Abstract
Most Rosaceae fruit trees such as Japanese plum and sweet cherry have a gametophytic self-incompatibility (GSI) system controlled by a single S locus containing at least two linked genes with multiple alleles, i.e., S-RNase as a pistil determinant and SFB (S-haplotype-specific F-box gene) as a candidate for the pollen S determinant. For identification of S genotypes, many methods based on polymerase chain reaction (PCR) utilizing polymorphism in length of the S-RNase and SFB gene have been developed. In this study, we developed two dot-blot analysis methods for S-haplotype identification utilizing allele-specific oligonucleotides based on the SFB-HVa region, which has high sequence polymorphism. Dot-blotting of allele-specific oligonucleotides hybridized with digoxigenin-labeled PCR products allowed S genotyping of plants with nine S haplotypes (S-a, S-b, S-c, S-e, S-f, S-h, S-k, S-7 and S-10) in Japanese plum and ten S haplotypes (S-1, S-2, S-3, S-4, S-4′, S-5, S-6, S-7, S-9 and S-16) in sweet cherry (dot-blot-S-genotyping). In addition, dot-blotting of PCR products of SFB probed with the allele-specific oligonucleotides, occasionally utilizing competitive hybridization, was successful in screening for a desirable S haplotype in sweet cherry (dot-blot-S-screening).
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Rosaceae fruits trees such as Prunus, Malus, and Pyrus are commercially important crops cultivated worldwide. Most of Rosaceae plants have a gametophytic self-incompatibility system (GSI) controlled by a single S locus with multiple alleles. Since the GSI inhibits self-fertilization resulting in a decrease of fruit set and the bearing of parthenocarpic fruits of low quality, pollination with optimum compatible pollen is essential. In commercial orchards, cross-compatible cultivars that flower simultaneously and harbor different S haplotypes are planted together to ensure cross-pollination. Therefore, utilization of information on S genotypes of cultivars is essential for fruit set and fruit quality. In breeding, new cultivars with self-compatibility have been preferred for labor efficiency. For example, self-compatible sweet cherries harboring the S haplotype mutated artificially by X-ray irradiation (Lewis 1949) have been developed. Since early screening of the mutated S haplotype enables efficient breeding of self-compatible cultivars, a reliable method by which the mutated S haplotype from the wild type S haplotype can be distinguished is necessary.
In Japanese pear, the pistil determinant of SI reaction has been identified as S-RNase, which is a basic glycoprotein with ribonuclease activity (Sassa et al. 1992). Similar subsequent studies have used many Rosaceae species such as almond (Tao et al. 1997; Ushijima et al. 1998), apple (Sassa et al. 1994), European pear (Zuccherelli et al. 2002), Japanese apricot (Yaegaki et al. 2001), Japanese plum (Yamane et al. 1999) and sweet cherry (Tao et al. 1999). Recently, a candidate for the pollen S-determinant has been characterized and designated as SFB (S-haplotype-specific F-box gene) in almond (Ushijima et al. 2003), sweet and sour cherry (Yamane et al. 2003a), Japanese plum (Zhang et al. 2007), apricot (Romero et al. 2004) or as SLF (S-locus F-box) in Japanese apricot (Entani et al. 2003). The term ‘SFB’ for the pollen S-determinant is hereafter used in this report.
S genotypes of cultivars in Rosaceae fruit trees are commonly determined by investigating seed/fruit set or pollen tube growth after pollination. However, since the availability of flower materials is necessary for these tests, a marker-assisted selection system, which is independent of flowering age and season, is required. The recent identification of the S-RNase gene and the SFB gene have enabled characterization of the S haplotypes in Rosaceae fruit trees by RFLP- or PCR-based analysis using vegetative tissue as material. In sweet cherry, S genotyping was initially begun with zymogram analysis of stylar RNase (Bošković et al. 1997), but such analysis has been replaced by the PCR-based method using consensus primers which can detect polymorphisms of first or second intron length of the S-RNase gene (Tao et al. 1999; Yamane et al. 2000). Allele-specific primers developed by Sonneveld et al. (2001, 2003) together with consensus primers have enabled the identification of S-1, S-2, S-3, S-4, S-5, S-6, S-7, S-9, S-10, S-12, S-13, S-14 and S-16 haplotypes in sweet cherry. In addition to these S haplotypes, techniques for discrimination of six (S-17 to S-22) and three (S-23 to S-25) additional S haplotypes, which were characterized by De Cuyper et al. (2005) and by Wünsch and Hormaza (2004), respectively, have been developed (Sonneveld et al. 2006; De Cuyper et al. 2005). Likewise, S genotyping has been developed for use in Japanese plum (Beppu et al. 2002, 2003; Sapir et al, 2004; Halász et al. 2007), Japanese apricot (Tao et al. 2002), sour cherry (Yamane et al. 2001), almond (Tamura et al. 2000; Ortega et al. 2005), apple (Janssens et al. 1995), Japanese pear (Ishimizu et al. 1998) and European pear (Moriya et al. 2007), resulting in a rapid and convenient technique.
On the other hand, alternative S-genotyping methods based on SFB have been developed by RFLP in sweet cherry (Yamane et al. 2003a; Ikeda et al. 2004a) and Japanese apricot (Yamane et al. 2003b). A technique based on PCR using allele-specific primers (S-1 to S-6) in sweet cherry (Ikeda et al. 2005) and one employing fragmentation analysis with consensus primers using a semi-automated sequencer (Vaughan et al. 2006) have also been developed. Ushijima et al. (2004) and Sonneveld et al. (2005) have revealed a 4-bp deletion in the region upstream of HVa of PaSFB4 in an S-4 mutated haplotype, denoted S-4′ herein, which is responsible for the loss of pollen activity for self-incompatibility. This finding led to the development of a technique for discrimination between S-4 and S-4′ using a designed dCAPS primer (Ikeda et al. 2004b)
The consensus primers for S-RNase and SFB are not satisfactory for discrimination in certain alleles because of the similar size of PCR products and weak secondary bands on the gel (Sonneveld et al. 2003; Vaughan et al. 2006). Although analyses with a semi-automated sequencer for precise size fragmentation have been recently reported to be available (Ortega et al. 2005; Sonneveld et al. 2006; Vaughan et al. 2006), such techniques would not be practical and convenient for researchers or breeders. In allele-specific PCR, individual PCR has to be carried out for each S haplotype, and internal control might be required for PCR to avoid the problem of false negatives as reported by Sonneveld et al. (2003). PCR-RFLP (CAPS) marker can reduce the possibility of the false negative. To our knowledge, however, a PCR-RFLP marker for S genotyping has been developed only in S-1 and S-3 to S-6 haplotypes of sweet cherry (Yamane et al. 2000), but not in Japanese plum.
More recently, Fujimoto and Nishio (2003) have developed two methods for identification of S haplotypes by dot-blot analysis using SP11 alleles in Brassica oleracea; one is blotting of plant genomic DNA probed with labeled SP11 m (mature protein region of SP11 cDNA), and the other is dot blotting of SP11 m DNA fragments amplified from each allele-cDNA hybridized with the SP11 coding region labeled by PCR using a template of plant genomic DNA. In the present study, rather than analyzing PCR products and genomic DNA blotted on the membrane, we applied dot-blot analysis to S genotyping in Japanese plum and sweet cherry utilizing allele-specific oligonucleotides based on the SFB-HVa region, which has high sequence polymorphism. As a result, our improved method allowed us to discriminate among nine S haplotypes (S-a, S-b, S-c, S-e, S-f, S-h, S-k, S-7and S-10) in Japanese plum and ten S haplotypes (S-1, S-2, S-3, S-4, S-4′, S-5, S-6, S-7, S-9 and S-16) in sweet cherry (dot-blot-S-genotyping). In addition, we examined a labeled oligonucleotide as a probe for screening of a desirable S haplotype (dot-blot-S-screening) according to Shirasawa et al. (2005, 2006).
Materials and methods
Plant materials
Thirteen Japanese plum cultivars, which were managed at Nanjing Agricultural University, and thirteen sweet cherry cultivars, which were provided by Sato farm at Yamanobe-machi in Yamagata Prefecture and by the National Institute of Fruit Tree Science (Morioka, Japan), were used (Table 1).
Design of allele-specific oligonucleotides
As shown in Table 2, nine and ten allele-specific oligonucleotides for Japanese plum and sweet cherry SFB alleles, respectively, were designed in the region covering hypervariable region a (HVa; Ikeda et al. 2004a). Tm values of all oligonucleotides were set at 70°C. In addition, PaSFBoli-4, −4′ and −6 (Table 2) were used as digoxigenin-labeled probes for dot-blot-S-screening experiments. For competitive hybridization experiments, digoxigenin-labeled and non-labeled oligonucleotides of PaSFBoli-4new and PaSFBoli-4′ new were designed (Table 2).
Dot-blot analysis with digoxigenin-labeled DNA: dot-blot-S-genotyping
The designed allele-specific oligonucleotide of 5 μM was dot-blotted onto a nylon membrane (Nytran, Whatman, Germany) by Multi-pin Blotter (Atto, Tokyo, Japan). The membrane was exposed to UV-light (0.12 J) for cross-linking of the oligonucleotides.
Genomic DNA was extracted from leaves (0.1 g) according to the method described by Doyle and Doyle (1987) with an extraction solution containing 2% 2-mercaptoethanol. The DNA was quantified by a spectrophotometer (Qubit\({\texttrademark}\) Fluorometer, Invitrogen\({\texttrademark}\), USA). In order to amplify digoxigenin-labeled DNA, two sets of degenerated consensus primers for a fragment of ca. 350 bp covering the HVa region of Japanese plum SFB and of sweet cherry SFB were designed (Table 3). Digoxigenin labeling of the ca. 350-bp fragments by PCR was performed in a 25-μl reaction mixture containing 10 ng of genomic DNA, 10 pmol of each primer, 1 × ExTaq buffer, 2.5 μl of PCR DIG labeling mix (Roche Diagnostics, Switzerland) and 1.25 U of ExTaq DNA polymerase (TaKaRa Biomedicals, Japan). The thermal cycling condition was 95°C for 30 s, 55°C for 30 s and 72°C for 3 min (45 cycles). The labeled DNA dissolved in 5 × SSC, 0.5% blocking reagent (Roche Diagnostics), 0.1% sodium N-lauroylsarcosine and 0.02% SDS was hybridized with the dot-blotted membrane at 40°C. After hybridization, the membrane was washed twice in 2 × SSC, 0.1% SDS at room temperature for 5 min and then twice in 1 × SSC, 0.1% SDS buffer at 40, 42, 45°C or 50°C for 20 min. Signals were detected using the DIG DNA Detection Kit (Roche Diagnostics) according to the manufacturer’s instructions.
Dot-blot analysis with digoxigenin-labeled oligonucleotide probes: dot-blot-S-screening
Amplification of ca. 350-bp fragments by PCR with the same degenerated consensus primers as those described in the former section was performed in a 25-μl reaction mixture containing 10 ng of genomic DNA, 10 pmol of each primer, 1 × ExTaq buffer, 200 μM dNTPs and 1.25 U ExTaq DNA polymerase. The thermal cycling condition was 95°C for 30 s, 55°C for 30 s and 72°C for 1 min (35 cycles). The amplified DNA was denatured in a solution of 0.4 N NaOH and 10 mM EDTA, and then dot-blotted onto a nylon membrane by Multi-pin Blotter (Atto, Japan). The digoxigenin-labeled oligonucleotide probe diluted in the same solution described in the above section was hybridized with the membrane at 60°C. The membrane was washed twice in 2 × SSC, 0.1% SDS at room temperature for 5 min and then twice in 0.1 × SSC, 0.1% SDS buffer at 60°C for 20 min. For competitive hybridization to discriminate between SFB-4 and SFB-4′ alleles, hybridization was carried out with a set of probe mixtures (PaSFB4-mix probe) containing 0.02 μM of the digoxigenin-labeled oligonucleotide for the SFB-4 allele (PaSFBoli-4; Table 2) and 0.2 μM of the unlabeled counterpart oligonucleotide for the SFB-4′ allele (PaSFBoli-4′) at 60°C, or with another set of probe mixtures (PaSFB4′-mix probe) containing 0.02 μM of the digoxigenin-labeled oligonucleotide for the SFB-4′ allele (PaSFBoli-4′new; Table 2) and 0.1 μM of the unlabeled counterpart oligonucleotide for the SFB-4 allele (PaSFBoli-4new) at 40°C. Washing was carried out with 0.1 × SSC, 0.1% SDS at 60°C for the PaSFB4-mix probe or with 1 × SSC, 0.1% SDS at 40°C for the PaSFB4′-mix probe. Signals were detected as previously described.
Results
S-genotyping with dot-blot analysis (dot-blot-S-genotyping) in Japanese plum
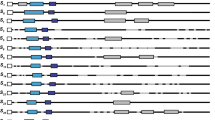
In Japanese plum, nineteen S-RNase alleles have been sequenced (Beppu et al. 2002, 2003; Sapir et al. 2004), while sequences of nine SFB alleles (SFB-a, -b, -c, -e, -f, -h, −7 and -10; Zhang et al. 2007, SFB-k; accession number EF490571) have been published to date. Since comparison of the HVa region of SFB indicated high sequence polymorphism among alleles (53–87% nucleotide identity), we designed allele-specific oligonucleotides for dot-blotting on the membrane by comparing the sequences of HVa and its adjacent regions. Using the cloned genes for SFB-b and –h alleles as templates, each digoxigenin-labeled DNA for SFB-b and –h was synthesized by PCR with consensus primers, resulting in amplification of ca. 350-bp products including the HVa region. Allele-specific signals for the SFB-b and SFB-h alleles were detected by hybridization of the allele-specific oligonucleotides blotted on the membrane with the digoxigenin-labeled DNAs (Fig. 1A). In addition, DNAs synthesized directly from genomic DNA of ‘Heibaoshi’ (S-b/S-h; Zhang et al. 2007) by PCR also detected the allele-specific signals for SFB-b and –h (Fig. 1B) consistent with the S-genotype of ‘Heibaoshi’.
Dot-blot analysis with SFB-allele-specific oligonucleotides in Japanese plum. Hybridization with digoxigenin-labeled DNAs synthesized from cloned SFB-b and SFB-h alleles by PCR (a), and digoxigenin-labeled DNAs synthesized from ‘Heibaoshi’ genomic DNA (b). Digoxigenin-labeled DNAs synthesized directly from genomic DNAs of 13 cultivars (c). a, b, c, e, f, h, k, 7 and 10 indicate allele-specific oligonucleotides designed from PsSFB-a, -b, -c, -e, -f, -h, -k, −7 and -10, respectively. Two dots were blotted for each allele in (c)
In the twelve other cultivars (Table 1), hybridization was carried out with each labeled DNA synthesized by PCR using each genomic DNA as a template. In ‘Meiguili’, ‘Nvgelei’, ‘Redgold’, ‘Weikeshun’, ‘Huangjiazuanshi’, ‘Kaiqinman’, ‘Younai’, ‘Huangpili’, ‘Akihime’ and ‘Ozarkpremier’, allele-specific signals consistent with their respective S genotypes were detected (Fig. 1C). Since the SFB-d gene has not yet been reported, only a signal for SFB-c was detected in ‘Oishiwasesumomo’ (S-c/S-d). Similarly, the DNA synthesized from ‘Dazhili’, in which one haplotype is S-10 and the other is unidentified, hybridized only with the oligonucleotide for SFB-10. These experiments suggest this dot-blotting method using sequence polymorphism of the SFB gene is useful for identification of all the nine S haplotypes used here.
Dot-blot-S-genotyping in sweet cherry
In sweet cherry, twenty-two S haplotypes have been identified in cultivated sweet cherry and wild cherry populations (Bošković and Tobutt 2001; Crane and Brown 1937; De Cuyper et al. 2005; Sonneveld et al. 2001, 2003), and S-RNase sequences of all these S haplotypes have been reported (Wünsch and Hormaza 2004; Sonneveld et al. 2001, 2003; De Cuyper et al. 2005). On the other hand, thirteen SFB alleles (SFB-1, −2, −3, −4, −4′, −5, −6, −7, −9, −10, −12, −13 and -16) among those S haplotypes have been characterized (Vaughan et al. 2006). Thirteen cultivars harboring ten alleles other than SFB-10, −12 and -13 were obtained as materials for this investigation (Table 1). Although the nucleotide identity of the HVa region between SFB-4 and -4′ is high (95%), because of only a 4-bp deletion in SFB-4′, comparison between the HVa sequences of the remaining alleles indicated high sequence polymorphism (55% −80% nucleotide identity). Therefore, we designed allele-specific oligonucleotides for dot-blotting on the membrane by comparing the sequences of HVa and those of adjacent regions as well as in Japanese plum (Table 2). Hybridization was carried out with each labeled DNA synthesized by PCR using consensus primers (Table 2) under the same condition as in Japanese plum. Labeled DNA synthesized from ‘Bigarreau de Jaboley’, ‘Early purple’, ‘Rodmersham seedling’, ‘Seneca’, ‘Vega’, ‘Satonishiki’, ‘Benisayaka’ and ‘Van’ detected allele-specific signals consistent with their respective S genotypes (Fig. 2A).
Dot-blot analysis with SFB-allele-specific oligonucleotides in sweet cherry. (a) Hybridization with digoxigenin-labeled DNAs synthesized from genomic DNAs of 13 cultivars by PCR. (b) Discrimination between S-4 and S-4′ haplotypes under different temperature conditions. 1, 2, 3, 4, 4′, 5, 6, 7, 9 and 16 indicate allele specific oligonucleotides designed from PsSFB-1, −2, −3, −4, −4′, −5, −6, −7, −9 and -16, respectively. Two dots were blotted for each allele
Because of the likelihood of identical sequence other than the 4-bp deletion causing cross-hybridization, signals for both SFB-4 and −4′ were detected in ‘Compact stella’, ‘Napoleon’, ‘Benishuho’, ‘Sam’ and ‘Taishonishiki’, together with the signals for the other SFB alleles. For distinct discrimination between SFB-4 and SFB-4′, the temperature condition in hybridization and washing was examined using ‘Napoleon’ harboring SFB-4 and ‘Compact Stella’ harboring SFB-4′. At 45°C, the signals for SFB-4 and SFB-4′ were preferentially detected in ‘Napoleon’ and ‘Compact Stella’, respectively (Fig. 2B). The contrast of signal intensities between these alleles was greater at 45°C than at 42°C (Fig. 2B). When the temperature was raised to 50°C, however, faint signals not allowing discrimination between these alleles were observed. Therefore, the 45°C condition for hybridization and washing is suggested to be optimum for individuals in which cross-hybridization has been observed.
S haplotype screening with digoxigenin-labeled allele-specific oligonucleotide probes (dot-blot-S-screening)
Although Fujimoto and Nishio (2003) and Shirasawa et al. (2005) dot-blotted genomic DNA on a membrane, low quality of genomic DNA can cause a weak signal (Shirasawa et al. 2005). Since it is difficult to extract genomic DNA of fruit trees with high quantity and quality due to impurities such as secondary metabolites or saccharide, we utilized PCR products as a dot-blotted DNA, as also done by Fujimoto and Nishio (2003) and Shirasawa et al. (2005).
The PCR product of ca. 350 bp for each sweet cherry cultivar, which is the same region used as a probe in the above experiments, was dotted on a membrane (Fig. 3A). The membrane was hybridized with a digoxigenin-labeled SFB-6 allele-specific oligonucleotide probe (Table 2) at 60°C. Signals were detected specifically in cultivars harboring an S-6 haplotype (Fig. 3A). In contrast, when hybridization was carried out with the digoxigenin-labeled SFB-4 allele-specific probe (PaSFBoli-4), signals were observed in cultivars harboring not only SFB-4 but also SFB4′, probably due to cross-hybridization (data not shown). A cross-hybridization signal was also observed in the case in which the digoxigenin-labeled SFB-4′ allele-specific probe (PaSFBoli-4′) was used. Shirasawa et al. (2006) reported a competitive hybridization method, which can detect sequence polymorphism such as SNP and insertion/deletion between one allele and a counterpart allele in rice. To improve the specific detection of SFB-4 or -4′, we utilized this competitive hybridization method. The addition of unlabeled PaSFBoli-4′ oligonucleotide to the hybridization buffer containing one-tenth the amount of the digoxigenin-labeled PaSFBoli-4 probe enabled specific detection of the SFB-4 allele (Fig. 3C). However, hybridization using the reverse set of probes detected a faint signal for SFB-4′ at 60°C (data not shown). In addition, lowering the temperature caused cross-hybridization signals (data not shown). Therefore, alternative oligonucleotides (PaSFBoli-4new and PaSFBoli-4′new; Table 2), which were shorter than the previous ones, were redesigned for the following competitive hybridization. The addition of the unlabeled PaSFBoli-4new oligonucleotide to the hybridization buffer containing one-fifth the amount of the digoxigenin-labeled PaSFBoli-4′new probe at 40°C resulted in detection of the specific signal of the SFB-4′ allele (Fig. 3D).
Screening of S haplotypes with allele-specific oligonucleotide probes. (A) The position for a PCR product of each cultivar dotted on membrane. a, Satonishiki; b, Napoleon; c, Taishonishiki; d, Benishuho; e, Benisayaka; f, Compact Stella; g, Bigarreau de Jaboley, h; Early Purple; i, Vega; j, Sam; k, Van; l, Seneca; m, Rodmersham Seedling (B) Hybridization with digoxigenin-labeled PaSFBoli-6 probe at 60°C followed by washing with 0.1 × SSC, 0.1% SDS at the same temperature. Competitive hybridization with the digoxigenin-labeled PaSFBoli-4 oligonucleotide mixed with the non-labeled PaSFBoli-4′ oligonucleotide (1:10) as a probe (C) or with digoxigenin-labeled PaSFBoli-4′-new mixed with non-labeled PaSFBoli-4-new (1:5) as a probe (D). Two dots were blotted for a PCR product of each cultivar
Discussion
Distinct S genotyping by dot-blotting
In this study, the dot-blot-S-genotyping technique was successful in discriminating between nine-haplotypes (nine SFB alleles) of Japanese plum and between ten S-haplotypes (ten SFB alleles) of sweet cherry. DNA amplification of the S-RNase and SFB alleles by PCR using the consensus primers in sweet cherry and Japanese plum sometimes results in the amplification of DNAs of similar size and is accompanied by the amplification of other DNA sequences, which makes S genotyping complicated (Tao et al. 1999; Yamane et al. 2000; Sonneveld et al. 2003; Beppu et al. 2003; Sapir et al. 2004; Vaughan et al. 2006). In the case of allele-specific primers, successful amplification of the alleles depends on several conditions such as the purity of template DNAs and the reaction temperature, and therefore, the reliability of S genotyping should be taken into consideration. False negative detection can be resolved by PCR-RFLP method. However, the method with S-RNase polymorphism has been developed only for S-1 and S-3 to S-6 haplotypes in sweet cherry (Yamane et al. 2000), but not yet in Japanese plum. Sakamoto et al. (2000) have identified twenty-one S haplotypes in a large number of cabbage and broccoli by their developed PCR-RFLP method, but the S haplotype identification has been confirmed by nucleotide sequencing because of the difficulty of reliable identification by PCR-RFLP, especially for heterozygotes. This implies that as PCR-RFLP markers in two Prunus species increase, great difficulty in S genotyping would be expected because most cultivars or selections in both fruit species are S heterozyote by nature. In contrast, the method developed in the present study showed distinct signals, demonstrating satisfactory reliability of S genotyping. Although Ortega et al. (2005), Sonneveld et al. (2006) and Vaughan et al. (2006) reported new methods for discriminating similar sizes of PCR products by a semi-automated sequencer, the present method (dot-blot-S-genotyping) enables distinct identification of S genotypes independent of DNA fragment sizes and, furthermore, is simpler and more cost-efficient than methods using a sequencer.
In sweet cherry, we were unable to test for the S-10, S-12 and S-13 haplotypes, whose SFB sequences have already been published, because of the unavailability of sweet cherry materials. However, preliminary experiments exhibited no cross-hybridization between designed oligonucleotides for S-10, S-12 and S-13 and the labeled PCR products of ca. 350 bp amplified from sweet cherry cultivars used in this study. Although the experiments using cultivars harboring S-10, S-12 and S-13 haplotypes are needed, in the future, the dot-blotting method herein presented would be more practical in S-genotyping for any S haplotypes.
According to the dot-blotting method reported by Fujimoto and Nishio (2003), the genomic DNAs or S-determinant PCR products of target haplotypes are necessary for dot-blotting on a membrane. Instead, we used oligonucleotides for target haplotypes, which resulted in satisfactory genotyping. Since the sequences of SP11s are highly variable between S haplotypes (Fujimoto and Nishio 2003), our method would be applicable to the identification of S haplotypes in Brassica species.
Dot-blotting useful for S screening
In order to select individuals harboring a target S haplotype, a membrane dotted with non-labeled PCR products for each individual was hybridized with the labeled-oligonucleotide probe in sweet cherry. The labeled-oligonucleotide probe for the S-6 haplotype enabled us to select cultivars harboring the S-6 haplotype (Fig. 3B). However, cross-hybridization between the SFB-4 and SFB-4′ alleles was observed with an independent probe for S-4 (PaSFBoli-4) or S-4′ (PaSFBoli-4′) (data not shown), possibly due to high sequence similarity. To date, Ikeda et al. (2004b) have succeeded in discrimination between SFB4 and SFB4′ by PCR using dCAPS primer sets that they designed, though PCR must be performed twice. Alternatively in this study, we were also successful in distinct and simple discrimination between these haplotypes utilizing the competitive hybridization method.
Although we used a cultivar population, S haplotypes of which have already been characterized, the dot-blot-S-screening method would be more practical than the method based on the PCR and would promote the screening of individuals that harbor the desirable S haplotype in a large-scale population.
Future application
In dot-blot-S-genotyping, we utilized a region of ca. 350 bp containing HVa. In our preliminary experiments, the longer sized fragments, ca. 500 bp, resulted in less intense signals, even when the temperature was below 40°C (data not shown), suggesting that the length of DNA fragments is important and that a length less than 500 bp is best for obtaining distinct signals. In Prunus, such as Japanese plum (Beppu et al. 2002, 2003; Sapir et al. 2004; Halász et al. 2007) and sweet cherry (Tao et al. 1999; Sonneveld et al. 2003), the first and second introns of S-RNase are highly variable in length. However, because the second introns of most alleles are more than 500 bp in length, they might not be suitable for dot-blot-S-genotyping. On the contrary, since the length of the first intron is less than 500 bp, dot-blot-S-genotyping might be applicable. The availability of allele specific oligonucleotides designed from S-RNase along with those from SFB for the dot-blot-S-genotyping method would enable more reliable S genotyping. As cloning of SFB alleles will progress not only in Prunoideae but also in Maloideae such as apple and Japanese pear (Cheng et al. 2006; Sassa et al. 2007), the diversity of the HVa sequence will be further elucidated. To deal with the larger number of alleles and the diversity of the SFB gene, methodological improvements contributing not only to agronomical utilization of the S haplotype but also to convenient analyses of S haplotypes for population genetics are needed. Thus, this alternative strategy for S genotyping and S screening herein presented would seem to be promising.
Abbreviations
- GSI:
-
gametophytic self-incompatibility
- SFB:
-
S-haplotype-specific F-box protein
- PCR:
-
polymerase chain reaction
- RFLP:
-
restriction fragment length polymorphism
- HVa:
-
hypervariable region a
References
Beppu K, Takemoto Y, Yamane H, Yaegaki H, Yamaguchi M, Kataoka I, Tao R (2003) Determination of S-haplotypes of Japanese plum (Prunus salicina Lindl.) cultivars by PCR and cross-pollination tests. J Hort Sci Biotechnol 78:315–318
Beppu K, Yamane H, Yaegaki H, Yamaguchi M, Kataoka I, Tao R (2002) Diversity of S-RNase genes and S-haplotypes in Japanese plum (Prunus salicina Lindl.). J Hort Sci Biotechnol 77:658–664
Bošković R, Russell K, Tobutt KR (1997) Inheritance of stylar ribonucleases in cherry progenies, and reassignment of incompatibility alleles to two incompatibility groups. Euphytica 95:221–228
Bošković R, Tobutt KR (2001) Genotyping cherry cultivars assigned to incompatibility groups, by analyzing stylar ribonucleases. Theor Apple Genet 103:475–485
Cheng J, Han Z, Xu X, Li T (2006) Isolation and identification of the pollen-expressed polymorphic F-box genes linked to the S-locus in apple (Malus × domestica). Sex Plant Reprod 19:175–183
Crane MB, Brown AG (1937) Incompatibility and sterility in the sweet cherry, Prunus avium L. J Pomol Hortic Sci 15:86–116
De Cuyper B, Sonneveld T, Tobutt KR (2005) Determining self-incompatibility genotypes in Belgian wild cherries. Mol Ecol 14:945–955
Doyle JJ, Doyle JL (1987) A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem Bull 19:11–15
Entani T, Iwano M, Shiba H, Che FS, Isogai A, Takayama S (2003) Comparative analysis of the self-incompatibility (S-) locus region of Prunus mume: identification of a pollen-expressed F-box gene with allelic diversity. Genes Cells 8:203–213
Fujimoto R, Nishio T (2003) Identification of S haplotypes in Brassica by dot-blot analysis of SP11 alleles. Theor Appl Genet 106:1433–1437
Halász J, Hegedüs A, Szabó Z, Nyéki J, Pedryc A (2007) DNA-based S-genotyping of Japanese plum and pluot cultivars to clarify incompatibility relationships. HortScience 42:5–184
Ikeda K, Igic B, Ushijima K, Yamane H, Hauck NR, Nakano R, Sassa H, Iezzoni AF, Kohn JR, Tao R (2004a) Primary structural features of the S haplotype-specific F-box protein, SFB, in Prunus. Sex Plant Reprod 16:235–243
Ikeda K, Watari K, Ushijima K, Yamane H, Hauck NR, Iezzoni AF, Tao R (2004b) Molecular makers for the self-compatible S-4′-haplotype, a pollen-part mutant in sweet cherry (Prunus avium L.). J Amer Soc Hort Sci 129:724–728
Ikeda K, Ushijima K, Yamane H, Tao R, Hauck NR, Sebolt AM, Iezzoni AF (2005) Linkage and physical distance between the S-haplotype S-RNase and SFB genes in sweet cherry. Sex Plant Reprod 17:289–296
Ishimizu T, Shinkawa T, Sakiyama F, Norioka S (1998) Primary structural features of rosaceous S-RNases with gametophytic self-incompatibility. Plant Mol Biol 37:931–941
Janssens GA, Goderis IJ, Broekaert F, Broothaerts W (1995) A molecular method for S-allele identification in apple based on allele-specific PCR. Theor Appl Genet 91:691–698
Lewis D (1949) Structure of the incompatibility gene. II. Heredity 3:339–355
Moriya Y, Yamamoto K, Okada K, Iwanami H, Bessho H, Nakanishi T, Takasaki T (2007) Development of a CAPS marker system for genotyping European pear cultivars harboring 17 S alleles. Plant Cell Rep 26:345–354
Ortega E, Sutherland BG, Dicenta F, Bošković R, Tobutt KR (2005) Determination of incompatibility genotypes in almond using first and second intron consensus primers: detection of new S alleles and correction of reported S genotypes. Plant Breeding 124:188–196
Romero C, Vilanova S, Burgos L, Martínez-Calvo J, Vicente M, Llácer G, Madenes ML (2004) Analysis of S-locus structure in Prunus armeniaca L: identification of S-haplotype specific S-RNase and F-box genes. Plant Mol Biol 56:145–157
Sakamoto K, Kusaba M, Nishio T (2000) Single-seed PCR-RFLP analysis for the identification of S haplotypes in commercial F1 hybrid cultivars of broccoli and cabbage. Plant Cell Rep 19:400–406
Sapir G, Stern RA, Eisikowitch DE, Goldway M (2004) Cloning of four new Japanese plum S-alleles and determination of the compatibility between cultivars by PCR analysis. J Hort Sci Biotechnol 79:223–227
Sassa H, Hirano H, Ikehashi H (1992) Self-incompatibility-related RNases in styles of Japanese pear (Pyrus serotina Rehd.). Plant Cell Physiol 33:811–814
Sassa H, Kakui H, Miyamoto M, Suzuki Y, Hanada T, Ushijima K, Kusaba M, Hirano H, Koba T (2007) S locus F-box brothers: multiple and pollen-specific F-box genes with S haplotype-specific polymorphisms in apple and Japanese pear. Genetics 175:1869–1881
Sassa H, Mase N, Hirano H, Ikehashi H (1994) Identification of self-incompatibility related glycoproteins in styles of apple (Malus x domestica). Theor Apple Genet 89:201–205
Shirasawa K, Kishitani S, Nishio T (2005) Dot-blot analysis for identification of japonica rice cultivars and genotyping of recombinant inbred lines. Breed Sci 55:187–192
Shirasawa K, Shiokai S, Yamaguchi M, Kishitani S, Nishio T (2006) Dot-blot-SNP analysis for practical plant breeding and cultivar identification in rice. Theor Appl Genet 113:147–155
Sonneveld T, Robbins TP, Bošković R, Tobutt KR (2001) Cloning of six cherry self-incompatibility alleles and development of allele-specific PCR detection. Theor Appl Genet 102:1046–1055
Sonneveld T, Robbins TP, Tobutt KR (2006) Improved discrimination of self-incompatibility S-RNase alleles in cherry and high throughput genotyping by automated sizing of first intron polymerase chain reaction products. Plant Breeding 125:305–307
Sonneveld T, Tobutt KR, Robbins TP (2003) Allele-specific PCR detection of sweet cherry self-incompatibility (S) alleles S 1 to S 16 using consensus and allele-specific primers. Theor Appl Genet 107:1059–1070
Sonneveld T, Tobutt KR, Vaughan SP, Robbins TP (2005) Loss of pollen-S function in two self-compatible selections of Prunus avium is associated with deletion/mutation of an S haplotype-specific F-box gene. Plant Cell 17:37–51
Tamura M, Ushijima K, Sassa H, Hirano H, Tao R, Gradziel TM, Dandekar AM (2000) Identification of self-incompatibility genotypes of almond by allele-specific PCR analysis. Theor Appl Genet 101:344–349
Tao R, Habu T, Namba A, Yamane H, Fuyuhiro F, Iwamoto K, Sugiura A (2002) Inheritance of S f-RNase in Japanese apricot (Prunus mume) and its relation to self-compatibility. Theor Appl Genet 105:222–228
Tao R, Yamane H, Sassa H, Mori H, Gradziel TM, Dandekar AM, Sugiura A (1997) Identification of stylar RNases associated with gametophytic self-incompatibility in almond (Prunus dulcis). Plant Cell Physiol 38:304–311
Tao R, Yamane H, Sugiura A, Murayama H, Sassa H, Mori H (1999) Molecular typing of S-alleles through identification, characterization and cDNA cloning for S-RNases in sweet cherry. J Am Soc Hortic Sci 124:224–233
Ushijima K, Sassa H, Tao R, Yamane H, Dandekar AM, Gradziel TM, Hirano H (1998) Cloning and characterization of cDNAs encoding S-RNases from almond (Prunus dulcis): primary structural features and sequence diversity of the S-RNases in Rosaceae. Mol Gen Genet 260:261–268
Ushijima K, Sassa H, Dandekar AM, Gradziel TM, Tao R, Hirano H (2003) Structural and transcriptional analysis of the self-incompatibility locus of almond: identification of a pollen-expressed F-box gene with haplotype-specific polymorphism. Plant Cell 15:771–781
Ushijima K, Yamane H, Watari A, Kakehi E, Ikeda K, Hauck NR, Iezzoni AF, Tao R (2004) The S haplotype-specific F-box protein gene, SFB, is defective in self-compatible haplotypes of Prunus avium and P. mume. Plant J 39:573–586
Vaughan SP, Russell K, Sargent DJ, Tobutt KR (2006) Isolation of S-locus F-box alleles in Prunus avium and their application in a novel method to determine self-incompatibility genotype. Theor Appl Genet 112:856–866
Wünsch A, Hormaza JI (2004) Cloning and characterization of genomic DNA sequences of four self-incompatibility alleles in sweet cherry (Prunus avium L.). Theor Appl Genet 108:299–305
Yaegaki H, Shimada T, Moriguchi T, Hayama H, Haji T, Yamaguchi M (2001) Molecular characterization of S-RNase genes and S-genotypes in the Japanese apricot (Prunus mume Sieb. et Zucc.). Sex Plant Reprod 13:251–257
Yamane H, Tao R, Murayama H, Sugiura A (2000) Determining the S-genotypes of several sweet cherry cultivars based on PCR-RFLP analysis. J Hort Sci Biotechnol 75:562–567
Yamane H, Tao R, Sugiura A (1999) Identification and cDNA cloning for S-RNases in self-incompatible Japanese plum (Prunus salicina Lindl. cv. Sordum). Plant Biotech 16:389–396
Yamane H, Tao R, Sugiura A (2001) Identification and characterization of S-RNases in tetraploid sour cherry (Prunus cerasus). J Am Soc Hortic Sci 126:661–667
Yamane H, Ikeda K, Ushijima K, Sassa H, Tao R (2003a) A pollen-expressed gene for a novel protein with an F-box motif that is very tightly linked to a gene for S-RNase in two species of cherry, Prunus cerasus and P. avium. Plant Cell Physiol 44:764–769
Yamane H, Ushijima K, Sassa H, Tao R (2003b) The use of the S haplotype-specific F-box protein gene, SFB, as a molecular marker for S-haplotypes and self-compatibility in Japanese apricot (Prunus mume). Theor Appl Genet 107:1357–1361
Zhang SL, Huang SX, Kitashiba H, Nishio T (2007) Identification of S-haplotype-specific F-box gene in Japanese plum (Prunus salicina Lindl.). Sex Plant Reprod 20:1–8
Zuccherelli S, Tassinari P, Broothaerts W, Tartarini S, Dondini L, Sansavini S (2002) S-allele characterization in self-incompatible pear (Pyrus communis L.). Sex Plant Reprod 15:153–158
Acknowledgements
This work was supported in part by a fund from the Dean of the Graduate School of Agricultural Science, Tohoku University.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kitashiba, H., Zhang, S.L., Wu, J. et al. S genotyping and S screening utilizing SFB gene polymorphism in Japanese plum and sweet cherry by dot-blot analysis. Mol Breeding 21, 339–349 (2008). https://doi.org/10.1007/s11032-007-9134-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11032-007-9134-6