Abstract
In this study a strategy based on using signal sequence and statistically optimized medium was applied for efficient extracellular production of the recombinant form of the therapeutic enzyme l-asparaginase. l-Asparaginase from E. coli is an important enzyme used in the cancer treatment, which has been produced and studied previously by other researchers. Nevertheless, to date no study has investigated the extracellular production of recombinant l-asparaginase in the culture medium optimized using response surface methodology. For this purpose, in this study at first, a complete gene of l-asparaginase II with its own signal sequence from a locally isolated E. coli was cloned and expressed in E. coli BL21 (DE3). Subsequently, the production of the active form of recombinant l-asparaginase was evaluated by asparaginase activity assay in the culture media 4 h after induction with IPTG in 250 mL shake flasks to evaluate the effect of eleven nutrient factors on the extracellular l-asparaginase production. Statistical design of experiments was employed in the screening stage and the most effective nutrients were selected for further optimization using central composite face design. Analysis of the results revealed a mature protein with correct N-terminal amino acids of l-asparaginase II in the culture medium. The highest enzyme activity of 17,386 U/L was resulted in the optimized medium consisting of 7.75 g/L tryptone, 9 g/L yeast extract, 5.25 g/L peptone and 0.6 g/L calcium chloride at shake flask level.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
l-Asparaginase (l-asparagine amidohydrolase, EC 3.5.1.1) is an important therapeutic enzyme used in the cancer treatment especially in acute lymphoblastic leukemia (ALL) (Müller and Boos 1998; Pieters et al. 2011). This enzyme catalyzes the conversion of the amino acid l-asparagine to l-aspartic acid with release of ammonia. The depletion of circulating asparagine after asparaginase administration affects certain neoplastic cells since these cells cannot synthesize sufficient amounts of l-asparagine and hence are dependent on an extracellular source of this amino acid (Broome 1968; Müller and Boos 1998). l-Asparaginase has been found naturally in many animals, plants and a wide variety of microorganisms (Müller and Boos 1998). However, only the enzymes isolated from Erwinia carotovora and E. coli strains have been used in cancer therapy (Duval et al. 2002; Pieters et al. 2011). E. coli contains two distinct isozymes of l-asparaginase; however, only l-asparaginase isozyme II has the properties required for therapeutic purposes (Campbell et al. 1967; Schwartz et al. 1966).
Production of recombinant proteins in the extracellular medium of gram negative bacterium E. coli and optimization of culture medium composition are important strategies to enhance the yield of biological active proteins. E. coli based expression systems are the most popular systems for recombinant protein production (Baneyx 1999; Lee 1996; Ni and Chen 2009). Based on the production strategy, heterologous proteins produced in E. coli may be found in the cytoplasm, periplasm, or extracellular space (Cornelis 2000; Hatti-Kaul and Mattiasson 2003). However, protein secretion into the culture medium offers several advantages in comparison to intracellular production such as simplified detection and purification, facilitated correct disulfide bonds formation, correct N-terminal amino acid residue after signal peptide cleavage, higher stability and solubility (Jonasson et al. 2002; Makrides 1996; Mergulhao et al. 2005). Furthermore, the fermentation medium provides an unlimited space to accumulate the released target protein with a relatively high yield. Despite the mentioned advantages of extracellular production, E. coli is considered as a poor producer of extracellular proteins (Ni and Chen 2009; Sandkvist and Bagdasarian 1996). In order to address this issue, use of novel metabolic engineering strategies along with optimization of culture medium composition can affect not only the cell growth and protein synthesis (Makrides 1996; Sivashanmugam et al. 2009) but also the secretion of recombinant protein into extracellular space (Mergulhao et al. 2005). Statistical design of experiments (DoE) is the method of choice to screen the important parameters for the production and optimization of the fermentation system. The DoE methodology is preferred to conventional methods such as ‘one-factor at a time’ since it can provide a time and cost-effective tool to investigate various parameters simultaneously and their interactions with reduced number of required experiments (Montgomery 2012; Myers et al. 2011). In such approaches, response surface methodology (RSM) is usually applied after the initial screening in order to optimize the process parameters for maximizing the protein production (Papaneophytou and Kontopidis 2014).
In recent years, production of E. coli l-asparaginase II using different sources, expression systems and production strategies was reported (Khushoo et al. 2005; Vidya et al. 2011; Wang et al. 2001). To date, no study has reported a statistically optimized medium for extracellular recombinant production of this enzyme. To achieve a high yield of recombinant l-asparaginase production in extracellular medium, therefore, we cloned an E. coli l-asparaginase II gene including its own signal sequence from a locally isolated strain with high asparaginase activity to construct a recombinant strain of E. coli. Consequently, screening of the important nutrient factors and growth medium optimization studies using design of experiments and response surface methodology were also carried out to enhance the production of the active form of recombinant l-asparaginase by determination of asparaginase activity in the culture medium.
Materials and Methods
Bacterial Strains and Plasmid
Escherichia coli YG 002 was isolated previously during a screening program using an optimized medium as described by Ghasemi et al. (2008). E. coli strain BL21 (DE3) and plasmid pET15b were purchased from Novagen, USA.
Enzymes and Chemicals
Restriction endonucleases, modifying enzymes and isopropyl β-d-1-thiogalactopyranoside (IPTG) were purchased from Fermentas, Lithuania. All other reagents were of analytical grade and obtained from commercial sources.
Cloning of l-Asparaginase II Gene
Escherichia coli YG 002 was grown in Luria–Bertani (LB) medium containing 10 g/L peptone, 5 g/L yeast extract and 10 g/L NaCl. After overnight incubation at 37 °C, the bacterial cells were harvested by centrifugation and genomic DNA was used as a template for PCR amplification of l-asparaginase II gene with the native signal sequence. The PCR was performed using the forward primer 5′-GCGCTCATATGGAGTTTTTCAAAAAGACG-3′ and reverse primer 5′-AGAGGATCCTTAGTACTGATTGAAGATCTGCTG-3′ which contained NdeI and BamHI restriction sites (underlined) at their 5′ end, respectively. The PCR procedure was composed of 30 cycles of 30 s at 94 °C, 30 s at 64 °C, and 90 s at 72 °C with a final extension of 72 °C for 10 min.
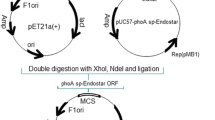
The resultant PCR amplicon was cleaved with NdeI and BamHI, and then inserted into the corresponding sites of expression vector pET-15b (Novagen, USA) which contains a T7 promoter under the control of the lac operator and a cleavable N-terminal His-tag coding sequence. The resulting recombinant plasmid, p15-ASP containing the insert was sequenced on both strands to confirm the identity of the construct. It was then transformed into E. coli BL21 (DE3) cells using a TransformAid™ Bacterial Transformation Kit (Fermentas, Lithuania).
Culture Conditions and Protein Expression
An overnight culture of the recombinant cells was used to inoculate TY medium containing 10 g/L tryptone and 5 g/L yeast extract. The culture was incubated at 37 °C until the optical density at 600 nm (OD600) reached 0.6. Then, 1 ml of this culture was added to all culture media that their compositions were determined based on statistical design of experiment. All cultures were grown at 37 °C for 3 h and then the protein expression was induced with 1 mM IPTG. After IPTG addition, the cells were incubated for 4 h at 34 °C. All cultivation processes were carried out on a rotary shaker at 160 rpm using 250 mL shake flask containing 50 mL culture medium with 100 μg/mL ampicillin.
SDS-PAGE Analysis
Twenty microliters of culture supernatant was collected after induction and analyzed by sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) on a 12 % polyacrylamide gel according to the method of Laemmli (1970). The gel was stained with Coomassie brilliant blue G-250. Unstained protein molecular weight marker (Fermentas, Lithuania) was used for molecular weight estimation.
N-Terminal Amino Acid Sequencing
The extracellular proteins separated in the SDS-PAGE gel (12 %) were transferred onto polyvinylidene fluoride (PVDF) membrane and stained with Coomassie brilliant blue G-250. The band of interest containing the major 34 kDa protein was then excised from the membrane and used for N-terminal sequencing by Edman degradation method.
Experimental Design
The statistical design of experiments using Modde software version 9 (Umetrics, Sweden) was applied in two main steps to optimize the medium composition for maximizing the production of extracellular recombinant l-asparaginase. The aim of the first step was to determine the important medium components with significant effect on asparaginase activity in culture medium. For this purpose, a fractional factorial design was used to screen the effect of eleven variables consisting of glucose, glycerol, sorbitol, sucrose, yeast extract, peptone, tryptone, ammonium chloride, sodium chloride, magnesium sulfate and calcium chloride as carbon, nitrogen and ion-sources in 16 experiments with three replicates in the central point.
The goal of the second step was to achieve an empirical model of optimization process to determine the optimum concentrations of selected medium components. In this stage, RSM with central composite face (CCF) design was employed to optimize the four effective factors (yeast extract, peptone, tryptone and calcium chloride) chosen from screening stage on the extracellular asparaginase activity. A set of 27 runs augmented with 3 replications at central point was performed to investigate the four independent variables at three different levels (normalized in −1, 0 and +1). The range of the variables and their real values used in the CCF design are described in Table 1.
Modde software version 9 (Umetrics, Sweden) was used to develop a model and estimate the optimum value of each component for maximizing l-asparaginase production. The fitness of regression model equation expressed as R2 and the significance of each effect and the interactions was determined by analysis of variance (ANOVA) test at 0.05 probability level.
Enzyme Activity Determination
Asparaginase activity was measured by direct determination of ammonia liberated from l-asparagine using the Nessler’s reaction (Imada et al. 1973). Briefly, 5.0 µL of culture supernatant was mixed with 0.01 M l-asparagine prepared in 0.05 M Tris–HCl, (pH 8.6) in a final volume of 1 mL and the mixture was incubated at 37 °C for 10 min. The reaction was terminated by addition of 0.1 mL of 15 % w/v trichloroacetic acid (TCA). To determine the amount of ammonia liberated, 0.2 mL Nessler’s reagent was added to the mixture and incubated at room temperature for 30 min. Ammonium sulfate solution was used as the standard and the absorbance was read against the blank that received TCA before culture supernatant addition at 480 nm using UV/Vis spectrophotometer (PG Instruments Ltd, USA). One unit of asparaginase is defined as the amount of enzyme required for the release of 1 µmol of ammonia per minute from l-asparagine at 37 °C.
Results and Discussion
Cloning of l-Asparaginase II Gene
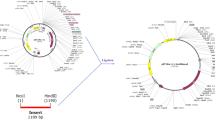
A full length of l-asparaginase II gene from a strain of E. coli was amplified by pfu DNA polymerase and was cloned into pET-15b expression vector. This recombinant plasmid was then used to transform E. coli BL21 (DE3). The nucleotide sequence analysis of the recombinant plasmid p15-ASP revealed a 1047-bp open reading frame (ORF) encoding a mature peptide of 326 amino acids in length fused to its native N-terminal signal peptide of 22 amino acids. The nucleotide sequence of this ORF was used as query in a BLASTN search of the NCBI database (http://blast.ncbi.nlm.nih.gov/Blast.cgi) and showed the highest identity (99 %) with l-asparaginase II sequence from other E. coli strains.
Protein Expression and Molecular Weight Determination
BL21 (DE3) cells were transformed with p15-ASP plasmid and induced with 1 mM of IPTG at 34 °C. The SDS-PAGE analysis of culture supernatant of E. coli BL21 (DE3) harboring p15-ASP after induction confirmed the presence of recombinant protein in the extracellular medium with a major band at about 34 kDa, which was in consistent with the expected molecular weight of the l-asparaginase mature peptide (Fig. 1).
Sequence of N-Terminal Amino Acid
The protein band of interest was transferred to PVDF membrane after SDS-PAGE separation and was examined for its N-terminal amino acid residues using Edman degradation method. The first seven residues of the l-asparaginase protein were determined to be NH2-Leu-Pro-Asn-Ile-Thr-Ile-Leu. This sequence was in accordance with the initial residues of the mature l-asparaginase revealing that the signal peptide had been cleaved during its secretion to the culture medium.
Effects of the Different Medium Components on Asparaginase Activity
Different carbon, nitrogen and ion sources were screened to investigate their effect on extracellular production of l-asparaginase. The results of initial screening experiments using fractional factorial design are summarized in Table 1. Statistical analyses of the selected variables are presented in Table 2 indicating that only 4 variables namely tryptone, yeast extract, peptone and calcium chloride had significant positive influence on l-asparaginase production (P < 0.05). The goodness of the model was assessed with the determination coefficient (R2) = 0.987 which means 98.7 % variability in the results could be explained by the model. Analysis of the results also indicated that in contrast to the three organic nitrogen sources, ammonium chloride significantly decreased the enzyme activity. In the case of ions effects tested, sodium chloride had no significant effect, whereas magnesium sulfate had a negative effect on the enzyme production. Among the studied carbon sources, sucrose, sorbitol and glycerol had no significant effect, while glucose revealed a negative impact on l-asparaginase production, which could be mainly due to the negative effects of glucose on the lac operon regulatory system resulting in reduced protein expression (Grossman et al. 1998; Studier 2005).
The effects of the variables on cell growth were also determined and analyzed at the end of the cultivation process. In accordance to the findings of the effective factors on the l-asparaginase production, the statistical analysis results indicated that all three organic nitrogen sources including yeast extract, tryptone and peptone significantly enhanced the cell growth; however, calcium chloride and other variables did not affect this response significantly.
Since the objective of the screening stage was to determine important factors affecting extracellular l-asparaginase production, tryptone, yeast extract, peptone and calcium chloride were selected for further optimization.
Optimization of l-Asparaginase Production
Determination of optimum concentrations of tryptone (X 1), yeast extract (X 2), peptone (X 3) and calcium chloride (X 4) to enhance the extracellular l-asparaginase production was carried out using the central composite face (CCF) design. The enzyme activity responses exhibited a range of 1340–16,980 U/L in different concentrations of variables in the culture media. The experimental design with the actual levels of the variables and the results of experiments are shown in Table 3. The observed results were fitted with a second order polynomial function for estimation of l-asparaginase production (Y) in terms of coded factors as follows:
where X 1, X 2, X 3 and X 4 are tryptone, yeast extract, peptone and calcium chloride concentrations, respectively.
Regression analysis of the experimental design for linear, quadratic and interaction terms was performed and listed in Table 4. The R2 value of 0.961 and the adjusted R2 value of 0.915 indicated a good fitness of the model which can explain 96.1 % of the variability in the response. The relationship between the observed extracellular asparaginase activity and the expected values by the model is also represented in Fig. 2. The results of ANOVA test (Table 5) with the model F value of 20.9427 and a very low probability value also indicated the statistical significance and adequacy of the quadratic model. As shown in Table 4, all single factor terms (X 1, X 2, X 3 and X 4), X 1 X 2 and X 2 X 3 were the main significant variables affecting l-asparaginase production (P < 0.05) while other interaction and quadratic model terms had no significant effect (P > 0.05) on l-asparaginase production.
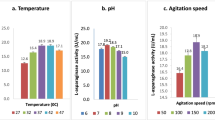
To visualize the effects of the variables on the response and evaluate the interactions and optimum levels of the variables for l-asparaginase production, the response surface plot was generated for each pair of experimental factors, while the other two were held constant at their middle levels. Figure 3 illustrates the surface plots of l-asparaginase production for the interactive effect of all possible pairwise combinations of variables. The analysis of the plots demonstrated that the highest asparaginase activity was achieved when the concentrations of yeast extract and calcium chloride were increased up to 9.0 and 0.6 g/L, respectively. The Interaction between tryptone and peptone demonstrated that the range of 7.0–9 g/L is the optimal tryptone concentration. It was also observed that the increase in peptone concentration up to 8.8 g/L enhanced the asparaginase activity but higher concentrations of peptone had no effect on the enzyme activity.
The highest asparaginase activity predicted by the model was 17,089 U/L with optimum values of tryptone (7.75 g/L), yeast extract (9 g/L), peptone (5.25 g/L) and calcium chloride (0.6 g/L). Consequently, the validation experiments were carried out under the optimized condition and the enzyme activity of 17,386 U/L was achieved which was very close to the value predicted by the model (with only 1.7 % error). This enzyme activity was obtained in the optimized medium with constant induction condition of 1 mM IPTG and 4 h post-induction at 34 °C. Although this induction condition was used as favorable condition in our preliminary study on l-asparaginase production in LB medium and based on the results obtained is adequate enough to get a valid model for the medium optimization, but it is interesting and valuable to evaluate the effect of different IPTG concentrations, post-induction time and temperature using DoE and RSM in the future studies to enhance the production of l-asparaginase in the optimized medium.
Conclusion
In this study we successfully used l-asparaginase II gene from the isolated E. coli with high asparaginase activity to produce a recombinant form of this enzyme at extracellular level. The expression system used in this investigation led to efficient production and release of the recombinant l-asparaginase as the majority of the total secreted proteins. Furthermore, the correct N-terminal amino acid residues and the product activity indicated the important role of l-asparaginase II signal peptide in this system. The results of this study also demonstrated the effective nutrients and their combinations on the recombinant l-asparaginase production with a validated model to determine the optimum conditions.
References
Baneyx F (1999) Recombinant protein expression in Escherichia coli. Curr Opin Biotechnol 10:411–421
Broome JD (1968) Studies on the mechanism of tumor inhibition by L-asparaginase. Effects of the enzyme on asparagine levels in the blood, normal tissues, and 6C3HED lymphomas of mice: differences in asparagine formation and utilization in asparaginase-sensitive and -resistant lymphoma cells. J Exp Med 127:1055–1072
Campbell HA, Mashburn LT, Boyse EA, Old LJ (1967) Two L-asparaginases from Escherichia coli B. Their separation, purification, and antitumor activity. Biochemistry 6:721–730. doi:10.1021/bi00855a011
Cornelis P (2000) Expressing genes in different Escherichia coli compartments. Curr Opin Biotechnol 11:450–454
Duval M et al (2002) Comparison of Escherichia coli-asparaginase with Erwinia-asparaginase in the treatment of childhood lymphoid malignancies: results of a randomized European Organisation for Research and Treatment of Cancer-Children’s Leukemia Group phase 3 trial. Blood 99:2734–2739
Ghasemi Y et al (2008) An optimized medium for screening of L-asparaginase production by Escherichia coli. Am J Biochem Biotechnol 4:422–424. doi:10.3844/ajbbsp.2008.422.424
Grossman TH, Kawasaki ES, Punreddy SR, Osburne MS (1998) Spontaneous cAMP-dependent derepression of gene expression in stationary phase plays a role in recombinant expression instability. Gene 209:95–103
Hatti-Kaul R, Mattiasson B (2003) Isolation and purification of proteins. Marcel Dekker, New York
Imada A, Igarasi S, Nakahama K, Isono M (1973) Asparaginase and glutaminase activities of micro-organisms. J Gen Microbiol 76:85–99
Jonasson P, Liljeqvist S, Nygren PA, Stahl S (2002) Genetic design for facilitated production and recovery of recombinant proteins in Escherichia coli. Biotechnol Appl Biochem 35:91–105
Khushoo A, Pal Y, Mukherjee KJ (2005) Optimization of extracellular production of recombinant asparaginase in Escherichia coli in shake-flask and bioreactor. Appl Microbiol Biotechnol 68:189–197. doi:10.1007/s00253-004-1867-0
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227:680–685
Lee SY (1996) High cell-density culture of Escherichia coli. Trends Biotechnol 14:98–105. doi:10.1016/0167-7799(96)80930-9
Makrides SC (1996) Strategies for achieving high-level expression of genes in Escherichia coli. Microbiol Rev 60:512–538
Mergulhao FJ, Summers DK, Monteiro GA (2005) Recombinant protein secretion in Escherichia coli. Biotechnol Adv 23:177–202. doi:10.1016/j.biotechadv.2004.11.003
Montgomery DC (2012) Design and analysis of experiments, 8th edn. Wiley, Hoboken
Müller HJ, Boos J (1998) Use of L-asparaginase in childhood ALL. Crit Rev Oncol Hematol 28:97–113
Myers RH, Montgomery DC, Anderson-Cook CM (2011) Response surface methodology: process and product optimization using designed experiments, 3rd edn. Wiley, Hoboken
Ni Y, Chen R (2009) Extracellular recombinant protein production from Escherichia coli. Biotechnol Lett 31:1661–1670. doi:10.1007/s10529-009-0077-3
Papaneophytou CP, Kontopidis G (2014) Statistical approaches to maximize recombinant protein expression in Escherichia coli: a general review. Protein Expr Purif 94:22–32. doi:10.1016/j.pep.2013.10.016
Pieters R et al (2011) L-asparaginase treatment in acute lymphoblastic leukemia: a focus on Erwinia asparaginase. Cancer 117:238–249. doi:10.1002/cncr.25489
Sandkvist M, Bagdasarian M (1996) Secretion of recombinant proteins by Gram-negative bacteria. Curr Opin Biotechnol 7:505–511. doi:10.1016/S0958-1669(96)80053-X
Schwartz JH, Reeves JY, Broome JD (1966) Two L-asparaginases from E. coli and their action against tumors. Proc Natl Acad Sci USA 56:1516–1519
Sivashanmugam A, Murray V, Cui C, Zhang Y, Wang J, Li Q (2009) Practical protocols for production of very high yields of recombinant proteins using Escherichia coli. Protein Sci 18:936–948. doi:10.1002/pro.102
Studier FW (2005) Protein production by auto-induction in high density shaking cultures. Protein Expr Purif 41:207–234
Vidya J, Vasudevan UM, Soccol CR, Pandey A (2011) Cloning, functional expression and characterization of L-asparaginase II from E. coli MTCC 739. Food Technol Biotechnol 49:286–290
Wang Y, Qian S, Meng G, Zhang S (2001) Cloning and expression of L-asparaginase gene in Escherichia coli. Appl Biochem Biotechnol 95:93–101. doi:10.1385/ABAB:95:2:093
Acknowledgments
This work was financially supported by the Research Council of Shiraz University of Medical Sciences, Shiraz, Iran.
Conflict of interest
The authors declare that they have no conflict of interest.
Human and Animal Rights and Informed Consent
This article does not contain any studies with human participants or animals performed by any of the authors.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Ghoshoon, M.B., Berenjian, A., Hemmati, S. et al. Extracellular Production of Recombinant l-Asparaginase II in Escherichia coli: Medium Optimization Using Response Surface Methodology. Int J Pept Res Ther 21, 487–495 (2015). https://doi.org/10.1007/s10989-015-9476-6
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10989-015-9476-6