Abstract
The thirteen-lined ground squirrel (Ictidomys tridecemlineatus) undergoes remarkable adaptive changes during hibernation. Interestingly, skeletal muscle remodelling occurs during the torpor-arousal cycle of hibernation to prevent net muscle loss despite inactivity. Reversible cardiomyocyte hypertrophy occurs in cardiac muscle, allowing the heart to preserve cardiac output during hibernation, while avoiding chronic maladaptive hypertrophy post-hibernation. We propose that calcium signalling proteins [calcineurin (Cn), calmodulin (CaM), and calpain], the nuclear factor of activated T cell (NFAT) family of transcription factors, and the NFAT targets myoferlin and myomaker contribute significantly to adaptations taking place in skeletal and cardiac muscle during hibernation. Protein-level analyses were performed over several conditions: euthermic room temperature (ER), euthermic cold room (EC), entrance into (EN), early (ET), and late torpor (LT) time points, in addition to early (EA), interbout (IA), and late arousal (LA) time points using immunoblotting and DNA–protein interaction (DPI) enzyme-linked immunosorbent assay (ELISAs). In skeletal and cardiac muscle, NFATc2 protein levels were elevated during torpor. NFATc4 increased throughout the torpor-arousal cycle in both tissues, and NFATc1 showed this trend in cardiac muscle only. NFATc3 showed an elevation in DNA-binding activity but not expression during torpor. Myoferlin protein levels dramatically increased during torpor in both skeletal and cardiac muscle. Myomaker levels also increased significantly in cardiac muscle during torpor. Cardiac Cn levels remained stable, whereas CaM and calpain decreased throughout the torpor-arousal cycle. Activation and/or upregulation of NFATc2, c3, myoferlin, and myomaker at torpor could be part of a stress-response mechanism to preserve skeletal muscle mass, whereas CaM and calpain appear to initiate the rapid reversal of cardiac hypertrophy during arousal through downregulation of the NFAT–Cn pathway.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The thirteen-lined ground squirrel, Ictidomys tridecemlineatus, is native to the central prairies of North America and survives winter by hibernating underground. During hibernation, the animal undergoes periods of torpor where it suppresses its metabolic rate (often to just 2–4 % of normal conditions) and drops its core body temperature (T b) to levels that match its surroundings (often as low as 0–5 °C) [1–4]. Prolonged periods of torpor are interspersed with brief periods of arousal back to where its metabolic rate and T b return to euthermic states. This strategy for conserving energy can save hibernating ground squirrels up to 88 % of the ATP expenditure that would otherwise be required to maintain euthermic physiological conditions over the course of winter months [4]. Metabolic rate suppression is a controlled process to allow for the maintenance of cellular homeostasis at colder T b, while simultaneously reprioritizing ATP use by different cell functions. These functions include cell preservation strategies such as antioxidant defenses and protein chaperones, as well as the expression of selected genes and/or proteins that support cell- or tissue-specific needs [2, 5–9].
Each organ/tissue of the hibernator must make specific adjustments that allow them to maintain or readjust physiological functions at low T b values to support long-term torpor. An issue confronting skeletal muscle is the potential for disuse-induced atrophy during hibernation; this process commonly occurs during periods of reduced use whereby muscle mass, strength, and relative amount of slow oxidative muscle fibre are reduced [10–13]. Interestingly, many studies have demonstrated a lack of skeletal muscle atrophy during hibernation, despite the prolonged period of mechanical unloading that occurs over the long winter of inactivity [14–16]. In summary, hibernators have unique mechanisms of muscle remodelling and muscle mass retention compared to non-hibernating mammals such as humans. Therefore, uncovering the molecular mechanisms of muscle preservation that occurs naturally in hibernators could be applied through the identification of novel targets for therapeutic intervention of muscle wasting and to assist in restoring muscle mass and function during rehabilitation.
Throughout hibernation, another tissue that undergoes physiological adaptations is the heart. Heart rate is reduced strongly during torpor, often to just 5–10 beats/min in comparison with the euthermic rates of 350–400 beats/min; this plus the increased viscosity of blood at low T b requires changes in heart dynamics [1, 17]. Therefore, the strength of each individual contraction must be significantly greater so cardiomyocyte hypertrophy is observed, which is defined as the enlargement of individual cardiomyocytes [3, 18, 19]. This adaptation is accompanied by enhanced protein synthesis, adjustments to the structure of sarcomeres, and changes in gene transcription [20–23]. In most mammals, cardiomyocyte hypertrophy is characterized by significant cardiac fibrosis, which reduces diastolic filling, and ultimately prevents the heart from pumping enough blood to meet body demands [20]. Interestingly, hibernators undergo reversible cardiac hypertrophy, where the heart hypertrophies during hibernation but is restored after hibernation to pre-hibernation dimensions, and this may be related to molecular adaptations that occur during hibernation, which are just beginning to be discovered [6, 22–24]. Understanding how these animals can rapidly reverse cardiac hypertrophy would provide novel insight into treating and preventing maladaptive cardiac hypertrophy.
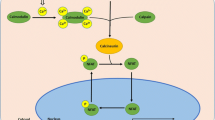
The present study focused on the regulation of the nuclear factor of activated T cells (NFAT) family of transcription factors and their downstream effectors that can play crucial roles in both cardiac and skeletal muscle remodelling. Currently, the NFAT family contains five members named NFAT 1-5 or NFAT c1-4 and NFAT 5 [25]. In its inactive state, NFAT proteins are phosphorylated and localized in the cytoplasm. NFATs are activated via dephosphorylation by calcineurin (Cn), a calmodulin-stimulated protein phosphatase, allowing NFATs to translocate to the nucleus and bind to transcriptional complexes of their target genes [26]. Calmodulin (CaM) is a calcium-binding protein that controls various signalling pathways, including the NFAT–Cn pathway. It does so by binding to the regulatory domain of Cn, known as the calcineurin A subunit (CnA), when the calcineurin B subunit is exposed to high calcium levels. The effect of activated or calcium-bound CaM binding to CnA is the activation of the CnA phosphatase activity, thus leading to NFAT nuclear translocation [27–29]. Another important protein that regulates the NFAT–Cn pathway is the calcium-dependent protease-calpain. μ-calpain (calpain1) causes the proteolysis of the autoinhibitory domain of CnA, which causes it to translocate to the nucleus and become constitutively active [30–33]. Therefore, calpain and CaM both play a vital role in regulation of the NFAT–Cn pathway and NFATs may contribute to the adaptive responses needed for skeletal and cardiac muscle remodelling during hibernation. For example, NFATs regulate cardiomyocyte atrophy, apoptosis, development, and growth [21, 34–37]. In another hibernation model, the use of isobaric tag for relative and absolute quantification (iTRAQ) technology with cardiac tissue of hibernating woodchucks (Marmota monax) demonstrated an up-regulation of the NFAT pathway during hibernation [21]. However, it is important to characterize the regulation of NFATs are their targets throughout hibernation as it is characterized by torpor-arousal bouts rather than a constant physiological state. In addition, it is necessary to study the expression of specific NFATs rather than the entire family during hibernation in skeletal and cardiac muscle as different NFATs play different roles in coordinating muscle remodelling during hibernation. For example, expression of constitutively active NFATc4 in cardiac tissue of transgenic mice induced cardiac hypertrophy [37]. With regard to skeletal muscle remodelling, NFAT have been implicated in multiple aspects including atrophy, hypertrophy, fibre-type switching, and myogenesis [38–42]. For instance, NFATc2-null mice have defective myoblast fusion and myogenesis, resulting in fibres with reduced size and delayed repair in response to injury [43]. Therefore, NFAT transcription factors can be considered central regulators of both skeletal and cardiac muscle remodelling.
Considering the unique tendencies of hibernators to undergo reversible cardiomyocyte hypertrophy and remodel their skeletal muscle over the course of the hibernation season, we hypothesized that the calcium signalling activates the NFAT–Cn pathway during hibernation, which plays a significant role in both the avoidance of disuse-induced muscle wasting and reversible cardiac hypertrophy. The NFAT–Cn pathway performs these roles by modulating the gene expression of selected proteins that are fundamental to muscle design. One such protein is myoferlin, a protein that is highly expressed in skeletal muscle and to a lesser degree in cardiac muscle [44]. However, myoferlin is highly expressed specifically in myoblasts undergoing fusion, where it localizes at the sites of apposed membranes undergoing fusion [45, 46]. Furthermore, myoferlin mRNA was up-regulated in human muscle affected by Duchenne muscular dystrophy [47]. In myoferlin-null mice, muscle fibre size was reduced due to impaired myoblast fusion, but the mice were still viable. However, myomaker, a membrane protein found in the muscle that also controls myoblast fusion results in the complete loss of myoblast fusion when it is mutated, resulting in the absence of all skeletal muscle, which leads to postnatal death in myomaker-null mice [48]. In addition, myomaker expression and increased myoblast fusion have been shown to occur in adult satellite cells following muscle injury [49]. These findings implicate both myomaker and myoferlin as novel candidates for muscular dystrophy or cardiomyopathy due to their roles in muscle formation and repair.
Myoferlin contains multiple NFAT-binding sites in its promoter, which drove high levels of myoferlin expression in vitro and in vivo. Furthermore, expression was elevated in response to muscle damage [50]. The regulation of myomaker is not well understood due to its recent discovery, with only two MRFs, muscle differentiation protein (MyoD), and myogenin (MyoG), being known to induce myomaker transcription [49]. However, NFAT has been shown to regulate the expression of MyoG cooperatively with MyoD [38]. In addition to affecting myomaker expression through its regulation of MyoG, we also used a DNA–protein interaction (DPI) enzyme-linked immunosorbent assay (ELISA) to test the ability of NFATs to bind to a novel putative NFAT-binding sequence in the myomaker promoter. The present study analysed the protein expression of Cn, CaM, calpain, NFATc1-4, myoferlin, and myomaker over the torpor-arousal cycle in the cardiac and skeletal muscle of thirteen-lined ground squirrels.
Materials and methods
Animals
Thirteen-lined ground squirrels (I. tridecemlineatus), weighing 150–300 g, were wild-captured and supplied by TLS Research, Bloomingdale, IL. Animals were transported to the Animal Hibernation Facility, National Institute of Neurological Disorders and Stroke, NIH, Bethesda, MD where all experiments were conducted at the laboratory of Dr. J. M. Hallenbeck as previously described [51]. Animals were individually housed in shoebox cages in a holding room with an ambient temperature of 21 °C under a 12 h light: 12 h dark cycle. Animals were fitted with a sensor chip (IPTT-300; Bio Medic Data Systems) injected subcutaneously, and were anaesthetized with 5 % isoflurane. Animals were fed water and standard rodent chow ad libitum until they gained sufficient lipid stores to enter hibernation. To enable a natural transition into torpor, animals were transferred to constant darkness in an environmental chamber at 4–5 °C. Body temperature (T b), time, and respiration rate were monitored and used to determine the stage of torpor-arousal cycle. All animals had been through several torpor-arousal bouts prior to sampling at a set stage. Animals were euthanized and the tissue samples were collected at seven different time points defined by T b, duration of torpor, and respiration rate: (1) Euthermic room temperature (ER); these animals were held in the holding room with an ambient temperature of 21 °C (T b = ~37 °C) and were capable of entering torpor. Skeletal muscle tissues were collected from animals at this time point after they had reached their plateau weight. (2) Euthermic cold room (EC); these squirrels had a stable T b of 34–37 °C for at least 3 days and were capable of entering torpor but had not done so in the past 72 h. (3) Early entrance into torpor (EN); T b falling with sampling between 31° and 18 °C. (4) Early torpor (ET); T b stable at ~5–8 °C for <24 h. (5) Late torpor (LT); T b stable at ~5–8 °C for >5 days. (6) Early arousal (EA); T b rising to at least ~12 °C with a rapid increase in respiration ≥60 breaths/min, (7) Late arousal (LA); respiration rate increased and T b reached 28–32 °C. Heart tissue was not collected from ER and LA, but it was collected at the additional sampling point—interbout arousal (IA)—where the animals were naturally aroused after the torpor phase of the hibernation bout and reached the respiratory rate, metabolic rate, and body temperature of fully aroused animals for 6 h after being in torpor for at least 5 days. These animals remain in the hibernaculum (4 °C) but their core body temperature is back to ~37 °C.
Total protein extraction
Samples of frozen skeletal muscle from n = 4 individuals from each of the seven time points in the torpor-arousal cycle (ER, EC, EN, ET, LT, EA, LA) were extracted as previously described [42]. Samples of frozen heart were extracted from six time points (EC, EN, ET, LT, EA, and IA). Aliquots of ~0.5 g tissue were quickly weighed, powdered into very small pieces under liquid nitrogen, and then homogenized 1:3 w:v using a Polytron PT10 in ice-cold homogenizing buffer (20 mM Hepes, 200 mM NaCl, 0.1 mM EDTA, 10 mM NaF, 1 mM Na3VO4, 10 mM β-glycerophosphate) with 1 mM phenylmethylsulfonyl fluoride (Bioshop) and 1 μL/mL protease inhibitor cocktail (Bioshop) added immediately before homogenization. Samples were centrifuged at 10,000 rpm (10 min, 4 °C) and then the supernatant was removed. Protein concentration was quantified by the Coomassie blue dye-binding using the BioRad reagent (BioRad Laboratories, Hercules, CA) at 595 nm on a MR5000 microplate reader. Samples were then adjusted to a constant of 10 μg/μL by addition of small amounts of homogenizing buffer and then aliquots were combined 1:1 v:v with 2 × SDS loading buffer (100 mM Tris-base, pH 6.8, 4 % w:v SDS, 20 % v:v glycerol, 0.2 % w:v bromophenol blue, 10 % v:v 2-mercaptoethanol) and then boiled. The final protein samples (5 μg/μL) were stored at −20 °C until use.
Preparation of nuclear protein extracts
Nuclear protein extracts were prepared as previously described [42] and were separately extracted from the skeletal muscle of four animals for each of the seven experimental stages described above for total protein extractions in skeletal muscle. Frozen skeletal muscle samples were homogenized 1:2 w:v using a Dounce homogenizer (5 piston strokes) in lysis buffer (10 mM HEPES, pH 7.9, 10 mM KCl, 10 mM EDTA, 20 mM β-glycerophosphate), with 10 μL of 100 mM DTT and 10 μL of protease inhibitor cocktail added immediately before homogenization. Samples were centrifuged for 10 min at 10,000 rpm and at 4 °C, and then the supernatants were removed, forming the cytoplasmic fraction. Pellets were resuspended in 147 μL of nuclear extraction buffer (20 mM HEPES, pH 7.9, 400 mM NaCl, 1 mM EDTA, 10 % v/v glycerol, 20 mM β-glycerophosphate) with 1.5 μL of 100 mM DTT and 1.5 μL of protease inhibitor cocktail added. Samples were incubated on ice with gentle rocking for 1 h and then centrifuged for 10 min at 10,000 rpm at 4 °C. Protein concentrations were determined with the BioRad protein assay, adjusted to 5 μg/μL, and the samples were stored at −80 °C until use.
Western blotting
Equal amounts of protein from each sample (25 μg) were loaded onto 6 % (NFATc1-4, myoferlin), 8 % (CnA, calpain) or 15 % (myomaker, CaM) polyacrylamide gels and were run at 180 V for 60–90 min. Proteins were then transferred to PVDF membranes by electroblotting at 320 mA for 1.5 h (CnA, CaM, calpain, myomaker) or 3 h (NFATc1-4, myoferlin) using a transfer buffer containing 25 mM Tris (pH 8.5), 192 mM glycine and 10 % v:v methanol at room temperature. Membranes were washed with 1× TBST (20 mM Tris-base, pH 7.6, 140 mM NaCl, 0.05 % v:v Tween-20, 90 % v:v ddH2O) for 3 × 5 min. Membranes were then blocked for 30 min with 2.5 (for NFATc1-4) or 5 % (for CnA, CaM, calpain1, myoferlin and myomaker) w:v milk in 1× TBST. After washing for 3 × 5 min again with 1x TBST, membranes were probed with specific primary antibodies at 4 °C overnight. Antibodies specific for mammalian NFAT-c1 (sc-13033), c2 (sc-13024), c3 (sc-8321), c4 (sc-13036), myoferlin (sc-134798) and myomaker (also known as TMEM8c, sc-2444460) were purchased from Santa Cruz Biotechnologies. All antibodies were used at a 1:500 v:v dilution in 1x TBST. A CaM (06-396) antibody from Upstate Biotechnology (Lake Placid, NY), as well as calcineurin A (GTX111039) and calpain1 (GTX102340) antibodies from Genetex (Irving, CA) were purchased and used at a 1:1000 v:v dilution in 1× TBST. After probing with primary antibody, membranes were washed for 3 × 5 min with 1× TBST and then incubated with HRP-conjugated secondary antibody for 20–30 min at room temperature. Membranes that had been probed with myomaker (TMEM8c) were incubated with HRP-linked anti-goat IgG secondary antibody (BioShop: 1:6000 v:v dilution). All other antibodies were detected using HRP-linked anti-rabbit IgG secondary antibody (Bioshop: 1:6000 v:v dilution). Bands were visualized by enhanced chemiluminescence (H2O2 and Luminol) and then blots were restained using Coomassie blue to visualize total protein levels. Immunoblot bands for ground squirrel proteins corresponded to the specified molecular weights, as confirmed by running PINK Plus Prestained Protein Ladder (FroggaBio) for myomaker, calpain, CnA, and CaM, or BLUeye Prestained Protein Ladder (FroggaBio) for NFATc1-4 and myoferlin.
Comparative sequence analysis
The total nucleotide sequences of human (Accession number: NM_001080483.2), mouse (Accession number: NM_025376.3), and 13-lined ground squirrel (Accession number: XM_005336956.1) myomaker were accessed from the NCBI Nucleotide database, and the sequence within 1500 base pairs (bp) upstream of the transcriptional start site was downloaded. These upstream sequences were compared using the Vista program (http://genome.lbl.gov/vista) with a window size of 100 bp and a minimum sequence identity of 70 %. NFAT transcription factor-binding sites were identified using the rVISTA program. Predictions were made based on the TRANSFAC Professional 9.3 library using the default core similarity value of 0.75 and the matrix similarity value of 0.70 as previously described [50].
The amino acid sequences of myomaker (human: NP_001073952.1, mouse: NP_079652.1, squirrel: XP_005337013.1), myoferlin (human: Q9NZM1, mouse: Q69ZN7, squirrel: XP_005333820.1), NFATc1 (human: O95644.3, mouse: O88942.1, squirrel: XP_005334428.1), NFATc2 (human: Q13469.2, mouse: Q60591.3, squirrel: I3ML24), NFATc3 (human: Q12968.1, mouse: P97305.1, squirrel: I3MSZ2), and NFATc4 (human: Q14934.2, mouse: Q8K120.2, squirrel: XP_005338768.1) from the NCBI protein database or the UniProtKB protein knowledge base were compared using the ClustalW2 multiple alignment tool at EMBL-EBI using default multiple sequence alignment and pairwise alignment options. Pairwise alignment scores (percent identity) between two sequences are calculated by taking the number of identities between two sequences, dividing by the length of the alignment, and representing this ratio as a percentage [52–54].
DNA–protein interaction (DPI) ELISA
DNA oligonucleotides were designed based on the DNA-binding elements of NFATc1-4 as previously described in the previous section, and were produced by Integrated DNA Technologies (Coralville, Iowa). The biotinylated probe (NFAT 5′-Biotin-GGGAAGGA AAGTGCGGGTGG-3′) and the complement probe (NFAT 5′-Biotin CCACCCGCACCCTTTTTCCC-3′) were first diluted in sterile water (500 pmol/μL) and mixed 1:1 v:v for a total of 20 μL. Probes were then placed in a thermocycler for 10 min at 94 °C and slowly cooled to room temperature. Double-stranded probes were diluted in phosphate-buffered saline (PBS; 137 mM NaCl, 2.7 mM KCL, 10 mM Na2HPO4, pH 7.4) and 50 μL of diluted DNA probe was added (40 pmol DNA/well) to streptavidin-coated wells on a microplate. Following a 1 h incubation, unbound probe was discarded and the wells were rinsed twice with wash buffer (1X PBS containing 0.1 % Tween-20), and a third time with 1X PBS. Transcription factor-binding buffer (10 mM HEPES, 50 mM KCL, 0.5 mM EDTA, 3 mM MgCl2, 10 % v/v glycerol, 0.5 mg/ml bovine serum albumin, 0.05 % NP-40, 0.5 mM DTT, 20 pg/μL Salmon Sperm DNA, 44 mM NaCl, pH 7.9) was vortexed and added to each well containing DNA probe. In addition, 27.5 μg of protein from nuclear protein preparation was added, with the exception of negative control wells which contained transcription factor-binding buffer without protein. Following 1 h incubation with gentle shaking, protein mixtures were discarded and the wells were washed three times with wash buffer.
Diluted NFAT primary antibody (1:1000) was then added (60 μL/well) for 1 h. Antibody was then discarded, and the wells were rinsed three times with wash buffer before incubation with diluted secondary (1:1000, 60 μL/well) for 1 h. This antibody was then discarded and the wells were rinsed four times with wash buffer. Primary and secondary antibodies were the same as those used for immunoblots. After secondary antibody incubation and washing, bound antibody was detected using tetramethylbenzidine (TMB) (Bioshop). A 60 μL aliquot of TMB was added to each well, colour was developed for 10–15 min, and then the reaction was stopped with 60 μL of 1 M HCl. Absorbance was measured at 450 nm (reference wavelength of 655 nm) using a Multiskan spectrophotometer. To control for background absorbance and non-specific binding, negative controls were run in duplicates with no probe, no protein, and no primary antibody added. Conditions were optimized such that negative control wells showed >50 % decreases in absorbance relative to sample wells before quantification runs of the entire torpor-arousal time course were conducted.
Quantification and statistics
Band densities on chemiluminescent immunoblots were visualized using a Chemi-Genius BioImaging system (Syngene, Frederick, MD) and quantified using the Gene Tools software. Immunoblot band density in each lane was standardized against the summed intensity of a group of Coomassie-stained protein bands in the same lane; this group of bands was chosen because they were not located close to the protein band of interest but were prominent and constant across all samples. This method of standardizing against a total protein loading control has been suggested to be more accurate in comparison with standardizing against housekeeping proteins such as tubulin [55]. To quantify DPI–ELISAs, absorbance values of negative control wells were subtracted from those of sample wells and the absorbance values were standardized relative to the average of EC absorbance values. Data were expressed as mean ± SEM, n = 4 independent samples from different animals. Statistical testing used the one-way ANOVA and Tukey post hoc tests on the GraphPad Prism software (San Diego, CA).
Results
Analysis of NFATc1-4 protein levels in skeletal muscle
NFAT c1-4 protein levels in skeletal muscle were analysed by immunoblotting; comparing animals sampled from seven different stages of the torpor-arousal cycle: ER, EC, EN, ET, LT, EA, and LA (Fig. 1). The antibodies cross-reacted with a single band on immunoblots at the expected molecular mass for NFATc3 of 130 kDa, in accordance with the protein size indicated on the Santa Cruz antibody specification sheet (sc-8321). For muscle NFATc1, c2, and c4, no distinct band was observed at the regions specified on the antibody specification sheets within 5–10 kDa. Because evolutionary changes may have occurred, the epitope for our antibodies may not have been conserved between 13-lined ground squirrels and the antibody host (rabbit). Therefore, protein sequences were identified in the NCBI protein database for NFAT c1 (XP_005334428) and NFATc4 (XP_005338768) for I. tridecemlineatus. Also, the protein sequence for NFATc2 (I3ML24) was found on the UniProt Knowledge Base (UniProtKB). The molecular weights of these protein sequences were subsequently calculated using the ExPASy Mw/pI tool (http://web.expasy.org/compute_pi/), and bands were detected at the calculated molecular weights (101, 92, and 85 kDa for NFATc1, c2, and c4, respectively).
Changes in the protein levels of NFAT transcription factors over the course of the torpor-arousal cycle in skeletal muscle of I. tridecemlineatus. NFATc1, c2, c3, and c4 total protein expression levels were visualized at seven sampling points: ER euthermic room temperature, EC euthermic cold room, EN entrance into torpor, ET early torpor, LT late torpor, EA early arousal, LA late arousal. See Materials and methods for more extensive definitions. Representative Western blots and Coomassie total protein loading controls are shown for selected pairs of sampling points that are labelled to the left and right of the gel. Sample numbers (lanes) are labelled along the top indicating 4 samples of one type (e.g. EC lanes 1, 2, 3, 4) and 4 samples of another (e.g. LA lanes 5, 6, 7, 8). Also shown are histograms with mean standardized band densities (±S.E.M., n = 4 independent protein isolations from different animals). Data were analysed using a one-way analysis of variance with a post hoc Tukey’s test (p < 0.05); for each parameter measured, values that are not statistically different from each other share the same letter notation
NFATc1 protein levels decreased during torpor, reaching its lowest levels at LT (78 % lower in comparison with EC, p < 0.05), before rising back up to EC levels during arousal. NFATc2 levels on the other hand increased during torpor, peaking at LT (1.75-fold higher in comparison with EN, p < 0.05). NFATc3 levels were the highest initially at ER (1.82-fold higher in comparison with EC, p < 0.05) before decreasing dramatically. Protein levels then decreased again during LA (55 % lower in comparison with LT). Finally, NFATc4 levels were highest levels at ER, then levels rose throughout the torpor-arousal cycle, peaking at LA (higher than EN by 1.97-fold, p < 0.05) (Fig. 1).
Analysis of NFATc1-4 protein levels in cardiac muscle
Analysis was performed on NFAT c1-4 protein levels by comparing immunoblots of cardiac muscle from EC, EN, ET, LT, EA, and IA animals at various stages of the torpor-arousal cycle with each other (Fig. 2). In cardiac tissue, NFATc1 levels did not change significantly over the course of the torpor-arousal time course. On the other hand, NFATc2 levels increased during torpor, peaking at ET (1.88-fold increase from EN, p < 0.05). NFATc3 protein levels were unchanged over the torpor-arousal cycle. Lastly, NFATc4 levels remained stable over torpor, but increased during arousal, peaking at IA (1.66-fold higher than ET, p < 0.05) (Fig. 2).
Changes in the protein levels of NFAT transcription factors over the course of the torpor-arousal cycle in cardiac muscle of I. tridecemlineatus. NFATc1, c2, c3, and c4 total protein expression levels were visualized at six sampling points: EC euthermic cold room, EN entrance into torpor, ET early torpor, LT late torpor, EA early arousal, IA interbout arousal. Other information as in Fig. 1
Analysis of myoferlin and myomaker (TMEM8c) protein levels
Antibodies for myoferlin and myomaker (TMEM8c) cross-reacted with single bands on immunoblots at the expected molecular weights of 235 and 25 kDa, respectively, in accordance with antibody specification sheets (sc-134798 and sc-244460 for myoferlin and myomaker, respectively). Myoferlin levels increased dramatically during torpor (EN, ET, LT increased by 3.45-, 4.75-, and 4.46-fold, respectively, in comparison with EC, p < 0.05) and then decreased upon entering arousal. Myomaker on the other hand showed constant protein levels from throughout the torpor-arousal cycle (Fig. 3). Similar to skeletal muscle, myoferlin levels in cardiac muscle rose dramatically at ET (2.68-fold higher in comparison with control, p < 0.05). In contrast to skeletal muscle levels, myomaker in cardiac muscle also peaked at ET (1.88-fold higher in comparison with control, p < 0.05) (Fig. 4).
Changes in myoferlin and myomaker total protein levels in skeletal muscle over the torpor-arousal cycle in I. tridecemlineatus. Other information as in Fig. 1
Changes in myoferlin and myomaker total protein levels in cardiac muscle over the torpor-arousal cycle in I. tridecemlineatus. Other information as in Fig. 2
Analysis of calcineurin A, calmodulin, and calpain1 protein levels in cardiac muscle
CnA, CaM, and calpain1 antibodies cross-reacted with single bands on immunoblots at the expected molecular weights of 59, 19, and 82 kDa, respectively. These molecular weights corresponded with molecular weights from antibody specification sheets for CnA (GTX111039) and calpain1 (GTX102340). For CaM, the molecular weight was determined using the 13-lined ground squirrel protein sequence found on NCBI (XP_005336620.1) and the ExPASy Mw/pI tool as previously described. CnA protein levels remained fairly stable throughout the torpor-arousal cycle, with the exception of IA, where levels decreased dramatically by 62 % from EA (p < 0.05). CaM levels were the highest at EC and EN, but levels began to decrease dramatically upon entering torpor, with the lowest levels being observed at EA (70 % decrease in comparison with EC, p < 0.05). Calpain levels showed a similar trend, with the highest levels being observed at EC and IA, and drastically lower levels throughout the torpor-arousal cycle. The lowest levels were observed during EN (85 % decrease from EC, p < 0.05) (Fig. 5).
Changes in calcineurin, calmodulin, and calpain total protein levels in cardiac muscle over the torpor-arousal cycle in I. tridecemlineatus. Other information as in Fig. 2
Analysis of NFATc1-4 relative binding to DNA
Based on the known role of NFAT proteins in muscle development and repair, the 1500 bp sequence upstream of the myomaker transcriptional start site was analysed for consensus NFAT-binding sites. Using rVista, all NFAT-DNA-binding sites were identified and pairwise alignments were conducted between myomaker upstream sequences in squirrel and mouse (Fig. S1a), and in squirrel and human (Fig. S1b). The program displayed red lines identifying potential NFAT-binding sites that were aligned, allowing a maximum core shift of 6 bp and only one gap of any length inside it. Squirrel-mouse and squirrel-human alignments had two NFAT-binding sites in common at 1095 and ~1440 bp upstream of the myomaker transcriptional start site, shown in Fig. S1 by the red lines. These two aligned NFAT-binding sites were cross-referenced using the conserved NFAT-binding sequence from the literature (GGAAA) [25, 56]. Only the 1095 bp upstream sequence remained a probable NFAT-myomaker binding site (Fig. S1c). Subsequently, a DNA oligonucleotide was designed for this putative binding site (NFAT 5′-Biotin-GGGAAGGAAAGTGCGGGTGG-3′) containing the general NFAT-binding sequence and the flanking region specific to the myomaker promoter.
To quantitatively measure the ability of NFATs to bind to its consensus sequence, specifically in the myomaker promoter, DNA–protein binding (DPI)–ELISAs were used because of its simplicity and robustness [57–59]. Findings indicated that only NFATc1 and c3 were able to bind specifically to the DNA oligonucleotide, as NFATc2 and c4 did not pass the test strip stage, with negative control wells showing high absorbance readings, despite persistent optimization. Relative binding to DNA (transcription factor activity) was measured for seven time points: ER, EC, EN, ET, LT, EA, and LA (Fig. 6). NFATc1 activity decreased by 58 and 54 % at EN and EA, respectively, relative to ER (p < 0.05), but there was a progressive increase throughout torpor from EN to LT. These decreases in DNA binding at EN and EA were seen in NFATc3 as well (46 and 30 % lower, respectively, relative to EC). In addition, there was a large increase in NFATc3-DNA binding at LT (3.99-increase relative to EN, p < 0.05).
Changes in binding of the transcription factors NFATc1 and c3 to a DNA-binding element designed for the myomaker promoter in the skeletal muscle of I. tridecemlineatus over the torpor-arousal cycle. Absorbance readings were corrected by subtraction of negative controls containing no protein and values were expressed relative to EC. Histograms show mean relative values ± S.E.M., n = 4 independent biological replicates for each of the five experimental conditions. Other information as in Fig. 1
Comparative amino acid sequence analysis of NFAT, myoferlin, and myomaker proteins
The amino acid sequences for myomaker, myoferlin, as well as NFATc1, c2, c3, and c4 in squirrel, mouse, and human were compared and analysed for evolutionary conservation using the multiple alignment tool ClustalW2 [52–54]. Multiple alignment of the myomaker sequence showed strong conservation with no gaps in the amino acid sequences between the three animals (Fig. S2a). Analysis of the degree of conservation of myomaker, myoferlin, and NFATc1-4 through pairwise alignments between human, mouse, and squirrel indicated that percent identity for all comparisons was strong (>80 %) (Fig. S2b).
Discussion
The present study aimed at furthering our understanding of the molecular mechanisms underlying muscle remodelling in both skeletal and cardiac muscle during hibernation in the 13-lined ground squirrel. The ground squirrel provides an excellent natural model system for studying muscle remodelling as the skeletal muscle appears to undergo alternating patterns of atrophy and hypertrophy without a net loss in muscle mass, and the cardiac muscle is known to undergo reversible cardiac hypertrophy [19, 21, 22, 60]. However, the molecular mechanisms that are responsible the preservation and/or change of muscle structure/function during hibernation are not well known. Therefore, the present study focuses on the family of transcription factors known as NFATs, which have been shown to regulate targets associated with skeletal muscle and cardiomyocyte atrophy, apoptosis, and development [21, 23, 34–42]. NFATs are regulated by calcineurin, a CaM-stimulated phosphatase that dephosphorylates NFATs, thus allowing them to translocate into the nucleus [26]. CnA is sensitive to changes in calcium levels and it is positively regulated by other calcium signalling proteins such as CaM and calpain [27–33]. Therefore, experiments were conducted to study the role of CnA, CaM, and calpain as upstream regulators of the NFAT pathway within the context of reversible cardiac hypertrophy during hibernation. Furthermore, the downstream targets myoferlin and myomaker were evaluated because of their newly identified and crucial role as muscle membrane proteins in muscle development and repair [46, 48–50].
The relationship between myoferlin and NFATc1-4 was established when it was observed that the myoferlin promoter contains multiple NFAT-binding sites that were sufficient to drive high levels of myoferlin expression, especially following muscle damage [50]. However, the link between myomaker and NFAT remains more of a mystery. Currently, only MyoD and MyoG have been shown to induce myomaker transcription due to its very recent discovery [49]. However, MyoD is a common co-factor of NFAT as they can bind to the same transcriptional complex, and they have been shown to synergistically regulate the transcription of myog [38]. Comparative sequence analysis of the myomaker promoter region for potential NFAT transcription factor-binding sites resulted in the identification of multiple sites with the consensus NFAT-binding sequence (GGAAA) from the literature [25, 56]. Following further diagnostic analysis of potential NFAT-binding sites using the rVista alignment tool, one NFAT-binding domain 1095 bp upstream of the myomaker transcriptional start site was identified as it contained the consensus sequence in addition to showing conservation in squirrel-mouse and squirrel-human pairwise alignments (Fig. S1). Therefore, comparative sequence analysis of the myomaker promoter region has resulted in the identification of a possible novel NFAT-myomaker binding site. Using DPI–ELISAs, NFATc1, and c3 were then shown to bind to this site differentially over the torpor-arousal cycle (Fig. 6). DPI–ELISAs represent a simple and efficient way to measure transcription factor-binding activity by quantitatively measuring the ability of transcription factors to bind to DNA from nuclear extracts. This is the first time that NFAT binding to the myomaker promoter has ever been identified, and this novel finding could lead to further studies that characterize NFAT regulation of this vital myogenic protein.
Comparative amino acid sequence analysis of NFATc1-4, myoferlin, and myomaker proteins was also conducted, showing that there is a high degree of conservation in all proteins analysed between 13-lined ground squirrel, mouse, and human (Fig. S2). This suggests that the subsequent immunoblotting analysis conducted using primary antibodies that were raised against hosts other than ground squirrels is still an accurate measure of protein levels in squirrel due to the conservation of the proteins of interest across mammals. Immunoblotting analysis showed elevations of NFATc2 at late torpor in the skeletal muscle of ground squirrels. Although all NFATs are expressed in skeletal muscle and are important for proper muscle function, NFATc2 and c3 are mostly commonly associated with myoblast fusion and muscle repair [38, 43, 50, 61]. Therefore, the increase in NFATc2 protein levels (Fig. 1) and the progressive increase in NFATc3-DNA binding throughout torpor (Fig. 6) are indicative of increased skeletal muscle regeneration and repair in an effort to maintain skeletal muscle mass, despite disuse-induced muscle wasting during torpor [60]. Similar to NFATc3-DNA binding, myoferlin protein levels increased significantly during torpor as well; rising upon entry into torpor (EN) and peaking at early (ET) and late (LT) torpors before decreasing to euthermic levels (Fig. 3). Therefore, despite not showing significant differences in protein levels, the large increase in NFATc3 activity during torpor may be driving the increase in myoferlin levels that maintain skeletal muscle mass [50]. This increase in myoferlin supports the hypothesis that there is an increase in muscle regeneration during torpor. Wu et al. also demonstrated that protein levels of forkhead box transcription factors of the O subclass 3a (FOXO3a) as well as its phosphorylated and activated forms also peaked during torpor [62]. FOXO3a regulates the expression of many ubiquitin ligases (i.e. MAFbx and Murf1) in the ubiquitin proteasomal system (UPS), which is the main pathway controlling muscle atrophy in skeletal and cardiac muscle, leading to heart failure and many muscle wasting diseases [63–66]. Therefore, the increase in both hypertrophic and atrophic factors in the skeletal muscle of ground squirrels during torpor suggests that there could be simultaneous degeneration and regeneration occurring to preserve skeletal muscle mass.
In cardiac muscle, the response of NFATc1-4 transcription factors as well as the downstream targets myoferlin and myomaker was enhanced during torpor (Figs. 2, 4). Similarly to skeletal muscle, NFATc2 was significantly elevated during torpor. Although myoferlin was elevated in both heart and skeletal muscle, myomaker remained stable in skeletal muscle throughout hibernation. In the heart, myomaker showed a spike at ET (1.88-fold higher in comparison with control, p < 0.05) (Fig. 4). After initial increases in NFATc2 as well their hypertrophic targets, NFAT c4 increased during arousal (Fig. 2). The regulator of NFATs, CnA decreased during arousal by 62 % from EA to IA (p < 0.05). One of the activators of CnA, CaM, also decreased during arousal. Similarly, calpain levels declined from euthermic control (EC) to EA by 61 % (p < 0.05). Calpain cleaves the autoinhibitory domain of calcineurin, thus activating it [30–33]. Therefore, these calcium signalling proteins (CaM and calpain) as well as the NFAT–CnA pathway and its downstream muscle targets (myoferlin and myomaker) could be part of the mechanisms behind reversible cardiac muscle hypertrophy. During torpor, NFATc2 could be causing up-regulation of myoferlin, thus promoting cardiac hypertrophy to increase contractility to the heart in response to the decrease in heart rate during hibernation [1, 17]. Then, the mechanism behind the rapid reduction of cardiac mass may be initiated by decreases in CnA protein levels and activity, which is the result of lower calpain and CaM levels during arousal. Subsequently, NFATc2 levels decrease and NFATc4 levels increase and this may be causing the downregulation of both myoferlin and myomaker, thus reversing the cardiac hypertrophic stimulus. Although the NFAT–Cn pathway has been studied in association with cardiac hypertrophy in the past, the present findings identify a novel mechanism of rapidly naturally reversing cardiac hypertrophy, whereby decreased calcium signalling results in a downregulation of muscle hypertrophy proteins targeted by NFATs [21, 34, 37].
In summary, the present study provides insight into some of the key proteins involved in skeletal and cardiac muscle remodelling during hibernation. Although more remains to be investigated, including the regulation of factors associated with the UPS, the results from this study indicate that calcium signalling proteins (calcineurin, calmodulin, calpain) regulating the activity of NFATc1-4 as well as the targets myoferlin and myomaker are important to the hypertrophy and preservation of skeletal and cardiac muscle size and function during hibernation. In addition, the identification of a novel NFAT-binding site in the promoter of myomaker suggests that NFAT could directly regulate the expression of this essential and recently discovered muscle fusion protein. This is also the first study that has found the myomaker protein to be expressed in the heart, although its specific role still remains unclear. Further elucidation of the role that the NFAT–Cn pathway plays in muscle growth and wasting in thirteen-lined ground squirrels will not only contribute to our understanding of muscle remodelling, but it could also potentially lead to the applications for treating chronic muscle wasting diseases and maladaptive cardiac hypertrophy. Specifically, the identification of calpain and CaM as initiators for reversing cardiac hypertrophy in squirrels shows promise. We also identified that measuring NFAT activity or DNA-binding using DPI–ELISAs in combination with protein levels allows for a more in-depth analysis of transcriptional regulation by NFATs. The unique and tissue-specific mechanisms in ground squirrel of preserving skeletal muscle and undergoing reversible cardiac hypertrophy during the torpor-arousal cycle make studying this animal biologically and clinically relevant.
References
Frerichs KU, Hallenbeck JM (1998) Hibernation in ground squirrels induces state and species-specific tolerance to hypoxia and aglycemia: an in vitro study in hippocampal slices. J Cereb Blood Flow Metab 18:168–175. doi:10.1097/00004647-199802000-00007
Storey KB (2010) Out cold: biochemical regulation of mammalian hibernation—a mini-review. Gerontology 56:220–230. doi:10.1159/000228829
Storey KB, Storey JM (2004) Metabolic rate depression in animals: transcriptional and translational controls. Biol Rev Camb Philos Soc 79:207–233. doi:10.1017/S1464793103006195
Wang LCH, Lee TF (1996) Torpor and hibernation in mammals: metabolic, physiological, and biochemical adaptations. In: Fregley MJ, Blatteis CM (eds) Handbook of physiology: environmental physiology, section 4, vol 1. Oxford University Press, New York, pp 507–532
Pier M, Ni Z, McMullen DC, Storey KB (2008) Expression of Nrf2 and its downstream gene targets in hibernating 13-lined ground squirrels, Spermophilus tridecemlineatus. Mol Cell Biochem 312:121–129. doi:10.1007/s11010-008-9727-3
Fahlman A, Storey JM, Storey KB (2000) Gene up-regulation in heart during mammalian hibernation. Cryobiology 40:332–342. doi:10.1006/cryo.2000.2254
Morin PJ, Storey KB (2007) Antioxidant defense in hibernation: cloning and expression of peroxiredoxins from hibernating ground squirrels, Spermophilus tridecemlineatus. Arch Biochem Biophys 461:59–65. doi:10.1016/j.abb.2007.01.035
Morin P, Storey KB (2006) Evidence for a reduced transcriptional state during hibernation in ground squirrels. Cryobiology 53:310–318. doi:10.1016/j.cryobiol.2006.08.002
Mamady H, Storey KB (2006) Up-regulation of the endoplasmic reticulum molecular chaperone GRP78 during hibernation in thirteen-lined ground squirrels. Mol Cell Biochem 292:89–98. doi:10.1007/s11010-006-9221-8
Malatesta M, Perdoni F, Battistelli S, Muller S, Zancanaro C (2009) The cell nuclei of skeletal muscle cells are transcriptionally active in hibernating edible dormice. BMC Cell Biol 10:19. doi:10.1186/1471-2121-10-19
Choi H, Selpides P-JI, Nowell MM, Rourke BC (2009) Functional overload in ground squirrel plantaris muscle fails to induce myosin isoform shifts. Am J Physiol Regul Integr Comp Physiol 297:R578–R586. doi:10.1152/ajpregu.00236.2009
Bassel-Duby R, Olson EN (2006) Signaling pathways in skeletal muscle remodeling. Annu Rev Biochem 75:19–37. doi:10.1146/annurev.biochem.75.103004.142622
Rourke BC, Yokoyama Y, Milsom WK, Caiozzo VJ (2004) Myosin isoform expression and MAFbx mRNA levels in hibernating golden- mantled ground squirrels (Spermophilus lateralis). Physiol Biochem 77:582–593
Cotton CJ, Harlow HJ (2015) Avoidance of skeletal muscle atrophy in spontaneous and facultative hibernators. Physiol Biochem Zool 83:551–560. doi:10.1086/650471
Gao YF, Wang J, Wang HP, Feng B, Dang K, Wang Q, Hinghofer-Szalkay HG (2012) Skeletal muscle is protected from disuse in hibernating dauria ground squirrels. Comp Biochem Physiol Part A 161:296–300. doi:10.1016/j.cbpa.2011.11.009
Xu R, Andres-Mateos E, Mejias R, MacDonald EM, Leinwand L, Merriman DK, Fink RHA, Cohn RD (2013) Hibernating squirrel muscle activates the endurance exercise pathway despite prolonged immobilization. Exp Neurol 247:392–401. doi:10.1016/j.expneurol.2013.01.005
Frerichs KU, Kennedy C, Sokoloff L, Hallenbeck JM (1994) Local cerebral blood flow during hibernation, a model of natural tolerance to “cerebral ischemia”. J Cereb Blood Flow Metab 14:193–205. doi:10.1038/jcbfm.1994.26
Nakipova OV, Zakharova NM, Andreeva LA, Chumaeva NA, Averin A, Kosarskii LS, Anufriev AI, von Lewinski D, Kockskamper J, Pieske B (2007) The seasonal peculiarities of force-frequency relationships in active ground squirrel Spermophilus undulatus ventricle. Cryobiology 55:173–181. doi:10.1016/j.cryobiol.2007.07.001
Wickler SJ, Hoyt DF, van Breukelen F (1991) Disuse atrophy in the hibernating golden-mantled ground squirrel, Spermophilus lateralis. Am J Physiol 261:R1214–R1217
Frey N, Katus HA, Olson EN, Hill JA (2004) Hypertrophy of the heart: a new therapeutic target? Circulation 109:1580–1589. doi:10.1161/01.CIR.0000120390.68287.BB
Li H, Liu T, Chen W, Jain MR, Vatner DE, Vatner SF, Kudej RK, Yan L (2013) Proteomic mechanisms of cardioprotection during Mammalian hibernation in woodchucks, Marmota Monax. J Proteome Res 12:4221–4229. doi:10.1021/pr400580f
Nelson OL, Rourke BC (2013) Increase in cardiac myosin heavy-chain (MyHC) alpha protein isoform in hibernating ground squirrels, with echocardiographic visualization of ventricular wall hypertrophy and prolonged contraction. J Exp Biol 216:4678–4690. doi:10.1242/jeb.088773
Tessier SN, Storey KB (2012) Myocyte enhancer factor-2 and cardiac muscle gene expression during hibernation in thirteen-lined ground squirrels. Gene 501:8–16. doi:10.1016/j.gene.2012.04.004
Yan L, Kudej RK, Vatner DE, Vatner SF (2015) Myocardial ischemic protection in natural mammalian hibernation. Basic Res Cardiol. doi:10.1007/s00395-015-0462-0
Rao A, Luo C, Hogan PG (1997) Transcription factors of the NFAT family: regulation and function. Annu Rev Immunol 15:707–747. doi:10.1146/annurev.immunol.15.1.707
Rusnak F, Mertz P (2000) Calcineurin: form and function. Physiol Rev 80:1483–1521. doi:10.1172/JCI57909.date
Shibasaki F, Hallin U, Uchino H (2002) Calcineurin as a multifunctional regulator. J Biochem 131:1–15
Yang S, Klee CB (2000) Low affinity Ca2+ -binding sites of calcineurin B mediate conformational changes in calcineurin A. Biochemistry 39:16147–16154
Al-Shanti N, Stewart CE (2009) Ca2+/calmodulin-dependent transcriptional pathways: potential mediators of skeletal muscle growth and development. Biol Rev 84:637–652. doi:10.1111/j.1469-185X.2009.00090.x
Burkard N (2005) Targeted proteolysis sustains calcineurin activation. Circulation 111:1045–1053. doi:10.1161/01.CIR.0000156458.80515.F7
Wu HY, Tomizawa K, Oda Y, Wei FY, Lu YF, Matsushita M, Li ST, Moriwaki A, Matsui H (2004) Critical role of calpain-mediated cleavage of calcineurin in excitotoxic neurodegeneration. J Biol Chem 279:4929–4940. doi:10.1074/jbc.M309767200
Lee SH, Choi J, Kim H, Lee DH, Roh GS, Kim HJ, Kang SS, Choi WS, Cho GJ (2014) FK506 reduces calpain-regulated calcineurin activity in both the cytoplasm and the nucleus. Anat Cell Biol 47:91–100. doi:10.5115/acb.2014.47.2.91
Shioda N, Moriguchi S, Shirasaki Y, Fukunaga K (2006) Generation of constitutively active calcineurin by calpain contributes to delayed neuronal death following mouse brain ischemia. J Neurochem 98:310–320. doi:10.1111/j.1471-4159.2006.03874.x
Lin Z, Murtaza I, Wang K, Jiao J, Gao J, Li P-F (2009) miR-23a functions downstream of NFATc3 to regulate cardiac hypertrophy. Proc Natl Acad Sci USA 106:12103–12108. doi:10.1073/pnas.0811371106
Liu Q, Wilkins BJ, Lee YJ, Ichijo H, Molkentin JD (2006) Direct interaction and reciprocal regulation between ASK1 and calcineurin–NFAT control cardiomyocyte death and growth. Mol Cell Biol 26:3785–3797. doi:10.1128/MCB.26.10.3785
Schubert W, Yang XY, Yang TTC, Factor SM, Lisanti MP, Molkentin JD, Rincon M, Chow C-W (2003) Requirement of transcription factor NFAT in developing atrial myocardium. J Cell Biol 161:861–874. doi:10.1083/jcb.200301058
Molkentin JD, Lu JR, Antos CL, Markham B, Richardson J, Robbins J, Grant SR, Olson EN (1998) A calcineurin-dependent transcriptional pathway for cardiac hypertrophy. Cell 93:215–228. doi:10.1016/S0092-8674(00)81573-1
Armand A-S, Bourajjaj M, Martínez-Martínez S, el Azzouzi H, da Costa Martins PA, Hatzis P, Seidler T, Redondo JM, De Windt LJ (2008) Cooperative synergy between NFAT and MyoD regulates myogenin expression and myogenesis. J Biol Chem 283:29004–29010. doi:10.1074/jbc.M801297200
Delling U, Tureckova J, Lim HAEW, Windt LJDE, Rotwein P, Molkentin JD (2000) A Calcineurin–NFATc3-dependent pathway regulates skeletal muscle differentiation and slow myosin heavy-chain expression. Mol Cell Biol 20:6600–6611
Hudson MB, Woodworth-Hobbs ME, Zheng B, Rahnert J, Blount MA, Gooch JL, Searles CD, Price SR (2014) miR-23a is decreased during muscle atrophy by a mechanism that includes calcineurin signaling and exosome-mediated export. Am J Physiol Cell Physiol 306:C551–C558. doi:10.1152/ajpcell.00266.2013
Schiaffino S, Sandri M, Murgia M (2007) Activity-dependent signaling pathways controlling muscle diversity and plasticity. Physiology (Bethesda) 22:269–278. doi:10.1152/physiol.00009.2007
Tessier SN, Storey KB (2010) Expression of myocyte enhancer factor-2 and downstream genes in ground squirrel skeletal muscle during hibernation. Mol Cell Biochem 344:151–162. doi:10.1007/s11010-010-0538-y
Horsley V, Friday BB, Matteson S, Kegley KM, Gephart J, Pavlath GK (2001) Regulation of the growth of multinucleated muscle cells by an NFATC2-dependent pathway. J Cell Biol 153:329–338. doi:10.1083/jcb.153.2.329
Davis DB, Delmonte AJ, Ly CT, McNally EM (2000) Myoferlin, a candidate gene and potential modifier of muscular dystrophy. Hum Mol Genet 9:217–226
Davis DB, Doherty KR, Delmonte AJ, McNally EM (2002) Calcium-sensitive phospholipid binding properties of normal and mutant ferlin C2 domains. J Biol Chem 277:22883–22888. doi:10.1074/jbc.M201858200
Doherty KR, Cave A, Davis DB, Delmonte AJ, Posey A, Earley JU, Hadhazy M, McNally EM (2005) Normal myoblast fusion requires myoferlin. Development 132:5565–5575. doi:10.1242/dev.02155
Haslett JN, Sanoudou D, Kho AT, Han M, Bennett RR, Kohane IS, Beggs AH, Kunkel LM (2003) Gene expression profiling of Duchenne muscular dystrophy skeletal muscle. Neurogenetics 4:163–171. doi:10.1007/s10048-003-0148-x
Millay DP, O’Rourke JR, Sutherland LB, Bezprozvannaya S, Shelton JM, Bassel-Duby R, Olson EN (2013) Myomaker is a membrane activator of myoblast fusion and muscle formation. Nature 499:301–305. doi:10.1038/nature12343
Millay DP, Sutherland LB, Bassel-duby R, Olson EN (2014) Myomaker is essential for muscle regeneration Myomaker is essential for muscle regeneration. Genes Dev 28(15):1641–1646. doi:10.1101/gad.247205.114
Demonbreun AR, Lapidos KA, Heretis K, Levin S, Dale R, Pytel P, Svensson EC, McNally EM (2010) Myoferlin regulation by NFAT in muscle injury, regeneration and repair. J Cell Sci 123:2413–2422. doi:10.1242/jcs.065375
McMullen DC, Hallenbeck JM (2010) Regulation of Akt during torpor in the hibernating ground squirrel, Ictidomys tridecemlineatus. J Comp Physiol B 180:927–934. doi:10.1007/s00360-010-0468-8
Larkin MA, Blackshields G, Brown NP, Chenna R, Mcgettigan PA, McWilliam H, Valentin F, Wallace IM, Wilm A, Lopez R, Thompson JD, Gibson TJ, Higgins DG (2007) Clustal W and Clustal X version 2.0. Bioinformatics 23:2947–2948. doi:10.1093/bioinformatics/btm404
Goujon M, McWilliam H, Li W, Valentin F, Squizzato S, Paern J, Lopez R (2010) A new bioinformatics analysis tools framework at EMBL-EBI. Nucleic Acids Res 38:695–699. doi:10.1093/nar/gkq313
McWilliam H, Li W, Uludag M, Squizzato S, Park YM, Buso N, Cowley AP, Lopez R (2013) Analysis tool web services from the EMBL-EBI. Nucleic Acids Res 41:597–600. doi:10.1093/nar/gkt376
Eaton SL, Roche SL, Llavero Hurtado M, Oldknow KJ, Farquharson C, Gillingwater TH, Wishart TM (2013) Total protein analysis as a reliable loading control for quantitative fluorescent western blotting. PLoS One 8:1–9. doi:10.1371/journal.pone.0072457
Hung H-F, Wang B-W, Chang H, Shyu K-G (2008) The molecular regulation of resistin expression in cultured vascular smooth muscle cells under hypoxia. J Hypertens 26:2349–2360. doi:10.1097/HJH.0b013e328311fa30
Brand LH, Kirchler T, Hummel S, Chaban C, Wanke D (2010) DPI–ELISA: a fast and versatile method to specify the binding of plant transcription factors to DNA in vitro. Plant Methods 6:25. doi:10.1186/1746-4811-6-25
Brand LH, Henneges C, Schüssler A, Kolukisaoglu HÃ, Koch G, Wallmeroth N, Hecker A, Thurow K, Zell A, Harter K, Wanke D (2013) Screening for protein-DNA interactions by automatable DNA-protein interaction ELISA. PLoS One 8:e75177. doi:10.1371/journal.pone.0075177
Jagelska E, Brázda V, Pospisilová S, Vojtesek B, Palecek E (2002) New ELISA technique for analysis of p53 protein/DNA binding properties. J Immunol Methods 267:227–235. doi:10.1016/S0022-1759(02)00182-5
Hindle AG, Otis JP, Epperson LE, Hornberger TA, Goodman CA, Carey HV, Martin SL (2014) Prioritization of skeletal muscle growth for emergence from hibernation. J Exp Biol 218:276–284. doi:10.1242/jeb.109512
Cho YY, Yao K, Bode AM, Bergen HR, Madden BJ, Oh SM, Ermakova S, Bong SK, Hong SC, Shim JH, Dong Z (2007) RSK2 mediates muscle cell differentiation through regulation of NFAT3. J Biol Chem 282:8380–8392. doi:10.1074/jbc.M611322200
Wu CW, Storey KB (2014) FoxO3a-mediated activation of stress responsive genes during early torpor in a mammalian hibernator. Mol Cell Biochem 1–11. doi:10.1007/s11010-014-1969-7
Foletta VC, White LJ, Larsen AE, Léger B, Russell AP (2011) The role and regulation of MAFbx/atrogin-1 and MuRF1 in skeletal muscle atrophy. Pflug Arch 461:325–335. doi:10.1007/s00424-010-0919-9
Schiaffino S, Dyar KA, Ciciliot S, Blaauw B, Sandri M (2013) Mechanisms regulating skeletal muscle growth and atrophy. FEBS J 280:4294–4314. doi:10.1111/febs.12253
Day SM (2013) The ubiquitin proteasome system in human cardiomyopathies and heart failure. Am J Physiol Heart Circ Physiol 304:H1283–H1293. doi:10.1152/ajpheart.00249.2012
de Andrade LHS, de Moraes WMAM, Matsuo Junior EH, de Orleans Carvalho de Moura E, Antunes HKM, Montemor J, Antonio EL, Bocalini DS, Serra AJ, Tucci PJE, Brum PC, Medeiros A (2015) Aerobic exercise training improves oxidative stress and ubiquitin proteasome system activity in heart of spontaneously hypertensive rats. Mol Cell Biochem 402:193–202. doi:10.1007/s11010-015-2326-1
Acknowledgments
We thank Dr. J. M. Hallenbeck and Dr. D.C. McMullen (NINDS, NIH, Bethesda) for providing the tissue samples for this study. Thanks also to J. M. Storey for editorial review of the manuscript. This work was supported by a Discovery grant from the Natural Sciences and Engineering Research Council (NSERC) of Canada (# 6793) and a grant from the Heart and Stroke Foundation of Canada (#G-14-0005874) to KBS. KBS holds the Canada Research Chair in Molecular Physiology; YZ held a postgraduate Queen Elizabeth II Graduate Scholarship in Science and Technology.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zhang, Y., Storey, K.B. Expression of nuclear factor of activated T cells (NFAT) and downstream muscle-specific proteins in ground squirrel skeletal and heart muscle during hibernation. Mol Cell Biochem 412, 27–40 (2016). https://doi.org/10.1007/s11010-015-2605-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11010-015-2605-x