Abstract
Seed number per pod (SNPP) and seed weight (SW) are two components of seed yield in rapeseed (Brassica napus). Here, a natural population of rapeseed was employed for genome-wide association analysis for SNPP and SW across multi-years. A total of 101 and 77 SNPs significantly associated with SNPP and SW with the phenotypic variances (R2) ranging from 1.35 to 29.47% and from 0.78 to 34.58%, respectively. And 43 and 33 homologs of known genes from model plants were located in the 65 and 49 haplotype blocks (HBs) for SNPP and SW, respectively. Notably, we found 5 overlapping loci and 3 sets of loci with collinearity for both SNPP and SW, of which 4 overlapping loci harbored the haplotypes with the same direction of genetic effects on SNPP and SW, indicating high possibility to simultaneously improve SNPP and SW in rapeseed. Our findings revealed both overlapping and independent loci controlling seed number per pod and seed weight in rapeseed.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Rapeseed (Brassica napus, AACC, 2n = 38) is derived from an interspecific cross between B. rape (AA, 2n = 20) and B. oleracea (CC, 2n = 18) (U N 1935). As an important oil crop, rapeseed is widely grown in China, Europe, North America, and Australia (Yang et al. 2017, 2018; Shahid et al. 2019), with the global seed production of 70 million tons per year (http://www.fao.org/faostat/en/#data/QC/visualize).
Seed yield per plant of rapeseed is determined by three components: pod number per plant (PN), seed number per pod (SNPP), and seed weight (SW), which are typical quantitative traits (Quijadaet al. 2006; Shi et al. 2015). In comparison with PN, SNPP and SW have a relatively high heredity (Lu et al. 2017; Shi et al. 2015). A slightly negative correlation was detected between SNPP and SW (Lu et al. 2011; Cai et al. 2014; Shi et al. 2015; Zhu et al. 2020), indicating a weak trade-off between SNPP and SW in rapeseed.
More than 100 quantitative trait loci (QTL) were detected for SNPP (Zhang et al. 2006; Radoev et al. 2008; Shi et al. 2009; Basunanda et al. 2010; Wang and Guan, 2010; Zhang et al. 2010; Chen et al. 2011; Zhang et al. 2011; Ding et al. 2012; Qi et al. 2014; Cai et al. 2014; Shi et al. 2015; Li et al. 2015; Yang et al. 2016; Lu et al. 2017; Zhu et al., 2020) and for SW (Udall et al. 2006; Shi et al. 2009, 2011, 2019; Fan et al. 2010; Chen et al. 2011; Zhang et al. 2011; Ding et al. 2012; Yang et al. 2012; Li et al. 2014; Fu et al. 2015; Liu et al. 2015; Wang et al. 2016a; Luo et al. 2017; Dong et al. 2018). Those QTL were distributed in almost all chromosomes of rapeseed. However, the positions of those QTL were seldom compared due to the differences of research materials and marker systems used in those studies. Moreover, the segregated population was derived from the cross between two parents in the majority of those studies, which only harbored the variances from two parents.
With the rapid development of sequencing technology, genome-wide association studies (GWAS) have been extensively used to dissect complex traits in crops. The releases of reference genomes of rapeseed and its two parental species, B. rapa and B. oleracea (Wang et al. 2011; Chalhoub et al. 2014; Liu et al. 2014; Sun et al. 2017; Bayer et al. 2017; Song et al. 2020), have ensured possibility to conduct collinearity analysis among QTL. In this study, we carried out whole-genome association analysis for SNPP and SW in a natural population of rapeseed, which composed of 157 varieties from three ecotype groups with distant diversity, and found 101 and 77 SNPs significantly associated with SNPP and SW, which located in 65 and 49 haplotype blocks, respectively, of which five overlapping loci and three pairs of loci with collinearity controlled both SNPP and SW. Our findings revealed both overlapping and independent loci controlling SNPP and SW, and provide the target loci for simultaneous improvement of SNPP and SW in rapeseed.
Materials and methods
Plant material and field experiment
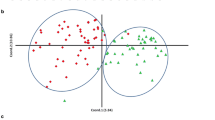
A natural population of rapeseed comprising of 52 spring inbred lines from North America, 54 winter inbred lines from Europe, and 51 semi-winter inbred lines from China (Table S1) was grown in Southwest University, China (Beibei, Chongqing), using a randomized complete block design with two replications. Each plot consisted of 30 plants, with 30 cm between rows and 20 cm within rows spacing. The field management followed the standard agriculture practice.
Trait evaluation and statistical analysis
At maturity, fifty well-developed siliques in the middle of inflorescence were collected from five individuals in the middle of each plot to investigate SNPP and SW across years (denoted as “trait-year”). SNPP was calculated as the average number of well-filled seeds per silique, and SW was the average weight of 1000 seeds in three replicates in each plot.
The data of SNPP was collected across 4 years (2015, 2016, 2018, 2019). The investigation of SW was reported across 4 years (2013–1016) in the previous study (Dong et al. 2018) and was extended to the other 2 years (2018–2019). The 6 years’ data of SW was merged for the following analysis.
Analysis of variance (ANOVA) and correlation analysis of each environment were performed using SAS version 9.3 (SAS Institute Inc.). Analysis of variance (ANOVA) was performed using SAS GLM procedure (Freund and Littell 1981); the Pearson’s correlation coeffecients were calculated by SAS CORR procedure. The broad-sense heritability was calculated according to the following formula: h2 = σ2G/(σ2G + σ2GE/e + σ2e/er), where σ2G, σ2GE, and σ2e are the variations of genetic, the interaction of the genotype by environment, and error, respectively. e and r are the numbers of environments and replications, respectively (Kowles 2001).
The best linear unbiased predictor (BLUP) value for each line was inferred across all years using the R package “LME4” by considering both the genotype and the environment as random effects (Lamprianou 2013).
Genome-wide association study
The genome of accessions in natural population was sequenced with a 5 × sequencing depth, producing total of 690,953 SNPs in the previous study, where population structure (Q) and relative kinship (K) analysis of natural population were calculated (Dong et al. 2018). Those SNPs were employed to detect associated signals for SNPP and SW with a multi-locus random-SNP-effect mixed linear model (Q + K) by using an R software of mrMLM v4.0 and integrating six GWAS methods (Zhang et al., 2020) , including mrMLM, FASTmrMLM, FASTmrEMMA, pLARmEB, pKWmEB, and ISIS EM-BLASSO.
The GWAS threshold for significant SNPs was set to − log10(P) > 5.83 (P = 1 / total SNP) for the models of mrMLM, FASTmrMLM, FASTmrEMMA, and pKWmEB, and the Manhattan plots and Q-Q plots were displayed using the R package “qqman” (Turner 2014). The GWAS threshold for significant SNPs was set to LOD > 3 for the models of pLARmEB and ISIS EM-BLASSO.
Co-location and synteny analyses of significant loci
The square of the correlation coefficient (r2) calculating with the R package “LDheatmap” was employed to estimate linkage disequilibrium (LD) between SNPs (Shin et al. 2006). The haplotype blocks (HBs) were determined by the average of r2 > 0.6 between adjacently significant SNPs on same chromosome (Qian et al. 2016), or by extending 50 kb on each side of the significantly associated SNPs outside of the HBs (Raman et al. 2016). The associated regions with the overlapped HB intervals for SNPP and SW were defined as genetic co-location loci. A chromosome-scale alignment of syntenic loci was performed using the large-scale genome synteny tool SYMAP version 4.2 (Soderlund et al. 2011).
To identify candidate genes in the haplotype blocks for SNPP and SW, the homologs of known genes associated with the two traits were annotated to the Darmor-bzh reference genome of rapeseed by BLASTP analysis. The SNPs and candidate genes of interest were integrated on the Circos diagram using Perl (Krzywinski et al. 2009). Haplotype maps were drawn in GraphPad Prism 8.
Results
Phenotypic variation of seed number per pod and seed weight
The SNPP and SW were investigated in a natural population across 4 years (2015, 2016, 2018, 2019) and 2 years (2018, 2019), respectively. A normal distribution of SNPP and SW was observed in most of years by Shapiro–Wilk normality test (Table 1). Wide variances were found for SNPP ranging from 4.40 to 31.63 seeds per silique, and for SW ranging from 1.82 to 5.53 g per 1000 seeds, indicating that SNPP and SW exhibit typically quantitative trait characterizations (Table 1 and Fig. S1).
In order to gain more information, the 4-year’s data of SW previously collected (Dong et al. 2018) was merged with the new data for the following analysis. ANOVA showed significant differences of genotype (G), environment (E), and interaction between genotype and environment (G × E) for both of SNPP and SW (Table S2). High broad-sense heritability was found for SNPP (0.854) and SW (0.922), in accordance with the previous studies (Cai et al. 2014; Lu et al. 2017), indicating that both SNPP and SW are mainly controlled by genotype. A high or middle significantly positive correlation was found for SNPP and SW across years (p < 0.001), but a slight correlation was found between SNPP and SW (Table 2).
Genome-wide association study
Genome-wide association analysis was carried out using the Q + K model with six methods to identify associated signals at whole-genome level. Manhattan plots and LOD score plots were shown in Fig. 1 using BLUP, and in supplementary Figure S2 and S3 using the data of each year for each trait. A total of 101 and 77 significantly associated SNPs for SNPP and SW were detected across years and methods, respectively (Table S3; Fig. S3), including 15 significant SNPs for SW detected in previous study (Dong et al. 2018). These significant SNPs which were unevenly distributed on all of the chromosomes, explained 1.35–29.47% and 0.78–34.58% of the phenotypic variances for SNPP and SW, respectively (Table 3 and Fig. 2).
Manhattan plots and LOD score plots of genome-wide association analysis for seed number per pod and seed weight. In the Manhattan plots, the x-axis indicates the physical positions of SNPs along each chromosome; the y-axis is the − log10P for the association; the horizontal dashed line indicates the significant threshold (− log10P = 5.83); each method is displayed by different colors. In the LOD score plots, the vertical red lines indicate significant SNPs with LOD > 3. SNPP (SW)-BLUP-number: (1) mrMLM; (2) FASTmrMLM; (3) FASTmrEMMA; (4) pKWmEB; (5) pLARmEB; (6) ISIS EM-BLASSO
Concentric circles of genetic intervals for seed number per pod (SNPP) and seed weight (SW) in Brassica napus. (a) Chromosomes. (b) Significant association SNPs with SW (blue) and SNPP (red). (c, d) Candidate genes for SW (purple) and SNPP (green). (e) Collinear regions on A and C subgenomes for SW and SNPP
Among these significant SNPs, 48 and 15 significantly associated SNPs were repeatedly detected for SNPP, and 37 and 9 SNPs were repeatedly detected for SW in different methods and years, respectively. Especially, 3 significantly associated SNPs (rs.A01.21808623, rs.A03.5434721, rs.C06.33564706) for SNPP and 7 significantly associated SNPs (rs.A04.15831065, rs.A04.15918524, rs.A04.15980541, rs.A04.18313809, rs.A06.21357638, rs.C06.30246876, rs.C07.40291083) for SW could be detected simultaneously in both different years and methods (Table S3).
In order to identify candidate genes, the homologs of known genes for SNPP and SW within the HB intervals were selected. In the LD analyses, 65 and 49 HBs were associated with SNPP and SW. Of which, 15 and 24 HBs for SNPP and SW were overlapped with the QTL previously detected, respectively (Table S4). It indicated that those overlapped loci had stable effects on SNPP and SW.
Homologs related with SNPP and SW
SNPP and SW are two important yield-related traits, which have been extensively researched in plant. More than 490 and 790 genes related with SNPP and SW were reported in model plants (Ge et al. 2019; Qu et al. 2015; Li et al. 2019), but only a few genes, BnaC9.SMG7b for SNPP (Li et al. 2015) and BnARF18, BnaA9.CYP78A9, BnaUPL3, and BnDA1 for SW (Liu et al. 2015; Shi et al. 2019; Miller et al. 2019; Wang et al. 2017a), were discovered in rapeseed partially due to its complex genome. We speculated that the homologs of those known genes from model plants may control SNPP and SW in rapeseed.
We aligned those known genes with the reference genome of rapeseed and found that 43 and 33 homologs were located in the HBs for SNPP and SW, respectively (Fig. 2 and Table S4). Those genes were involved in the processes of gamete development, double fertilization, and seed development. For example, BnaA05g27620D, BnaA06g33830D, and BnaC07g34400D are homologous to Arabidopsis AtMYB65, AtSPP, and AtRAB1A, which were involved in pollen development; BnaC05g17010D and BnaC05g18750D are homologous to AtEMB1968 and AtINO, which were involved in ovule development; BnaC01g40440D and BnaC08g16910D are homologous to AtEDA30 and AtRH36, which were involved in embryo sac development; BnaC01g40560D and BnaC08g35530D are homologous to AtUNE7 and AtEC1.3, which were involved in double fertilization; and BnaA06g04380D and BnaC05g06470D are homologous to AtAPX1 and AtPIAL1, which were involved in embryo development (Yang et al. 2016). BnaC01g00190D and BnaC09g07510D are homologous to AtBUPS1 and AtRALF34, which were required for pollen tube integrity (Ge et al. 2019) (Table S4). Those homologs of the known genes within HB intervals are important candidate genes for SNPP and SW in rapeseed, which might contribute to the differences on SNPP and SW in natural population of rapeseed.
Collinearity and genetic co-location analyses of associated loci for SNPP and SW
There are wildly homologous fragments of chromosome between the A and C subgenomes of rapeseed (Chalhoub et al. 2014), harboring possibly homologous QTL for SNPP and SW. We compared HB intervals for SNPP and SW at the whole-genome level and found 2 sets of loci with collinearity for SNPP (one set of associated region: A03. 4,768,780–4,789,504 bp and C03. 6,403,467–6,441,508 bp and the other set of associated region: A03.5436489–5,484,721 bp and C03.7165828-7245967 bp), and 3 sets of loci with collinearity for SNPP and SW (harboring associated region A03. 5,414,635–5,472,966 bp for SNPP vs C03. 7,105,825–7,205,825 bp for SW; A05. 1,054,698–1,076,186 bp for SNPP vs C04. 1,164,692–1,187,029 bp for SW; C06. 4,084,407–4,184,407 bp for SNPP vs A06. 1,743,417–1,779,766 bp for SW) (Table 4 and Fig. 2).
Genetic linkage and pleiotropy are common phenomena in plant (Wagner and Zhang, 2011; Yang et al. 2016). By screening possibly pleiotropic loci for SNPP and SW, we found 5 overlapping association regions for both SNPP and SW (A05. 9,622,605–9,700,016 bp, C03. 7,165,828–7,205,825 bp, C05. 10,738,902–10,761,872 bp, C07. 40,241,083–40,340,965 bp, and C08. 20,687,111–20,731,514 bp) (Table 4 and Fig. 2).
The genetic effect of haplotypes was calculated in those overlapping association regions for SNPP and SW in natural population. It was interesting that the same direction of genetic effect on SNPP and SW was found among haplotypes in those overlapping association regions, except an overlapping association region (C03. 7,167,118–7,173,145 bp) with opposite direction of genetic effect on SNPP and SW among haplotypes (Fig. 3). Those findings indicate pleiotropic effects on SNPP and SW in those overlapping association regions.
Genetic effects on seed number per pod and seed weight among haplotypes in overlapping association regions in a natural population of rapeseed. Linkage disequilibrium analysis and haplotype analysis of 5 associated regions (a–e). Top, the horizontal axis represents haplotypes and the vertical axis represents the phenotypic values of seed number per pod and seed weight. The red and blue dots represent the average performance of SNPP and SW across years among 157 accessions in a natural population. ***,**, *: Significance at P < 0.001, P < 0.01, and P < 0.05, respectively. Bottom, pairwise LD estimates in the different haplotype block
Discussion
Seed number per pod and seed weight are two important components of seed yield per plant in rapeseed. In this study, a total of 101 and 77 significant SNPs for SNPP and SW located in 65 and 49 haplotype blocks, which were identified in the whole-genome level. Of which, five loci were overlapped for SNPP and SW, and three pairs of loci exhibited collinearity for SNPP and SW, indicating both overlapping and independent loci underlying SNPP and SW in rapeseed. To the best of our knowledge, it is the first study to conduct a comparative QTL study on seed number per pod and seed weight in rapeseed. Our findings not only discovered target loci for improvement of SNPP and SW, but also showed high possibility to simultaneously improve SNPP and SW by manipulating those loci with the same direction of genetic effect.
A slight correlation between SNPP and SW was detected in this study, in accordance with the previous studies (Lu et al. 2011; Cai et al. 2014; Shi et al. 2015; Zhu et al. 2020), indicating diverse regulation mechanisms for SNPP and SW. Seed number per pod is related with the processes of fertilization and seed development, such as the number of ovules per ovary, the proportion of fertile ovules, the proportion of ovules fertilized, and the proportion of fertilized ovules that develop into seeds (Yang et al. 2016; Shi et al. 2015), while seed weight is regulated by the signals of maternal and zygotic tissues, involving the ubiquitin–proteasome pathway, G-protein signaling, mitogen-activated protein kinase (MAPK) signaling, phytohormone perception and homeostasis, and some transcriptional regulators (Li et al. 2019). The diverse regulation mechanisms for SNPP and SW were supported by the researches of carbon partitioning, which is vital for seed growth and development. The leaf photosynthesis acts on SNPP, while silique wall photosynthesis alone acts on the SW in Arabidopsis (Zhu et al. 2018). Silique photoassimilation was a major contributor to seed weight in rapeseed (Hua et al. 2012).
However, the five overlapping loci and three loci with collinearity for SNPP and SW which were detected in this study, and the QTL of qSN.A6 with antagonistic pleiotropy on SNPP and SW which was detected in the previous study (Yang et al. 2016), seemed to show that there were same regulators or pathways to control both SNPP and SW in rapeseed. Similar observations were documented in other species. For example, the expression of AtMINI3 and AtIKU2, the two downstream genes of the SHB1-MINI3-IKU2 cascade in endosperm proliferation and embryo development pathway, were particularly reduced by the overexpression of AtRAV1, resulting in reduced SNPP and SW in Arabidopsis (Shin and Nam 2018). AtPGI1 which participates in GA-mediated reproductive development and storage reserve biosynthesis positively regulates Arabidopsis SNPP and SW (Bahaji et al. 2018). The loss of function of AtGRDP1 which is involved in abiotic stress response showed a diminished number of seeds per pod and a reduction of seed weight in Arabidopsis (Rodríguez-Hernández et al. 2017). Downregulation of OsOTUB1, which encodes a deubiquitinating enzyme to interact with the E2 ubiquitin-conjugating protein OsUBC13 and transcription factor OsSPL14 (Wang et al. 2017b), and overexpression of OsSGL, which regulates stress tolerance and cell growth (Wang et al. 2016b), can enhanced grain number and seed weight in rice. OsGSN1 encodes the mitogen-activated protein kinase phosphatase OsMKP1, and the GSN1-MAPK module coordinates the trade-off between grain number and grain size by integrating localized cell differentiation and proliferation (Guo et al. 2018). Overexpression of OsDEP1, which is involved in regulating the carbon–nitrogen metabolic balance, can decrease grain weight and increase grain number in rice (Zhao et al. 2019). OsSPL18 controls grain weight and grain number in the OsmiR156k-OsSPL18-DEP1 pathway in rice (Yuan et al. 2019).
Taken together, our findings revealed both overlapping and independent loci controlling SNPP and SW, and high possibility to simultaneously improve SNPP and SW by manipulating those loci with same direction of genetic effect in rapeseed.
Data availability
The data sets supporting the results of this article are included within the article and its additional files.
Code availability
The code and materials analyzed during the current study are available from the corresponding author on reasonable request.
Change history
27 August 2021
A Correction to this paper has been published: https://doi.org/10.1007/s11032-021-01243-y
References
Bahaji A, Almagro G, Ezquer I, Gámez-Arcas S, Sánchez-López ÁM, Muñoz FJ, Barrio RJ, Sampedro MS, Diego ND, Spíchal L, Doležal K, Tarkowská D, Caporali E, Mendes MA, Baroja-Fernández E, Pozueta-Romero J (2018) Plastidial phosphoglucose isomerase is an important determinant of seed yield through its involvement in gibberellin-mediated reproductive development and storage reserve biosynthesis in Arabidopsis. Plant Cell 30(9):2082–2098
Basunanda P, Radoev M, Ecke W, Friedt W, Becker HC, Snowdon RJ (2010) Comparative mapping of quantitative trait loci involved in heterosis for seedling and yield traits in oilseed rape (Brassica napus L.). Theor Appl Genet 120(2):271–281
Bayer PE, Hurgobin B, Golicz AA, Chan CKK, Yuan YX, Lee HT, Renton M, Meng JL, Li RY, Long Y, Zou J, Bancroft I, Chalhoub B, King GJ, Batley J, Edwards D (2017) Assembly and comparison of two closely related Brassica napus genomes. Plant Biotechnol J 15(12):1602–1610
Cai DF, Xiao YJ, Yang W, Ye W, Wang B, Younas M, Wu JS, Liu KD (2014) Association mapping of six yield-related traits in rapeseed (Brassica napus L.). Theor Appl Genet 127(1):85–96
Chalhoub B, Denoeud F, Liu S, Parkin IAP, Tang HB, Wang XY, Chiquet J, Belcram H, Tong CB, Samans B, Corréa M, Silva CD, Just J, Falentin C, Koh CS, Clainche IL, Bernard M, Bento P, Noel B, Labadie K et al (2014) Early allopolyploid evolution in the post-Neolithic Brassica napus oilseed genome. Sci 345(6199):950–953
Chen W, Zhang YS, Yao JB, Ma CZ, Tu JX, Fu TD (2011) Quantitative trait loci mapping for two seed yield component traits in an oilseed rape (Brassica napus) cross. Plant Breeding 130:640–646
Ding GD, Zhao ZK, Liao Y, Hu YF, Shi L, Long Y, Xu FS (2012) Quantitative trait loci for seed yield and yield-related traits, and their responses to reduced phosphorus supply in Brassica napus. Ann Bot 109(4):747–759
Dong HL, Tan CD, Li YZ, He Y, Wei S, Cui YX, Chen YG, Wei DY, Fu Y, He YJ, Wan HY, Liu Z, Xiong Q, Lu K, Li JN, Qian W (2018) Genome-wide association study reveals both overlapping and independent genetic loci to control seed weight and silique length in Brassica napus. Front Plant Sci 9:921
Fan CC, Cai GQ, Qin J, Li QY, Yang MY, Wu JZ, Fu TD, Liu KD, Zhou YM (2010) Mapping of quantitative trait loci and development of allele-specific markers for seed weight in Brassica napus. Theor Appl Genet 121(7):1289–1301
Freund RJ, Littell RC (1981) SAS for linear models: a guide to the ANOVA and GLM procedures. SAS Institute Inc, Cary (SAS users guide: Basics)
Fu Y, Wei DY, Dong HL, He YJ, Cui YX, Mei JQ, Wan HF, Li JN, Snowdon R, Friedt W, Li XR, Qian W (2015) Comparative quantitative trait loci for silique length and seed weight in Brassica napus. Sci Rep 5:14407
Ge Z, Cheung AY, Qu LJ (2019) Pollen tube integrity regulation in flowering plants: insights from molecular assemblies on the pollen tube surface. New Phytol 222(2):687–693
Guo T, Chen K, Dong NQ, Shi CL, Ye WW, Gao JP, Shan JX, Lin HX (2018) GRAIN SIZE AND NUMBER1 negatively regulates the OsMKKK10-OsMKK4-OsMPK6 cascade to coordinate the trade-off between grain number per panicle and grain size in Rice. Plant Cell 30(4):871–888
Hua W, Li RJ, Zhan GM, Liu J, Li J, Wang XF, Liu GH, Wang HZ (2012) Maternal control of seed oil content in Brassica napus: the role of silique wall photosynthesis. Plant J 69(3):432–444
Kowles R (2001) Solving Problems in Genetics. Springer-Verlag, New York
Krzywinski M, Schein J, Birol I, Connors J, Gascoyne R, Horsman D, Jones SJ, Marra MA (2009) Circos: an information aesthetic for comparative genomics. Genome Res 19(9):1639–1645
Lamprianou I (2013) Application of single-level and multi-level Rasch models using the lme4 package. J Appl Meas 14(1):79–90
Li N, Shi JQ, Wang XF, Liu GH, Wang HZ (2014) A combined linkage and regional association mapping validation and fine mapping of two major pleiotropic QTLs for seed weight and silique length in rapeseed (Brassica napus L.). BMC Plant Biol 14:114
Li N, Xu R, Li Y (2019) Molecular networks of seed size control in plants. Annu Rev Plant Biol 70:435–463
Li SP, Chen L, Zhang LW, Li X, Liu Y, Wu ZK, Dong FM, Wan LL, Liu KD, Hong DF, Yang GS (2015) BnaC9. SMG7b functions as a positive regulator of the number of seeds per silique in Brassica napus by regulating the formation of functional female gametophytes. Plant Physiol 169(4):2744–2760
Liu SY, Liu YM, Yang XH, Tong CB, Edwards D, Parkin IAP, Zhao MX, Ma JX, Yu JY, Huang SM, Wang XY, Wang JY, Lu K, Fang ZY, Bancroft I, Yang TJ, Hu Q, Wang XF, Yue Z, Li HJ et al (2014) The Brassica oleracea genome reveals the asymmetrical evolution of polyploid genomes. Nat Commun 5:3930
Liu J, Hua W, Hu ZY, Yang HL, Zhang L, Li RJ, Deng LB, Sun XC, Wang XF, Wang HZ (2015) Natural variation in ARF18 gene simultaneously affects seed weight and silique length in polyploid rapeseed. Proc Natl Acad Sci USA 112(37):E5123–E5132
Lu GY, Zhang F, Zheng PY, Cheng Y, Liu FL, Fu GP, Zhang XK (2011) Relationship among yield components and selection criteria for yield improvement in early rapeseed (Brassica napus L.). Agricultural Sci in China 10(7):997–1003
Lu K, Peng L, Zhang C, Lu JH, Yang B, Xiao ZC, Liang Y, Xu XF, Qu CM, Zhang K, Liu LZ, Zhu QL, Fu ML, Yuan XY, Li JN (2017) Genome-wide association and transcriptome analyses reveal candidate genes underlying yield-determining traits in Brassica napus. Front Plant Sci 8:206
Luo ZL, Wang M, Long Y, Huang YJ, Shi L, Zhang CY, Liu X, Fitt BDL, Xiang JX, Mason AS, Snowdon RJ, Liu PF, Meng JL, Zou J (2017) Incorporating pleiotropic quantitative trait loci in dissection of complex traits: seed yield in rapeseed as an example. Theor Appl Genet 130(8):1569–1585
Miller C, Wells R, McKenzie N, Trick M, Ball J, Fatihi A, Dubreucq B, Chardot T, Lepiniec L, Bevan MW (2019) Variation in expression of the HECT E3 ligase UPL3 modulates LEC2 levels, seed size, and crop yields in Brassica napus. Plant Cell 31(10):2370–2385
Qi LP, Mao L, Sun CM, Pu YY, Fu TD, Ma CZ, Shen JX, Tu JX, Yi B, Wen J (2014) Interpreting the genetic basis of silique traits in Brassica napus using a joint QTL network. Plant Breeding 133:52–60
Qian L, Qian W, Snowdon RJ (2016) Haplotype hitchhiking promotes trait coselection in Brassica napus. Plant Biotechnol J 14(7):1578–1588
Qu LJ, Li L, Lan Z, Dresselhaus T (2015) Peptide signalling during the pollen tube journey and double fertilization. J Exp Bot 66(17):5139–5150
Quijada PA, Udall JA, Lambert B, Osborn TC (2006) Quantitative trait analysis of seed yield and other complex traits in hybrid spring rapeseed (Brassica napus L.): 1. Identification of genomic regions from winter germplasm. Theor Appl Genet 113(3):549–561
Radoev M, Becker HC, Ecke W (2008) Genetic analysis of heterosis for yield and yield components in rapeseed (Brassica napus L.) by quantitative trait locus mapping. Genetics 179(3):1547–1558
Raman H, Raman R, Coombes N, Song J, Prangnell R, Bandaranayake C, Tahira R, Sundaramoorthi V, Killian A, Meng J, Dennis ES, Balasubramanian S (2016) Genome-wide association analyses reveal complex genetic architecture underlying natural variation for flowering time in canola. Plant Cell Environ 39(6):1228–1239
Rodríguez-Hernández AA, Muro-Medina CV, Ramírez-Alonso JI, Jiménez-Bremont JF (2017) Modification of AtGRDP1 gene expression affects silique and seed development in Arabidopsis thaliana. Biochem Biophys Res Commun 486(2):252–256
Shahid M, Cai GQ, Zu F, Zhao Q, Qasim MU, Hong YY, Fan CC, Zhou YM (2019) Comparative transcriptome analysis of developing seeds and silique wall reveals dynamic transcription networks for effective oil production in Brassica napus L. Int J Mol Sci 20(8):1982
Shi JQ, Li RY, Qiu D, Jiang CC, Long Y, Morgan C, Bancroft I, Zhao JY, Meng JL (2009) Unraveling the complex trait of crop yield with quantitative trait loci mapping in Brassica napus. Genetics 182(3):851–861
Shi JQ, Li RY, Zou J, Long Y, Meng JL (2011) A dynamic and complex network regulates the heterosis of yield-correlated traits in rapeseed (Brassica napus L.). PLoS One 6(7):e21645
Shi JQ, Zhan JP, Yang YH, Ye J, Huang SM, Li RY, Wang XF, Liu GH, Wang HZ (2015) Linkage and regional association analysis reveal two new tightly-linked major-QTLs for pod number and seed number per pod in rapeseed (Brassica napus L.). Sci Rep 5:14481
Shi LL, Song JR, Guo CC, Wang B, Guan ZL, Yang P, Chen X, Zhang QH, King GJ, Wang J, Liu KD (2019) A CACTA-like transposable element in the upstream region of BnaA9.CYP78A9 acts as an enhancer to increase silique length and seed weight in rapeseed. Plant J 98(3):524–539
Shin HY, Nam KH (2018) RAV1 negatively regulates seed development by directly repressing MINI3 and IKU2 in Arabidopsis. Mol Cells 41(12):1072–1080
Shin JH, Blay S, Mcneney B, Graham J (2006) LDheatmap: An R function for graphical display of pairwise linkage disequilibria between single nucleotide polymorphisms. J Stat Soft 16:1–9
Soderlund C, Bomhoff M, Nelson WM (2011) SyMAP v3.4: a turnkey synteny system with application to plant genomes. Nucleic Acids Res 39(10):e68
Song JM, Guan ZL, Hu JL, Guo CC, Yang ZQ, Wang S, Liu DX, Wang B, Lu SP, Zhou R, Xie WZ, Cheng YF, Zhang YT, Liu KD, Yang QY, Chen LL, Guo L (2020) Eight high-quality genomes reveal pan-genome architecture and ecotype differentiation of Brassica napus. Nat Plants 6(1):34–45
Sun FM, Fan GY, Hu Q, Zhou YM, Guan M, Tong CB, Li JN, Du DZ, Qi CK, Jiang LC, Liu WQ, Huang SM, Chen WB, Yu JY, Mei DS, Meng JL, Zeng P, Shi JQ, Liu KD, Wang X (2017) The high-quality genome of Brassica napus cultivar ‘ZS11’ reveals the introgression history in semi-winter morphotype. Plant J 92(3):452–468
Turner SD (2014) qqman: An R package for visualizing GWAS results using QQ and manhattan plots. BioRxiv. https://doi.org/10.1101/005165
UN (1935) Genome analysis in Brassica with special reference to the experimental formation of B. napus and peculiar mode of fertilization. Jap J Bot 7:389–452
Udall JA, Quijada PA, Lambert B, Osborn TC (2006) Quantitative trait analysis of seed yield and other complex traits in hybrid spring rapeseed (Brassica napus L.): 2. Identification of alleles from unadapted germplasm. Theor Appl Genet 113(4):597–609
Wagner GP, Zhang JZ (2011) The pleiotropic structure of the genotype-phenotype map: the evolvability of complex organisms. Nat Rev Genet 12(3):204–213
Wang F, Guan CY (2010) Molecular mapping and identification of quantitative trait loci for yield components in rapeseed (Brasscia napus L.). Hereditas 32(3):271–277
Wang JL, Tang MQ, Chen S, Zheng XF, Mo HX, Li SJ, Wang Z, Zhu KM, Ding LN, Liu SY, Li YH, Tan XL (2017a) Down-regulation of BnDA1, whose gene locus is associated with the seeds weight, improves the seeds weight and organ size in Brassica napus. Plant Biotechnol J 15(8):1024–1033
Wang ML, Lu XD, Xu GY, Yin XM, Cui YC, Huang LF, Rocha PSCF, Xia XJ (2016a) OsSGL, a novel pleiotropic stress-related gene enhances grain length and yield in rice. Sci Rep 6:38157
Wang SS, Wu K, Qian Q, Liu Q, Li Q, Pan YJ, Ye YF, Liu XY, Wang J, Zhang JQ, Li S, Wu YJ, Fu XD (2017b) Non-canonical regulation of SPL transcription factors by a human OTUB1-like deubiquitinase defines a new plant type rice associated with higher grain yield. Cell Res 27(9):1142–1156
Wang XW, Wang HZ, Wang J, Sun RF, Wu J, Liu SY, Bai YQ, Mun JH, Bancroft I, Cheng F, Huang SW, Li XX, Hua W, Wang JY, Wang XY, Freeling M, Pires JC, Paterson AH, Chalhoub B, Wang B (2011) The genome of the mesopolyploid crop species Brassica rapa. Nat Genet 43(10):1035–1039
Wang XD, Chen L, Wang AN, Wang H, Tian JH, Zhao XP, Chao HB, Zhao YJ, Zhao WG, Xiang J, Gan JP, Li MT (2016b) Quantitative trait loci analysis and genome-wide comparison for silique related traits in Brassica napus. BMC Plant Biol 16:71
Yang P, Shu C, Chen L, Xu JS, Wu JS, Liu KD (2012) Identification of a major QTL for silique length and seed weight in oilseed rape (Brassica napus L.). Theor Appl Genet 125(2):285–296
Yang YH, Shi JQ, Wang XF, Liu GH, Wang HZ (2016) Genetic architecture and mechanism of seed number per pod in rapeseed: elucidated through linkage and near-isogenic line analysis. Sci Rep 6:24124
Yang Y, Shen YS, Li SD, Ge XH, Li ZY (2017) High density linkage map construction and QTL detection for three silique-related traits in Orychophragmus violaceus derived Brassica napus population. Front Plant Sci 8:1512
Yang Y, Zhu KY, Li HL, Han SQ, Meng QW, Khan SU, Fan CC, Xie KB, Zhou YM (2018) Precise editing of CLAVATA genes in Brassica napus L. regulates multilocular silique development. Plant Biotechnol J 16(7):1322–1335
Yuan H, Qin P, Hu L, Zhan SJ, Wang SF, Gao P, Li J, Jin MY, Xu ZY, Gao Q, Du AP, Tu B, Chen WL, Ma BT, Wang YP, Li SG (2019) OsSPL18 controls grain weight and grain number in rice. J Genet Genomics 46(1):41–51
Zhang LW, Liu PW, Hong DF, Huang AQ, Li SP, He QB, Yang GS (2010) Inheritance of seeds per silique in Brassica napus L. using joint segregation analysis. Field Crop Res 116:58–67
Zhang LW, Yang GS, Liu PW, Hong DF, Li SP, He QB (2011) Genetic and correlation analysis of silique-traits in Brassica napus L. by quantitative trait locus mapping. Theor Appl Genet 122(1):21–31
Zhang SF, Fu TD, Zhu JC, Wang JP, Wen YC, Ma CZ (2006) QTL mapping and epistasis analysis for yield and its components in Brassica napus L. Acta Agron Sin 32:1135–1142
Zhang YW, Tamba CL, Wen YJ, Li P, Ren WL, Ni YL, Gao J, Zhang YM (2020) mrMLM v4.0: An R platform for multi-locus genome-wide association studies. Genomics Proteomics Bioinformatics. https://doi.org/10.1016/j.gpb.2020.06.006
Zhao MZ, Zhao MH, Gu S, Sun J, Ma ZB, Wang LL, Zheng WJ, Xu ZJ (2019) DEP1 is involved in regulating the carbon-nitrogen metabolic balance to affect grain yield and quality in rice (Oriza sativa L.). PLoS One 14(3):e0213504
Zhu XY, Zhang L, Kuang C, Guo Y, Huang CQ, Deng LB, Sun XC, Zhan GM, Hu ZY, Wang HZ, Hua W (2018) Important photosynthetic contribution of silique wall to seed yield-related traits in Arabidopsis thaliana. Photosynth Res 137(3):493–501
Zhu YY, Ye J, Zhan JP, Zheng XX, Zhang JJ, Shi JQ, Wang XF, Liu GH, Wang HZ (2020) Validation and characterization of a seed number per silique quantitative trait locus qSN A7 in rapeseed (Brassica napus L.). Front Plant Sci 11:68
Funding
This study was funded by National Key Research and Development Program (2016YFD0100202), National Program on Key Basic Research Project of China (2015CB150201), National Nature Science Foundation of China (31471529), and the Project of Chongqing Science and Technology Commission (cstc2019jcyj-zdxmX0012, XmT2018081, and cstc2019jcyj-bshX0055).
Author information
Authors and Affiliations
Contributions
WQ, SX, and HD conceived and designed the study. SX, HD, LY, DH, FZ, YC, SW, and JL participated in the phenotyping of seed number per pod and seed weight and performed the experiments. SX, HD, YH, HW, ZL, and XL contributed to data analysis and interpretation. SX, HD, and WQ wrote the paper. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Xin, S., Dong, H., Yang, L. et al. Both overlapping and independent loci underlie seed number per pod and seed weight in Brassica napus by comparative quantitative trait loci analysis. Mol Breeding 41, 41 (2021). https://doi.org/10.1007/s11032-021-01232-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11032-021-01232-1