain quality components. QTLs identified in this study for protein fractions and grain protein content would be an advantageous outcome for developing highly nutritious rice varieties and provide food nutritional security in rice-consuming communities.
Data availability
The data matrixes produced during the current investigation are only accessible from the corresponding author on a justifiable request.
References
Adachi T, Izumi H, Yamada T, Tanaka K, Takeuchi S, Nakamura R, Matsuda T (1993) Gene structure and expression of rice seed allergenic proteins belonging to the α-amylase/trypsin inhibitor family. Plant Mol Biol 21:239–248
Aluko G, Martinez C, Tohme J, Castano C, Bergman C, Oard J (2004) QTL mapping of grain quality traits from the interspecific cross Oryza sativa× O. glaberrima. Theor Appl Genet 109(3):630–639
Bhullar NK, Gruissem W (2013) Nutritional enhancement of rice for human health: The contribution of biotechnology. Biotechnol Adv 31(1):50–57
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72(1–2):248–254
Brar DS, Khush GS (2018) Wild relatives of rice: a valuable genetic resource for genomics and breeding research. The Wild Oryza Genomes, Compendium of Plant Genomes, pp 1–25
Bruno E, Choi Y-S, Chung IK, Kim KM (2017) QTLs and analysis of the candidate gene for amylose, protein, and moisture content in rice (Oryza sativa L.). 3 Biotech 7:1–8
Chattopadhyay K, Behera L, Bagchi TB, Sardar SS, Moharana N, Patra NR, Chakraborti M, Das A, Marndi BC, Sarkar A (2019) Detection of stable QTLs for grain protein content in rice (Oryza sativa L.) employing high throughput phenotyping and genotyping platforms. Sci Rep 9(1):1–16
Chen E, Huang X, Tian Z, Wing RA, Han B (2019) The genomics of Oryza species provides insights into rice domestication and heterosis. Annu Rev Plant Biol 70:639–665
Chen P, Shen Z, Ming L, Li Y, Dan W, Lou G, Peng B, Wu B, Li Y, Zhao D (2018) Genetic basis of variation in rice seed storage protein (Albumin, Globulin, Prolamin, and Glutelin) content revealed by genome-wide association analysis. Front Plant Sci 9:612
Cheng L, Xu Q, Zheng T, Ye G, Luo C, Xu J, Li Z (2013) Identification of stably expressed quantitative trait loci for grain yield and protein content using recombinant inbred line and reciprocal introgression line populations in rice. Crop Sci 53(4):1437–1446
Chin JH, Chu S-H, Jiang W, Cho Y-I, Basyirin R, Brar DS, Koh H-J (2011) Identification of QTLs for hybrid fertility in inter-subspecific crosses of rice (Oryza sativa L.). Genes Genomics 33:39–48
Hamaker BR, Griffin VK (1993) Effect of disulfide bond-containing protein on rice starch gelatinization and pasting. Cereal Chem 70(4):377–380
Harushima Y, Nakagahra M, Yano M, Sasaki T, Kurata N (2001) A genome-wide survey of reproductive barriers in an intraspecific hybrid. Genetics 159(2):883–892
He W, Wang L, Lin Q, Yu F (2021) Rice seed storage proteins: Biosynthetic pathways and the effects of environmental factors. J Integr Plant Biol 63(12):1999–2019
Hillerislambers D, Rutger J, Qualset C, Wiser W (1973) Genetic and environmental variation in protein content of rice (Oryza sativa L.). Euphytica 22(2):264–273
Hu Z-L, Li P, Zhou M-Q, Zhang Z-H, Wang L-X, Zhu L-H, Zhu Y-G (2004) Mapping of quantitative trait loci (QTLs) for rice protein and fat content using doubled haploid lines. Euphytica 135(1):47–54
Ina S, Ninomiya K, Mogi T, Hase A, Ando T, Matsukaze N, Ogihara J, Akao M, Kumagai H, Kumagai H (2016) Rice (Oryza sativa japonica) albumin suppresses the elevation of blood glucose and plasma insulin levels after oral glucose loading. J Agric Food Chem 64(24):4882–4890
Jang S, Han J-H, Lee YK, Shin N-H, Kang YJ, Kim C-K, Chin JH (2020) Mapping and validation of QTLs for the amino acid and total protein content in brown rice. Front Genet 11:240
Kashiwagi T, Munakata JJE (2018) Identification and characteristics of quantitative trait locus for grain protein content, TGP12, in rice (Oryza sativa L.). Euphytica 214(9):165
Kawakatsu T, Hirose S, Yasuda H, Takaiwa F (2010) Reducing rice seed storage protein accumulation leads to changes in nutrient quality and storage organelle formation. Plant Physiol 154(4):1842–1854
Kawakatsu T, Takaiwa F (2010) Cereal seed storage protein synthesis: fundamental processes for recombinant protein production in cereal grains. Plant Biotechnol J 8(9):939–953
Kawakatsu T, Yamamoto MP, Hirose S, Yano M, Takaiwa F (2008) Characterization of a new rice glutelin gene GluD-1 expressed in the starchy endosperm. J Exp Bot 59(15):4233–4245
Kepiro J, McClung A, Chen M-H, Yeater K, Fjellstrom R (2008) Mapping QTLs for milling yield and grain characteristics in a tropical japonica long grain cross. J Cereal Sci 48(2):477–485
Kinoshita N, Kato M, Koyasaki K, Kawashima T, Nishimura T, Hirayama Y, Takamure I, Sato T, Kato K (2017) Identification of quantitative trait loci for rice grain quality and yield-related traits in two closely related Oryza sativa L. subsp. japonica cultivars grown near the northernmost limit for rice paddy cultivation. Breed Sci 67(3):191–206
Kong X, Kasapis S, Bao J (2015) Viscoelastic properties of starches and flours from two novel rice mutants induced by gamma irradiation. LWT-Food Sci Technol 60(1):578–582
Kosambi DD (2016) The estimation of map distances from recombination values. In: Ramaswamy R (ed) D.D. Kosambi. Springer, New Delhi
Kumamaru T, Satoh H, Iwata N, Omura T, Ogawa M, Tanaka K (1988) Mutants for rice storage proteins. Theor Appl Genet 76(1):11–16
Li H, Yang J, Yan S, Lei N, Wang J, Sun B (2019) Molecular causes for the increased stickiness of cooked non-glutinous rice by enzymatic hydrolysis of the grain surface protein. Carbohyd Polym 216:197–203
Lin R, Luo Y, Liu D, Huang C (1993) Determination and analysis on principal qualitative characters of rice germplasm. Rice germplasm resources in China Agricultural Science and Technology Publisher of China, Beijing:83–93
Lincoln S, Daly M, Lander E (1992) Mapping genes controlling quantitative traits with MAPMAKER/QTL, version 1.1. In: A tutorial and reference manual, 2nd edn. Whitehead Institute Technical Report 46, Cambridge
Liu X, Wan X, Ma X, Wan J (2011) Dissecting the genetic basis for the effect of rice chalkiness, amylose content, protein content, and rapid viscosity analyzer profile characteristics on the eating quality of cooked rice using the chromosome segment substitution line population across eight environments. Genome 54(1):64–80
Long X, Chunmin G, Lin W, Liting J, Xiangjin F, Qinlu L, Zhengyu H, Chun L (2023) Rice storage proteins: Focus on composition, distribution, genetic improvement and effects on rice quality. Rice Sci 30:207−221
Lou J, Chen L, Yue G, Lou Q, Mei H, Xiong L, Luo L (2009) QTL mapping of grain quality traits in rice. J Cereal Sci 50(2):145–151
Martin M, Fitzgerald M (2002) Proteins in rice grains influence cooking properties. J Cereal Sci 36(3):285–294
Nan J, Feng X, Wang C, Zhang X, Wang R, Liu J, Yuan Q, Jiang G, Lin S (2018) Improving rice grain length through updating the GS3 locus of an elite variety Kongyu 131. Rice 11:1–9
Park S-G, Park H-S, Baek M-K, Jeong J-M, Cho Y-C, Lee G-M, Lee C-M, Suh J-P, Kim C-S, Kim S-M (2019) Improving the glossiness of cooked rice, an important component of visual rice grain quality. Rice 12:1–13
Paterson AH, Brubaker CL, Wendel JF (1993) A rapid method for extraction of cotton (Gossypium spp.) genomic DNA suitable for RFLP or PCR analysis. Plant Mol Biol Rep 11:122–127
Peng B, Kong H, Li Y, Wang L, Zhong M, Sun L, Gao G, Zhang Q, Luo L, Wang G (2014) OsAAP6 functions as an important regulator of grain protein content and nutritional quality in rice. Nat Commun 5(1):1–12
Pradhan SK, Pandit E, Pawar S, Baksh SY, Mukherjee AK, Mohanty SP (2019) Development of flash-flood tolerant and durable bacterial blight resistant versions of mega rice variety ‘Swarna’through marker-assisted backcross breeding. Sci Rep 9(1):1–15
Ren Y, Wang Y, Liu F, Zhou K, Ding Y, Zhou F, Wang Y, Liu K, Gan L, Ma W (2014) GLUTELIN PRECURSOR ACCUMULATION3 encodes a regulator of post-Golgi vesicular traffic essential for vacuolar protein sorting in rice endosperm. Plant Cell 26(1):410–425
Shewry PR (2007) Improving the protein content and composition of cereal grain. J Cereal Sci 46(3):239–250
Swamy B, Rahman MA, Inabangan-Asilo MA, Amparado A, Manito C, Chadha-Mohanty P, Reinke R, Slamet-Loedin IH (2016) Advances in breeding for high grain zinc in rice. Rice 9(1):1–16
Tan Y, Sun M, Xing Y, Hua J, Sun X, Zhang Q, Corke H (2001) Mapping quantitative trait loci for milling quality, protein content and color characteristics of rice using a recombinant inbred line population derived from an elite rice hybrid. Theor Appl Genet 103:1037–1045
Wang L, Zhong M, Li X, Yuan D, Xu Y, Liu H, He Y, Luo L, Zhang Q (2008) The QTL controlling amino acid content in grains of rice (Oryza sativa) are co-localized with the regions involved in the amino acid metabolism pathway. Mol Breed 21:127–137
Wang S (2006) Windows QTL cartographer 2.5. http://statgenncsuedu/qtlcart/WQTLCarthtm. Accessed 1 Nov 2018
Wang X, Pang Y, Zhang J, Wu Z, Chen K, Ali J, Ye G, Xu J, Li Z (2017) Genome-wide and gene-based association mapping for rice eating and cooking characteristics and protein content. Sci Rep 7(1):17203
Wang Y, Zhu S, Liu S, Jiang L, Chen L, Ren Y, Han X, Liu F, Ji S, Liu X (2009) The vacuolar processing enzyme OsVPE1 is required for efficient glutelin processing in rice. Plant J 58(4):606–617
Wu K, Wang S, Song W, Zhang J, Wang Y, Liu Q, Yu J, Ye Y, Li S, Chen J (2020a) Enhanced sustainable green revolution yield via nitrogen-responsive chromatin modulation in rice. Science 367(6478):eaaz2046
Wu Y-B, Li G, Zhu Y-J, Cheng Y-C, Yang J-Y, Chen H-Z, Song X-J, Ying J-Z (2020b) Genome-wide identification of QTLs for grain protein content based on genotyping-by-resequencing and verification of qGPC1-1 in rice. Int J Mol Sci 21(2):408
Xiong Q, Sun C, Wang R, Wang R, Wang X, Zhang Y, Zhu J (2023) The key metabolites in Rice quality formation of conventional japonica varieties. Curr Issues Mol Biol 45(2):990–1001
Yamagata H, Sugimoto T, Tanaka K, Kasai Z (1982) Biosynthesis of storage proteins in developing rice seeds. Plant Physiol 70(4):1094–1100
Yang Y, Guo M, Li R, Shen L, Wang W, Liu M, Zhu Q, Hu Z, He Q, Xue Y (2015) Identification of quantitative trait loci responsible for rice grain protein content using chromosome segment substitution lines and fine mapping of qPC-1 in rice (Oryza sativa L.). Mol Breed 35:1–9
Yang Y, Guo M, Sun S, Zou Y, Yin S, Liu Y, Tang S, Gu M, Yang Z, Yan C (2019) Natural variation of OsGluA2 is involved in grain protein content regulation in rice. Nat Commun 10(1):1949
Ye G, Liang S, Wan J (2010) QTL mapping of protein content in rice using single chromosome segment substitution lines. Theor Appl Genet 121:741–750
Yu Y-H, Li G, Fan Y-Y, Zhang K-Q, Min J, Zhu Z-W, Zhuang J-Y (2009) Genetic relationship between grain yield and the contents of protein and fat in a recombinant inbred population of rice. J Cereal Sci 50(1):121–125
Yun B-W, Kim M-G, Handoyo T, Kim K-M (2014) Analysis of rice grain quality-associated quantitative trait loci by using genetic mapping. Am J Plant Sci 5:1125–1132
Zhang H, Jang S-G, Lar SM, Lee A-R, Cao F-Y, Seo J, Kwon S-W (2021) Genome-wide identification and genetic variations of the starch synthase gene family in rice. Plants 10(6):1154
Zhang W, Bi J, Chen L, Zheng L, Ji S, Xia Y, Xie K, Zhao Z, Wang Y, Liu L (2008) QTL mapping for crude protein and protein fraction contents in rice (Oryza sativa L.). J Cereal Sci 48(2):539–547
Zhao L, Zhao C-F, Zhou L-H, Yao S, Zhao Q-Y, Chen T, Zhu Z, Zhang Y-D, Wang C-L (2022) Mapping QTLs for rice (Oryza sativa L.) grain protein content via chromosome segment substitution lines. Cereal Res Commun 50:699–708
Zheng L, Zhang W, Chen X, Ma J, Chen W, Zhao Z, Zhai H, Wan J (2011) Dynamic QTL analysis of rice protein content and protein index using recombinant inbred lines. J Plant Biol 54:321–328
Zheng L, Zhang W, Liu S, Chen L, Liu X, Chen X, Ma J, Chen W, Zhao Z, Jiang L, Wan J (2012) Genetic relationship between grain chalkiness, protein content, and paste viscosity properties in a backcross inbred population of rice. J Cereal Sci 56(2):153–160
Zhong M, Wang L, Yuan D, Luo L, Xu C, He Y (2011) Identification of QTL affecting protein and amino acid contents in rice. Rice Sci 18(3):187–195
Zhou L, Liu Q, Zhang C, Xu Y, Tang S, Gu M (2009) Variation and distribution of seed storage protein content and composition among different rice varieties. Acta Agron Sin 35(5):884–891
Acknowledgements
The authors extend their highest appreciation for the support from the National Key Laboratory of Crop Genetic Improvement, Huazhong Agricultural University.
Funding
This work was supported by grants from the National Natural Science Foundation of China (U21A20211), the Ministry of Science and Technology (2021YFF1000200, 2022YFD1200100), AgroST Project (NK20220501), and China Agriculture Research System (CARS-01–01).
Author information
Authors and Affiliations
Contributions
MA and YH designed this experiement. MA, GJ, YW, and JC conducted this experiemnt and collected all phenotypic data. MA and GL performed data analaysis. MA, GL,YH, YZ, and SL contributed in the graphical representaion. YH supervised all investiations. MA have wrote this manusript. All authors have read and authorized this manuscript for publication.
Corresponding author
Ethics declarations
Conflict of interest
All authors declare that there is no conflict of interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Alam, M., Wang, Y., Chen, J. et al. QTL detection for rice grain storage protein content and genetic effect verifications. Mol Breeding 43, 89 (2023). https://doi.org/10.1007/s11032-023-01436-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11032-023-01436-7