Abstract
Rho GTPases are molecular switches that play an important role in regulating the behavior of a variety of tumor cells. RhoA GTPase-activating protein 26 (ARHGAP26) is a GTPase-activating protein and inhibits the activity of Rho GTPases by promoting the hydrolytic ability of Rho GTPases. It also affects tumorigenesis and progression of various tumors through several methods, including formation of abnormal fusion genes and circular RNA. This review summarizes the biological functions and molecular mechanisms of ARHGAP26 in different tumors, proposes the potential clinical value of ARHGAP26 in cancer treatment, and discusses current issues that need to be addressed.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
RhoA GTPase-activating protein 26 (ARHGAP26) was first discovered in avian tissues by Hildebrand et al. in 1996 when they studied focal adhesion kinase (FAK) in integrin-mediated signaling. As such, this protein was initially named as GTPase regulator associated with FAK (GRAF) [1]. As a type of GTPase-activating protein (GAP), ARHGAP26 enhances the hydrolysis of GTPases and converts GTPases from an active form to an inactive form, thereby inhibiting signaling transduction [2]. Numerous studies have demonstrated that ARHGAP26 expression and involvement in tumorigenesis and tumor progression are not the same in different tumors. For example, ARHGAP26 expression is significantly reduced in acute myeloid leukemia (AML) [3], chronic myeloid leukemia (CML) [3, 4], and ovarian cancer [5], while the transcription factor activity of ARHGAP26 is significantly increased and expression of ARHGAP26 is upregulated in prostate cancer [6]. In gastric cancer, ARHGAP26 is fused with claudin-18 gene (CLDN18), and the translated abnormal fusion protein regulates the development of gastric cancer [7,8,9], while circular RNA ARHGAP26 (circ-ARHGAP26) modulates microRNA through a “sponge” mechanism to affect the progression of gastric cancer [10,11,12]. In glioblastoma, ARHGAP26 acts as an important executive molecule downstream of the integrin complex to promote tumor invasion [13]. This article reviews the biological functions, molecular mechanisms, and clinical characteristics of ARHGAP26 in different tumors (Table 1), proposes its potential clinical applications, and explores the research directions and unsolved issues in related fields.
Role of ARHGAP26 in tumors and its potential clinical value
ARHGAP26 in gastric cancer
A study of 295 cases of gastric cancer in The Cancer Genome Atlas (TCGA) sample database showed that gastric cancer can be divided into four subtypes based on the molecular characterization. Expression of the CLDN18-ARHGAP26 fusion gene was present in patients with a genomically stable subtype (Fig. 1) [7], which is the first report of the ARHGAP26 fusion gene. Since then, multiple studies have shown the existence of this abnormal fusion gene [9, 14,15,16]. Based on research data, 151 out of 1908 (7.9%) patients with gastric cancer had this gene fusion [17]. Analysis of clinical characteristics showed that the CLDN18-ARHGAP26 fusion gene is more common in female patients [9], young patients, and patients with diffuse-type gastric cancer (Lauren’s classification) [15] and is associated with poor prognosis [9, 14]. In vitro experiments have shown that the presence of the CLDN18-ARHGAP26 fusion gene increases the migration and invasion ability of gastric cancer cells [8, 9, 15], as well as the resistance of tumor cells to chemotherapy drugs [9].
Schematic diagram of CLDN18-ARHGAP26 fusion gene and fusion protein. In the upper panel, the CLDN18 gene is fused with the ARHGAP26 gene, which initiates the translation of the fusion protein CLDN18-ARHGAP26 at a cryptic splice site. In the lower panel, the CLDN18-ARHGAP26 fusion protein contains the transmembrane domain of CLDN18 and the GAP and SH3 domains of ARHGAP26. CLDN18, claudin 18; ARHGAP26, RhoA GTPase-activating protein 26; GAP GTPase-activating protein, SH3 src homology 3, UTR untranslated region
It is noteworthy that the high affinity of the CLDN18-ARHGAP26 fusion protein for the cell membrane, as well as its exclusive expression in gastric cancer cells provide new opportunities for targeted therapies [16]. CLDN18 protein is involved in the formation of tight junctions in epithelial cells [18]. A CLDN18 protein subtype, CLDN18.2, is the specific CLDN18 expressed in the stomach and is an important component of the gastric acid barrier [19,20,21,22]. Zolbetuximab, formerly ideal monoclonal antibody 362 (IMAB362), is a new type of immunoglobulin G1 antibody that is highly specific for the CLDN18.2 protein [22,23,24]. Thus, it has the potential to be used in targeted therapy for gastric cancer patients with the CLDN18-ARHGAP26 fusion gene. This drug is currently undergoing clinical trials to evaluate its efficacy and safety [25, 26].
Through circular RNA microarrays, Shao et al. showed that the top ten upregulated circular RNAs in gastric cancer tissues included hsa_cirs_0074362, i.e., circ-ARHGAP26 (7.58-fold change, p = 0.01156) [27]. Normally, the ARHGAP26 mRNA is transcribed from 46 kb of the genomic DNA at chromosome region 5q31.3, containing 23 exons. However, the circ-ARHGAP26 only contains the transcript from exon 5 to exon 11 in this chromosome region. Due to its structure, Xie et al. used divergent primers for circ-ARHGAP26 amplification and showed by RT-PCR that the expression of circ-ARHGAP26 was significantly lower in gastric cancer tissues than in paired normal adjacent tissues [12]. The expression of circ-ARHGAP26 was also lower in five gastric cancer cell lines (AGS, BGC-823, HGC-27, MGC-803, and SGC-7901) than that in the normal gastric epithelial GES-1 cell line. In addition, the expression of circ-ARHGAP26 in gastric tissues of patients with mild and moderate gastritis was also significantly lower than that of the normal gastric tissues, but higher than that of gastric cancer tissues [12]. In view of the contradictory results of these studies [12, 27], Lv et al. assessed the levels of circ-ARHGAP26 in gastric cancer cells and normal gastric epithelial cells and showed that circ-ARHGAP26 level was significantly lower in gastric cancer cell lines than in normal gastric epithelial cells [11]. In functional experiments, downregulation of circ-ARHGAP26 resulted in a decline in the proliferation of gastric cancer epithelial cells and an increase in apoptosis. Based on the above results, the expression of circ-ARHGAP26 in gastric cancer should be verified further. Meanwhile, more investigations are warranted to explore the underlying mechanism through which circ-ARHGAP26 affects tumorigenesis and progression of gastric cancer.
ARHGAP26 in myeloid malignancies
Previous studies have shown that the expression of ARHGAP26 in AML was significantly lower than that in the control group [3, 4]. Further studies showed that AML patients with relatively high expression of ARHGAP26 have a longer overall survival. The expression of ARHGAP26 has no significant correlation with clinical characteristics, the French-American-British (FAB) classification, or cytogenetic risk subgroups of AML patients [3]. Similarly, the expression of ARHGAP26 in CML was also significantly lower than that in the controls. In addition, the expression of ARHGAP26 significantly decreased with the progression of CML. No significant difference in the expression of ARHGAP26 was found between CML patients in the remission phase and chronic phase [4]. In terms of the mechanism, a significant increase in the methylation ratio of the ARHGAP26 promoter in patients is an important reason for the decrease in ARHGAP26 expression [28]. Another study showed that the transcription level of ARHGAP26 was significantly lower in AML patients compared with the normal population, regardless of whether the ARHGAP26 promoter was methylated. In addition, patients with a methylated ARHGAP26 promoter had a lower ARHGAP26 transcription level, indicating that hypermethylation of the ARHGAP26 promoter was an early event in AML progression [29].
Gene fusion is a common chromosomal structural abnormality in patients with acute leukemias. The majority of translocations that occur at 11q23 disrupt mixed lineage leukemia gene (MLL) and fuse it to many different partner genes [30]. Studies have shown that the MLL/ARHGAP26 fusion is a chromosomal abnormality in infants and young children with AML. Through case summary analysis, it was found that infants with the MLL/ARHGAP26 fusion responded well to treatment [30,31,32].
ARHGAP26 in glioblastoma
Studies have shown that the integrin complex CD151-α3β1 significantly promotes the invasion and migration of glioblastoma, and ARHGAP26 is a key molecule downstream of the complex [13]. Moreover, antibodies against ARHGAP26 were first discovered in patients with subacute cerebellar ataxia [33, 34]. Subsequent studies have shown that these antibodies are related to cognitive impairment and dyslexia [35,36,37,38]. Interestingly, researchers also detected other systemic tumors in patients with such neurological diseases who were positive for ARHGAP26 antibodies, suggesting that anti-ARHGAP26 antibodies may be a potential tumor predictor in such patients [38, 39]. Nevertheless, the role that these antibodies play in the tumorigenesis of other cancers remains unclear. Large-scale clinical trials should be carried out to investigate the value of using anti-ARHGAP26 antibodies to identify patients with the aforementioned characteristics.
ARHGAP26 in prostate cancer
An analysis of different genes in a cohort from the Gene Expression Omnibus (GEO) database, which contained 18 prostate cancer tissues and 21 normal tissues, showed that the activity of transcription factor specificity protein 1 (SP1), which significantly upregulates the expression of ARHGAP26, was significantly upregulated in prostate cancer [6]. However, no further in vivo or in vitro studies were conducted to clarify the impact of ARHGAP26 on the phenotype of prostate cancer. In addition, the correlation between the clinicopathological characteristics of prostate cancer patients and upregulation of ARHGAP26 expression is also worth exploring.
ARHGAP26 in ovarian cancer
A previous study showed that expression of ARHGAP26 in ovarian cancer tissues was significantly reduced and related to poor prognosis of the patients [5]. Cytological experiments have shown that upregulation of ARHGAP26 leads to a decrease in cell proliferation, migration, and invasion and downregulation of the downstream molecules, including GTP-RhoA, β-catenin, vascular endothelial growth factor (VEGF), matrix metallopeptidase (MMP)2, and MMP7, whereas downregulation of ARHGAP26 leads to the opposite cellular phenotypes—all of which can be inhibited by the β-catenin inhibitor, DKK1. Animal experiments have shown that upregulation of ARHGAP26 reduces the lung metastasis of ovarian cancer cells. Interestingly, upregulation of ARHGAP26 in SKOV3 ovarian carcinoma cells was shown to effectively inhibit the migration and invasion of tumor cells due to the upregulation of smad ubiquitination regulatory factor 1 (SMURF1). As an E3 ubiquitination ligase, SMURF1 degrades ARHGAP26 through the ubiquitination pathway. Thus, researchers believe that SMURF1-dependent regulation of ARHGAP26 ubiquitination promotes the invasion and migration of ovarian cancer cells through the β-catenin pathway [5].
Mechanism of ARHGAP26 in cancers
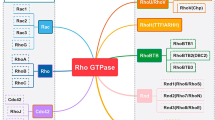
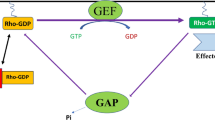
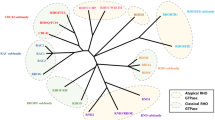
Rho GTPases are a group of signaling proteins belonging to the Ras GTPases superfamily [40, 41]. There are 22 types of mammalian Rho GTPases [42]. Among them, RhoA, rac family small GTPase 1 (Rac1), and cell division cycle 42 (Cdc42) are the most commonly studied. Rho GTPases are molecular switches that regulate signaling pathways through GTP-loading/GTP-hydrolysis (Fig. 2), namely GAPs, guanine nucleotide exchange factors (GEFs), and Rho GDP dissociation inhibitors (GDIs) [43]. GAPs promote GTPase hydrolysis to inhibit signaling transduction. GEFs catalyze the GTPase-loading reaction to stimulate signaling transduction, and GDIs inhibit the dissociation of GDP by binding to GDP-bound Rho GTPase, thereby inhibiting the activation of signaling pathways [43, 44]. Rho GTPases play an important role in cellular processes, including cell adhesion and polarity, cell morphology and movement, exchanges between vesicles and cell membranes, cell cycle, cell division, and cell differentiation (Fig. 2) [44,45,46]. In addition, Rho GTPases are closely related to tumorigenesis and tumor progression.
The cycle of the GTP-binding and GTP-hydrolysis and signaling functions of Rho GTPases involved in cells. The schematic shows how GEFs, GAPs, and GDIs regulate the cycle of the Rho GTPase signaling pathway and the associated cell behaviors. GEF guanine nucleotide exchange factor, GAP GTPase-activating protein; GDI Rho GDP dissociation inhibitor
ARHGAP26 is a GAP, and it is specific for only Cdc42 and Rho A [1, 2]. ARHGAP26 converts both proteins from the GTP to GDP forms, which inactivates them, thereby regulating downstream molecules [44]. The ARHGAP26 molecule includes the GAP domain in the central region, the src homology 3 (SH3) domain in the C segment, and the Bin/amphiphysin/Rvs (BAR) and pleckstrin homology (PH) domains at the N-terminal. These domains are highly conserved among GAPs and GEFs [47, 48]. The GAP domain mainly inactivates Rho GTPases by enhancing the hydrolysis of GTPases [1, 2]. The SH3 domain is combined by a variety of regulatory molecules to regulate the activity of the GAP domain of ARHGAP26 [1, 2, 49]. The BAR and PH domains of ARHGAP26 regulate endocytosis by binding and inducing membrane curvature [50,51,52,53]. Numerous studies have shown that Rho GTPases play important, but different roles in a variety of tumors [54,55,56]. Thus, this partly explains the difference in the role of their upstream regulatory molecule, ARHGAP26, in different tumors.
Abnormal fusion of ARHGAP26 with other genes was first reported in 2000 [30]. ARHGAP26 was fused to MLL in a unique t (5; 11) (q31; q23) fusion in infants with myelodysplastic syndrome (MDS) and AML. Since then, several cases with similar fusion genes have been reported; however, MLL/ARHGAP26 fusion is relatively rare in patients with hematological malignancies [31, 32]. In 2014, the fusion of ARHGAP26 and CLDN18 (CLDN18-ARHGAP26 fusion) was first reported in gastric cancer patients [7]. There are many types of CLDN18-ARHGAP26 fusion. Among them, CLDN18/exon5-ARHGAP26/exon12 fusion is the most common [17]. The CLDN18-ARHGAP26 fusion translates into the CLDN18-ARHGAP26 fusion protein by activating a cryptic splice site within exon 5 of CLDN18 (Fig. 1) [14, 15, 18]. The fusion protein retains the GAP domain located in the central region of ARHGAP26 and the SH3 domain located at the C-terminal [8, 17], due to which the fusion protein retains some functions, including regulating its downstream RhoA pathway and mediating the integrin signaling pathway [1, 2, 49]. The N-terminal BAR and PH domains of wild-type ARHGAP26 are missing from the fusion protein, so the GAP domain activity of the fusion protein cannot be regulated by its upstream molecules, and the ability of wild-type ARHGAP26 to regulate the endocytosis pathway is also affected [51, 57,58,59]. CLND18 protein is involved in the formation of tight junctions in epithelial cells and plays a crucial role in the gastric acid barrier [19,20,21,22]. The CLDN18-ARHGAP26 fusion protein only has the transmembrane domain of CLDN18 and lacks the cytoplasmic part at the C-terminus. Therefore, the adhesion function of epithelial cells expressing the fusion protein is impaired, which, in turn, leads to dysfunction of the gastric mucosal barrier [7, 8, 16, 17, 21]. The two abnormal functions of the fusion protein that promote the invasion and migration of gastric cancer cells also partly explain the decreased survival rates and worse prognosis of patients with diffuse gastric cancer with the CLDN18-ARHGAP26 fusion compared with those without the CLDN18-ARHGAP26 fusion.
Circ-ARHGAP26 was first discovered in human gastric cancer tissues by Shao et al. in 2017 [27]. Circular RNA, also known as circRNA, is a long-chain, non-coding precursor RNA (pre-RNA) formed by non-canonical splicing. Its most important biological function is to regulate gene expression [60]. Post-transcriptional regulation has been an extensively recognized method for circRNA to regulate gene expression, (i.e., using circRNA as a “microRNA sponge” or “microRNA reservoir” for sequestering and regulating microRNA) [61, 62]. Studies have shown that regulation of circRNA plays an important role in tumorigenesis and tumor progression [60, 63]. Moreover, it has been reported that circ-ARHGAP26 is closely related to the expression of the tumor marker carbohydrate antigen 19–9 (CA19-9) [12], which may provide evidence for the involvement of circ-ARHGAP26 in the expression and regulation of tumor-related proteins.
Issues to be solved
First, the upstream regulators responsible for the abnormal expression of ARHGAP26 in tumors are the most important issues that need to be addressed. The regulatory mechanisms of ARHGAP26 in different tumors vary, including changes in the expression level of ARHGAP26, abnormal fusion of ARHGAP26 with other genes, and microRNA sponge mechanism of circ-ARHGAP26. Understanding the cause of the abnormal expression of ARHGAP26 aids in evaluation of the tumor risk and implementation of pre-intervention.
Second, downstream mechanisms responsible for the abnormal expression of ARHGAP26 in tumors still need to be further investigated, including the expression of circ-ARHGAP26 in gastric cancer patients. The mechanism of regulation of microRNAs by circ-ARHGAP26 to alter the expression of downstream genes still needs to be explored. The underlying mechanism of the effects of CLDN18-ARHGAP26 fusion protein on the downstream molecules to cause tumorigenesis and progression of gastric cancer also needs to be further studied.
Lastly, although the clinical trials using the Zolbetuximab monoclonal antibodies against CLDN18.2 have reported encouraging results [25, 26], there are still problems that need to be resolved. A question worth exploring is whether CLDN18-ARHGAP26 fusion has a positive correlation with the CLDN18.2 protein expression. The answer to this question will determine whether this fusion gene can be used as a specific indicator of tumors that would be sensitive to Zolbetuximab-targeted therapy [17]. In addition, given the differences between the Eastern and Western populations, Zolbetuximab in combination with other chemotherapies exerts a greater toxicity in Asian people than in other populations [22]. Therefore, the tolerability and therapeutic effect of the combination of Zolbetuximab and other chemotherapeutics in the Asian population still needs to be evaluated.
Conclusion
ARHGAP26, a GAP regulating the Rho GTPases, plays a crucial role in tumorigenesis and progression of human cancers. The expression level and pattern of ARHGAP26 in various tumors are quite different. However, for a given type of cancer, it can be used as a biomarker because of its altered expression compared with normal tissues or cells. Moreover, the CLDN18-ARHGAP26 fusion protein may became a novel target for the treatment of gastric cancer patients in the future. Notably, although many studies have explored the biological functions and molecular mechanisms of ARHGAP26, there are still many contradictions and problems worth exploring (Fig. 3). Therefore, more basic experiments and large-scale clinical trials are needed in this field.
Data availability
All data were included in the manuscript.
References
Hildebrand JD, Taylor JM, Parsons JT (1996) An SH3 domain-containing GTPase-activating protein for Rho and Cdc42 associates with focal adhesion kinase. Mol Cell Biol 16:3169–78. https://doi.org/10.1128/mcb.16.6.3169
Taylor JM, Macklem MM, Parsons JT (1999) Cytoskeletal changes induced by GRAF, the GTPase regulator associated with focal adhesion kinase, are mediated by Rho. J Cell Sci 112(Pt 2):231–42
Aly RM, Ghazy HF (2014) High expression of GTPase regulator associated with the focal adhesion kinase (GRAF) is a favorable prognostic factor in acute myeloid leukemia. Blood Cells Mol Dis 53:185–8. https://doi.org/10.1016/j.bcmd.2014.07.004
Qian Z, Qian J, Lin J, Yao DM, Chen Q, Ji RB, Li Y, Xiao GF, Li JY (2010) GTPase regulator associated with the focal adhesion kinase (GRAF) transcript was down-regulated in patients with myeloid malignancies. J Exp Clin Cancer Res 29:111. https://doi.org/10.1186/1756-9966-29-111
Chen X, Chen S, Li Y, Gao Y, Huang S, Li H, Zhu Y (2019) SMURF1-mediated ubiquitination of ARHGAP26 promotes ovarian cancer cell invasion and migration. Exp Mol Med 51:1–12. https://doi.org/10.1038/s12276-019-0236-0
Li J, Xu YH, Lu Y, Ma XP, Chen P, Luo SW, Jia ZG, Liu Y, Guo Y (2013) Identifying differentially expressed genes and small molecule drugs for prostate cancer by a bioinformatics strategy. Asian Pac J Cancer Prev 14:5281–6. https://doi.org/10.7314/apjcp.2013.14.9.5281
Cancer Genome Atlas Research Network (2014) Comprehensive molecular characterization of gastric adenocarcinoma. Nature 513:202–9. https://doi.org/10.1038/nature13480
Yao F, Kausalya JP, Sia YY, Teo AS, Lee WH, Ong AG, Zhang Z, Tan JH, Li G, Bertrand D, Liu X, Poh HM, Guan P, Zhu F, Pathiraja TN, Ariyaratne PN, Rao J, Woo XY, Cai S, Mulawadi FH, Poh WT, Veeravalli L, Chan CS, Lim SS, Leong ST, Neo SC, Choi PS, Chew EG, Nagarajan N, Jacques PÉ, So JB, Ruan X, Yeoh KG, Tan P, Sung WK, Hunziker W, Ruan Y, Hillmer AM (2015) Recurrent fusion genes in gastric cancer: CLDN18-ARHGAP26 induces loss of epithelial integrity. Cell Rep 12:272–85. https://doi.org/10.1016/j.celrep.2015.06.020
Shu Y, Zhang W, Hou Q, Zhao L, Zhang S, Zhou J, Song X, Zhang Y, Jiang D, Chen X, Wang P, Xia X, Liao F, Yin D, Chen X, Zhou X, Zhang D, Yin S, Yang K, Liu J, Fu L, Zhang L, Wang Y, Zhang J, An Y, Cheng H, Zheng B, Sun H, Zhao Y, Wang Y, Xie D, Ouyang L, Wang P, Zhang W, Qiu M, Fu X, Dai L, He G, Yang H, Cheng W, Yang L, Liu B, Li W, Dong B, Zhou Z, Wei Y, Peng Y, Xu H, Hu J (2018) Prognostic significance of frequent CLDN18-ARHGAP26/6 fusion in gastric signet-ring cell cancer. Nat Commun 9:2447. https://doi.org/10.1038/s41467-018-04907-0
Song X, Zhang N, Han P, Moon BS, Lai RK, Wang K, Lu W (2016) Circular RNA profile in gliomas revealed by identification tool UROBORUS. Nucleic Acids Res 44:e87. https://doi.org/10.1093/nar/gkw075
Wangxia LV, Fang Y, Liu Y, Zhao Y, Shi Z, Zhong H (2019) Circular RNA ARHGAP26 is over-expressed and its downregulation inhibits cell proliferation and promotes cell apoptosis in gastric cancer cells. Saudi J Gastroenterol 25:119–125. https://doi.org/10.4103/sjg.SJG_283_18
Xie Y, Shao Y, Sun W, Ye G, Zhang X, Xiao B, Guo J (2018) Downregulated expression of hsa_circ_0074362 in gastric cancer and its potential diagnostic values. Biomark Med 12:11–20. https://doi.org/10.2217/bmm-2017-0114
Zhou P, Erfani S, Liu Z, Jia C, Chen Y, Xu B, Deng X, Alfáro JE, Chen L, Napier D, Lu M, Huang JA, Liu C, Thibault O, Segal R, Zhou BP, Kyprianou N, Horbinski C, Yang XH (2015) CD151-α3β1 integrin complexes are prognostic markers of glioblastoma and cooperate with EGFR to drive tumor cell motility and invasion. Oncotarget 6:29675–29693. https://doi.org/10.18632/oncotarget.4896
Nakayama I, Shinozaki E, Sakata S, Yamamoto N, Fujisaki J, Muramatsu Y, Hirota T, Takeuchi K, Takahashi S, Yamaguchi K, Noda T (2019) Enrichment of CLDN18-ARHGAP fusion gene in gastric cancers in young adults. Cancer Sci 110:1352–1363. https://doi.org/10.1111/cas.13967
Yang H, Hong D, Cho SY, Park YS, Ko WR, Kim JH, Hur H, Lee J, Kim SJ, Kwon SY, Lee JH, Park DY, Song KS, Chang H, Ryu MH, Cho KS, Kang JW, Kook MC, Thiessen N, He A, Mungall A, Han SU, Kim HK (2018) RhoGAP domain-containing fusions and PPAPDC1A fusions are recurrent and prognostic in diffuse gastric cancer. Nat Commun 9:4439. https://doi.org/10.1038/s41467-018-06747-4
Tanaka A, Ishikawa S, Ushiku T, Yamazawa S, Katoh H, Hayashi A, Kunita A, Fukayama M (2018) Frequent CLDN18-ARHGAP fusion in highly metastatic diffuse-type gastric cancer with relatively early onset. Oncotarget 9:29336–29350. https://doi.org/10.18632/oncotarget.25464
Zhang WH, Zhang SY, Hou QQ, Qin Y, Chen XZ, Zhou ZG, Shu Y, Xu H, Hu JK (2020) The Significance of the CLDN18-ARHGAP Fusion Gene in Gastric Cancer: A Systematic Review and Meta-Analysis. Front Oncol 10:1214. https://doi.org/10.3389/fonc.2020.01214
Li J, Zhang Y, Hu D, Gong T, Xu R, Gao J (2020) Analysis of the expression and genetic alteration of CLDN18 in gastric cancer. Aging 12:14271–14284. https://doi.org/10.18632/aging.103457
Niimi T, Nagashima K, Ward JM, Minoo P, Zimonjic DB, Popescu NC, Kimura S (2001) claudin-18, a novel downstream target gene for the T/EBP/NKX2.1 homeodomain transcription factor, encodes lung- and stomach-specific isoforms through alternative splicing. Mol Cell Biol 21:7380–90. https://doi.org/10.1128/MCB.21.21.7380-7390.2001
Linares GR, Brommage R, Powell DR, Xing W, Chen ST, Alshbool FZ, Lau KH, Wergedal JE, Mohan S (2012) Claudin 18 is a novel negative regulator of bone resorption and osteoclast differentiation. J Bone Miner Res 27:1553–65. https://doi.org/10.1002/jbmr.1600
Kage H, Flodby P, Zhou B, Borok Z (2019) Dichotomous roles of claudins as tumor promoters or suppressors: lessons from knockout mice. Cell Mol Life Sci 76:4663–4672. https://doi.org/10.1007/s00018-019-03238-7
Zhang J, Dong R, Shen L (2020) Evaluation and reflection on claudin 18.2 targeting therapy in advanced gastric cancer. Chin J Cancer Res 32:263–270. https://doi.org/10.21147/j.issn.1000-9604.2020.02.13
Singh P, Toom S, Huang Y (2017) Anti-claudin 18.2 antibody as new targeted therapy for advanced gastric cancer. J Hematol Oncol 10:105. https://doi.org/10.1186/s13045-017-0473-4
Dhakras P, Uboha N, Horner V, Reinig E, Matkowskyj KA (2020) Gastrointestinal cancers: current biomarkers in esophageal and gastric adenocarcinoma. Transl Gastroenterol Hepatol 5:55. https://doi.org/10.21037/tgh.2020.01.08
Sahin U, Schuler M, Richly H, Bauer S, Krilova A, Dechow T, Jerling M, Utsch M, Rohde C, Dhaene K, Huber C, Türeci Ö (2018) A phase I dose-escalation study of IMAB362 (Zolbetuximab) in patients with advanced gastric and gastro-oesophageal junction cancer. Eur J Cancer 100:17–26. https://doi.org/10.1016/j.ejca.2018.05.007
Türeci O, Sahin U, Schulze-Bergkamen H, Zvirbule Z, Lordick F, Koeberle D, Thuss-Patience P, Ettrich T, Arnold D, Bassermann F, Al-Batran SE, Wiechen K, Dhaene K, Maurus D, Gold M, Huber C, Krivoshik A, Arozullah A, Park JW, Schuler M (2019) A multicentre, phase IIa study of zolbetuximab as a single agent in patients with recurrent or refractory advanced adenocarcinoma of the stomach or lower oesophagus: the MONO study. Ann Oncol 30:1487–1495. https://doi.org/10.1093/annonc/mdz199
Shao Y, Li J, Lu R, Li T, Yang Y, Xiao B, Guo J (2017) Global circular RNA expression profile of human gastric cancer and its clinical significance. Cancer Med 6:1173–1180. https://doi.org/10.1002/cam4.1055
Bojesen SE, Ammerpohl O, Weinhäusl A, Haas OA, Mettal H, Bohle RM, Borkhardt A, Fuchs U (2006) Characterisation of the GRAF gene promoter and its methylation in patients with acute myeloid leukaemia and myelodysplastic syndrome. Br J Cancer 94:323–32. https://doi.org/10.1038/sj.bjc.6602939
Qian J, Qian Z, Lin J, Yao DM, Chen Q, Li Y, Ji RB, Yang J, Xiao GF, Wang YL (2011) Abnormal methylation of GRAF promoter Chinese patients with acute myeloid leukemia. Leuk Res 35:783–786. https://doi.org/10.1016/j.leukres.2010.10.013
Borkhardt A, Bojesen S, Haas OA, Fuchs U, Bartelheimer D, Loncarevic IF, Bohle RM, Harbott J, Repp R, Jaeger U, Viehmann S, Henn T, Korth P, Scharr D, Lampert F (2000) The human GRAF gene is fused to MLL in a unique t(5;11)(q31;q23) and both alleles are disrupted in three cases of myelodysplastic syndrome/acute myeloid leukemia with a deletion 5q. Proc Natl Acad Sci USA 97:9168–73. https://doi.org/10.1073/pnas.150079597
Wilda M, Perez AV, Bruch J, Woessmann W, Metzler M, Fuchs U, Borkhardt A (2005) Use of MLL/GRAF fusion mRNA for measurement of minimal residual disease during chemotherapy in an infant with acute monoblastic leukemia (AML-M5). Genes Chromosom Cancer 43:424–6. https://doi.org/10.1002/gcc.20182
Panagopoulos I, Kitagawa A, Isaksson M, Mörse H, Mitelman F, Johansson B (2004) MLL/GRAF fusion in an infant acute monocytic leukemia (AML M5b) with a cytogenetically cryptic ins(5;11)(q31;q23q23). Genes Chromosom Cancer 41:400–4. https://doi.org/10.1002/gcc.20097
Jarius S, Wandinger KP, Horn S, Heuer H, Wildemann B (2010) A new Purkinje cell antibody (anti-Ca) associated with subacute cerebellar ataxia: immunological characterization. J Neuroinflammation 7:21. https://doi.org/10.1186/1742-2094-7-21
Dahm L, Ott C, Steiner J, Stepniak B, Teegen B, Saschenbrecker S, Hammer C, Borowski K, Begemann M, Lemke S, Rentzsch K, Probst C, Martens H, Wienands J, Spalletta G, Weissenborn K, Stöcker W, Ehrenreich H (2014) Seroprevalence of autoantibodies against brain antigens in health and disease. Ann Neurol 76:82–94. https://doi.org/10.1002/ana.24189
Doss S, Nümann A, Ziegler A, Siebert E, Borowski K, Stöcker W, Prüss H, Wildemann B, Endres M, Jarius S (2014) Anti-Ca/anti-ARHGAP26 antibodies associated with cerebellar atrophy and cognitive decline. J Neuroimmunol 267:102–4. https://doi.org/10.1016/j.jneuroim.2013.10.010
Bartels F, Prüss H, Finke C (2018) Anti-ARHGAP26 autoantibodies are associated with isolated cognitive impairment. Front Neurol 9:656. https://doi.org/10.3389/fneur.2018.00656
Naskar T, Faruq M, Banerjee P, Khan M, Midha R, Kumari R, Devasenapathy S, Prajapati B, Sengupta S, Jain D, Mukerji M, Singh NC, Sinha S (2018) Ancestral variations of the PCDHG gene cluster predispose to dyslexia in a multiplex family. EBioMedicine 28:168–179. https://doi.org/10.1016/j.ebiom.2017.12.031
Jarius S, Martínez-García P, Hernandez AL, Brase JC, Borowski K, Regula JU, Meinck HM, Stöcker W, Wildemann B, Wandinger KP (2013) Two new cases of anti-Ca (anti-ARHGAP26/GRAF) autoantibody-associated cerebellar ataxia. J Neuroinflammation 10:7. https://doi.org/10.1186/1742-2094-10-7
Jarius S, Wildemann B (2015) “Medusa head ataxia”: the expanding spectrum of Purkinje cell antibodies in autoimmune cerebellar ataxia. Part 2: Anti-PKC-gamma, anti-GluR-delta2, anti-Ca/ARHGAP26 and anti-VGCC. J Neuroinflammation 12:167. https://doi.org/10.1186/s12974-015-0357-x
Settleman J, Albright CF, Foster LC, Weinberg RA (1992) Association between GTPase activators for Rho and Ras families. Nature 359:153–4. https://doi.org/10.1038/359153a0
Hall A (2012) Rho family GTPases. Biochem Soc Trans 40:1378–82. https://doi.org/10.1042/BST20120103
Rojas AM, Fuentes G, Rausell A, Valencia A (2012) The Ras protein superfamily: evolutionary tree and role of conserved amino acids. J Cell Biol 196:189–201. https://doi.org/10.1083/jcb.201103008
Cherfils J, Zeghouf M (2013) Regulation of small GTPases by GEFs, GAPs, and GDIs. Physiol Rev 93:269–309. https://doi.org/10.1152/physrev.00003.2012
Zandvakili I, Lin Y, Morris JC, Zheng Y (2017) Rho GTPases: anti- or pro-neoplastic targets. Oncogene 36:3213–3222. https://doi.org/10.1038/onc.2016.473
Ridley AJ (2013) RhoA, RhoB and RhoC have different roles in cancer cell migration. J Microsc 251:242–9. https://doi.org/10.1111/jmi.12025
Etienne-Manneville S, Hall A (2002) Rho GTPases in cell biology. Nature 420:629–35. https://doi.org/10.1038/nature01148
Ohta K, Goto H, Matsumoto Y, Yumine N, Tsurudome M, Nishio M (2016) Graf1 controls the growth of human parainfluenza virus type 2 through Inactivation of RhoA signaling. J Virol 90:9394–405. https://doi.org/10.1128/JVI.01471-16
Lucken-Ardjomande Häsler S, Vallis Y, Jolin HE, McKenzie AN, McMahon HT (2014) GRAF1a is a brain-specific protein that promotes lipid droplet clustering and growth, and is enriched at lipid droplet junctions. J Cell Sci 127:4602–19. https://doi.org/10.1242/jcs.147694
Shibata H, Oishi K, Yamagiwa A, Matsumoto M, Mukai H, Ono Y (2001) PKNbeta interacts with the SH3 domains of Graf and a novel Graf related protein, Graf2, which are GTPase activating proteins for Rho family. J Biochem 130:23–31. https://doi.org/10.1093/oxfordjournals.jbchem.a002958
Doherty GJ, Åhlund MK, Howes MT, Morén B, Parton RG, McMahon HT, Lundmark R (2011) The endocytic protein GRAF1 is directed to cell-matrix adhesion sites and regulates cell spreading. Mol Biol Cell 22:4380–9. https://doi.org/10.1091/mbc.E10-12-0936
Doherty GJ, Lundmark R (2009) GRAF1-dependent endocytosis. Biochem Soc Trans 37:1061–5. https://doi.org/10.1042/BST0371061
Lundmark R, Doherty GJ, Howes MT, Cortese K, Vallis Y, Parton RG, McMahon HT (2008) The GTPase-activating protein GRAF1 regulates the CLIC/GEEC endocytic pathway. Curr Biol 18:1802–8. https://doi.org/10.1016/j.cub.2008.10.044
Eberth A, Lundmark R, Gremer L, Dvorsky R, Koessmeier KT, McMahon HT, Ahmadian MR (2009) A BAR domain-mediated autoinhibitory mechanism for RhoGAPs of the GRAF family. Biochem J 417:371–7. https://doi.org/10.1042/BJ20081535
Orgaz JL, Herraiz C, Sanz-Moreno V (2014) Rho GTPases modulate malignant transformation of tumor cells. Small GTPases 5:e29019. https://doi.org/10.4161/sgtp.29019
Jansen S, Gosens R, Wieland T, Schmidt M (2018) Paving the Rho in cancer metastasis: Rho GTPases and beyond. Pharmacol Ther 183:1–21. https://doi.org/10.1016/j.pharmthera.2017.09.002
Haga RB, Ridley AJ (2016) Rho GTPases: regulation and roles in cancer cell biology. Small GTPases 7:207–221. https://doi.org/10.1080/21541248.2016.1232583
Kim S, Nahm M, Kim N, Kwon Y, Kim J, Choi S, Choi EY, Shim J, Lee C, Lee S (2017) Graf regulates hematopoiesis through GEEC endocytosis of EGFR. Development 144:4159–4172. https://doi.org/10.1242/dev.153288
Holst MR, Vidal-Quadras M, Larsson E, Song J, Hubert M, Blomberg J, Lundborg M, Landström M, Lundmark R (2017) Clathrin-independent endocytosis suppresses cancer cell blebbing and invasion. Cell Rep 20:1893–1905. https://doi.org/10.1016/j.celrep.2017.08.006
Elvers M, Beck S, Fotinos A, Ziegler M, Gawaz M (2012) The GRAF family member oligophrenin1 is a RhoGAP with BAR domain and regulates Rho GTPases in platelets. Cardiovasc Res 94:526–36. https://doi.org/10.1093/cvr/cvs079
Zhao ZJ, Shen J (2017) Circular RNA participates in the carcinogenesis and the malignant behavior of cancer. RNA Biol 14:514–521. https://doi.org/10.1080/15476286.2015.1122162
Cheng DL, Xiang YY, Ji LJ, Lu XJ (2015) Competing endogenous RNA interplay in cancer: mechanism, methodology, and perspectives. Tumour Biol 36:479–88. https://doi.org/10.1007/s13277-015-3093-z
Guil S, Esteller M (2015) RNA-RNA interactions in gene regulation: the coding and noncoding players. Trends Biochem Sci 40:248–56. https://doi.org/10.1016/j.tibs.2015.03.001
Shang X, Li G, Liu H, Li T, Liu J, Zhao Q, Wang C (2016) Comprehensive circular RNA profiling reveals that hsa_circ_0005075, a new circular RNA biomarker, is involved in hepatocellular crcinoma development. Medicine 95:e3811. https://doi.org/10.1097/MD.0000000000003811
Funding
None.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
All authors declared that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhang, L., Zhou, A., Zhu, S. et al. The role of GTPase-activating protein ARHGAP26 in human cancers. Mol Cell Biochem 477, 319–326 (2022). https://doi.org/10.1007/s11010-021-04274-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11010-021-04274-3