Abstract
Context
Conversion of forest ecosystems to human-modified landscapes threatens the persistence of forest-specialist species. However, the local and landscape drivers of population abundance and genetic diversity of these species are largely unknown, especially for elusive and critically endangered species, such as the salamander Pseudoeurycea robertsi—a species microendemic to the Nevado de Toluca volcano, Mexico.
Objectives
We assessed the relative influence of local- and landscape-scale habitat amount and habitat spatial configuration on the number of individuals and genetic diversity of P. robertsi. Given its low vagility, we expected stronger responses to local habitat amount than to landscape variables, with habitat configuration showing the weakest effects on all responses.
Methods
Using multiscale and multimodel inference approaches, we tested the relative effect of local habitat amount (fallen logs volume), landscape habitat amount (forest cover) and landscape configuration (forest edge density and forest fragmentation per se) on the number of salamanders and its genetic diversity.
Results
The number of individuals was more strongly related to local and landscape variables than genetic diversity. As predicted, local habitat amount showed stronger positive effects on number of individuals and number of alleles than forest cover. In addition, all response variables increased in landscapes with lower edge density. Fragmentation per se showed weak influence on all responses.
Conclusions
Fallen logs volume is a major driver of this forest-specialist salamander. Yet, edge density also shapes salamander populations, especially the number of individuals. Retaining fallen logs in forests and increasing forest core areas are critical for salamander conservation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Productive activities such as agriculture, cattle ranching, timber extraction, and urbanization have transformed a large proportion of the planet’s land surface (Foley et al. 2005). In fact, deforestation between 2000 and 2012 resulted in the loss of 2.3 million km2 of tree cover (Hansen et al. 2013). These land use changes are threatening the maintenance of global biodiversity (Newbold et al. 2016), and is causing rapid changes in the composition and configuration of terrestrial landscapes (Fahrig et al. 2011). Landscape changes are also typically followed by a number of local disturbances, such as edge effects (i.e., biotic and abiotic changes at habitat edges) and resource exploitation (e.g., hunting, logging) by human populations (Laurance et al. 2002; Tuff et al. 2016; Arroyo-Rodríguez et al. 2017). These local and landscape disturbances are expected to have strong effects on forest-specialist species (Laurance 1991; Kouki 2001; Tuff et al. 2016; Pfeifer et al. 2017), but this topic remains poorly understood, especially for elusive species, such as most amphibians.
Amphibians are among the most threatened vertebrates on Earth (Catenazzi 2015), and their populations are rapidly declining worldwide, mainly due to the loss and degradation of their natural habitats (Stuart et al. 2004; Eigenbrod et al. 2008). Salamanders, like other low-vagility ectotherms, are highly vulnerable to local-scale disturbances, such as the loss of thermally suitable habitat (Nowakowski et al. 2018). Salamanders are important as top-down controls of many invertebrates, and can also be a source of high energy prey for other predators (Davic and Welsh 2004). They can also provide an important indirect regulatory role in the processing of detritus-litter by ingestion of detritivore prey (Davic and Welsh 2004). Although many species are narrowly distributed, salamanders can represent an important proportion of the vertebrate biomass in old-growth forests (Davic and Welsh 2004). In temperate ecosystems, terrestrial salamanders prefer areas with high relative moisture like Abies, Pinus and Pinus-Quercus forests, which provide important microhabitats, such as fallen logs, which are particularly used by the Plethodontidae family for shelter (Sánchez-Jasso et al. 2013). Landscape-scale disturbances can also shape amphibian communities (e.g., Skelly et al. 1999; Lowe and Bolger 2002; Van Buskirk 2005; Russildi et al. 2016), but the relative effect of local versus landscape disturbances on salamanders is not well understood. In fact, although some studies of salamanders assess the association between landscape structure and population genetics (Spear et al. 2005; Wang et al. 2009; Savage et al. 2010; Velo-Antón et al. 2013), they are focused on genetic responses to landscape connectivity, overlooking other potential local and landscape predictors of genetic diversity, such as habitat amount. As genetic diversity can determine the capacity of populations to adapt to and cope with environmental changes (Frankham et al. 2002), filling this gap of information is critically needed to improve conservation strategies.
Mexico is within the five biologically richest countries in the world (Groombridge and Jenkins 2000), but it is suffering rapid and extensive land use changes (Mas et al. 2004), especially for the expansion of agricultural lands (Arredondo-León et al. 2008). Between 1976 and 2000, more than 20,000 km2 of temperate forests were cleared in Mexico (annual deforestation rate = 0.25%; Mas et al. 2004). Temperate forests are distributed in high elevation areas, such as the Trans-Mexican Volcanic Belt, which preserves one of the most species-rich herpetological communities in Mexico, and the most important in terms of endemic amphibian species (Flores-Villela et al. 2010). Unfortunately, it is one of the most disturbed regions of the country due to the expansion of big cities, such as Mexico City, Puebla and Toluca (CONAPO 2010; González-Fernández et al. 2018).
Here, we assessed the relative influence of local habitat amount (i.e., fallen logs volume), landscape habitat amount (Abies forest cover) and habitat spatial configuration (i.e., the spatial arrangement of remaining habitat) on the number of individuals and genetic diversity of Pseudoeurycea robertsi. Habitat configuration was measured with two metrics: the effective number of forest patches (i.e., forest fragmentation independent of forest cover; “fragmentation per se”, sensu Fahrig 2003), and forest edge density (a proxy of the amount of edge-dominated forest). This little-known salamander of the Plethodontidae family is classified as Critically Endangered, and is microendemic to the Nevado de Toluca volcano (SEMARNAT 2010; IUCN SSC Amphibian Specialist Group 2016), located in the Trans-Mexican Volcanic Belt. This species inhabits fallen logs, where it finds refuge and food (Bille 2009). Given its low vagility, we expect stronger responses to local habitat amount than to landscape structure (Miguet et al. 2016). Also, as this is a forest-specialist species (Davic and Welsh 2004), when assessing the effect of landscape-scale patterns, we expect stronger responses to landscape habitat amount than to habitat spatial configuration (Fahrig 2003; Jackson and Fahrig 2016). If forest fragmentation per se is relevant for this species, we could expect either positive or negative effects on the number of salamanders and its genetic diversity (Fahrig et al. 2019). Positive effects are predicted because mean inter-patch isolation distance typically decreases with increasing fragmentation per se (Fahrig 2003, 2017; Fahrig et al. 2019), and thus, it can favor inter-patch dispersal movements and mating, potentially favoring genetic diversity (Cushman et al. 2012). However, fragmentation per se can also increase the extent of edge-affected habitats in the landscape (Fahrig et al. 2019), and this forest-interior specialist can be negatively affected by the abiotic changes (e.g., lower relative humidity and higher temperature) that usually occur at forest edges (Arroyo-Rodríguez et al. 2017). In this sense, more fragmented landscapes with higher edge density may sustain fewer individuals.
Methods
Study area
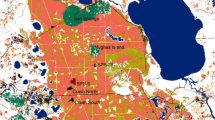
We conducted this study in the Nevado de Toluca volcano (18°59′–19°18′N, 99°40′–99°59′ W), the fourth highest peak in Mexico (De la Torre 1971), located in the Trans-Mexican Volcanic Belt at 22 km of southwest Toluca city (Fig. 1). The region has a temperate semi-cold climate, with precipitations during the summer and an average annual temperature ranging between 5 and 12 °C. Annual precipitation averages 1200–1800 mm (CONABIO 2000). The dominant land cover types of the volcano are old-growth and secondary forest patches of fir (Abies religiosa) and pine (Pinus hartwegii and P. pseudostrobus). There are also broad-leaved forests (oak and alder) in a lower extent, at the eastern part of the volcano, and alpine grasslands at the highest elevations (Fig. 1). There is an important extension of agricultural lands at northeastern part of the volcano, including cattle pastures (Franco-Maass et al. 2006) and human settlements (Toscana-Aparicio and Granados-Ramírez 2015). Between 1972 and 2000, Abies forests experienced an important recovery, whereas Pinus forest cover had decreased by 40% due to timber extraction for commercial purposes (Franco-Maass et al. 2006). The Nevado de Toluca volcano is considered a priority terrestrial region for biodiversity conservation (CONABIO 2000), and was declared a National Park in 1936, although the Mexican Government has recently changed this highly restrictive protection category to a less restrictive one (DOF 2013)—a controversial decision that could lead to further degradation of the last well-preserved Abies forests (Mastretta-Yanes et al. 2014).
Location of the study forest sites in the Nevado de Toluca volcano, located in the Trans-Mexican Volcanic Belt, Mexico. Circles within the main panel represent the 14 landscapes selected in each region, considering the largest buffer size (500 ha). Landscape metrics were measured in 6 different-sized and concentric buffers from the center of each sample site (see example in the top left corner)
Study design and landscape variables
We used a site-landscape approach; i.e., number of individuals and genetic samples were obtained from 14 forest sites (10 ha each, see below), and landscape variables were measured within different-sized and concentric buffers from the geographic center of each sample site (Fahrig 2013; Fig. 1). Sample sites were distributed across the volcano, between 2850 and 3450 m asl. Site selection was arbitrary, but trying to cover the entire gradient of Abies-Pinus forest cover. This means that we first identified the range of variation of forest cover, and arbitrarily (i.e., without using any field information) selected a site from the SIG. We considered buffers of 10, 50, 100, 200, 300, 400 and 500 ha to assess the scale of effect, i.e., the spatial extent that yields the strongest response-landscape relationship (Fahrig 2013; Jackson and Fahrig 2015; Miguet et al. 2016). The smallest buffer is the size of the study site, and was selected because plethodontid salamanders have very small home ranges (Petranka et al. 1993), which can decrease the scale of effect (Miguet et al. 2016). The largest buffer was selected following previous studies about the scale of landscape effect on other anuran species (Vos and Chardon 1998; Eigenbrod et al. 2008). To avoid pseudoreplication problems, we located each study site isolated enough to avoid spatial overlap between landscapes (Eigenbrod et al. 2011). The average distance between sample sites (center to center) was 15.14 km (Fig. 1).
Using ArcGis 10.5 software and satellite images (SPOT 6/7) of very high resolution (1.5 m) for year 2015, we made a supervised classification considering six land cover types: Pinus forest, Abies forest, grasslands, agriculture, urban areas, and water sources. Within each buffer size, we then calculated two metrics of landscape composition: (i) the percentage of forest cover; and (ii) the percentage of Abies cover (from the total forest cover), and two metrics of landscape configuration: (iii) forest fragmentation per se, and (iv) forest edge density; the latter two metrics considering both Abies and Pinus forest. We calculated forest fragmentation with the splitting index (S) proposed by Jaeger (2000): \(S = A_{t}^{2} /\mathop \sum \limits_{i = 1}^{n} A_{i}^{2}\), where At is the area covered by forest in the landscape and Ai is the area of the forest patch i. This index represents the ‘effective number of forest patches’ and is a measure of fragmentation per se because it is independent of the total forest cover in the landscape (Fahrig 2003). Forest edge density was measured as the length of all forest borders divided by the total area of the landscape (expressed as m/ha). Forest cover was computed using ArcGis software, and both configuration variables were calculated with FRAGSTATS software (McGarigal et al. 2012). At the local scale (i.e., within each 10 ha site), we also evaluated (v) fallen logs volume. To do this, we used the formula of the cylinder volume: V = hπr2, where h is the sum of the length of all logs found in the site (the same logs where we surveyed the salamanders), and r is the mean radius of these logs (all fallen logs showed a similar diameter).
Sampling method and tissue collection
Within each 10 ha site, we sampled salamanders via visual encounter surveys (sensu Crump and Scott 1994). As there is no information on the timing of peak of Preudoeurycea robertsi abundance, we carry out a pilot study to identify the annual season with higher number of salamanders in Abies and Pinus forests. In particular, we surveyed three study sites from April to October 2015 and visited each site once per month. We did not record any individual in April and October. In May and September, the number of individuals was twice lower than in June–August. We therefore recorded the number of individuals in all 14 sites from mid-June to mid-August 2016, visiting each site once. During each visit 2 people looked for salamanders under the bark of all fallen logs with a diameter of ≥ 5 cm and length of ≥ 30 cm. We started at 9:00 am and spent two effective hours of search (i.e., excluding stops).
The tissue collection was carried out at the same time, but we re-visited some sites in September to collect additional genetic samples, but in these extra-surveys we did not record the number of individuals. We sampled 2 mm of tail tips of adult salamanders for DNA extraction (license number SGPA/DGVS/05701/16). This methodology is a low-invasive method that does not affect the survival or growth of salamanders (Arntzen et al. 1999; Polich et al. 2013). We released all individuals immediately after data collection in the same place where they were found.
DNA extraction and genetic diversity metrics
Tail tissues were preserved in 90% ethanol and then frozen at − 20 °C until processed. We extracted the DNA according to the commercial kit protocol “GF-1 Tissue DNA” of “Vivantis” mark. We used nine fluorescently labelled microsatellite primers developed for Pseudoeurycea leprosa by Velo-Antón et al. (2009). We multiplexed and run PCR microsatellite products on an ABI Prism3730xl (Applied Biosystems), with Rox-500 as an internal size standard by a commercial laboratory (Roy. J. Carver Biotechnology Center in Illinois University, USA). We obtained allele sizes with PEAKSCANNER 1.0 software (Applied Biosystems) and the fragment lengths with TANDEM 1.08 (Matschiner and Salzburger 2009). To avoid wrong data interpretation, we tested the presence of null alleles and other typing errors in the software MICROCHECKER 2.2.3 (Van Oosterhout et al. 2004). In POPPR 2.4.1 (Kamvar et al. 2014) for the R software (version 3.4.0; R Development Core Team 2017), we made an analysis to create a genotype accumulation curve, useful to determining the minimum number of loci necessary to discriminate between individuals in a population (Kamvar et al. 2014). This function will randomly sample loci without replacement and count the number of observed multilocus genotypes (Kamvar et al. 2014). The genotype accumulation curve showed that the minimum number of loci necessary to discriminate between individuals in this population is eight (Online Appendix 1). Therefore, we can assume that our study has enough microsatellite primers (N = 9).
We used up to 16 samples per site (150 samples in total) but in some sites, we found less than 16 individuals (Online Appendix 2). We finally excluded three forest sites where we found less than four genetic samples (i.e., for genetic diversity analyses we considered 11 sites, with 13.6 ± 4.3 samples per site, mean ± SD). To estimate the accuracy of allelic accounts, we used the coverage estimator recommended by Chao and Jost (2012), which is recommended to estimate the accuracy of species inventories. Sample coverage was high in all sites (> 0.91), indicating that our sampling effort was adequate to estimate diversity metrics within each site (Chao and Jost 2012). However, we calculated the expected values of allele richness based on coverage extrapolations performed with the entropart package (Marcon and Hérault 2015) for R 3.0.1 (R Development Core Team 2017) to avoid any potential bias in our results due to differences in sample coverage (0.91 to 1) among sites (Chao and Jost 2012). We calculated genetic diversity using true diversity measures (Hill 1973): \({\text{Diversity}}\;\Delta \equiv \left( {\mathop \sum \limits_{i = 1}^{k} p_{i}^{q} } \right)^{{1/\left( {1 - q} \right)}}\), where pi is the population frequency of the i-th allele and the exponent q determines the measure’s sensitivity to allele frequencies. When q = 0, the equation gives the allele number (Na), and when q = 2 it gives Kimura and Crow’s (1964) effective number of alleles (Ne) (Jost 2008). We calculated all diversity metrics with the entropart package (Marcon and Hérault 2015) for R (version 3.4.0; R Development Core Team 2017). Na is not sensitive to allele abundances, and thus gives disproportional weight to rare alleles that can appear due to migration. Yet, Ne can be interpreted as the number of dominant alleles in the population, which are the result of the parental inheritance on the site. Thus, both metrics provide complementary information. These true diversity measures are easily interpretable metrics, and meet the ‘‘doubling’’ property (sensu Hill 1973), while heterozygosity and Shannon entropy do not (Jost 2008).
Data analyses
We first assessed the scale of effect of each landscape variable following the protocol proposed in previous studies (Fahrig 2013; Jackson and Fahrig 2015). In brief, we ran generalized linear models between each landscape metric and each response, and assessed the strength of each relationship with the parameter estimate (slope). We carried out such analysis for each of the seven buffer sizes to identify the spatial extent at which the strongest associations between each response variable and each predictor were observable. These optimal scales were used in the statistical analyses that are described below.
We used generalized linear models to assess the effects of local and landscape attributes (measured at the scale of effect) on each response variable. We fixed a Gaussian error distribution for continuous response variables (expected values of Na and Ne) after verifying that model’s residuals followed a Normal distribution (Shapiro–Wilk test). Number of individuals (count-dependent variable) was assessed by fixing a Poisson error distribution. To assess collinearity among predictor variables we estimated their variance inflation factors (VIF) using the car package for R version 3.0.1. A VIF > 4 suggests possible collinearity, and a VIF > 10 indicates severe collinearity (Neter et al. 1996). We found severe collinearity between forest cover (i.e., Abies and Pinus forest) and splitting index in all models. Thus, we excluded forest cover from the models, and included only Abies forest cover, which was not collinear with the splitting index. In fact, Abies forest is probably the main habitat of the species, as in our pilot survey we did not record any individual in Pinus forest. Although elevation varied among sites from 2850 to 3450 m asl, we did not included this variable in the analyses because elevation was not significantly correlated to number of salamanders, allele number or effective number of alleles (p < 0.05, in all cases).
We used a multimodel inference approach to assess the relative effect of each predictor on each response variable (Burnham and Anderson 2002). To obtain model-averaged parameter estimates we used Akaike weights (wi) (Burnham and Anderson 2002). We also used Akaike weights to assess the relative importance of each explanatory variable, by summing the Akaike weights for each model in which each explanatory variable appears. The sum of Akaike weights (∑wi) of each predictor can be interpreted as equivalent to the probability that that predictor is a component of the best model for the data (Burnham and Anderson 2002). Yet, to be more conservative in our selection of important explanatory variables, we considered that a given variable was important for a given response if accomplishing these three conditions simultaneously: (i) the explanatory variable showed a relatively high sum of Akaike weights; (ii) the model-averaged unconditional variance was lower than the model-averaged parameter estimate, and (iii) the goodness-of-fit of the model (i.e., the percentage of deviance explained by the complete model compared with the null model) was relatively high (Russildi et al. 2016; Sánchez-de-Jesús et al. 2016). All models were built using the package glmulti for R version 3.0.1 (Calcagno and Mazancourt 2010).
Results
In total, we recorded 185 individuals and 33 alleles (from a total of 150 genotyped individuals). The number of individuals within each site varied from 0 to 46 (mean ± SD, 13.2 ± 12.7 individuals). The extrapolated Na per site ranged from 23.6 to 31.3 (27.5 ± 2.3 alleles), and the Ne was 15.5 to 21.5 (19.1 ± 1.9 effective alleles) (Online Appendix 3). Fallen logs volume ranged from 1.3 to 19.6 m3 (9.7 ± 4.3 m3).
The percentage of Abies forest cover was slightly higher when measured in the smallest landscape size, and gradually decreased with increasing landscape size (Online Appendix 4). In contrasts, fragmentation per se and forest edge density showed similar values across scales. Regarding the scale of effect of each landscape metric, all response variables were more strongly related to edge density when measured at the largest scale (500 ha); however, the scale of effect of Abies forest cover and fragmentation per se differed among response variables (Online Appendix 5).
Both local and landscape variables showed a relatively high explanatory power, being higher for the number of individuals (75% of explained deviance by the complete model) than for genetic diversity (44–56%) (Fig. 2). As predicted, local habitat amount (fallen logs volume) was relatively more important than landscape habitat amount (Abies forest cover), positively affecting the number of individuals (Fig. 2a) and Na (Fig. 2b) (also see Online Appendix 6). Contrary to our expectations, forest cover generally showed a weaker effect than forest spatial configuration. In particular, the number of individuals responded negatively to forest edge density (∑wi = 1.0; Fig. 2a). Genetic diversity (Na and Ne) also responded negatively to increasing edge density (Fig. 2b and c), but such responses were weaker as the null model showed the lowest AICc (Online Appendix 6). All response variables were also weakly related to forest fragmentation per se, but such relationships were consistently positive.
Local and landscape predictors of the abundance, allele number (Na) and effective number of alleles (Ne) of Pseudoeurycea robertsi in the Nevado de Toluca volcano, Mexico. The sum of Akaike weights (∑wi) is indicated (panels in the left side). Panels in the right side show the values of model-averaged parameter estimates (β) and unconditional variance of information-theory-based model selection and multimodel inference. The sign (±) of parameter estimates represents a positive or negative effect of each predictor on the response variable. The percentage of deviance explained by each complete model is also indicated in each panel as a measure of goodness-of-fit
Discussion
This study assesses the relative effect of local- and landscape-scale habitat amount and habitat spatial configuration on the number of individuals and genetic diversity of Pseudoeurycea robertsi—a critically endangered salamander microendemic to the Nevado de Toluca volcano, Mexico. Our findings suggest that explanatory variables better predict the number of salamanders than its genetic diversity. As expected, local habitat amount (i.e., fallen logs volume) has stronger positive effects on number of salamanders and allele number than landscape-scale habitat amount (i.e., Abies forest cover). However, contrary to our expectations, landscape-scale habitat amount shows a relatively weaker impact on salamanders than forest spatial configuration. In particular, all response variables increased in landscapes with lower edge density (i.e., smaller extent of edge-dominated forest), but the influence of fragmentation per se tended to be weak. As discussed below, these findings have important theoretical and conservation implications.
The fact that all explanatory variables have stronger effects on the number of salamanders than on its genetic diversity is not surprising. The colonization or loss of a given allele in the population can require many generations, and thus, genetic diversity is expected to be regulated by a larger number of generations than abundance (Jackson and Fahrig 2014). There is no information about the generation length (average age at reproduction) for this species, but in other salamanders varies between 4 and 8 years (Flageole and Leclair 1992; Gamble et al. 2009; Plunkett 2009). In this sense, as most of the forested area in the study region was converted to agricultural lands in the second half of the nineteenth century (Mastretta-Yanes et al. 2014), the history of land-use change is probably not long enough to allow the full spectrum of local and landscape effects on genetic diversity be exhibited. This helps to explain why the null model was the best model for the genetic data (Online Appendix 6). Therefore, additional long-term monitoring studies may be necessary before we can draw stronger conclusions about the effect of habitat disturbance on the genetic diversity of this species, including traditional landscape genetic studies (e.g., Coulon et al. 2004; Cushman et al. 2006; Garrido-Garduño and Vázquez-Domínguez 2013) on gen flow between sites.
As predicted, the number of individuals was positively related to fallen logs volume. This species inhabits forest areas with high relative moisture (Sánchez-Jasso et al. 2013), especially fallen logs, where it finds refuge and food (Bille 2009). Thus, as salamanders are mainly found under the bark of fallen logs, fallen logs volume seems to be an adequate proxy of local habitat amount, with stronger effects on salamanders than landscape-scale habitat amount (i.e., Abies forest cover). Such stronger effects can be explained by its relatively low vagility, which can limit the interaction with landscape patterns at larger spatial scales (Jackson and Fahrig 2012; Miguet et al. 2016).
Surprisingly, forest edge density shows a higher predictive power than the percentage of Abies forest cover. We expected stronger responses to Abies forest cover because this is a forest-specialist species (Davic and Welsh 2004), and evidence suggests that landscape-scale habitat amount has stronger effects on biodiversity than habitat configuration (Fahrig 2003, 2017). Yet, the loss of individuals in forest sites surrounded by higher edge density can be related to the loss of ‘core forest areas’ in landscapes with higher edge density (Ewers and Didham 2006). This is particularly relevant when considering relatively large scales (500 ha buffers), probably because core areas are better recorded at this scale (Fig. 1). This salamander species, as other ectotherms, can be negatively impacted by the abiotic changes (e.g., lower relative humidity and higher temperature) that typically occur at forest edges (Arroyo-Rodríguez et al. 2017; Nowakowski et al. 2018). Thus, as has been reported for other salamander and anuran species (deMaynadier and Hunter 1998), our findings suggest that this species can be particularly dependent on the availability of core forest areas in the landscape. In fact, changes in microclimatic conditions at forest edges, especially the loss of humidity, can have strong negative impacts on plethodontids, even stronger than on other amphibians, because salamanders from this family do not have lugs and relay on cutaneous respiration (Petranka et al. 1993; deMaynadier and Hunter 1998). Moreover, drier edge conditions also slow wood decomposition (Kapos et al. 1993), which may reduce the availability of suitable refuges for salamanders.
Forest fragmentation per se showed low explanatory power. This is consistent with two global reviews on the topic, which demonstrate that fragmentation per se generally has weak effects on biodiversity (Fahrig 2003, 2017). However, Fahrig (2017) finds that when significant, the effects of fragmentation are mostly positive, not negative. This is consistent with our findings, as we also found that, although weak, all responses to fragmentation were consistently positive. A plausible mechanism that can explain such positive responses is that mean inter-patch isolation distance decreases in landscapes with a higher number of forest patches, potentially facilitating inter-patch animal movements and patch colonization, which can increase population and metapopulation persistence (Fahrig 2003, 2017; Jackson and Fahrig 2016).
Ecological and conservation implications
Our findings suggest that retaining fallen logs in the forest is critically needed to preserve P. robertsi populations. Many saproxylic insects (i.e., insects dependent on dying or dead trees) and vertebrates depend on the amount of dead wood within their movement range, and thus, leaving dead wood in the forest and avoiding clear-cutting can be critical to preserve not only salamanders, but a large number of forest-dwelling species (Petranka et al. 1993; deMaynadier and Hunter 1998; Kouki 2001; Grove 2002). Maintaining and increasing forest core areas (i.e., areas unaffected by forest edges) is also paramount, not only for P. robertsi populations, but also for other forest-core specialist species (Haddad et al. 2015; Pfeifer et al. 2017). For example, Marsh et al. (2004) found that at 100 m from forest edges salamander abundance was higher than at 25 m from the edge and Pfeifer et al. (2017) demonstrate that vertebrate species (amphibians, reptiles, birds and mammals) that inhabit forest cover areas and that are more likely to be included in the IUCN Red List of Threatened Species show higher abundances at sites farther than 200–400 m from forest edges. Therefore, we should prevent the loss of the largest forest patches in the region, and avoid deviations from circularity in patch shapes to increase the amount of core habitat (Ewers and Didham 2006). Although selective logging is preferably to deforestation when considering ecosystem function, services or biodiversity, from the perspective of edge creation, it may pose a greater threat to forest sustainability than deforestation as logging extended more deeply into the interior core of remaining intact forest areas (Broadbent et al. 2008). Thus, selective logging can also be detrimental, especially for forest specialist species, like P. robertsi. These management strategies need to be urgently considered under the context of the recent change of protection level of the Nevado de Toluca volcano. Originally decreed as a National Park, this reserve was recently (2013) decreed as a Flora and Fauna Protected Area—a much less restrictive category that allows forest harvesting practices with commercial proposes in almost all Abies forest extension (Mastretta-Yanes et al. 2014). Therefore, we should be very careful with this productive activity if we are to attain an effective conservation of this endemic and critically endangered species.
References
Arntzen JW, Smithson A, Oldham RS (1999) Marking and tissue sampling effects on body condition and survival in the newt Triturus cristatus. J Herpetol 33:567–576
Arredondo-León C, Muñoz-Jiménez J, García-Romero A (2008) Recent changes in landscape-dynamics trends in tropical highlands, central Mexico. Interciencia 33:569–577
Arroyo-Rodríguez V, Saldaña-Vázquez RA, Fahrig L, Santos BA (2017) Does forest fragmentation cause an increase in forest temperature? Ecol Res 32:81–88
Bille T (2009) Field observations on the salamanders (Caudata: Ambystomatidae, Plethodontidae) of Nevado de Toluca, Mexico. Salamandra 45:155–164
Broadbent EN, Asner GP, Keller M, Knapp DE, Oliveira PJ, Silva JN (2008) Forest fragmentation and edge effects from deforestation and selective logging in the Brazilian Amazon. Biol Conserv 141:1745–1757
Burnham KP, Anderson DR (2002) Model selection and multimodel inference. A practical information-theoretic approach. Springer, New York
Calcagno V, Mazancourt C (2010) Glmulti: an R package for easy automated model selection with (generalized) linear models. J Stat Softw 34:1–29
Catenazzi A (2015) State of the world’s amphibians. Annu Rev Environ Resour 40:91–119
Chao A, Jost L (2012) Coverage-based rarefaction and extrapolation: standardizing samples by completeness rather than size. Ecology 93:2533–2547
CONABIO, Comisión Nacional para el Conocimiento y Uso de la Biodiversidad (2000) Regionalización. http://www.conabio.gob.mx/conocimiento/regionalizacion/doctos/rtp_109.pdf. Accessed 5 Nov 2018
CONAPO (2010) Delimitación de las zonas metropolitanas de México http://www.conapo.gob.mx/en/CONAPO/Zonas_metropolitanas_2010. Accessed 5 Nov 2018
Coulon A, Cosson JF, Angibault JM, Cargnelutti B, Galan M, Morellet N, Petit E, Aulagnier S, Hewison AJ (2004) Landscape connectivity influences gene flow in a roe deer population inhabiting a fragmented landscape: an individual-based approach. Mol Ecol 13:2841–2850
Crump ML, Scott NJ Jr (1994) Visual encounter Surveys. In: Heyer WR, Donnelly MA, McDiarmid RW, Hayek LC, Foster MC (eds) Measuring and monitoring biological diversity: standard methods for amphibians. Smithsonian Institution Press, Washington, pp 84–92
Cushman SA (2006) Effects of habitat loss and fragmentation on amphibians: a review and prospectus. Biol Conserv 128:231–240
Cushman SA, Shirk A, Landguth EL (2012) Separating the effects of habitat area, fragmentation and matrix resistance on genetic differentiation in complex landscapes. Landscape Ecol 27:369–380
Davic RD, Welsh HH Jr (2004) On the ecological roles of salamanders. Annu Rev Ecol Evol Syst 35:405–434
De la Torre Y (1971) Volcanes de México. 2nd edition - Aguilar, México D.F
deMaynadier PG, Hunter ML (1998) Effects of silvicultural edges on the distribution and abundance of amphibians in Maine. Conserv Biol 12:340–352
DOF, Diario Oficial de la Federación Mexicana (2013): Decreto que reforma, deroga y adiciona diversas disposiciones del diverso publicado el 25 de enero de 1936, por el que se declaró Parque Nacional la montaña denominada “Nevado de Toluca” que fue modificado por el diverso publicado el 19 de febrero de 1937. http://dof.gob.mx/nota_detalle_popup.php?codigo=5315889. Accessed 5 Nov 2018
Eigenbrod F, Hecnar SJ, Fahrig L (2008) The relative effect of road traffic and forest cover on anuran populations. Biol Conserv 141:35–46
Eigenbrod F, Hecnar SJ, Fahrig L (2011) Sub-optimal study design has major impacts on landscape-scale inference. Biol Conserv 144:298–305
Ewers RM, Didham RK (2006) Confounding factors in the detection of species responses to habitat fragmentation. Biol Rev 81:117–142
Fahrig L (2003) Effects of habitat fragmentation on biodiversity. Annu Rev Ecol Evol Syst 34:487–515
Fahrig L (2013) Rethinking patch size and isolation effects: the habitat amount hypothesis. J Biogeogr 40:1649–1663
Fahrig L (2017) Ecological responses to habitat fragmentation per se. Annu Rev Ecol Evol Syst 48:1–23
Fahrig L, Baudry J, Brotons L, Burel FG, Crist TO, Fuller RJ, Sirami C, Siriwardena GM, Martin JL (2011) Functional landscape heterogeneity and animal biodiversity in agricultural landscapes. Ecol Lett 14:101–112
Fahrig L, Arroyo-Rodríguez V, Bennett J, Boucher-Lalonde V, Cazeta E, Currie D, Eigenbrod F, Ford A, Jaeger J, Koper N, Martin A, Metzger JP, Morrison P, Rhodes J, Saunders D, Simberloff D, Smith A, Tischendorf L, Vellend M, Watling J (2019) Is habitat fragmentation bad for biodiversity? Biol Conserv 230:179–186
Flageole S, Leclair R (1992) Demography of a salamander (Ambystoma maculatum) population studied by skeletochronology. CanJ Zool 70:740–749
Flores-Villela O, Canseco-Márquez L, Ochoa-Ochoa L (2010) Geographic distribution and conservation of the herpetofauna of the highlands of Central Mexico. In: Wilson LD, Towsend JH, Johnson JD (eds) Conservation of Mesoamerican Amphibians and Reptiles. Eagle Mountain Publishing Co., Utah, pp 303–321
Foley JA, DeFries R, Asner GP, Barford C, Bonan G, Carpenter SR, Chapin FS, Coe MT, Daily GC, Gibbs HK, Helkowski JH, Holloway T, Howard EA, Kucharik CJ, Monfreda C, Patz JA, Prentice IC, Ramankutty N, Snyder PK (2005) Global consequences of land use. Science 309:570–574
Franco-Maass S, Regil-García HH, González-Esquivel C, Nava-Bernal G (2006) Cambio de uso del suelo y vegetación en el Parque Nacional Nevado de Toluca, México, en el periodo 1972-2000. Invest Geog 61:38–57
Frankham R, Briscoe DA, Ballou JD (2002) Introduction to conservation genetics. Cambridge University Press, Cambridge
Gamble LR, McGarigal K, Sigourney DB (2009) Timm BC (2009) Survival and breeding frequency in marbled salamanders (Ambystoma opacum): implications for spatio-temporal population dynamics. Copeia 2:394–407
Garrido-Garduño T, Vázquez-Domínguez E (2013) Métodos de análisis genéticos, espaciales y de conectividad en genética del paisaje. Rev Mex Biodivers 84:1031–1054
González-Fernández A, Manjarrez J, García-Vázquez U, D’Addario M, Sunny A (2018) Present and future ecological niche modeling of garter snake species from the Trans-Mexican Volcanic Belt. PeerJ 6:e4618
Groombridge B, Jenkins M (2000) Global Biodiversity. Earth’s Living Resources in the 21st Century. World Conservation Monitoring Centre, Cambridge, U.K
Grove SJ (2002) Saproxylic insect ecology and the sustainable management of forests. Annu Rev Ecol Syst 33:1–23
Haddad NM, Brudvig LA, Clobert J, Davies KF, Gonzalez A, Holt RD, Lovejoy TE, Sexton JO, Austin MP, Collins CD, Cook WM (2015) Habitat fragmentation and its lasting impact on Earth’s ecosystems. Sci Adv 1:e1500052
Hansen MC, Potapov PV, Moore R, Hancher M, Turubanova SAA, Tyukavina A, Thau D, Stehman SV, Goetz SJ, Loveland TR, Kommareddy A, Egorov A, Chini L, Justice CO, Townshend JRG (2013) High-resolution global maps of 21st-century forest cover change. Science 342:850–853
Hill M (1973) Diversity and evenness: a unifying notation and its consequences. Ecology 54:427–432
IUCN SSC Amphibian Specialist Group (2016) Pseudoeurycea robertsi. The IUCN Red List of Threatened Species. https://www.iucnredlist.org/species/59393/53983925. Accessed 12 Feb 2018
Jackson HB, Fahrig L (2012) What size is a biologically relevant landscape? Landscape Ecol 27:929–941
Jackson ND, Fahrig L (2014) Landscape context affects genetic diversity at a much larger spatial extent than population abundance. Ecology 95:871–881
Jackson HB, Fahrig L (2015) Are ecologists conducting research at the optimal scale? Glob Ecol Biogeogr 24:52–63
Jackson ND, Fahrig L (2016) Habitat amount, not habitat configuration, best predicts population genetic structure in fragmented landscapes. Landscape Ecol 31:951–968
Jaeger JA (2000) Landscape division, splitting index, and effective mesh size: new measures of landscape fragmentation. Landscape Ecol 15:115–130
Jost L (2008) G(ST) and its relatives do not measure differentiation. Mol Ecol 17:4015–4026
Kamvar ZN, Tabima JF, Grünwald NJ (2014) Poppr: an R package for genetic analysis of populations with clonal, partially clonal, and/or sexual reproduction. PeerJ 2:e281
Kapos V, Ganade G, Matsui E, Victoria RL (1993) ∂13C as an indicator of edge effects in tropical rainforest reserves. J Ecol 81:425–432
Kimura M, Crow J (1964) The number of alleles that can be maintained in a finite population. Genetics 49:725–738
Kouki J, Löfman S, Martikainen P, Rouvinen S, Uotila A (2001) Forest fragmentation in Fennoscandia: linking habitat requirements of wood-associated threatened species to landscape and habitat changes. Scand J Forest Res 16:27–37
Laurance WF (1991) Edge effects in tropical forest fragments: application of a model for the design of nature reserves. Biol Conserv 57:205–219
Laurance WF, Lovejoy TE, Vasconcelos HL, Bruna EM, Didham RK, Stouffer PC, Gascon C, Bierregaard RO, Laurance SG, Sampaio E (2002) Ecosystem decay of Amazonian forest fragments: a 22-year investigation. Conserv Biol 16:605–618
Lowe WH, Bolger DT (2002) Local and landscape-scale predictors of salamander abundance in New Hampshire headwater streams. Conserv Biol 16:183–193
Marcon E, Hérault B (2015) entropart: An R package to measure and partition diversity. http://cran.r-project.org/package=entropart
Marsh DM, Thakur KA, Bulka KC, Clarke LB (2004) Dispersal and colonization through open fields by a terrestrial, woodland salamander. Ecology 85:3396–3405
Mas J, Velázquez A, Díaz-Gallegos J, Mayorga-Saucedo R, Alcántara C, Bocco G, Castro R, Fernández T, Pérez-Vega A (2004) Assessing land use/cover changes: a nationwide multidate spatial database for Mexico. Int J Appl Earth Obs Geoinf 5:249–261
Mastretta-Yanes A, Cao R, Nicasio-Arzeta S, Quadri P, Escalante-Espinosa T, Arredondo L, Piñero D (2014) ¿Será exitosa la estrategia del cambio de categoría para mantener la biodiversidad del Nevado de Toluca? Oikos 12:7–17
Matschiner M, Salzburger W (2009) TANDEM: integrating automated allele binning into genetics and genomics workflows. Bioinformatics 25:1982–1983
McGarigal K, SA Cushman, E Ene (2012) FRAGSTATS v4: Spatial Pattern Analysis Program for Categorical and Continuous Maps. Computer software program produced by the authors at the University of Massachusetts, Amherst. http://www.umass.edu/landeco/research/fragstats/fragstats.html
Miguet P, Jackson HB, Jackson ND, Martin AE, Fahrig L (2016) What determines the spatial extent of landscape effects on species? Landscape Ecol 31:1177–1194
Neter J, Kutner MH, Nachtsheim CJ, Wasserman W (1996) Applied linear statistical models. Irwin, Chicago
Newbold T, Hudson LN, Arnell AP, Contu S, De Palma A, Ferrier S (2016) Has land use pushed terrestrial biodiversity beyond the planetary boundary? A global assessment. Science 353:288–291
Nowakowski AJ, Watling JI, Thompson ME, Brusch GA, Catenazzi A, Whitfield SM, Kurz DJ, Suárez-Mayorga A, Aponte-Gutiérrez A, Donnelly MA, Todd BD (2018) Thermal biology mediates responses of amphibians and reptiles to habitat modification. Ecol Lett 21:345–355
Petranka JW, Eldridge ME, Haley KE (1993) Effects of timber harvesting on southern Appalachian salamanders. Conserv Biol 7:363–377
Pfeifer M, Lefebvre V, Peres CA, Wearn O, Marsh C, Banks-Leite C, Butchart S, Arroyo-Rodríguez V, Barlow J, Cerezo A, Cisneros L, D’Cruze N, Faria D, Hadley A, Klingbeil B, Kormann U, Lens L, Rangel GM, Morante-Filho JC, Olivier P, Peters S, Pidgeon A, Ribeiro D, Scherber C, Schneider-Maunoury L, Struebig M, Urbina-Cardona N, Watling JI, Willig M, Wood E, Ewers R (2017) Creation of forest edges has a global impact on forest vertebrates. Nature 551:187–191
Plunkett EB (2009) Conservation implications of a marbled salamander, Ambystoma opacum, metapopulation model. Master’s Thesis, Department of Environmental Conservation, University of Massachusetts Amherst
Polich RL, Searcy CA, Shaffer HE (2013) Effects of tail clipping on survivorship and growth of larval salamanders. J Wildl Manage 77:1420–1425
R Development Core Team (2017) R: A language and environment for statistical computing. R Foundation for statistical computing, Vienna, Austria. http://www.r-project.org)
Russildi G, Arroyo-Rodríguez V, Hernández-Ordóñez O, Pineda E, Reynoso VH (2016) Species-and community-level responses to habitat spatial changes in fragmented rainforests: assessing compensatory dynamics in amphibians and reptiles. Biodivers Conserv 25:375–392
Sánchez-de-Jesús HA, Arroyo-Rodríguez V, Andresen E, Escobar F (2016) Forest loss and matrix composition are the major drivers shaping dung beetle assemblages in a fragmented rainforest. Landscape Ecol 31:843–854
Sánchez-Jasso JM, Aguilar-Miguel X, Medina-Castro JP, Sierra-Domínguez G (2013) Species richness of vertebrates in a reforested woodland of the Nevado de Toluca National Park, Mexico. Rev Mex Biodivers 84:360–373
Savage WK, Fremier AK, Bradley Shaffer H (2010) Landscape genetics of alpine Sierra Nevada salamanders reveal extreme population subdivision in space and time. Mol Ecol 19:3301–3314
SEMARNAT (2010) Norma Oficial Mexicana NOM-059-SEMARNAT-2010, Protección ambiental-especies nativas de México de flora y fauna silvestres-Categorías de riesgo y especificaciones para su inclusión, exclusión o cambio. Lista de especies en riesgo. Diario Oficial de la Federación. http://dof.gob.mx/nota_detalle.php?codigo=5173091&fecha=30/12/2010. Accessed 5 Nov 2018
Skelly DK, Werner EE, Cortwright SA (1999) Long-term distributional dynamics of a Michigan amphibian assemblage. Ecology 80:2326–2337
Spear SF, Peterson CR, Matocq MD, Storfer A (2005) Landscape genetics of the blotched tiger salamander (Ambystoma tigrinum melanostictum). Mol Ecol 14:2553–2564
Stuart SN, Chanson JS, Cox NA, Young BE, Rodrigues ASL, Fischman DL, Waller RW (2004) Status and trends of amphibian declines and extinctions worldwide. Science 306:1783–1786
Toscana-Aparicio A, Granados-Ramírez R (2015) Recategorización del Parque Nacional Nevado de Toluca. Polit Cult 44:79–105
Tuff KT, Tuff T, Davies KF (2016) A framework for integrating thermal biology into fragmentation research. Ecol Lett 19:361–374
Van Buskirk J (2005) Local and landscape influence on amphibian occurrence and abundance. Ecology 86:1936–1947
Van Oosterhout C, Hutchinson WF, Wills DPM, Shipley P (2004) MICRO-CHECKER: Software for identifying and correcting genotyping errors in microsatellite data. Mol Ecol Notes 4:535–538
Velo-Antón G, Windfield JC, Zamudio K, Parra-Olea G (2009) Microsatellite markers for Pseudoeurycea leprosa, a plethodontid salamander endemic to the Transmexican Neovolcanic Belt. Conserv Genet Resour 1:5–7
Velo-Antón G, Parra JL, Parra-Olea G, Zamudio KR (2013) Tracking climate change in a dispersal-limited species: reduced spatial and genetic connectivity in a montane salamander. Mol Ecol 22:3261–3278
Vos CC, Chardon JP (1998) Effects of habitat fragmentation and road density on the distribution pattern of the moor frog Rana arvalis. J Appl Ecol 35:44–56
Wang IJ, Savage WK, Bradley Shaffer H (2009) Landscape genetics and least-cost path analysis reveal unexpected dispersal routes in the California tiger salamander (Ambystoma californiense). Mol Ecol 18:1365–1374
Acknowledgements
We thank IGECEM for the SPOT images and Francisco Reyna-Sáenz, Fabiola Judith Gandarilla-Aizpuro and Carmen Galán-Acedo for their help with SPOT images processing. A.G.-F. obtained a scholarship from CONACyT. A.S received financial support from the Secretary of Research and Advanced Studies (SYEA) of the Universidad Autónoma del Estado de México (Grant No. 4732/2019CIB).
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
González-Fernández, A., Arroyo-Rodríguez, V., Ramírez-Corona, F. et al. Local and landscape drivers of the number of individuals and genetic diversity of a microendemic and critically endangered salamander. Landscape Ecol 34, 1989–2000 (2019). https://doi.org/10.1007/s10980-019-00871-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10980-019-00871-2