Abstract
Concerns on increasing trends of multidrug resistance (MDR) around the world have triggered the need to investigate and develop new therapeutic strategies and potent antibacterial drugs. Polymyxins, a class of polycationic antimicrobial peptides, have been regarded as the last-line therapy against Gram-negative bacteria due to limited new antibiotics and phenylalanine-arginine β-naphthylamide (PAβN), a peptidomimetic compound has been characterised as an efflux pump inhibitor (EPI) that have been investigated to overcome efflux-mediated multidrug resistance. In this work, the antibacterial activity of two Schiff base ligands derived from the condensation of S-benzyl dithiocarbazate with 4-carboxybenzaldehye (SB4CB) and 4-formyl-3-hydroxybenzoic acid (SBFH) their copper(II) complexes (Cu(SB4CB)2 and Cu(SBFH)2) were tested individually, and the most promising compound was tested in combination with polymyxin B (POLY) and PAβN against different bacteria, such as antibiotic-susceptible strains: Acinetobacter baumannii ATCC 19606, Escherichia coli ATCC 25922, Pseudomonas aeruginosa ATCC 27853, and Staphylococcus aureus ATCC 35923; multidrug-resistant strains: A. baumannii ATCC BAA-1797, E. coli BAA-196, P. aeruginosa ATCC BAA-2108 and S. aureus ATCC 43300. Initial minimum inhibition concentration (MIC) results showed the Cu(II) complexes Cu(SB4CB)2 and Cu(SBFH)2, demonstrated obvious antibacterial activity as compared to the ligand alone. Fractional inhibitory concentration (FIC) index showed improved MIC values with additivity and synergistic effect for Cu(SBFH)2 in combination with POLY and PAβN. From the in silico molecular docking investigation, Cu(SBFH)2 was shown to engage in hydrophobic interactions via its phenyl rings with surrounding hydrophobic residues in the binding pocket of S. aureus NorA, E. coli AcrB, P. aeruginosa MexB and A. baumannii AdeB efflux pumps. The phenyl rings of the ligand could also form π-π stacking with adjacent residues in the binding site of A. baumannii AdeB. Besides, hydrogen bonding and π-cation interactions were also observed via the carboxyl group, hydroxyl group and phenyl ring of SBFH moiety, respectively with nearby residues in the E. coli AcrB binding pocket. This study indicates that the combination strategy of Cu(SBFH)2 with POLY and PAβN enhances therapeutic potential and sheds light on the binding pockets inside bacteria efflux pumps and the binding interactions of ligand in the binding site.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
The World Health Organization (WHO) has noted a rising trend of multidrug resistance (MDR) exhibited by bacteria which reduces the potency and efficacy of many antibiotics. Around 23,000 cases of death were caused by antibiotic-resistance reported in the US annually and Malaysia also observed a steady increasing trend of antibiotic-resistance (World Health Organisation 2014; Meer Ahmad 2019). The patterns of antibiotic-resistance circulate around several bacteria which consist of Escherichia coli (E. coli), Klebsiella pneumoniae (K. pneumoniae), Staphylococcus aureus (S. aureus), and many more (Meer Ahmad 2019). Due to the wide spreads of resistances, scientists and healthcare researchers are urgently putting their efforts for the discovery of new strategies and potent drugs that could potentially tackle the MDR challenge.
Polymyxins firstly isolated in 1947 from different species of Bacillus polymyxa, are cationic polypeptides (Dai et al. 2020), sharing similar structure and mechanism of action with cationic antimicrobial peptides (CAMPs) like defensins, which target and disrupt the integrity of the outer membrane of the Gram-negative bacteria, causing it to be susceptible to antibiotics. The polymyxins family has five members in it, which are polymyxin A, B, C, D and E (Samal et al. 2021; Fazly Bazzaz et al. 2021). However, only polymyxin B (available as sulfate salt) and polymyxin E (also called colistin) are used clinically, as last line therapy to treat life-threatening infections caused by Gram-negative bacteria, particularly Pseudomonas aeruginosa (P. aeruginosa) and Acinetobacter baumannii (A. baumannii) which are identified as the “Priority 1: Critical” pathogens in the 2017 WHO Priority Pathogen List (Nang et al. 2021).
Efflux pumps are membrane proteins that can extrude harmful cell substances such as antibiotics, toxins and waste metabolites from the bacteria into the external environment, hence playing a role in antimicrobial resistance. There are five superfamilies of efflux pumps which are involved in MDR: multidrug and toxin extrusion (MATE), small multidrug resistance (SMR), major facilitator superfamily (MFS), ATP-binding cassette (ABC) and resistance-nodulation division (RND) (Alav et al. 2018; Auda et al. 2020). Over the years, researchers found out that efflux pumps of Gram-negative strains such as P. aeruginosa and E. coli belong to the RND family. While MFS (for example, NorA in S. aureus) and ABC transporters are one of the most frequent efflux pumps found in Gram-positive bacteria (Soto 2013). Phenylalanine-arginine β-naphthylamide (PAβN), one of the most studied efflux pump inhibitors (EPIs) has proven to be able to inhibit bacterial efflux pumps from eluting out the antibiotics through several mechanisms. It can act as a competitive inhibitor by binding to the efflux pumps or alternatively steric hindrance generated during binding can impair antibiotics from binding to its affinity site (Jamshidi et al. 2017).
Combination therapy has shown various advantages such as preventing resistance development and reducing doses of respective drug or treatment periods (Zusman et al. 2017). An example of this would be the utilisation of polymyxin in combination with other agents due to its ability to disrupt the outer membrane integrity of Gram-negative bacteria and therefore enhancing the activity of other drugs during combination (Lenhard et al. 2016; Nang et al. 2021). Low et al. (2014) studied that with the presence of polymyxin B nonapeptide in combination with Schiff bases derived from S-benzyl dithiocarbazate (SBDTC) such as SB4CB and Cu(SB4CB)2, the antibacterial properties was shown to increase against A. baumannii ATCC 19606, K. pneumoniae ATCC 11296, P. aeruginosa PA01, Salmonella enterica SL696 and S. aureus SA1199. On the other hand, Lamers et al. (2013) also reported improvement on antibacterial activities of other antibiotics when used in combination with PAβN. In the presence of 25 µg/mL PAβN, the MIC values of vancomycin reduced from > 256 to 96 µg/mL for the wild type and dacB strains of P. aeruginosa. The MIC values of vancomycin further reduced to 32 µg/mL when treated the bacteria with 50 µg/mL PAβN. Interestingly, the researchers found out that the susceptibility change was not entirely due to efflux inhibition by PAβN, the reduction of MICs was also potentially due to its permeabilising ability across the bacterial outer membrane (Lamers et al. 2013; Rampioni et al. 2017).
In 1864, synthetic chemist Hugo Schiff successfully synthesised and introduced a class of compounds containing the functional groups azomethine or imine, known as Schiff bases (Tidwell 2008; Singh and Barman 2021). These Schiff bases can be synthesised by using the condensation method of primary amine with an aldehyde or a ketone. Numerous publications were reported over the decades, highlighting the potential of Schiff base and their metal complexes for the design of novel antibacterial therapeutic drugs (Djoko et al. 2015; Singh and Barman 2021). Among the metal ions used in metal complexes, copper (Cu) ions received tremendous interest because of its excellent potential in therapeutic applications. Dhahagani et al. (2018) reported the Cu(II) complexes of morpholine derived Schiff base ligands displayed better antibacterial activities compared to corresponding Schiff base ligands as well as standard drug amikacin, indicating the importance of chelation in facilitating metal complexes to cross the cell membranes. Similar enhancement upon coordination was also observed for Cu(II) complexes with 5-nitroimidazole-derived Schiff bases that exhibit potent antimicrobial activity against pathogenic anaerobic bacteria (Oliveira et al. 2018). In addition, Cu(II) Schiff base complexes have also been explored for their anticancer (Singh et al. 2020; Foo et al. 2019; Deng et al. 2018), anti-inflammatory (Choo et al. 2018), antifungal (Lima et al. 2018) and positron emission tomography (PET) hypoxia imaging (Brown et al. 2017) properties.
Therefore, the antibacterial potentials of synthesised Schiff bases and their Cu(II) complexes were evaluated in this work by determining their activities against all eight bacteria isolates, including both susceptible and resistant strains of S. aureus, P. aeruginosa, E. coli and A. baumannii. As an extension of previous work (Low et al. 2014) on S-benzyl dithiocarbazate (SBDTC) derived Schiff base ligands and Cu(II) complexes where small structural modification can affect the bioactivity, 4-formyl-3-hydroxybenzoic acid with an addition hydroxyl, –OH group was of interest to form SBFH and Cu(SBFH)2 as past study reported that compounds with hydroxyl group demonstrated good antimicrobial activity (Hejchman et al. 2019). Combination tests were further carried out using the most promising compound Cu(SBFH)2 with POLY and PAβN. Furthermore, to support in vitro antibacterial study results, the molecular docking studies were carried out to investigate the binding modes and interactions of the compound within the active site of efflux pumps, and also to understand the molecular recognition of ligands within the binding pocket.
Materials and methods
Synthesis
Materials and instrumentations
All chemicals and solvents were of analytical grade: Cu(II) acetate monohydrate (Cu(OAc)2·H2O) (Merck), 4-formyl-3-hydroxybenzoic acid (Sigma-Aldrich), dimethyl sulfoxide (DMSO) (Fisher Chemicals) and acetonitrile (ACN) (Fisher Chemical). The Fourier-transform infrared (FTIR) spectra were recorded in the range of 400–4000 cm−1 on Shimadzu IRAffinity-1S FTIR spectrophotometer in ATR mode and under 20 total scans with four resolution settings. Spectra of compounds were saved and replotted using ORIGIN software. Elemental analysis results were obtained by using the LECO MicroTruspec CHNS Elemental Analyzer which is available in Universiti Malaya. Perkin Elmer Lambda 25 UV/VIS spectrophotometer using quartz cuvette with 1 cm optical path were used to record UV–Visible spectra. 1H Nuclear Magnetic Resonance (NMR) and 13C NMR were recorded on Bruker DRX300.
Preparation of SBDTC, SB4CB and Cu(SB4CB)2
SBDTC used in this project was previously synthesised at University Putra Malaysia (UPM) by reacting carbon disulfide, hydrazine and benzyl chloride. SB4CB and Cu(SB4CB)2 were reprepared in International Medical University (IMU) following previously reported procedure (Low et al. 2014). Briefly, to a solution of SBDTC in ACN, an equimolar amount of 4-carboxybenzaldehyde in the same solvent was added dropwise. The mixture was heated to reduce the volume to about one third of the original volume. The SB4CB products formed were filtered and dried. For Cu(SB4CB)2, a solution containing a half-molar amount of Cu(OAc)2·H2O, dissolved in acetonitrile was added dropwise to a solution of the ligand, SB4CB in ACN. The resulting mixture was heated to reduce the volume to about one third of the original volume and then let to cool to room temperature. The precipitates were filtered and dried. FTIR spectroscopy and elemental analysis were used to confirm the compounds in comparison to reported data.
Preparation of SBFH
A total 0.129 g (0.6254 mmol, 1 equiv.) of SBDTC was dissolved in 20 mL hot ACN. Next, 0.0940 g (0.6254 mmol, 1 equiv.) of 4-formyl-3-hydroxybenzoic acid predissolved in 40 mL hot ACN was added dropwise slowly into the solution of SBDTC. The mixture was stirred and heated on an electric stirrer hotplate for 1.5 h until solvent has reduced to one third of its original volume. Yellow precipitate was filtered out, dried and collected to get 0.108 g of product (Yield = 50%). Elemental analysis for C16H14N2O3S2: Calcd. C 55.48, H 4.07, N 8.09; Found C 55.81, H 3.69, N 7.76.
Preparation of Cu(SBFH)2
A total 0.0346 g (0.1 mmol, 1 equiv.) of SBFH was dissolved in 20 mL hot ACN. 0.0104 g (0.05 mmol, 0.5 equiv.) of Cu(OAc)2·H2O predissolved in 10 mL hot ACN was transferred dropwise slowly into the solution of SBFH. The mixture was heated and stirred for 1 h until solvent has reduced to one third of its original volume. Brown precipitate was filtered, dried, and collected to get 0.0110 g of product (Yield = 29%). Elemental analysis for C32H26CuN4O6S4: Calcd. C 50.95, H 3.47, N 7.43; Found C 50.60, H 3.04, N 7.21.
Antibacterial properties
Materials and instrumentations
The materials used in antibacterial assessments are Muller-Hinton broth (Oxoid), Muller-Hinton agar (Oxoid), sodium chloride (Sigma), calcium chloride dihydrate (Merck), magnesium chloride (Calbiochem), gentamicin sulfate (Sigma), ampicillin (Sigma), polymyxin B sulfate (Sigma) and phe-arg-β-naphthylamide dihydrochloride (Sigma). The bacteria strains used in this study were obtained from the American Type Culture Collection (ATCC): A. baumannii ATCC 19606, E. coli ATCC 25922, P. aeruginosa ATCC 27853, S. aureus ATCC 35923, A. baumannii ATCC BAA-1797, E. coli ATCC BAA-196, P. aeruginosa ATCC BAA-2108 and S. aureus ATCC 43300. The bacterial glycerol stock (10%) was made for each of strains, and stored at -80 °C. These bacterial stocks were streaked and grown at 37 °C on MHA plates for overnight cultivation prior to any assay. Instrumentation used to quantify the bacteria amount was based on absorbance which is measured by SpectraMax® M3 Multi-Mode Microplate Reader at 625 nm wavelength. The value of OD625 ≈ 0.08–0.13 is approximately to 108 CFU/mL that equivalence to 0.5 McFarland.
Evaluation of minimum inhibition concentration (MIC) on susceptible and resistant strains
The microdilution procedures were adapted based on the Standard Operating Procedures (SOP) drafted by Monash University Facility for Anti-Infective Drug Development and Innovation (FADDI) and Clinical and Laboratory Standards Institute (CLSI) guidelines (Grace et al. 2016). The stock solutions of the compounds and antibiotics were prepared in concentration of 1.024 × 10–2 M by dissolving the solid powder in 100% DMSO and ultrapure water, respectively. The stock solutions were then serially diluted two-fold to obtain final concentrations in the 96 wells plate with the range of 0.0625 to 128 µM. The final DMSO concentration was maintained at not more than 1.25%. The 0.5 McFarland was prepared via direct colony suspension from the overnight culture plate. Briefly, the pure bacteria colonies were aseptically picked and re-suspended into 1 mL sterile normal saline solution (0.9% NaCl), the absorbance was then measured using the microplate reader of the brand of SpectraMax at wavelength 625 nm. As described, optical density (OD625) range between 0.08 and 0.13, was used as it is yielding 0.5 McFarland turbidity. A total 200 µL of this 0.5 McFarland bacterial suspension was added to the 19.8 mL of cation-adjusted Mueller–Hinton broth (CAMHB). A volume of 100 µL synthesised compounds followed by 100 µL bacteria suspension were pipetted carefully into each well of 96 wells U-bottomed plate and incubated for 18 h at 37 °C. MIC is defined as the lowest compound concentration that exhibited complete inhibition of microbial growth. All assays were performed in at least duplicate.
Evaluation of combination test between Cu(SBFH)2 with POLY and PAβN on susceptible and resistant strains
Cu(SBFH)2 due to its promising antibacterial activity was selected for combination tests alongside POLY and PAβN against all bacteria. Hence, a checkerboard method was used to generate combinations between Cu(SBFH)2 with POLY and PAβN, respectively (Hsieh et al. 1993). Procedures during combination was similar to MIC evaluation, except for each well, a combination of 50 µL Cu(SBFH)2 + 50 µL POLY/PAβN + 100 µL bacteria suspension was added to obtain the final concentrations. The plates were then incubated in 37 °C for 18 h. Fractional inhibitory concentration (FIC) index was calculated. FIC index is a mathematical formula expression to represent the degree of interaction between two drugs (Fig. 1).
In Fig. 1, the (X) represents concentration of drug X in one of the wells along the border line of the clear-cloudy or growth-no-growth wells; meanwhile the MICX represents control MIC of drug X alone against the bacteria. FICX is the fractional inhibitory concentration of drug X that is calculated using MICs of drug X in the combination divided by the MICs of drug X alone. For drug Y, MICY and FICY are determined and calculated like drug X as described previously. FIC index is the sum of FICX and FICY. “Synergism” is defined when FIC index is of ≤ 0.5, “Additivity” when between > 0.5 and ≤ 1, “Indifference” is of > 1 to ≤ 4 and “Antagonism” when FIC index is of > 4 (Hsieh et al. 1993; Walsh et al. 1995). Only additive or synergistic effects were shown in the following result tables.
Molecular docking
Protein preparation
The efflux pump protein structure of E. coli, P. aeruginosa and A. baumannii AcrB (PDB:1T9Y) (Aparna et al. 2014), MexB (PDB: 2V50) (Aparna et al. 2014) and AdeB (PDB: 7KGG) (Morgan et al. 2021), respectively were retrieved from the Protein Data Bank (PDB). Due to unavailability of crystal structure of S. aureus NorA, homology model was constructed from glycerol 3-phosphate transporter pump of E. coli (PDB ID: IPW4) as previously reported (Singh et al. 2017). Maestro of Schrödinger Suite was used to run the in silico molecular docking studies. The crystal structures of AcrB, MexB, AdeB and NorA were prepared using the protein preparation wizard module. Missing atoms or incomplete residues of the active site were fixed prior to the docking. In addition, crystallographic waters were removed from the crystal structures. Hydrogen bonding networks were automatically optimized, and the resultant protein structure was energy-minimised prior to docking.
Ligand preparation
2D sketcher in Maestro was used to draw the 2D chemical structures of all compounds. Except for the Cu(II) Schiff base complexes, a structure-cleaning step utilising LigPrep was carried out to convert two-dimensional (2D) structures of the compounds to three-dimensional (3D), to generate stereoisomers, and to determine the most probable ionization state at neutral pH 7; conformers for each compound were generated through ConfGen by applying the OPLS-2005 force field method.
Docking simulation
Docking grid for AcrB and MexB was mapped onto each other due to the absence of an inhibitor-MexB co-crystal PDB structure of P. aeruginosa and hence AcrB-MC-207110 complex structure (PDB ID:1T9Y) from E. coli was used to identify the binding site information of P. aeruginosa MexB. Docking grid of AdeB was centred on the binding site of reference substrate, ethidium with grid coordinates of X: 179.34, Y: 159.2, Z: 166.54. For NorA, the grid coordinates are X: 33.35, Y: 20.68 and Z: -37.65. Glide docking program of Schrödinger suite was used for the docking studies of all compounds on AcrB, MexB, AdeB and NorA efflux pumps. Standard precision (SP) docking calculation was performed on the compounds in the AcrB, MexB and NorA, while Extra precision (XP) docking calculation was used for the compounds in the AdeB. Binding poses and interactions of each compound with adjacent residues of binding pocket were analysed.
Results and discussion
Synthesis and characterisations
The synthesis of the Schiff bases (SB4CB and SBFH) and their Cu(II) complexes metal complexes is shown in general Scheme 1. The empirical formula and purity of each compound were established by elemental analysis. The analytical data agreed well with the formulations proposed. All compounds synthesised were found to be stable under room temperature and at atmospheric conditions. SBFH and Cu(SBFH)2 being new compounds were also characterised using FTIR, UV–Vis and NMR (Supporting Information). The stability of SB4CB and Cu(SB4CB)2 under HPLC aqueous acidic conditions were previously reported in Low et al. (2014). Aromatic Schiff bases and complexation to Cu(II) ions significantly increased stability. SBFH and Cu(SBFH)2 are also expected to display similar stability as their only difference from SB4CB and Cu(SB4CB)2 is the additional –OH group.
FTIR
Strong bond structured bands stretching from 3260 to 3300 cm−1 in the IR spectra of SBFH ligand and its corresponding Cu(II) complex indicates νOH, νNH and νCH overlapping stretching vibrations. SBFH possess bands ν(C=O) at 1699 cm−1, ν(C=N) at 1597 cm−1, ν(C=S) at 1036 cm−1 and ν(CSS) band at 959 cm−1. Upon complexation with Cu(II) ions, ν(C=S) disappeared and ν(CSS) split into two peaks due to deprotonation and coordination via the thiolate form of sulphur atoms. ν(C=N) band shifted to a lower wavenumber further confirming the coordination of azomethine nitrogen atom of the Schiff base ligand (Low et al. 2014; Crouse et al. 2004).
UV–Vis
Both SBFH and its Cu(II) complex were scanned through UV–Vis spectrometer at the concentrations of 25 \(\mu\)M and 1 mM (metal complex only) in DMSO. SBFH showed two bands from ca. 360 nm to 450 nm which indicates \(\pi \to \pi\)* and n \(\to \pi\)* transitions. The λ max of SBFH is 370 nm. For copper complex, the intraligand band at 300 nm to 400 nm are assigned to n \(\to \pi\)* and \(\pi \to \pi\)* transition of the aromatic ring. This is because of the lone pair of electrons from the aromatic ring is being donated to the metal resulting in the occurrence of the coordination of azomethine group (Latheef and Kurup 2008). There is d-d transition observed at 600 nm for Cu(SBFH)2 which indicates to have Jahn–Teller distortion, a distorted square planar environment around the Cu(II) ion. The Cu(II) complex also showed ligand-to-metal charge transfer (LMCT) band (400–450 nm) arising from S \(\to\) Cu(II) interaction at concentration of 25 \(\mu\)M (Crouse et al. 2004). This proves that the Cu(II) complex formed is coordinated with sulphur atom (Low et al. 2016, 2014). UV–Vis titrations were also carried out to confirm the formation of expected 2:1 ligand–metal complex system. The titration was performed in acetate buffer pH 6 for aqueous solution studies with the concentration of ligand at 10–5 M. After addition of Cu(OAc)2, the changes of spectrum were observed with a wide absorption band which ascribed from the formed compound at the region of 350–400 nm with λmax ≈ 353 nm (log ε = 4.20) while the intensity of the band of ligand at λmax ≈ 353 nm (log ε = 4.20) decreases. The spectrum shows clear isosbestic point which illustrate a single complexation event and clear end-point at 0.5 equivalent of Cu which confirmed the formation of 2:1 ligand–metal complex (Low et al. 2014; How et al. 2008; Akbar Ali et al. 2004).
NMR
The 1H NMR and 13C NMR spectra of SBFH were recorded in DMSO-d6 solvent. The spectra obtained were assigned to each hydrogen present in the ligand as expected from the drawn structure. The NH signals occurred above 12 ppm illustrated that the predominant isomer is in Z-configuration (Rebolledo et al. 2005). The signal at 8.5 ppm was assigned to CH=N while the multiplets at 7.28–7.78 ppm were attributed to the overlapping of aromatic hydrogens from both SBDTC and 4-formyl-3-hydroxybenzoic acid moiety. The other characteristics signal of SBFH was the singlet signal at 4.5 ppm which was assigned to S-CH2 of SBFH. 13C NMR was carried out in Attached Proton Test (APT) in which the positive signals yield the CH and CH3 signals whereas the negative signals are assigned to quaternary C and CH2 signals. Each signal is ascribed from each carbon signal from the expected structure of SBFH. The signal occurred at ca. 196.26 ppm was assigned to –CSS– peak. This signal demonstrated that the thione form predominates in the solution (Low et al. 2016, 2014). The presence of –C=N– at ca. 142.68 ppm was due to the formation of hydrazone bond during the reaction of 4F3H and SBDTC. The spectrum shows carbon (C–OH) occurred at ca. 155 ppm. The aromatic carbon signals have appeared at the region of ca. 116.94–136.68 ppm.
MIC determination
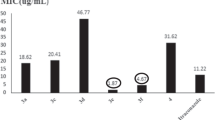
Referring to Tables 1 and 2, Cu(SBFH)2 and Cu(SB4CB)2 showed promising inhibitory activity against S. aureus ATCC 35923 (methicillin-susceptible S. aureus, MSSA) and S. aureus ATCC 43300 (methicillin-resistant S. aureus, MRSA) bacterial strains with lower MIC values compared to the remaining tested bacteria in our experiments. Cu(SBFH)2 was reported to have the lowest MIC value at 4 µM, followed by Cu(SB4CB)2 at 8 µM when tested on MSSA. Meanwhile for MRSA, MIC values of Cu(SBFH)2 and Cu(SB4CB)2 were 8 µM and 4 µM, respectively. These MIC values were closely compared with the commercialised antibiotics and drugs such as gentamicin sulfate, ampicillin, POLY and PAβN. After MICs determination of all compounds, Cu(SBFH)2 and Cu(SB4CB)2 possessed similar effects against Gram-positive bacteria S. aureus, hence, only Cu(SBFH)2 was selected for combination tests as it possessed overall better antibacterial activity (MIC = 128 µM) against Gram-negative bacteria susceptible strain A. baumannii ATCC 19606, resistant strain P. aeruginosa ATCC BAA-2108 and resistant strain A. baumannii ATCC BAA-1797 compared to Cu(SB4CB)2 with MIC values of > 128 µM. A study conducted by Said et al. (2020) in synthesising Schiff base compounds showed strong antimicrobial activity against Gram-positive and Gram-negative bacteria. While most compounds with a hydroxy group shows good antibacterial property, one of the compounds synthesised was a Schiff base with three hydroxyl groups, is of particular interest, due to its superior activity compared to the other compounds synthesised (Said et al. 2020). The antimicrobial activities of SBDTC-derived Schiff base metal complexes were also supported via studies by Low et al. (2014), Adly and El-shafiy (2019) as well as Damit et al. (2021) in which the complexes have higher activities as compared to the organic ligand and showed highest activity towards the Gram-positive bacteria. The compounds were more active against Gram-positive than Gram-negative bacteria due to the presence of an additional outer layer of lipid membrane in Gram-negative bacteria which affects the drug uptake as described by other studies (Bolla et al. 2011; Meer Ahmad 2019).
Combination test
Although Cu complexes have been reported to enhance antimicrobial properties against several bacterial strains (Dhahagani et al. 2018; Nair et al. 2012; Kremer et al. 2006; Claudel et al. 2020), Cu(SBFH)2 only showed potent activities towards S. aureus from the MIC determination. Also, there are research findings about development of tolerance to Cu ions in bacteria via several mechanisms such as active efflux systems and detoxifications by copper oxidase enzymes (Breijyeh et al. 2020). As there are studies supporting that combinations could help to improve antibacterial activities (Low et al. 2014; Lamers et al. 2013), Cu(SBFH)2 was combined with a membrane permeabilising agent, such as POLY and an efflux pumps inhibitor, such as PAβN, hoping to improve uptake of the compound under investigation and consequently affect its antimicrobial efficiency in a positive manner. Different sets of ratio for combination test of Cu(SBFH)2 with POLY and Cu(SBFH)2 with PAβN had been used and the results were tabulated in Tables 3 and 4, respectively.
Based on the results shown in Table 3, combination of Cu(SBFH)2 and POLY showed additive interaction, against both susceptible and resistant strains of P. aeruginosa as well as A. baumannii. The FIC value shown when tested against susceptible and resistant strains of P. aeruginosa is 1.0000, whereas when it was tested against both susceptible and resistant strains of A. baumannii, the FIC value ranged from 0.5625 to 1.0000 and 0.5313 to 1.0000 respectively. For P. aeruginosa susceptible and resistant strains, there was a twofold decrease in MIC value of Cu(SBFH)2 when 0.5 µM and 1 µM of POLY were added. However, the additive interaction was more prominent when tested against both susceptible and resistant strains of A. baumannii, noting a twofold decrease in MIC value of Cu(SBFH)2 when POLY were used in susceptible and resistant strains respectively. Interestingly, there was a fourfold decrease in both susceptible and resistant strains in MIC value of Cu(SBFH)2 when concentration of POLY was used at 0.5 µM and 1 µM, respectively. The additive effect of combining POLY with other antimicrobial drugs was consistent with several studies which showed similar effect. Based on a study by Yoon et al. (2004), a combination of POLY with other antibiotics (rifampin or imipenem) was shown to exhibit either synergistic (FIC ≤ 0.5) or additive (FIC > 0.5–1.0) effect against 2 isolates of A. baumannii resistant strains. A report by Manikal et al. (2000) reinforced this, where it was reported that a combination between POLY and azithromycin via chequerboard tested against 24 A. baumannii isolates (both resistant and susceptible strains) and showed synergistic or additive interaction. Combination of POLY and azithromycin showed synergistic (FIC range ≤ 0.18 − 0.5) and additive (FIC value range 0.52 − 1.0) interaction against 4 and 20 strains respectively (Manikal et al. 2000). In another study by Guelfi et al. (2008), drug combination between POLY and meropenem was more effective against A. baumannii strains (with 8 out of 10 strains showing additive interaction) with FIC ranging ≤ 0.625 − 0.75 compared to P. aeruginosa strains (with 2 out of 10 strains showing additive interaction) with FIC range ≤ 0.75–1. Apart from combining POLY with FDA-approved antibitoics, the additive effect of POLY with novel compounds was noted as well. One such study conducted by Figueiredo et al. (2019) had shown that one of the 5-hydrazinylethylidenepyrimidine compound exhibited additive interaction when combining it with POLY, with an FIC value ≤ 1.0 when tested against a resistant strain of A. baumannii (Figueiredo et al. 2019). It was hypothesised that the additive or synergistic interaction of the drug combination can be attributed to the rapid permeabilization of the outer membrane of the Gram-negative bacteria by POLY, thus enhancing antibiotics penetration into the bacteria (Yoon et al. 2004; Zharkova et al. 2019; Guelfi et al. 2008; Manikal et al. 2000). In addition to that, it was shown that drug combination of POLY with other drugs when tested, was more effective against A. baumannii compared to P. aeruginosa. This was supported by the minimum FIC shown in study by Guelfi et al. (2008), with FICmin of 0.75 in P. aeruginosa when compared to FICmin of 0.625 in A. baumannii. Our results were consistent with this as well, with our FICmin of 1 in P. aeruginosa when compared to FICmin of 0.5313 in A. baumannii (Guelfi et al. 2008).
Our combination results as shown in Table 4 demonstrated that there was no significant improvement, on MICs of Cu(SBFH)2, neither synergism nor additivity, when it is used in combination with PAβN against S. aureus, E. coli and A. baumannii (Supporting Information). A possible explanation would be there is presence of complicated drug resistance mechanisms in these bacteria, of which efflux pumps are only part of them (Charkhi et al. 2020; Li and Nikaido 2009). On the other hand, interestingly, there was synergistic or additive effects against both susceptible and resistant strains of P. aeruginosa, with FIC values ranging from 0.3125 to 1.000, when PAβN was used in combination with Cu(SBFH)2. For P. aeruginosa susceptible strain, there was more than fourfold decrease in MIC value of Cu(SBFH)2 when 64 µM of PAβN was added. P. aeruginosa resistant strain showed similar trend as the susceptible strain but with better activities. Surprisingly, 16-fold decrease in MIC value of Cu(SBFH)2 was observed in the presence of 32 µM of PAβN, when treated against P. aeruginosa resistant strain. The ability of PAβN to inhibit efflux pumps in P. aeruginosa was evaluated and supported via a study by Lamers et al. (2013). There was a fourfold and 16-fold reduction in MICs of erythromycin in the presence of 51.76 µM (25 µg/mL) and 103.52 µM (50 µg/mL) of PAβN respectively, when treated against wild type PAO1, a strain of P. aeruginosa expressing wild type levels of MexAB-OprM. P. aeruginosa has 12 resistance nodulation division (RND) type efflux systems, which MexAB-OprM being the most characterised (Ferrer-Espada et al. 2019). Another study by Lomovskaya et al. (2001) recorded synergistic effect of levofloxacin and PAβN against wild-type P. aeruginosa. Similarly, a research by Mbaveng et al. (2016) reported reduction in MIC of hydrazinoselenazoles 16 from 498.05 µM (128 µg/mL) to 31.13 µM (8 µg/mL) in the presence of 41.41 µM (20 µg/mL) of PAβN when tested against P. aeruginosa. It should be noted that PAβN used in our experiment had no effect on the survival of P. aeruginosa at concentrations up to 128 uM, showing that PAβN in our experimental setting did not have bacterial killing effect by itself. Clearly, the enhancement of antibacterial activities was due to the efflux pump inhibition by PAβN or in other words, efflux pump was likely one of the mechanisms that modulates the susceptibility of P. aeruginosa in our experiment (Kuete et al. 2010). Efflux pumps will reduce the intracellular drug concentration and consequently their antimicrobial activities against bacteria (Lacmata et al. 2012; Vergalli et al. 2020). Hence, by blocking the efflux activity via PAβN, our compounds can restore their normal intracellular concentration to kill the bacteria (Nikaido and Pagès 2012).
The antimicrobial activity of copper complex in combination with other drugs in determining their drug interaction had also been reported by other studies. In another recent report from our group, Cu(SBFH)2 demonstrated additive and synergistic interaction when combined with oxacillin with FIC value 0.63 and 0.19–0.38 respectively when tested against S. aureus. It is also worth noting that when Cu(SBFH)2 was tested with the oxacillin against MRSA, additive effect was also shown, with FIC value ranging from 0.31 to 0.56 (Chung et al. 2021). The synergism between novel copper complex and other antibiotics have also been extensively researched and assessed, with one such study by Glišić et al. (2016), supplements this rationale in combining novel Cu(II) complex with piperacillin or ceftazidime in testing against P. aeruginosa. In combining the Cu(II) complex with piperacillin or ceftazidime, there was a twofold and fourfold reduction of MIC values when the concentration of the Cu(II) complex was at 1645 µM (500 µg/mL) and 3290 µM (1000 µg/mL) respectively (Glišić et al. 2016). Leite et al. (2019) has also reported novel Cu(II) complexes of naphthyl derived 3-hydroxy-4-pyridinone chelators (naph1pp) which are found to display antimicrobial activity against both MDR Gram-positive and Gram-negative bacteria. At concentration of 6.88 µM (4 µg/mL) of Cu(naph1pp)2 complex with ciprofloxacin, it shows both synergistic interactions against resistant strain of E. faecalis with FIC value of 0.38. Concentrations of Cu(naph1pp)2 complex ranging from 13.75 to 110 µM (8–64 µg/mL) results in additive effect against both resistant strains of E. faecalis and S. aureus with FIC value ranging from 0.63 to 1.00 (Leite et al. 2019). The antimicrobial activity was hypothesised to be attributed to the redox cycling that happens between Cu(II) and Cu(I) ion which results in the formation of highly reactive hydroxyl radicals that is involve with the denaturation of enzymes, proteins as well as other biomolecules in S. aureus (Chung et al. 2021). Other possible explanation for the good antimicrobial activity of copper complex can be explained via Tweedy's Chelation theory, which suggest that the chelation of the Cu(II) ion reduces its polarity due to the overlap of the ligand orbital and partial sharing of the positive charge of the Cu(II) ion with the donor groups and possible π-electron delocalization over the whole chelate ring, which enhances the lipophilicity of the complexes and the penetration of the complexes into the lipid membranes (Leite et al. 2019). The cytotoxicity investigation of Cu(SB4CB)2 and Cu(SBFH)2 alone against normal cell lines MRC5 (normal lung tissue) using the MTT assay has been recently reported in another related work by Chung et al. (2021). The IC50 values for Cu(SB4CB)2 and Cu(SBFH)2 were found to be 62 µM (45 µg/mL) and 69 µM (52 µg/mL), respectively indicated that Cu(SBFH)2 and its additive and synergetic combinations were toxic against selected bacteria at concentrations lower than the IC50 concentration for MRC5 cells.
Molecular docking studies
The binding interactions of ligand Cu(SBFH)2 in binding pocket of S. aureus NorA efflux pump, E. coli AcrB efflux pump, P. aeruginosa MexB efflux pump and A. baumannii AdeB efflux pump were analysed and depicted in Figs. 2A, B, 3A, B, 4A, B and 5A, B, respectively. In the binding pocket of S. aureus NorA efflux pump, the phenyl rings interacted with the adjacent hydrophobic residues, such as Ile256, Met308, Ile309, Pro311, Met338 and Pro344. Polar residues including Asn137, Asn315 and Thr314 formed polar interactions with the polar nitrogen atoms of the ligand. For the A. baumannii AdeB efflux pump, the ligand was docked at the distal binding site of AdeB. The phenyl rings of the ligand formed π-π stacking with adjacent aromatic ring side chain of Phe168 and Trp568. Additionally, the phenyl rings were also surrounded by hydrophobic residues in vicinity, such as Phe136, Met570, Ile607, Phe612, Phe623, Pro660, Pro661 and Ile663. As in the E. coli AcrB efflux pump, the ligand engaged in a variety of interactions with nearby residues. The carboxyl group of one of the SBFH moieties formed two hydrogen bonding with backbone of Ile38 and Ala465; another hydrogen bonding was formed between the adjacent hydroxyl group and Arg468. A π-cation interaction was also observed between the phenyl ring of the SBFH moiety and Arg468. Other hydrophobic residues like Ala384, Ala385, Ala386, Ala457 and Ile472 were found to interact with the phenyl rings of ligand. Nevertheless, the binding interactions of Cu(SBFH)2 in S. aureus NorA, A. baumannii AdeB and E. coli AcrB efflux pumps do not seem to affect its antibacterial activity as according to the finding from combination study of these bacteria.
A Binding interaction of Cu(SBFH)2 with adjacent residues in E. coli AcrB efflux pump through SP docking. Green dotted line: π-cation interaction; yellow dotted line: hydrogen bonding. B 2D ligand interaction diagram of Cu(SBFH)2 with adjacent residues. Green curvy ribbon: hydrophobic interaction; blue curvy ribbon: polar interaction; purple line: hydrogen bonding; red line: π-cation interaction
In the binding pocket of P. aeruginosa MexB efflux pump, there were mainly hydrophobic interactions between the phenyl rings of the ligand and hydrophobic residues in proximity, such as Phe386, Phe388, Phe458, Ile472, Val475, Ala479 and Leu480. Interestingly, the combination study showed that the presence of efflux pump inhibitor has resulted in the additive and synergistic effect towards antibacterial activity of Cu(SBFH)2. This finding suggests that the Cu(SBFH)2 could be a substrate of the MexB efflux pump and the interaction between the ligand and efflux pump may ultimately affect its antibacterial activity. Nonetheless, this inference has to be further confirmed by other specific efflux pump assay.
Conclusion
To conclude this work, four compounds have been successfully evaluated for their potential antibacterial properties against susceptible and resistant bacteria. New compounds SBFH and Cu(SBFH)2 have been successfully characterised using elemental analysis, FTIR, UV–Vis and NMR. The result of MIC tests showed better antibacterial properties against S. aureus strains for Cu(SB4CB)2 and Cu(SBFH)2 as compared to their ligands alone. Combination of Cu(SBFH)2 with phenylalanine-arginine β-naphthylamide (PAβN) resulted in enhanced antibacterial activity against P. aeruginosa ATCC 27853 and P. aeruginosa ATCC BAA-2108 with additive or synergistic effects. In addition to that, combination of Cu(SBFH)2 with polymyxin B sulfate (POLY) showed enhanced antibacterial activity against A. baumannii ATCC 19606, A. baumannii ATCC BAA-1797, P. aeruginosa ATCC 27853 and P. aeruginosa ATCC BAA-2108 with additive effects. Interaction of Cu(SBFH)2 with efflux pumps was studied using the computer modelling software Maestro Schrödinger to understand the binding pockets inside bacteria efflux pumps and the binding interactions of ligand in the binding site. The exact mechanism of action for the compound and its interaction with efflux pumps will be further explored in future studies. Taking into consideration the seriousness of MDR, the combination strategy highlighted in this work involving novel Cu(II) Schiff base complex with POLY/PAβN is potentially useful for the development of new therapeutic agent and strategy to treat bacterial infections.
References
Adly OMI, El-shafiy HF (2019) New metal complexes derived from S-benzyldithiocarbazate (SBDTC) and chromone-3-carboxaldehyde: synthesis, characterization, antimicrobial, antitumor activity and DFT calculations. J Coord Chem 72(2):1–16. https://doi.org/10.1080/00958972.2018.1564912
Akbar Ali M, Huq mirza A, Ai Fong G (2004) Synthesis, characterization and X-ray crystal structures of the bis-ligand zinc(II) and cadmium(II) complexes of the methylpyruvate Schiff base of S-methyldithiocarbazate. Transit Met Chem 29:613–619. https://doi.org/10.1007/s11243-004-2511-7
Alav I, Sutton JM, Rahman KM (2018) Role of bacterial efflux pumps in biofilm formation. J Antimicrob Chemother 73(8):2003–2020. https://doi.org/10.1093/jac/dky042
Aparna V, Dineshkumar K, Mohanalakshmi N, Velmurugan D, Hopper W (2014) Identification of natural compound inhibitors for multidrug efflux pumps of Escherichia coli and Pseudomonas aeruginosa using in silico high-throughput virtual screening and in vitro validation. PLoS ONE 9(7):e101840. https://doi.org/10.1371/journal.pone.0101840
Auda IG, Ali Salman IM, Odah JG (2020) Efflux pumps of Gram-negative bacteria in brief. Gene Reports 20:100666. https://doi.org/10.1016/j.genrep.2020.100666
Bolla JM, Alibert Franco S, Handzlink J (2011) Strategies for bypassing the membrane barrier in multidrug resistant gram-negative bacteria. Febsletters 585(11):1682–2690. https://doi.org/10.1016/j.febslet.2011.04.054
Breijyeh Z, Jubeh B, Karaman R (2020) Resistance of Gram-negative bacteria to current antibacterial agents and approaches to resolve it. Molecules 25(6):1340. https://doi.org/10.3390/molecules25061340
Brown OC, Torres JB, Holt KB, Blower PJ, Went MJ (2017) Copper complexes with dissymmetrically substituted bis(thiosemicarbazone) ligands as a basis for PET radiopharmaceuticals: control of redox potential and lipophilicity. Dalton Trans 46(42):14612–14630. https://doi.org/10.1039/C7DT02008B
Charkhi P, Haghshenas MR, Mirzaei B, Davoodi L, Bazgir ZN, Goli HR (2020) Comparison of the effect of phenylalanine arginine beta naphthylamide (PAΒN) and curcumin on minimum inhibitory concentration of aminoglycosides on Pseudomonas aeruginosa clinical isolates. J Adv Med Biomed Res 28(127):105–110. https://doi.org/10.30699/jambs.28.127.105
Choo XY, Liddell JR, Huuskonen MT, Grubman A, Moujalled D, Roberts J, Kysenius K, Patten L, Quek H, Oikari LE, Duncan C, James SA, McInnes LE, Hayne DJ, Donnelly PS, Pollari E, Vähätalo S, Lejavová K, Kettunen MI, Malm T, Koistinaho J, White AR, Kanninen KM (2018) CuII (atsm) attenuates neuroinflammation. Front Neurosci 12:668. https://doi.org/10.3389/fnins.2018.00668
Chung PY, Khoo REY, Liew HS, Low ML (2021) Antimicrobial and antibiofilm activities of Cu (II) Schiff base complexes against methicillin-susceptible and resistant Staphylococcus aureus. Ann Clin Microbiol Antimicrob 20(1):67. https://doi.org/10.1186/s12941-021-00473-4
Claudel M, Schwarte JV, Fromm KM (2020) New antimicrobial strategies based on metal complexes. Chemistry 2(4):849–899. https://doi.org/10.3390/chemistry2040056
Crouse KA, Chew K-B, Tarafder MT, Kasbollah A, Ali A, Yamin B, Fun HK (2004) Synthesis, characterization and bio-activity of S-2-picolyldithiocarbazate (S2PDTC), some of its schiff bases and their Ni(II) complexes and X-ray structure of S-2-picolyl-β-N-(2-acetylpyrrole)dithiocarbazate. Polyhedron 23(1):161–168. https://doi.org/10.1016/J.POLY.2003.09.025
Dai C, Wan Y, Sharma G, Shen J, Velkov T, Xiao X (2020) Polymyxins–curcumin combination antimicrobial therapy: safety implications and efficacy for infection treatment. Antioxidants 9(6):506. https://doi.org/10.3390/antiox9060506
Damit NSHH, Hamid MHSA, Rahman NSRHA, Ilias SNHH, Keasberry NA (2021) Synthesis, structural characterisation and antibacterial activities of lead(II) and some transition metal complexes derived from quinoline-2-carboxaldehyde 4-methyl-3-thiosemicarbazone. Inorg Chim Acta 527(14):120557. https://doi.org/10.1016/j.ica.2021.120557
Deng J, Yu P, Zhang Z, Wang J, Cai J, Wu N, Sun H, Liang H, Yang F (2018) Designing anticancer copper (II) complexes by optimizing 2-pyridine-thiosemicarbazone ligands. Eur J Med Chem 158:442–452. https://doi.org/10.1016/j.ejmech.2018.09.020
Dhahagani K, Kesavan MP, Kumar GGV, Ravi L, Rajagopal G, Rajesh J (2018) Crystal structure, optical properties, DFT analysis of new morpholine based Schiff base ligands and their copper(II) complexes: DNA, protein docking analyses, antibacterial study and anticancer evaluation. Mater Sci Eng C 90:119–130. https://doi.org/10.1016/j.msec.2018.04.032
Djoko KY, Goytia MM, Donnelly PS, Schembri MA, Shafer WM, McEwan AG (2015) Copper(II)-bis(thiosemicarbazonato) complexes as antibacterial agents: insights into their mode of action and potential as therapeutics. Antimicrob Agents Chemother 59(10):6444–6453. https://doi.org/10.1128/aac.01289-15
Fazly Bazzaz BS, Seyedi S, Hoseini Goki N, Khameneh B (2021) Human antimicrobial peptides: spectrum, mode of action and resistance mechanisms. Int J Pept Res Ther 27:801–816. https://doi.org/10.1007/s10989-020-10127-2
Ferrer-Espada R, Shahrour H, Pitts B, Stewart PS, Sánchez-Gómez S, Martínez-de-Tejada G (2019) A permeability-increasing drug synergizes with bacterial efflux pump inhibitors and restores susceptibility to antibiotics in multi-drug resistant Pseudomonas aeruginosa strains. Sci Rep. https://doi.org/10.1038/s41598-019-39659-4
Figueiredo J, Serrano JL, Soares M, Ferreira S, Domingues FC, Almeida P, Silvestre S (2019) 5-Hydrazinylethylidenepyrimidines effective against multidrug-resistant Acinetobacter baumannii: synthesis and in vitro biological evaluation of antibacterial, radical scavenging and cytotoxic activities. Eur J Pharm Sci 137:104964. https://doi.org/10.1016/j.ejps.2019.104964
Foo JB, Ng LS, Lim JH, Tan PX, Lor YZ, Loo JSE, Low ML, Chan LC, Beh CY, Leong SW, Yazan LS, Tor YS, How CW (2019) Induction of cell cycle arrest and apoptosis by copper complex Cu (SBCM) 2 towards oestrogen-receptor positive MCF-7 breast cancer cells. RSC Adv 9(32):18359–18370. https://doi.org/10.1039/C9RA03130H
Glišić BĐ, Aleksic I, Comba P, Wadepohl H, Ilic-Tomic T, Nikodinovic-Runic J, Djuran MI (2016) Copper (II) complexes with aromatic nitrogen-containing heterocycles as effective inhibitors of quorum sensing activity in Pseudomonas aeruginosa. RSC Adv 6(89):86695–86709. https://doi.org/10.1039/C6RA19902J
Grace JL, Huang JX, Cheah SE, Truong NP, Cooper MA, Li J, Davis TP, Quinn JF, Velkov T, Whittaker MR (2016) Antibacterial low molecular weight cationic polymers: dissecting the contribution of hydrophobicity, chain length and charge to activity. RSC Adv 6(19):15469–15477. https://doi.org/10.1039/C5RA24361K
Guelfi KC, Tognim MCB, Cardoso CL, Gales AC, Carrara-Marrone FE, Garcia LB (2008) In vitro evaluation of the antimicrobial activity of meropenem in combination with polymyxin B and gatifloxacin against Pseudomonas aeruginosa and Acinetobacter baumannii. J Chemother 20(2):180–185. https://doi.org/10.1179/joc.2008.20.2.180
Hejchman E, Kruszewska H, Maciejewska D, Sowirka-Taciak B, Tomczyk M, Sztokfisz-Ignasiak A, Jankowski J, Młynarczuk-Biały I (2019) Design, synthesis, and biological activity of Schiff bases bearing salicyl and 7-hydroxycoumarinyl moieties. Monatsh Chem 150(17):255–266. https://doi.org/10.1007/s00706-018-2325-5
How FN-F, Crouse KA, Tahir MIM, Tarafder MTH, Cowley AR (2008) Synthesis, characterization and biological studies of S-benzyl-β-N-(benzoyl) dithiocarbazate and its metal complexes. Polyhedron 27(15):3325–3329. https://doi.org/10.1016/j.poly.2008.07.022
Hsieh MH, Yu CM, Yu VL, Chow JW (1993) Synergy assessed by checkerboard a critical analysis. Diagn Microbiol Infect Dis 16(4):343–349. https://doi.org/10.1016/0732-8893(93)90087-N
Jamshidi S, Sutton JM, Rahman KM (2017) Computational study reveals the molecular mechanism of the interaction between the efflux inhibitor PAβN and the AdeB transporter from Acinetobacter baumannii. ACS Omega 2(6):3002–3016. https://doi.org/10.1021/acsomega.7b00131
Kremer E, Facchin G, Estévez E, Alborés P, Baran EJ, Ellena J, Torre MH (2006) Copper complexes with heterocyclic sulfonamides: synthesis, spectroscopic characterization, microbiological and SOD-like activities: crystal structure of [Cu(sulfisoxazole)2(H2O)4]·2H2O. J Inorg Biochem 100(7):1167–1175. https://doi.org/10.1016/j.jinorgbio.2006.01.042
Kuete V, Ngameni B, Tangmouo JG, Bolla JM, Alibert-Franco S, Ngadjui BT, Pagès JM (2010) Efflux pumps are involved in the defense of Gram-negative bacteria against the natural products isobavachalcone and diospyrone. Antimicrob Agents Chemother 54(5):1749–1752. https://doi.org/10.1128/AAC.01533-09
Lacmata ST, Kuete V, Dzoyem JP, Tankeo SB, Teke GN, Kuiate JR, Pages JM (2012) Antibacterial activities of selected Cameroonian plants and their synergistic effects with antibiotics against bacteria expressing MDR phenotypes. Evid Based Complem Alternat Med 2012:623723. https://doi.org/10.1155/2012/623723
Lamers RP, Cavallari JF, Burrows LL (2013) The efflux inhibitor phenylalanine-arginine beta-naphthylamide (PAβN) permeabilizes the outer membrane of gram-negative bacteria. PLoS ONE 8(3):e60666. https://doi.org/10.1371/journal.pone.0060666
Latheef L, Kurup MRP (2008) Spectral and structural studies of copper(II) complexes of thiosemicarbazones derived from salicylaldehyde and containing ring incorporated at N(4)-position. Spectrochim Acta Part A 70(1):86–93. https://doi.org/10.1016/j.saa.2007.07.015
Leite A, Bessa LJ, Silva AMG, Gameiro P, de Castro B, Rangel M (2019) Antibacterial activity of naphthyl derived bis-(3-hydroxy-4-pyridinonate) copper (II) complexes against multidrug-resistant bacteria. J Inorg Biochem 197:110704. https://doi.org/10.1016/j.jinorgbio.2019.110704
Lenhard JR, Nation RL, Tsuji BT (2016) Synergistic combinations of polymyxins. Int J Antimicrob Agents 48(6):607–613. https://doi.org/10.1016/j.ijantimicag.2016.09.014
Li XZ, Nikaido H (2009) Efflux-mediated drug resistance in bacteria: an update. Drugs 69(12):1555–1623. https://doi.org/10.2165/11317030-000000000-00000
Lima FC, Silva TS, Martins CHG, Gatto CC (2018) Synthesis, crystal structures and antimicrobial activity of dimeric copper (II) complexes with 2-hydroxyphenyl-ethylidene-dithiocarbazates. Inorg Chim Acta. https://doi.org/10.1016/j.ica.2018.08.032
Lomovskaya O, Warren MS, Lee A, Galazzo J, Fronko R, Lee M, Blais J, Cho D, Chamberland S, Renau T, Leger R, Hecker S, Watkins W, Hoshino K, Ishida H, Lee VJ (2001) Identification and characterization of inhibitors of multidrug resistance efflux pumps in Pseudomonas aeruginosa: novel agents for combination therapy. Antimicrob Agents Chemother 45(1):105–116. https://doi.org/10.1128/AAC.45.1.105-116.2001
Low ML, Maigre L, Tahir MIM, Tiekink ERT, Dorlet P, Guillot R, Ravoof TB, Rosli R, Pagès J-M, Policar C, Delsuc N, Crouse KA (2016) New insight into the structural, electrochemical and biological aspects of macroacyclic Cu(II) complexes derived from S-substituted dithiocarbazate schiff bases. Eur J Med Chem 14(120):1–12. https://doi.org/10.1016/j.ejmech.2016.04.027
Low ML, Maigre L, Dorlet P, Guillot R, Pagès J-M, Crouse KA, Policar C, Delsuc N (2014) Conjugation of a new series of dithiocarbazate Schiff base copper(II) complexes with vectors selected to enhance antibacterial activity. Bioconjug Chem 25(12):2269–2284. https://doi.org/10.1021/bc5004907
Manikal VM, Landman D, Saurina G, Oydna E, Lal H, Quale J (2000) Endemic carbapenem-resistant Acinetobacter species in Brooklyn, New York: citywide prevalence, interinstitutional spread, and relation to antibiotic usage. Clin Infect Dis 31(1):101–106. https://doi.org/10.1086/313902
Mbaveng AT, Ignat AG, Ngameni B, Zaharia V, Ngadjui BT, Kuete V (2016) In vitro antibacterial activities of p-toluenesulfonyl-hydrazinothiazoles and hydrazinoselenazoles against multi-drug resistant Gram-negative phenotypes. BMC Pharmacol Toxicol. https://doi.org/10.1186/s40360-016-0046-0
Meer Ahmad AM (2019) Antibiotic resistance in Malaysia, and its public health implications. J Drug Deliv Therap 9(2):534–541. https://doi.org/10.22270/jddt.v9i2.2427
Morgan CE, Glaza P, Leus I, Trinh A, Su C-C, Cui M, Zgurskaya HI, Yu EW (2021) Cyroelectron microscopy structures of AdeB illuminate mechanisms of simultaneous binding and exporting of substrates. J Clin Microbiol. https://doi.org/10.1128/mBio.03690-20
Nair MS, Arish D, Joseyphus RS (2012) Synthesis, characterization, antifungal, antibacterial and DNA cleavage studies of some heterocyclic Schiff base metal complexes. J Saudi Chem Soc 16(1):83–88. https://doi.org/10.1016/j.jscs.2010.11.002
Nang SC, Azad MAK, Velkov T, Zhou Q, Li J (2021) Rescuing the last-line polymyxins: achievements and challenges. Pharmacol Rev 73(2):679–728. https://doi.org/10.1124/pharmrev.120.000020
Nikaido H, Pagès JM (2012) Broad-specificity efflux pumps and their role in multidrug resistance of Gram-negative bacteria. FEMS Microbiol Rev 36(2):340–363. https://doi.org/10.1111/j.1574-6976.2011.00290.x
Oliveira AA, Oliveira APA, Franco LL, Ferencs MO, Ferreira JFG, Bachi SMPS, Speziali NL, Farias LM, Magalhães PP, Beraldo H (2018) 5-Nitroimidazole-derived Schiff bases and their copper (II) complexes exhibit potent antimicrobial activity against pathogenic anaerobic bacteria. Biometals 31(4):571–584. https://doi.org/10.1007/s10534-018-0106-6
Rampioni G, Pillai CR, Longo F, Bondì R, Baldelli V, Messina M, Imperi F, Visca P, Leoni L (2017) Effect of efflux pump inhibition on Pseudomonas aeruginosa transcriptome and virulence. Sci Rep. https://doi.org/10.1038/s41598-017-11892-9
Rebolledo AP, Vieites M, Gambino D, Piro OE, Castellano EE, Zani CL, Souza-Fagundes EM, Teixeira LR, Batista AA, Beraldo H (2005) Palladium(II) complexes of 2-benzoylpyridine-derived thiosemicarbazones: spectral characterization, structural studies and cytotoxic activity. J Inorg Biochem 99(3):698–706. https://doi.org/10.1016/j.jinorgbio.2004.11.022
Said MA, Al-Harbi WS, Shanmugam M, Aljohani FS, Bouqellah NA, Al-Kaff NS (2020) Synthesis, XRD, HAS, in silico molecular docking studies and biological assessment of novel Schiff base compounds as anti-cancer and antimicrobial agents. J Taibah Univ Sci 14(1):1590–1603. https://doi.org/10.1080/16583655.2020.1849492
Samal S, Mishra SB, Patra SK, Rath A, Dash A, Nayak B, Mohanty D (2021) Polymyxin monotherapy vs. combination therapy for the treatment of multidrug-resistant infections: a systematic review and meta-analysis. Indian J Crit Care Med 25(2):199–206. https://doi.org/10.5005/jp-journals-10071-23720
Singh A, Barman P (2021) Recent advances in Schiff base ruthenium metal complexes: synthesis and applications. Top Curr Chem 379(4):29. https://doi.org/10.1007/s41061-021-00342-w
Singh NK, Kumbhar AA, Pokharel YR, Yadav PN (2020) Anticancer potency of copper (II) complexes of thiosemicarbazones. J Inorg Biochem 210:111134. https://doi.org/10.1016/j.jinorgbio.2020.111134
Singh S, Kalia N, Joshi P, Kumar A, Sharma P, Kumar A, Bharate S, Inshad K (2017) Boeravinone B, A novel dual inhibitor of NorA Bacterial efflux pump of Staphylococcus aureus and human P-glycoprotein, reduces the biofilm formation and intracellular invasion of bacteria. Front Microbiol 8:1868. https://doi.org/10.3389/fmicb.2017.01868
Soto SM (2013) Role of efflux pumps in the antibiotic resistance of bacteria embedded in a biofilm. Virulence 4(3):223–229. https://doi.org/10.4161/viru.23724
Tidwell TT (2008) Hugo (Ugo) Schiff, Schiff bases, and a century of beta-lactam synthesis. Angew Chem Int Ed 47(6):1016–1020. https://doi.org/10.1002/anie.200702965
Vergalli J, Atzori A, Pajovic J, Dumont E, Malloci G, Masi M, Vargiu AV, Winterhalter M, Réfrégiers M, Ruggerone P, Pagès JM (2020) The challenge of intracellular antibiotic accumulation, a function of fluoroquinolone influx versus bacterial efflux. Commun Biol. https://doi.org/10.1038/s42003-020-0929-x
Walsh TJ, Peter J, McGough DA, Fothergill AW, Rinaldi MG, Pizzo PA (1995) Activities of amphotericin B and antifungal azoles alone and in combination against Pseudallescheria boydii. Antimicrob Agents Chemother 39(6):1361–1364. https://doi.org/10.1128/AAC.39.6.1361
World Health Organisation (2014) Antimicrobial resistance: global report on surveillance. https://apps.who.int/iris/handle/10665/112642
Yoon J, Urban C, Terzian C, Mariano N, Rahal JJ (2004) In vitro double and triple synergistic activities of polymyxin B, imipenem, and rifampin against multidrug-resistant Acinetobacter baumannii. Antimicrob Agents Chemother 48(3):753–757. https://doi.org/10.1128/AAC.48.3.753-757.2004
Zharkova MS, Orlov DS, Golubeva OY, Chakchir OB, Eliseev IE, Grinchuk TM, Shamova OV (2019) Application of antimicrobial peptides of the innate immune system in combination with conventional antibiotics—a novel way to combat antibiotic resistance? Front Cell Infect Microbiol 9:128. https://doi.org/10.3389/fcimb.2019.00128
Zusman O, Altunin S, Koppel F, Dishon Benattar Y, Gedik H, Paul M (2017) Polymyxin monotherapy or in combination against carbapenem-resistant bacteria: systematic review and meta-analysis. J Antimicrob Chemother 72(1):29–39. https://doi.org/10.1093/jac/dkw377
Acknowledgements
The authors would like to thank the School of Postgraduate Studies and the Institute for Research, Development and Innovation (IRDI) of the International Medical University for the support and funding (Project ID No: MAPC I/2019(2)-IR118) provided.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Data availability
All data generated or analysed during this study are included in this published article and its supplementary information file.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Gan, W.K., Liew, H.S., Pua, L.J.W. et al. Novel Cu(II) Schiff Base Complex Combination with Polymyxin B/Phenylalanine-Arginine β-Naphthylamide Against Various Bacterial Strains. Int J Pept Res Ther 28, 60 (2022). https://doi.org/10.1007/s10989-021-10358-x
Accepted:
Published:
DOI: https://doi.org/10.1007/s10989-021-10358-x