Abstract
Hydrazoic acid (HN3) and its deprotonated form azide ion (N3−) (AHA) are toxic because they inhibit the cytochrome c oxidase complex IV (CoX IV) embedded in the inner mitochondrial membrane that forms part of the enzyme complexes involved in cellular respiration. Critical to its toxicity is the inhibition of CoX IV in the central nervous system and cardiovascular system. Hydrazoic acid is an ionizable species and its affinity for membranes, and the associated permeabilities, depend on the pH values of aqueous media on both sides of the membranes. In this article, we address the permeability of AHA through the biological membrane. In order to understand the affinity of the membrane for the neutral and ionized form of azide, we measured the octanol/water partition coefficients at pH values of 2.0 and 8.0, which are 2.01 and 0.00034, respectively. Using a Parallel Artificial Membrane Permeability Assay (PAMPA) experiment, we measured the effective permeability through the membrane, which is logPe − 4.97 and − 5.26 for pH values of 7.4 and pH 8.0, respectively. Experimental permeability was used to validate theoretical permeability, which was estimated by numerically solving a Smoluchowski equation for AHA diffusion through the membrane. We demonstrated that the rate of permeation through the cell membrane of 8.46·104 s−1 is much higher than the rate of the chemical step of CoX IV inhibition by azide of 200 s−1. The results of this study show that transport through the membrane does not represent the rate-limiting step and therefore does not control the rate of CoX IV inhibition in the mitochondria. However, the observed dynamics of azide poisoning is controlled by circulatory transport that takes place on a time scale of minutes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Sodium azide (NaN3) is a white to colourless, crystalline solid that is highly water-soluble, odourless and tasteless [1]. It has a variety of uses: in agriculture, for the production of pesticides and herbicides, as well as for pest control; in biomedicine, as a chemical preservative in aqueous diagnostic reagents; and in hospitals and laboratories, as a chemical preservative in biological fluids. Moreover, it is used for military purposes for the production of detonators and other explosives [2, 3]. The most common use of sodium azide is as a propellant in air bags in vehicles. In this capacity, sodium azide is thermally unstable and it rapidly decomposes to nitrogen gas when it reaches a temperature of 300 °C, causing rapid expansion of the air bag [4]. In pharmacy, NaN3 is used as precursor in the synthesis of numerous compounds such as antiviral drugs, antibiotics [5], and a wide range of tetrazole derivatives, such as sartans. Sartan medicines (azilsartan medoxomil, azilsartan, candesartan cilexetil, candesartan, eprosartan, irbesartan, losartan, EXP3174, olmesartan medoxomil, olmesartan, telmisartan, valsartan [6]) are used as a long-term treatment for patients with arterial hypertension and those with certain heart or kidney diseases. Their use is well-documented in a recent review of the pharmacological treatment of arterial hypertension [7]. To a lesser extent, sartans are used concurrently in the treatment of diabetes, anticoagulant therapy, and cholesterol therapy [4]. Sartan medicines are known as antagonists of angiotensin II receptors and consequently block the action of angiotensin II, a hormone that constricts blood vessels and causes blood pressure to rise [8]. The active pharmaceutical ingredients of sartans and sartan medicines, contain residues of sodium azide in ppm levels as a result of the synthesis processes.
Although the genotoxic properties of sodium azide have been extensively studied, its in vivo genotoxic and mutagenic effects in mammalian cells are still controversial. The European Chemical Agency (ECHA) reports that sodium azide is a non-carcinogenic and non-mutagenic molecule [9] in mammalian cells based on a two-year rat study conducted in 1991. The U.S. Occupational Health and Safety Administration (OSHA) has classifed sodium azide as TLV-A4, “not classifiable as a human carcinogen” [10]. However, there are some studies that have demonstrated the genotoxicity or mutagenicity of sodium azide in mammalian cells. A study on Drosophila melanogaster larvae conducted in 1996 reports the induction of mitotic recombination in wing somatic cells of Drosophila melanogaster larvae after chronic exposure to sodium azide [11]. Raicu and Mixich reported in 1992 that sodium azide encapsulated in liposomes produced chromosomal aberrations when introduced into human heteroploid HFp-2 cells [12]. Jones et al. examining azide mutagenicity towards several rodent cell lines concluded that sodium azide is a mutagen for rodent cells [13]. The European Medicines Agency, a regulatory body, places the Threshold of Toxicological Concern (TTC) at 1.5 µg day−1 per person for intakes of genotoxic or mutagenic impurities, which is considered to be an acceptable risk level (excess cancer risk of < 1 in 100,000 over a lifetime of exposure) for most pharmaceuticals [14]. The United States Pharmacopoeia (USP) sets a concentration of NMT (not more than) 10 ppm of azide for sartans [15].
Chang et al. [16] pointed out that hypotension is the most common symptom of poisoning with sodium azide. Acute poisonings can occur with ingestion of sodium azide or with inhalation of vapours of hydrazoic acid (HN3) that are formed when sodium azide is dissolved in an acidic aqueous solution. At lower heart rates, tissues are less well-supplied with oxygen. Tissues that consume the largest amount of oxygen are muscles (20%), liver (20%), brain (20%), heart (10–12%), and kidney (7.2%) [17]. Damage to neurons is particularly critical since, in contrast to most other cell types, they do not easily regenerate and are not simply replaced after damage.
Sodium azide is primarily a mitochondrial toxin, as it binds within the electron transport chain (ETC), thus inhibiting oxidative phosphorylation [18]. The azide anion (N3−) has a greater affinity for the heme complex than O2 [19]. Therefore, it covalently binds to the fully oxidized form of iron (Fe3+), in place of O2, in the binuclear complex of heme a3 \({(\text{F}\text{e}}_{\text{h}\text{e}\text{m}\text{e} \text{a}3}^{3+}-{\text{C}\text{u}}_{\text{B}}^{2+})\) in the cytochrome c oxidase complex IV (CoX IV) bound to the inner mitochondrial membrane [20]. In CoX IV, the azide anion and trivalent iron form a bridging structure of \({\text{F}\text{e}}_{\text{h}\text{e}\text{m}\text{e}\,\text{a}3}^{3+}-\text{N}=\text{N}=\text{N}-{\text{C}\text{u}}_{\text{B}}^{2+}\) [21]. The resulting reduction in adenosine triphosphate (ATP) production, even in the presence of oxygen, results in metabolic failure (See Fig. S1, Fig. S2 and Fig. S3 under Supplementary Materials for a schematic representation of the normal ETC in Complex IV and a representation of the azide anion binding to the CoX IV complex in ETC [22, 23] )
When sodium azide dissolves in water or in an acidic medium it dissociates to sodium and azide ions. In acidic solution, the azide ion binds the proton and forms a neutral, non-ionizable and volatile hydrazoic acid (HN3). The chemical reaction takes place according to the following mechanism:
HN3 is a weak acid with a pKa value of 4.65 [18]. In acidic solution, neutral HN3 distributes preferentially to the octanol phase rather than to water. In basic solution, azide takes the deprotonated form of the azide anion, N3− , which is distributed mainly in the aqueous phase when an octanol/water equilibrium is established. The contribution of ionic species to Kow (octanol/water partition coefficient) is basically negligible in the case of HN3/N3−, while it becomes significant in the case of lipophilic species. For example, the local anesthetic bupivacaine, a lipophilic molecule, prefers the octanol environment over the acidic aqueous environment in its protonated form [24].
In this study, we addressed the permeability of AHA through the biological membrane. In order to understand the affinity of the membrane for the neutral and ionized form of azide we measured the octanol/water partition coefficients at pH values of 2.0 and 8.0, respectively. We measured the effective permeability through the membrane using a PAMPA experiment. By numerically solving the diffusion equation, we demonstrated that the measured permeability is at least to some extent dependent on diffusion through the unstirred water layer on each side of the membrane. Based on these experiments, we tried to determine whether diffusion through the membrane is the limiting step in the inhibition of the complex.
2 Materials and methods
2.1 Determination of the Octanol/Water Partition Coefficients (Kow) of Azide Ion/Hydrazoic Acid for Different pH Values Using Reversed-Phase Liquid Chromatography and UV Detection
In our study, we considered experimental octanol water partition coefficients for azidic acid and azide ion, respectively. In principle, one can apply one of the developed constant pH simulation algorithms [25, 26], but it should be noted that the HN3 pKa value at the interface is highly dependent on the coordinate and is computationally as demanding as determining pKa values in the protein interior [27, 28] and in contrast to proteins inaccessible to experiment [29].
Since hydrazoic acid is an ionizable species its octanol/water partition coefficient (Kow) depends on pH. We therefore decided to measure Kow for two pH values and apply an extrapolation approach to determine the rest of the pH values [30]. The approach of Hodges et al. [31] is very similar to the approach of Strichartz et al. and both approaches consider contributions of both neutral and ionized species to the octanol/water partition coefficient. The determination of octanol/water (octanol/water) partition coefficients (Kow) of AHA was carried out using two different pH values of azide solutions (pH 2.0 and 8.0) and a reversed-phase liquid chromatography with UV detection. The pH-dependent partition coefficients Kow(pH) were calculated from the experimental data using the equation:
where [A]oct represents total azide concentration (sum of neutral HN3 and deprotonated N3− species) in the octanol organic phase and [A]aq represents the total azide concentration in the aqueous phase.
Since the HPLC method does not distinguish between neutral and ionizable species of hydrazoic acid and azide anion, the partition coefficient Kow(pH) below can be expressed as a distribution coefficient Q(pH). Azide exists in its neutral or protonated form both in aqueous solution and in octanol. The ratio of concentrations of azide in octanol (the organic phase) and in water regardless of its protonation state at the given pH value of the environment is described by the distribution coefficient Q. Q(pH) is an equilibrium constant and a measure of free energy. We define Q(pH) as:
where [A0]org stands for the concentration of the neutral form of azide (HN3, hydrazoic acid) in the organic phase (octanol), [A0]aq for the concentration in the aqueous solution, [A¯]org for the concentration of the deprotonated form of azide (N3¯) in the organic phase, and [A¯]aq for the concentration of azide in the aqueous solution.
By expanding the numerator and denominator of the right-hand side of the Eq. (2) by factor 1/[A−]aq one obtains
If a new variable P0 is introduced as the partition coefficient of a neutral species, then P0 = [A0]org/[A0]aq, and P¯ as the partition coefficient of deprotonated species is P¯ = [A¯]org/[A¯]aq, meaning the equation simplifies to a more compact form:
[A0]aq/[A−]aq is the equilibrium constant of neutral and deprotonated species in aqueous solution that depends both on the pKa value of azide and the aqueous phase pH value.
The free energy for azide deprotonation (ΔG) with a given pKa in a solution with a given pH value is:
where kB is Boltzmann’s constant and equals 1.987 kcal (mol K)−1 and T is the absolute temperature in Kelvins (310 K under physiological conditions). Equation 5 implies that an ionizable group with a certain pKa value is in contact and in equilibrium with the aqueous solution at a certain pH value. There is a one-to-one correspondence between the equilibrium constant and free energy [32], which is expressed as
By combining Eqs. 5 and 6 and by considering the equation \({\text{ln}}x = {\text{ln}}10 \cdot {\text{log}}x\), and introducing the relationship \(\text{log}\beta =pH-{pK}_{a}\) or \(\beta ={10}^{{\text{p}\text{K}}_{\text{a}}-\text{p}\text{H}}\), it is possible to write an equation for the azide distribution coefficient as a function of pH and pKa.
The distribution coefficient Q is the equilibrium constant for azide distribution between the organic phase and the aqueous phase regardless of the azide protonation state. The corresponding free energy ΔG associated with azide transfer from the aqueous solution with a certain pH value to the membrane is:
Since extracellular fluid (ECF) and cytoplasm (CYT) have different pH values of 7.4 and 7.2, respectively, the free energy for the transfer of AHA is not zero. Please note that both pH values refer to human cells [33, 34]. AHA tends to concentrate in the compartment with a more favourable solvation free energy. To calculate the relative population of AHA in the cell membrane, the appropriate Q for a given pH value is used:
The same equation can be used for the equilibrium between the cytoplasm and the membrane. By combining these two relative populations of AHA in the cytoplasm and the extracellular fluid (ECF), we get:
The relative AHA population in the cytoplasm is therefore simply the ratio of both Q.
By inserting Eq. 7, we can derive a simplified formula:
As seen in Eq. 11, the overall AHA concentration in the cytoplasm is pKa independent and depends only on the pH value difference of the cytoplasm and ECF.
2.2 Determination of Effective Permeability, P e (pH) Using the PAMPA Method
Simulation of azide partition between the extracellular fluid and biological membrane was performed using the PAMPA (Parallel Artificial Membrane Permeability Assay) method, the schematics of which are shown in Fig. 1. PAMPA is an in vitro tool for high-throughput prediction of in vivo drug permeability and is also useful for the assessment of passive transport mechanisms [35]. Our experiments were performed using the Corning BioCoat Pre-Coated PAMPA Plate System with 96-well insert system with a 0.45 μm PVDF (polyvinylidene fluoride) filter plate which has been pre-coated with structured layers of phospholipids and a matched receiver microplate. The thickness d of the phospholipid membrane was 125 μm = 1,250,000 Å.
Schematic representation of a single well in the PAMPA sandwich. 200 µL of solvent per well was pipetted into upper donor plate and 300 µL of sample per well was pipetted into acceptor plate. The upper donor plate with sample was sealed with a label to prevent hydrazoic acid from evaporating and cross-contaminating other wells. We placed the filter plate on the receiver plate and incubated the assembly at room temperature for 5 h without stirring
PAMPA experimental conditions are described in Table S3 under Supplementary Materials. Prior to use, the pre-coated PAMPA plate system was warmed to room temperature for 60 min. The compound solutions of NaN3 with concentrations of 0.241 and 0.120 mg mL−1 were prepared by dilution of stock solutions (1.2 mg mL−1) into a PBS buffer adjusted to two pH values of 7.4 and 8.0 using 1 M NaOH. 300 µL of solvent per well was pipetted into the acceptor plate and 200 µL of sample per well was pipetted into the upper donor plate. The upper donor plate with sample was sealed with the Sealing Tape for 96-Well Plates to prevent hydrazoic acid from evaporating and contaminating other wells. We placed the filter plate on the receiver plate and incubated the assembly at room temperature for 5 h without stirring. After that, we separated the plates and determined the compound concentrations in both plates using HPLC with UV detection; the chromatographic conditions are listed in Table S4 under Supplementary Materials.
Determination of effective permeability, Pe(pH) for azide (cm s−1) at pH values of 7.4 and 8.0 was calculated as follows:
We calculated Mass Retention (%) using Eq. 13:
where C0 is the initial azide concentration in the donor well (mM), CD(t) is the azide concentration in the donor well at time t (mM), CA(t) is the azide concentration in acceptor well at time t (mM), VD is the donor well volume of 0.2 mL (including compound in buffer), VA is the acceptor well volume of 0.3 mL (including buffer), S is the filter area of 0.3 cm2, and t is the incubation time (5 h = 18,000 s). Equilibrium concentration, Cequilibrium (µM), was calculated as
2.3 Theoretical Prediction of Permeability (Pt)
Herman J.C. Berendsen was among the first to perform molecular simulations of membranes to address their structure, stability, and transport properties [36,37,38,39,40,41], followed by his disciples who included simulation of membrane embedded transporters [42,43,44,45,46].
Diffusion is a passive process of the net movement of a substance driven by a concentration gradient. Diffusion is described by the following diffusion equation in n-dimensions:
where u is the coordinate and time-dependent concentration, D is the diffusion coefficient, and \({\nabla }^{2}\) is the Laplace operator. It must be noted that in our calculation D is coordinate- and time- independent. Furthermore, in complex environments, such as those in a nerve cell, the concentration difference is not the only force driving diffusion. Therefore, we include the coordinate-dependent chemical potential (\(\mu\)) in Eq. 15, which alters the time-dependent concentration profiles. This is a special case of the more general Fokker-Planck equation, called the Smoluchowski equation [47], which reads:
where \(\mu\) is the coordinate-dependent chemical potential, kb is the Boltzmann constant, T is the absolute temperature, and \(\nabla\) is the gradient of a function.
Since the diffusion equation with coordinate-dependent chemical potential described above does not allow for analytical solution, we proceeded to solve it numerically on a grid. Algorithms for numerical solutions of such a diffusion equation have appeared only recently [48]. The carefully performed computational study of oxygen diffusion through membranes by Ghysels et al. is particularly illustrative [49]. We have used the finite differences numerical method. Two numerical simulations were made, one with the dimensions of a realistic cell membrane with thickness of 50 Å and one with real PAMPA membrane thickness of 125 μm according to the manufacturer. The first experiment used grid spacing of 1 Å and a time step of 0.1 ns, while the second experiment used grid spacing of 1 μm and time step of 1 µs. In both simulations a diffusion coefficient D of 820 µm2 s− 1 from Huang and co-workers [50] was used. Moreover, both simulations used the free energy profiles of the transition into and out of the membrane, which were experimentally determined earlier in this study.
The permeability across the membrane is described by the following equation:
where j is the flux, P is the membrane permeability, and \(\varDelta c\) is the difference in concentration on both sides of the membrane. Flux was calculated numerically from the diffusion equation with coordinate-dependent chemical potential, while the concentration difference was initially 1.
2.4 Preparation of Solutions
Solvents for the sample solutions were prepared by mixing acetonitrile and water in ratio 1:10 (V/V). Solvents were adjusted to pH 2.0 with 20% H3PO4 and to pH 8.0 with 0.01 M NaOH before filling to the calibration mark.
First, the calibration curves were prepared at five concentration levels: these were from 0.12 to 1.5 mg mL−1 of NaN3 (corresponding to 0.08 to 1.0 mg mL−1 of HN3) for a pH value of 2.0, and from 0.0019 to 0.12 mg mL−1 of NaN3 (corresponding to 0.0013 to 0.08 mg mL−1 of HN3) for a pH value of 8.0.
Aqueous sample solutions of NaN3 with pH values of 2.0 and 8.0 were also prepared. The concentration of the sample solution with a pH value of 2.0 was 1.5 mg mL−1 of NaN3 (equal to 1.0 mg mL−1 of HN3) and the sample solution with a pH value of 8.0 was 12 mg mL−1 (equal to 8.0 mg mL−1 of HN3). Equal volumes, 15 mL of the solution of NaN3 and 15 mL of 1-octanol, were then pipetted into a 50 mL separatory funnel and mixed well every 10 min for one hour to equilibrate. Finally, the upper organic and lower aqueous phase were sampled. The aqueous phase with a pH value of 8.0 was diluted by a factor of 1250. All solutions for the calibration curves and final solutions of the organic and aqueous phases were injected into an HPLC system.
3 Results
3.1 Determination of octanol/water Partition Coefficients (Kow)
The octanol/water partition coefficient is defined as the ratio of equilibrium concentrations of a given species between octanol and water. For ionizable species such as azide, the ratio depends on the pH value of the aqueous solution.
The octanol/water partition coefficients (Kow) were calculated from the experimentally determined concentrations of AHA in both phases: the organic phase of octanol and the water phase of a 10% solution of acetonitrile. Kow was 2.01 for the pH value of 2.0 and 0.00034 for pH value of 8.0. The R-squared of the calibration curves with solutions of sodium azide were 0.974 for a pH value of 2.00 and 1.0 for a pH value of 8.0. We used Eq. 7 to interpolate and extrapolate experimental data and to calculate Kow(pH) = Q(pH).
Experimental results show that in an acid environment (pH 2.0) in which azide mostly takes its neutral form (HN3), 67% of azide is in the octanol phase. In a basic environment with a pH above pKa 4.65, the azide mostly exists in its ionic form N3ˉ, so a lower partition coefficient and concentration in octanol is expected and indeed the experimentally determined value is 0.00034 (pH of 8.0) of azide in octanol. The experimental values of the corresponding calculated free Gibbs energies of activation ΔG‡(pH) for the transition of AHA from the aqueous to the organic phase are − 0.413 kcal mol−1 (pH 2.0) and 3.29 kcal mol−1 (pH 8.0) and were calculated using the equation ΔG‡ = ∆Gow = − kBT ∙ ln Kow. Extrapolating from the experimental results, the distribution coefficients, Q, and free energies, ΔG, for the transfer of AHA were calculated and are listed in Table 1.
Please refer to Fig. 2 to aid with understanding of the local environment around the cell membrane.
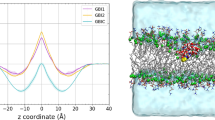
Free energy profile for transfer of AHA from extracellular fluid (ECF) to cytoplasm (CYT). Please note that the free energy difference between ECF and CYT is due to the different pH values on both sides of the membrane. Relative concentrations of AHA in the ECF, cell membrane and CYT are represented by the blue line. ECF is an endless reservoir with the relative concentration of AHA set to 1 and a pHECF value of 7.4. The pHCYT value of the cytoplasm is 7.2. The red line represents the free energy profile for transfer of AHA from the ECF to CYT under physiological conditions. Total ΔG is the change in free energy between the two compartments (0.25 kcal mol−1). All values are given in kcal mol−1. Note that the free energy profile was constructed solely from the experimental data
3.2 Experimental Determination of Effective Permeability, Pe (pH)
The results of the effective permeability of the artificial membrane for AHA were obtained using the following approaches: experimentally, by using the PAMPA method, and, theoretically, by numerically solving the diffusion equation.
Experimentally, we obtained a value of Pe of 11.2 · 10−6 cm s−1 at a pH value of 7.4 and 5.5 · 10−6 cm s−1 at a pH value of 8.0 (see Table 2). Nearly 11% of azide crossed the membrane into the acceptor solution with a pH value of 7.4 and 9% into the acceptor solution with pH value of 8.0. Mass retention, calculated on the donor and acceptor side was on average − 0.15% and − 0.03% for pH 7.4 and 8.0, respectively, which indicates a minor analytical error.
3.3 Computational Prediction of Permeability (Pt)
By solving the diffusion equation numerically, we obtained two permeability values Pt (logPt) for the AHA transition from the ECF to CYT (Fig. 2) of 4.23 ·10−2 cm s−1 (− 1.73) and 22.3 ·10−6 cm s−1 (− 4.65), for 50 Å and 125 μm membrane, respectively.
Experimental (− 4.97) and theoretical values (− 4.65) of permeability at a physiological pH of 7.4 for the 125 μm PAMPA membrane differ only slightly. However, the realistic 50 Å membrane has a much higher permeability (104 times). Similarly, we can calculate the theoretical rate constant of permeation through the membrane, which is 8.46 · 104 s−1 and 1.55 · 10−3 s−1 for the 50 Å and 125 μm membrane, respectively.
4 Discussion
Current opinion is that azide enters the cell through the plasma membrane rather than via any of the transporters. In this respect it is very likely that it enters in its neutral form of HN3, since its deprotonated form (N3−) has a too-favourable free energy of hydration, consequently the free energy cost for entering the membrane would be too high. The free energy for protonation of ionized species with a given pKa value in aqueous medium with a given pH value is \(\varDelta G=-{ln}\left(10\right){k}_{B}T\left(pH-p{K}_{a}\right)\), where \(\varDelta G\) is the free energy, \({k}_{B}\) is the Boltzmann constant, T is the absolute temperature (310 K for human body) and the pKa value for hydrazoic acid is 4.65. Hence, the free energy for azide protonation at pH of 7.4 is \(- 2.303 \cdot 0.593\,{\text{kcal mol}}^{{ - 1}} \cdot (7.4 - 4.65) = - 3.75\,{\text{kcal mol}}^{{ - 1}}\). On the other hand, the energy cost for HN3 deprotonation in media with a pH value of 2.0 is 3.62 kcal mol− 1. Since deprotonated ionic species (N3−) have a more favourable hydration energy in more basic media, equilibrium populations of HN3 are higher in the cytoplasm (CYT, with a pH value of 7.2) than in the extracellular fluid (ECF, with a pH value of 7.4) or in the mitochondria (with a pH value of 8) and equilibrium populations of N3ˉ are highest in the mitochondria.
At lower pH, azide mostly exists in the protonated form of hydrazoic acid (HN3), which is hydrophobic to some extent and preferentially distributes to the gas phase or lipid membrane from aqueous solution. Hydrazoic acid has a relatively small dipole moment of 0.847 ± 0.005 Debye [51]. Observing the measured octanol/water partition coefficient supports this. At pH 2, there is more AHA in the nonpolar solvent (2:1), while at pH 8 there is significantly more AHA in the water phase (1:2940). Consequently, within the physiological pH range, azide is always hydrophilic and predominantly in ionized form (N3−). According to the Overton rule [52] more hydrophilic substances are less membrane permeable [53, 54]. Consequently, one could predict that AHA will not permeate easily through the cell membrane. However, its small molecular size and the spread charge in its ionized form yield a moderately low free energy barrier of 3.29 kcal mol−1 for entry into the membrane. By comparison, MMB4, a popular drug that crosses the blood-brain barrier, has a free energy barrier of 28 kcal mol−1 [55], morphine has a free energy barrier of 7 kcal mol−1 [56], and carbon dioxide has a free energy barrier of around 1 kcal mol−1 [57]. AHA diffuses from the region with a higher pH value (ECF) to the region with a lower pH value. More azide molecules are in protonated form (HN3) in cytoplasm with a pH value of 7.2, than in the ECF at a higher pH value of 7.4, consequently the equilibrium free energy difference for permeation from the ECF to the cytoplasm is 0.25 kcal mol−1, which is solely due to the difference in pH. Local acidosis or alkalosis would change this. More interesting is that the equilibrium concentration ratio between the cytoplasm and the ECF changes depending on the pH difference. In the case of physiological pH values, less AHA is in cells than in the ECF (0.238), while in the case of local acidosis, additional AHA would accumulate in cells. For example, poisoning with azide disturbs the aerobic metabolism of the cell, shifting it to anaerobic, which produces acidic lactic acid, thus lowering the ECF pH and starting a positive feedback loop or death spiral.
Our experimental data for the permeability of AHA using the PAMPA method gave a permeability value (logPe) of − 4.97, which is still considered fast, according to Bennion et al., who estimate that compounds with logPe < − 5.5 have low permeability [55]. Bodor et al. studied the correlation between the octanol/water partition ratio (Q) and blood-brain barrier permeability [58]. AHA with a Q of 0.00391 would have a low blood-brain barrier permeability similar to urea. It should be noted that PAMPA is specifically designed to study intestinal absorption of drugs, but we study permeation through the cell membrane, which is about 25,000 times thinner. Therefore, we conducted two simulations. The results of numerical solutions for the diffusion equation predict the experimental permeability for a membrane of 125 μm reasonably well. In fact, a surprisingly good agreement between the experimental and calculated values was found. However, currently there are no reliable experimental methods that would yield the permeability of a cell membrane with a thickness of approximately 50 Å. Since the computational predictions proved to be valid for a 125 μm thick membrane we firmly believe that the permeability predictions for a 50 Å membrane are equal reliable.
Given that the rate constant of permeation through the cell membrane for AHA is 8.46 ·104 s−1 and the rate constant for the association of AHA with the Cytochrome c oxidase (CoX IV) is 200 s−1 as measured by Antalis et al. [59] for association with myoglobin, we have good evidence that the diffusion through the membrane is not the rate limiting step and does not control the rate of CoX IV inhibition by AHA. The chemical step, which is the reaction of the heme group when AHA forms a Michaelis complex with CoX IV, controls the overall rate of inhibition. The myoglobin association rate is similar to CoX IV because both have heme groups, which are targeted by the azide ion. The rate limiting step is therefore the association with heme, since AHA is a small molecule that can easily permeate the membrane regardless of its hydrophilic nature. It is worth stressing that for most enzymatic reactions in aqueous solution diffusion is rarely the rate-limiting step. The “average enzyme” is far from kinetic perfection as it exhibits a kcat of ∼10 s−1 [60]. We are aware that the observed dynamics of azide poisoning is however controlled by circulatory transport that takes place on a time scale of minutes and to some extent depends on the route of administration (intravenous (IV), inhalation, intramuscular (IM)).
5 Conclusions
By measuring the octanol/water partition coefficients for the azide ion/hydrazoic acid (AHA) we have demonstrated that, at physiological pH values of extracellular fluid, AHA prefers extracellular fluid over cytoplasm, while membrane and other lipophilic compartments have the lowest relative populations.
The experimental values for effective permeability obtained using the PAMPA method were compared with the results of numerical solution of the diffusion equation through the membrane. For a 50 Å thick membrane we estimated the permeability of AHA to be 8.46 · 104 s−1 while the rate constant for the association of AHA with the heme group is 200 s−1 as measured by Antalis et al. [59]. One can therefore conclude that CoX IV inhibition by azide ion is controlled by the chemical step rather than by diffusion through the membrane. It should be emphasized that this study focused on the events that follow complete embedding of the cell in the azide ion/hydrazoic acid solution. However, the observed dynamics of azide poisoning are controlled by transport in the circulation, which takes place on a time scale of minutes.
Data Availability
All data and code can be obtained from the authors upon request.
References
Gričar M, Andrenšek S (2016) Determination of azide impurity in sartans using reversed-phase HPLC with UV detection. J Pharm Biomed Anal 125:27–32. https://doi.org/10.1016/j.jpba.2016.03.020
Kubota N (2015) Propellants and Explosives: thermochemical Aspects of Combustion. Wiley, New York
Hirata N, Matsuda N, Kubota N (2000) Combustion of NaN3 Based Energetic Pyrolants. Propell, Explosives, Pyrotech 25(5):217–9
Ciesla MM, Calello DP, Nelson LS (2018) When the poisoned risk poisoning others: fatal sodium azide overdose. Emerg Med. https://doi.org/10.12788/emed.2018.0093
Vinković K, Drevenkar V (2008) Ion chromatography of azide in pharmaceutical protein samples with high chloride concentration using suppressed conductivity detection. J Chromatogr B Analyt Technol Biomed Life Sci 864(1–2):102–108. https://doi.org/10.1016/j.jchromb.2008.01.043
Michel MC, Foster C, Brunner HR, Liu L (2013) A systematic comparison of the properties of clinically used angiotensin II type 1 receptor antagonists. Pharmacol Rev 65(2):809–848. https://doi.org/10.1124/pr.112.007278
European Medicines Agency SMH (2019) : Sartan medicines: companies to review manufacturing processes to avoid presence of nitrosamine impurities. https://www.ema.europa.eu/en/documents/referral/angiotensin-ii-receptor-antagonists-sartans-article-31-referral-sartan-medicines-companies-review_en.pdf Accessed
Williams B, Mancia G, Spiering W, Agabiti Rosei E, Azizi M, Burnier M et al (2018) 2018 ESC/ESH guidelines for the management of arterial hypertension. Eur Heart J 39(33):3021–3104. https://doi.org/10.1093/eurheartj/ehy339
Agency EC (2022) : Sodium azide. https://echa.europa.eu/registration-dossier/-/registered-dossier/14174/7/8 Accessed 20.12.2022
Administration OSaH (2022) : SODIUM AZIDE. https://www.osha.gov/chemicaldata/331 Accessed 20.12.2022
González-César E, Ramos-Morales P (1997) Sodium azide induces mitotic recombination in Drosophila melanogaster larvae. Mutat Res/Genet Toxicol Environ Mutagen 389(2):157–165. https://doi.org/10.1016/S1383-5718(96)00142-8
Raicu P, Mixich F (1992) Cytogenetic effects of sodium azide encapsulated in liposomes on heteroploid cell cultures. Mutat Res Lett 283(3):215–219. https://doi.org/10.1016/0165-7992(92)90110-4
Jones JA, Starkey JR, Kleinhofs A (1980) Toxicity and mutagenicity of sodium azide in mammalian cell cultures. Mutat Res 77(3):293–299. https://doi.org/10.1016/0165-1218(80)90064-6
European Medicines Agency SMH (2015) : ICH guideline M7(R1) on assessment and control of DNA reactive (mutagenic) impurities in pharmaceuticals to limit potential carcinogenic risk. https://www.ema.europa.eu/en/documents/scientific-guideline/ich-guideline-m7r1-assessment-control-dna-reactive-mutagenic-impurities-pharmaceuticals-limit_en.pdf Accessed
Pharmacopoeia US USP-43-NF38–2402. https://www.uspnf.com/sites/default/files/usp_pdf/EN/USPNF/usp-nf-commentary/usp-43-nf-38-index.pdf Accessed
Chang S, Lamm SH (2003) Human health effects of sodium azide exposure: a literature review and analysis. Int J Toxicol 22(3):175–186. https://doi.org/10.1080/10915810305109
Significance of Brain Tissue Oxygenation and the Arachidonic Acid Cascade in Stroke (2011) Antioxid Redox Signal 14(10):1889–1903. https://doi.org/10.1089/ars.2010.3474
Agency USEP (1987) : Sodium azide; CASRN 26628-22-8. https://iris.epa.gov/static/pdfs/0191_summary.pdf Accessed
Casarett LJ, Doull J, Klaassen CD (2008) Toxicology: the basic science of poisons. McGraw-Hill, New York
Leavesley HB, Li L, Prabhakaran K, Borowitz JL, Isom GE (2007) Interaction of Cyanide and nitric oxide with cytochrome c oxidase: implications for acute cyanide toxicity. Toxicol Sci 101(1):101–111. https://doi.org/10.1093/toxsci/kfm254
Tsubaki M, Mogi T, Hori H (1999) Fourier-transform infrared studies on azide-binding to the binuclear center of the Escherichia coli bo-type ubiquinol oxidase11This work was supported in part by Grants-in-aid for Scientific Research on Priority Areas (10129226 to MT, 08249106 to TM and HH) and for Scientific Research (C) (09833002 to MT, 09680651 to HH) from the Ministry of Education, Science, Sports and Culture, Japan. FEBS Lett 449(2):191–195. This is the paper XXXVIII in the series ‘Structure-function studies on the E. coli cytochrome bo’https://doi.org/10.1016/S0014-5793(99)00423-8
Zhao RZ, Jiang S, Zhang L, Yu ZB (2019) Mitochondrial electron transport chain, ROS generation and uncoupling (review). Int J Mol Med 44(1):3–15. https://doi.org/10.3892/ijmm.2019.4188
Berg JM, Tymoczko JL, Stryer L, Information NCfB (2002) Oxydative Phosphorylation. Biochemistry, Fifth Edition. vol pts. 1–34: W. H. Freeman; p. 491–526
Kavcic H, Umek N, Vintar N, Mavri J (2021) Local anesthetics transfer relies on pH differences and affinities toward lipophilic compartments. J Phys Org Chem. https://doi.org/10.1002/poc.4275
Donnini S, Ullmann RT, Groenhof G, Grubmüller H (2016) Charge-neutral constant pH Molecular Dynamics Simulations using a parsimonious Proton buffer. J Chem Theory Comput 12(3):1040–1051. https://doi.org/10.1021/acs.jctc.5b01160
Bennett WFD, Chen AW, Donnini S, Groenhof G, Tieleman DP (2013) Constant pH simulations with the coarse-grained MARTINI model — application to oleic acid aggregates. Can J Chem 91(9):839–846. https://doi.org/10.1139/cjc-2013-0010
Sham YY, Chu ZT, Warshel A (1997) Consistent calculations of pKa’s of Ionizable residues in Proteins: semi-microscopic and microscopic approaches. J Phys Chem B 101(22):4458–4472. https://doi.org/10.1021/jp963412w
Repič M, Purg M, Vianello R, Mavri J (2014) Examining Electrostatic Preorganization in Monoamine Oxidases A and B by Structural comparison and pKa calculations. J Phys Chem B 118(16):4326–4332. https://doi.org/10.1021/jp500795p
Czerwinski RM, Harris TK, Massiah MA, Mildvan AS, Whitman CP (2001) The structural basis for the perturbed pKa of the catalytic base in 4-oxalocrotonate tautomerase: kinetic and structural effects of mutations of Phe-50. Biochemistry 40(7):1984–1995. https://doi.org/10.1021/bi0024714
Strichartz GR, Sanchez V, Arthur GR, Chafetz R, Martin D (1990) Fundamental properties of local anesthetics. II. Measured octanol:buffer partition coefficients and pKa values of clinically used drugs. Anesth Analg 71(2):158–170. https://doi.org/10.1213/00000539-199008000-00008
Hodges G, Eadsforth C, Bossuyt B, Bouvy A, Enrici M-H, Geurts M et al (2019) A comparison of log Kow (n-octanol–water partition coefficient) values for non-ionic, anionic, cationic and amphoteric surfactants determined using predictions and experimental methods. Environ Sci Europe 31(1):1. https://doi.org/10.1186/s12302-018-0176-7
Atkins P, DePaula J, Keeler J (2018) Atkins’ Physical Chemistry. 11th Edition. Oxford: Oxford University Press;
Madshus IH (1988) Regulation of intracellular pH in eukaryotic cells. Biochem J 250(1):1–8. https://doi.org/10.1042/bj2500001
Hopkins E, Sanvictores T, Sharma S, Physiology (2022) Acid base balance. StatPearls [Internet]. StatPearls Publishing, Treasure Island (FL)
Di L, Kerns E (2008) Drug-like Properties: concepts, structure design and methods. Academic Press, USA
Tieleman DP, Marrink SJ, Berendsen HJC (1997) A computer perspective of membranes: molecular dynamics studies of lipid bilayer systems. Biochimica et Biophysica Acta (BBA) - reviews on Biomembranes. ;1331(3):235–70. doi: https://doi.org/10.1016/S0304-4157(97)00008-7
Marrink S-J, Berendsen HJC (1994) Simulation of water transport through a lipid membrane. J Phys Chem 98(15):4155–4168. https://doi.org/10.1021/j100066a040
Marrink SJ, Berendsen HJC (1996) Permeation process of small molecules across lipid membranes studied by Molecular Dynamics Simulations. J Phys Chem 100(41):16729–16738. https://doi.org/10.1021/jp952956f
Berendsen HJC, Marrink S-J (1993) Molecular dynamics of water transport through membranes: water from solvent to solute. Pure Appl Chem 65(12):2513–2520. https://doi.org/10.1351/pac199365122513
Vanbuuren A, Berendsen HJC, MOLECULAR-DYNAMICS SIMULATIONS OF CARBOHYDRATE-BASED (1994) SURFACTANTS IN SURFACTANT WATER OIL SYSTEMS. Langmuir 10:1703–1713
Ahlstrom P, Berendsen HJC (1993) A molecular dynamics study of lecithin monolayers. J Phys Chem 97(51):13691–13702. https://doi.org/10.1021/j100153a044
Liu Y, de Vries AH, Pezeshkian W, Marrink SJ (2021) Capturing membrane phase separation by Dual Resolution Molecular Dynamics Simulations. J Chem Theory Comput 17(9):5876–5884. https://doi.org/10.1021/acs.jctc.1c00151
Liu Y, De Vries AH, Barnoud J, Pezeshkian W, Melcr J, Marrink SJ (2020) Dual resolution membrane simulations using virtual Sites. J Phys Chem B 124(19):3944–3953. https://doi.org/10.1021/acs.jpcb.0c01842
Marrink SJ, de Vries AH, Tieleman DP (2009) Lipids on the move: simulations of membrane pores, domains, stalks and curves. Biochimica et Biophysica Acta (BBA). - Biomembr 1788(1):149–168. https://doi.org/10.1016/j.bbamem.2008.10.006
Wennberg CL, van der Spoel D, Hub JS (2012) Large influence of cholesterol on Solute partitioning into lipid membranes. J Am Chem Soc 134(11):5351–5361. https://doi.org/10.1021/ja211929h
Kopec W, Rothberg BS, de Groot BL (2019) Molecular mechanism of a potassium channel gating through activation gate-selectivity filter coupling. Nat Commun 10(1):5366. https://doi.org/10.1038/s41467-019-13227-w
Smoluchowski MV (1916) Drei Vortrage uber Diffusion, Brownsche Bewegung und Koagulation von Kolloidteilchen. Z Angew Phys 17:557–585
Livshits AI (2016) The effect of spatial variation in potential energy on the diffusion in heterogeneous media. Phys Lett A 380(22):1891–1894. https://doi.org/10.1016/j.physleta.2016.03.042
Ghysels A, Venable RM, Pastor RW, Hummer G (2017) Position-dependent diffusion tensors in Anisotropic Media from Simulation: Oxygen Transport in and through membranes. J Chem Theory Comput 13(6):2962–2976. https://doi.org/10.1021/acs.jctc.7b00039
Huang T, Li B, Wang H, Granick S (2022) Reply to “Comment on ‘Molecules, the Ultimate Nanomotor: linking Chemical reaction intermediates to their molecular Diffusivity’. ACS Nano 16(7):9977–9981. https://doi.org/10.1021/acsnano.2c04698
Amble E, Dailey BP (1950) The structure and Dipole Moment of Hydrazoic Acid. J Chem Phys 18(10):1422. https://doi.org/10.1063/1.1747504
Overton E (1899) Vierteljahrsschr Naturforsch Ges Zurich 1899(44):88–135
Missner A, Pohl P (2009) 110 years of the Meyer-Overton rule: predicting membrane permeability of gases and other small compounds. ChemPhysChem 10(9–10):1405–1414. https://doi.org/10.1002/cphc.200900270
Al-Awqati Q (1999) One hundred years of membrane permeability: does Overton still rule? Nat Cell Biol 1(8):E201–E202. https://doi.org/10.1038/70230
Bennion BJ, Be NA, McNerney MW, Lao V, Carlson EM, Valdez CA et al (2017) Predicting a drug’s membrane permeability: a computational model validated with in Vitro Permeability Assay Data. J Phys Chem B 121(20):5228–5237. https://doi.org/10.1021/acs.jpcb.7b02914
Lomize AL, Pogozheva ID (2019) Physics-based method for modeling Passive membrane permeability and translocation pathways of Bioactive Molecules. J Chem Inf Model 59(7):3198–3213. https://doi.org/10.1021/acs.jcim.9b00224
Wang Y, Gallagher E, Jorgensen C, Troendle EP, Hu D, Searson PC et al (2019) An experimentally validated approach to calculate the blood-brain barrier permeability of small molecules. Sci Rep 9(1):6117. https://doi.org/10.1038/s41598-019-42272-0
Bodor N, Buchwald P (1999) Recent advances in the brain targeting of neuropharmaceuticals by chemical delivery systems. Adv Drug Deliv Rev 36(2–3):229–254. https://doi.org/10.1016/s0169-409x(98)00090-8
Antalis TM, Palmer G (1982) Kinetic characterization of the interaction between cytochrome oxidase and cytochrome c. J Biol Chem 257(11):6194–6206
Bar-Even A, Noor E, Savir Y, Liebermeister W, Davidi D, Tawfik DS et al (2011) The moderately efficient enzyme: evolutionary and physicochemical trends shaping enzyme parameters. Biochemistry 50(21):4402–4410. https://doi.org/10.1021/bi2002289
Acknowledgements
We would like to thank Ms. Charlotte C. W. Taft for careful reading of the manuscript and linguistic corrections.
Funding
Slovenian Research Agency Grant P1-0012 (JM).
Author information
Authors and Affiliations
Contributions
SLH, SZ, VS, MSD, JM: Conceptualization. SLH, SZ, VS: Methodology. SLH, SZ, VS: Investigation. SLH, VS: Visualization. JM: Funding acquisition. MSD, VS, JM: Project administration. SZ, VS, MSD, JM: Supervision. SLH, VS, JM: Writing – original draft. SLH, SZ, VS, MSD, JM: Writing – review & editing.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
This article is dedicated to the memory of Herman J.C. Berendsen. One of us (JM) is very fortunate that Herman was his teacher and coworker for three years.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hartl, S.L., Žakelj, S., Dolenc, M.S. et al. How Azide Ion/Hydrazoic Acid Passes Through Biological Membranes: An Experimental and Computational Study. Protein J 42, 229–238 (2023). https://doi.org/10.1007/s10930-023-10127-3
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10930-023-10127-3