Abstract
Fast magic-angle spinning and partial sample deuteration allows direct detection of 1H in solid-state NMR, yielding significant gains in mass sensitivity. In order to further analyze the spectra, 1H detection requires assignment of the 1H resonances. In this work, resonance assignments of backbone HN and Hα are presented for HET-s(218–289) fibrils, based on the existing assignment of Cα, Cβ, C’, and N resonances. The samples used are partially deuterated for higher spectral resolution, and the shifts in resonance frequencies of Cα and Cβ due to the deuterium isotope effect are investigated. It is shown that the deuterium isotope effect can be estimated and used for assigning resonances of deuterated samples in solid-state NMR, based on known resonances of the protonated protein.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
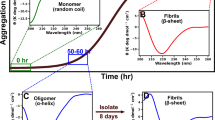
Amyloid fibrils are protein aggregates, which form long fibers that are rich in β-sheets and are stabilized by a network of hydrogen bonds between layers. Fibrils appear in many diseases including Alzheimer’s, Hungtington’s, and Parkinson’s (Knowles et al. 2014). However, they also serve functional roles such as hormone storage (Maji et al. 2009) and can provide structure and pigmentation (Fowler et al. 2007). The HET-s prion, the topic of this study, provides a means of programmed cell death, known as heterokaryon incompatability, in Podospora anserina (Glass and Kaneko 2003; Saupe 2000). The HET-s(218–289) prion is useful as a model amyloid, yielding at physiological conditions only one polymorph (Meier and Böckmann 2015) and producing well-resolved solid-state NMR spectra (Siemer et al. 2005). The assignment (13C, 15N resonances) and structure of HET-s(218–289) have been solved (Siemer et al. 2006; Van Melckebeke et al. 2010; Wasmer et al. 2008), excluding ~14 residues found in the termini and a loop that do not give strong NMR signals. Recently, we have also investigated site-specific dynamics of the HET-s amyloids, for which the assignment presented in this paper was used (Smith et al. 2016).
In this study, we expand the existing HET-s(218–289) assignment to include backbone 1H’s (Hα, HN). Fast magic-angle spinning (MAS, >50 kHz), in conjunction with partially-deuterated samples, gives sufficient resolution in the 1H dimension such that 1H detected experiments become useful (Agarwal et al. 2014; Huber et al. 2011; Knight et al. 2011; Lewandowski et al. 2011; Smith et al. 2016). For constant sample size, one then gains approximately eight times the total signal-to-noise ratio as compared to 13C detection, with relatively small losses due to an extra CP transfer step (Penzel et al. 2015). For HET-s(218–289) fibrils, it is possible to resolve most residues using Hα–Cα 2D correlation experiments or H–N correlation experiments, with partial protonation and fast MAS (60 kHz). For assignment of the Hα resonances, we have used partial protonation at the Cα, using 25% protonation, which has been shown to be adequate for randomly protonated samples (Asami et al. 2012). For assignment of the HN resonances, we used a fully deuterated sample, with 100% back-protonation on the HN and exchangeable side-chain protons. We furthermore re-assign the Cα, Cβ, C’, and N resonances that have shifted for the different samples—due to the deuterium isotope effect for 13C resonances, with additional shifting for 15N resonances due to pH and temperature. We calculate the deuterium isotope effect for 13C (Hansen 1988; Jameson and Osten 1986; Maciel et al. 1967), and use it for re-assignment of resonances. We also compare the deuterium isotope effect among different proteins and between solution- and solid-state NMR, and discuss use of calculations of the isotope effect for assignment of NMR spectra.
Theory
The existing resonance assignment of the 13C in fully protonated HET-s(218–289) (Siemer et al. 2006; Van Melckebeke et al. 2010), BMRB accession numbers 11028, 11064, allows us to utilize the deuterium isotope effect to estimate the 13Cα and 13Cβ resonance frequencies in partially deuterated samples. The deuterium isotope effect can be estimated as by (Jameson and Osten 1984).
Here, \({{\sigma }_{0}}\) and \(\sigma _{0}^{*}\) are the chemical shielding of some nucleus, with and without isotope substitution, respectively, and \(\left\langle \Delta {{r}_{i}} \right\rangle\) and \({{\left\langle \Delta {{r}_{i}} \right\rangle }^{*}}\) are the mean bond distance, before and after isotopic substitution. Then, the \({{(\partial \sigma /\partial \Delta {{r}_{i}})}_{e}}\) are the derivatives of the chemical shielding with respect to some bond distance, Δr i , in the molecule, and are, in the Born–Oppenheimer approximation, mass independent. As indicated, the total isotope effect is additive, and the \({{(\partial \sigma /\partial \Delta {{r}_{i}})}_{e}}\) are usually negative, and become smaller in magnitude for bonds further away from the nucleus of interest—so that when some bond is less extended, then the nucleus is more shielded. In this form, it is clear that the deuterium isotope effect is then the result of a decrease of the average bond distances, \(\left\langle \Delta {{r}_{i}} \right\rangle\), when a proton is substituted with the heavier deuterium atom, yielding increased chemical shielding (decrease in chemical shift).
Because the isotope effect is approximately additive, it is possible to reformulate Eq. (1) for the total shift for Cα or Cβ with the following formula (Hansen 1988, 2000):
Here, TΔC(D) is the total shift of the 13C due to the deuterium isotope effect (by convention, TΔC(D) is the difference of deuterated and the protonated sample, and is negative), nΔC(D) is the shift due to a deuterium n bonds away from the 13C, and d n is the number of deuterium atoms n bonds away (where contributions for n > 3 are negligible). Note that in this formulation \(^{n}\Delta \text{C}(\text{D})=-{{(\partial \sigma /\partial \Delta {{r}_{i}})}_{e}}\left[ {{\langle \Delta {{r}_{i}}\rangle }^{*}}-\langle \Delta {{r}_{i}}\rangle \right]\), where the ith bond is n bonds away. By assuming that the nΔC(D) values are approximately the same for different residues in a given protein, it is possible to fit the residue specific deuterium isotope effect for that protein to Eq. (2), in order to obtain the nΔC(D) (Venters et al. 1996). Note that the assumption that the nΔC(D) are the same for different residues is not entirely correct, with even structural information being contained in the isotope effect of the Cα resulting from deuteration at the amide position (Ottiger and Bax 1997), and further correlation between the isotope effect resulting from deuteration at the Cα position and structure (LeMaster et al. 1994; Morris and Murray 1975).
Materials and methods
Sample preparation of partially-protonated, [U–13C, 15N] HET-s fibrils
Partial Deuteration
Samples in this study were prepared to yield partial deuteration, with protonation of the Cα (referred to as “Hα sample”), the backbone N (referred to as “HN sample”), or protonation at both positions (referred to as “HαHN sample”). Obtaining the desired protonation for the HN sample is straightforward: before fibrilization, HET-s(218–289) is unstructured (Balguerie et al. 2003; Berthelot et al. 2010), and so expression in deuterated media followed by fibrilization in fully protonated solvent yields protonation at all exchangeable sites. Partial protonation at the Cα with reduced protonation elsewhere can be achieved by expression in 2H,13C glucose as the sole carbon source, and a mixture of D2O/H2O (Asami et al. 2010; Lundström et al. 2009). We use this approach, with a 75/25 ratio of D2O to H2O. This yields 25% protonation at the Cα. However, it will also cause variability in the protonation level at the nearby carbons. In Table 1, the protonation fraction at the Cβ is given for different residues, and also the probability that at least one proton occurs at the Cβ (Lundström et al. 2009). Although experiments used with the Hα and HαΗΝ samples select for Cα that are protonated, the variability in protonation level on nearby carbons will also give some variability in the chemical shift of those carbons and their neighbors due to the isotope effect.
Expression and purification
The Hα and HαHN samples are prepared using partial protonation at non-exchangeable sites using the above method, with fibrilization in D2O for the Hα sample to yield deuteration at all exchangeable sites, and fibrilization in H2O for the HαHN sample to yield protonation at all exchangeable sites. The HN sample is prepared with full deuteration except at exchangeable sites. All samples are uniformly 13C, 15N labeled, and all preparations follow standard expression steps that have been previously described (Balguerie et al. 2003), with variations in sample protonation. For all samples, Histidine6-tagged HET-s(218–289) was recombinantly expressed in Escherichia coli BL21 in M9 minimal medium, containing 2H,13C glucose (2.5 g/L) as the sole carbon source and 15NH4Cl (1 g/L) as the sole nitrogen source. Purification steps are always performed in H2O buffers. After expression, the cells were lysed in 150 mM NaCl and 50 mM Tris–HCl, pH = 8 and disrupted using a microfluidizer (Microfluidics). The lysate was centrifuged for 90 min at 8250 g. The pellet was resuspended in buffer (150 mM NaCl and 50 mM Tris–HCl, pH = 8). Guanidinium HCl powder was added until the solution had doubled in volume, and the sample was incubated overnight at 60 °C. The sample was then centrifuged for 3 h at 186,000 g until the supernatant was clear, and subsequently filtered (0.2 μm pore-size). Equilibrated Ni-Sepharose was added to bind to the Histidine-tagged sample and the protein was allowed to bind over 1 h to the resin. The sample was transferred to a column and the resin was washed, using a buffer (6 M guanidinium HCl, 20 mM Imidazole, 0.5 M NaCl, 50 mM Tris–HCl, pH = 8) with eight times the column volume. The protein was eluted using elution buffer (6 M guanidinium HCl, 500 mM Imidazole, 0.5 M NaCl, 50 mM Tris–HCl, pH = 8), while taking 1 mL fractions. The fractions with protein were pooled in one Falcon tube.
Fibrilization
For all samples, the buffer was exchanged with 150 mM acetic acid pH = 2.5 (the Hα sample uses acetic acid in D2O), using a HiPrep 26/10 desalting column (GE Healthcare). The Hα sample was lyophilized after elution and resuspended in D2O. This was repeated two more times, in order to remove all exchangeable protons from the Hα sample. For all samples, the pH was adjusted to pH = 7.4 by addition of 3 M Tris at 25 °C, triggering fibrilization. Aggregation of HET-s is visible after about 10 min, but is allowed to fully fibrilize for 3 days. Fibrils were then washed three times with D2O for the Hα sample, and with H2O for the other samples. Fibrils were finally washed with 50 mM citric acid at pH 5 (in D2O for the Hα sample and in H2O for the other samples). All samples were centrifuged into 1.3 mm (Bruker) rotors (Böckmann et al. 2009).
NMR spectroscopy
Solid-state NMR spectroscopy
All spectra were acquired on a Bruker Avance III 850 MHz spectrometer with MAS at 60 kHz, using a Bruker 1.3 mm triple-resonance probe. Sample temperature was 23.3 ± 0.3 °C for all spectra, determined by the resonance frequency of water, using referencing to DSS (Böckmann et al. 2009). Three-dimensional (3D) correlation experiments were used in this study, shown in Fig. 1. Transfer methods were chosen based on the work of Penzel et al. (Penzel et al. 2015), where it has been shown that for fast MAS (~90 kHz) that H–X and backbone C–N transfers are typically most efficient using adiabatic cross-polarization (CP)(Hediger et al. 1995), whereas C–C transfers are more efficient using J-coupling based transfers. Figure 1a shows the pulse sequence for HACACB correlation. The initial CP step transfers from Hα to Cα, followed by a J-coupled transfer from Cα (in-phase) to Cβ (anti-phase), while simultaneously evolving on the Cα spin. After a π/2 pulse, magnetization then evolves on Cβ, followed by transfer back to Cα, water suppression using the MISSISSIPPI sequence (Zhou and Rienstra 2008), and finally CP from Cα to Hα. Figure 1b shows the pulse sequence for HACAN correlation ((H)NCAHA), which uses CP transfers at each step, and Fig. 1c shows the pulse sequence for HNCA and HNCO correlation [(H)CANH, (H)CONH], which also uses CP transfers at each step. Experimental parameters for each experiment are shown in Table 2. The signal to noise and the number of assigned peaks for each 3D experiment are also given in Table 2.
3D pulse sequences a is used for HACACB correlation. t 1, t 2, and t 3 correspond to evolution on Cα, Cβ, and Hα, respectively. Selective pulses are all set to excite the aliphatic carbons (Cα/Cβ in particular) without excitation of C’ b is used for HACAN correlation, with t 1, t 2, and t 3 corresponding to evolution on N, Cα, and Hα. c is used for HNCA and HNCO correlation, with t 1, t 2, and t 3 corresponding to evolution on Cα or CO, N, and HN
Assignment
The chemical shifts of the Cα and Cβ resonances were estimated by calculating the expected change (TΔC(D)) of resonance frequency with respect to the protonated protein for each atom using Eq. (2). Values used for the parameters nΔC(D) were those determined for solution-state α-synuclein, an intrinsically unstructured protein (Maltsev et al. 2012). For the Hα and HαHN sample, d 1 is zero for Cα: although there is only 25% protonation at the Cα position, the pulse sequence only correlates Cα which are protonated and therefore have no one-bond isotope effect. Also, d 2 and d 3 for Cα in the Hα sample include contributions from one deuterium each from the backbone amides, in addition to contributions from deuterium on the side chain. Note that in this study, we do not explicitly account for residual side chain protonation since variation in deuteration of the side chain did not yield resolved peaks. In principle, it might be possible to use fractional values of the d n to calculate average chemical shifts, however the transfer efficiency of the pulse sequences will be reduced at higher protonation levels, complicating the calculation of the d n [see (Nietlistpach et al. 1996) for discussion of averaged quantities in cases of partial deuteration]. The HN sample is deuterated except at exchangeable positions so that the Cα d 1 is one, and d 2 and d 3 only have contributions from the side chain. The TΔC(D) were added to the 13C resonances found in the assignment of protonated HET-s(218–289), given in BMRB ascension number 11064 (Siemer et al. 2006; Van Melckebeke et al. 2010). 15N resonances were estimated by shifting the existing resonance assignment by −1 ppm (shifts due to a combination of the isotope effect, in addition to pH and temperature differences).
Using these estimates, it was then possible to search in the acquired 3D spectra for resonances near to the estimated values. This procedure allowed identification of the isotopically shifted 15N and 13C resonances and assignment of the Hα and HN resonances. The HACACB correlation spectrum (Fig. 1a) was used to re-assign most Cα and Cβ resonances for the Hα and HαHN samples, along with assignment of the Hα resonances. Glycine Hα and Cα assignments as well as several ambiguous assignments were determined for the Hα and HαHN samples using the HACAN correlation spectrum (Fig. 1b). The HACAN spectrum was also used to assign the N resonances in the Hα and HαHN samples. The Hα and Cα resonance assignments are plotted on a 2D HACA correlation spectrum shown in Fig. 2, where peaks are marked at the assigned position [average of positions from 2D and 3D spectra, as calculated by CCPN (Vranken et al. 2005)]. Some peaks are not resolved in the 2D spectra, and near the termini or loops, may appear only weakly. In this case, the position has been taken only from the 3D spectra, but is also marked on the 2D spectrum (signal to noise and number of assigned peaks in the 3D spectra is included in Table 2). Note that several glycines are assigned to two peaks in the spectrum, due to different resonance frequencies for each of the two Hα protons. The HNCA correlation spectrum (Fig. 1c) was used to re-assign Cα and N resonances for the HN sample, and determine the HN assignment. The HN and N resonance assignments are plotted on a 2D HN correlation spectrum shown in Fig. 3, where peaks are marked at the assigned position. An HNCO experiment was finally used to re-assign the C’ resonances in the HN sample, although no systematic change of the C’ resonances is expected, since no significant isotope affect has been seen in solution-state NMR (Gardner and Kay 1998). The full assignment has been deposited in the BMRB under ascension number 26913.
Results and discussion
By using an estimation of the 13Cα and 13Cβ shifts using Eq. (1), as well as a constant offset of 15N shifts, we were able to re-assign the partially deuterated samples without using a backbone walk. In all, 93% of residues in the Hα and HαΗΝ samples were assigned and 83% of residues in the HN sample residues were assigned, for which the backbone resonances are known from the assignment of protonated HET-s(218–289). The remaining peaks were not visible in the 3D experiments, and so could not be assigned. The full assignment plot is shown in Fig. 4.
Assignment plot for HN and Hα resonances. In a, the resonance assignment is shown for the assignment of the HN, along with re-assignment of N, Cα, and C’ for the partially deuterated sample (HN sample). In b, the resonance assignment is shown for the assignment of Hα, along with re-assignment of the N, Cα, and Cβ resonances (Hα and HαHN samples). Note that parts of the termini and loop (218–220, 250–259, and 288–289) are excluded from the plot
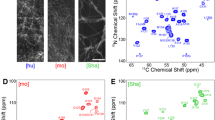
The estimated 13Cα and 13Cβ deuterium isotope effect is compared with the experimentally measured isotope effect in Fig. 5a–c. To further characterize the deuterium isotope effect, the experimentally determined change in the 13Cα and 13Cβ resonance frequencies were then re-fitted to Eq. (2) to obtain new values of the nΔC(D). Resonances from the Hα and HN samples were included in the fit, yielding 1ΔC(D) = −290 ± 20 ppb, 2ΔC(D) = −90 ± 10 ppb, and 3ΔC(D) = −10 ± 8 (95% confidence interval given). One further notes in Fig. 5d, that there is no clear trend in the change of C’ resonances due to deuteration, therefore when calculating the above nΔC(D), only the experimental deuterium isotope effect of the Cα and Cβ nuclei were included. For 15N resonances (Fig. 5e, f), there is a large overall change in resonance frequency, although this is larger than expected due to only the deuterium isotope effect, and is likely due to differences in pH and temperature as compared to the assignment of protonated HET-s.
Changes in chemical shift. a–d Show the changes in 13C chemical shift resulting primarily from partial deuteration of the sample (colored bars). a and c Show shifts for the Hα sample, for which Cα shifts due to deuteration two or more bonds away (pulse sequence only selects protonated Cα), and Cβ shifts both due to one- and multi-bond isotope effect. b Shows the Cα shift in the HN sample, for which Cα shift also includes a one-bond isotope effect. d Shows the shift of C’ in the HN sample, where only small shifts are observed. Predicted isotope effect is shown in a–c using Eq. (2) with values of nΔC(D) taken from α-synuclein, shown as black dots. e and f Show the shifts for 15N, however these are affected by pH and temperature in addition to the isotope effect and so were not fitted
The values of the nΔC(D) determined from HET-s(218–289) are similar to those previously determined for other proteins using solution-state or solid-state NMR. Table 3 compares the values of the nΔC(D) obtained for HET-s(218–289) to those previously determined for solution-state α-synuclein (Maltsev et al. 2012), Human carbonic anhydrase (Venters et al. 1996), and SH2 C-terminal domain (Gardner et al. 1997). In order to determine if there is any clear distinction between the isotope effect in the solution-state and the solid-state, we also calculate the nΔC(D) using previously published data for Ubiquitin (Penzel et al. 2015; Schubert et al. 2006) and GB1(Tang et al. 2010). In fact, one finds that the values for the different proteins cannot be said to be different at the 95% confidence level (with the exception that 3ΔC(D) for HET-s and Human carbonic anhydrase are significantly different). Furthermore, there is no apparent difference in the deuterium isotope effect for solid- and solution-state NMR. Note that previous studies have shown that some structural information is contained in the deuterium isotope effect, but the accuracy obtained here is insufficient to conclusively show correlation between structure and the isotope effect (Albildgaard et al. 2009; LeMaster et al. 1994; Morris and Murray 1975; Ottiger and Bax 1997).
The similarity of the nΔC(D) for multiple proteins suggests that the approach used here should be a fairly robust method for re-assignment of partially deuterated proteins in solid-state NMR. In our assignment, we first estimated the Cα and Cβ chemical shifts using the nΔC(D) determined from α-synuclein. However, given some variability for different proteins, it is not clear how important is the choice of reference values for the nΔC(D). In Fig. 6, we use the nΔC(D) for each protein in Table 3, and plot the offset from the calculated Cα and Cβ shifts to the correct value for all assigned residues. As expected, when using the values derived from HET-s(218–289), the deviation from the correct value is smallest. However, using values derived from α-synuclein, SH2 C-terminal domain, Ubiquitin, and GB1 yields only slightly higher error. Values from Human carbonic anhydrase do deviate significantly, although it is not clear why this is the case. This indicates that for most proteins, the choice of values for the nΔC(D) is not critical. Also in Fig. 6, for each assigned residue we plot the offset in the Cα and Cβ dimensions to the nearest neighboring Cα/Cβ resonance. Therefore, when the distance to the nearest neighbor is smaller than errors arising in the prediction of the deuterium isotope effect, it is possible for assignment ambiguity to arise—but the plot does not indicate significant change in ambiguity depending on the reference protein chosen.
Scatter plots of Cα and Cβ resonance estimation accuracy. In each plot, the HET-s(218–289) Cα and Cβ shifts for the Hα sample are estimated using (2) and the nΔC(D) derived from each of the six proteins in Table 3. The difference of the correct shifts and the calculated shifts are plotted on the x- and y-axes for Cα and Cβ shifts, respectively, for each assigned residue (red circles). a Also plots the difference of the shifts of the partially deuterated sample and the protonated sample without correction (blue +). Each plot also shows the distance of each resonance to the nearest neighboring resonance (black x), so that one expects assignment ambiguities where the nearest neighbors are closer to the origin than some of the estimated shifts. Note that although the nΔC(D) from HET-s (a) give the best estimation, it is not much better than estimations using the other protein data (excepting Human carbonic anhydrase)
Thus far, we have neglected the statistical distribution of deuteration in the Hα sample (see Table 1). As stated above, the pulse sequence selects only protonated Cα, but deuteration at Cβ (≤25% protonation) and more distant carbons will vary (Lundström et al. 2009). The consequence is that the same 13C nucleus on different HET-s molecules will have different numbers of bonded protons, having potentially zero, one, or two bonded protons. Each of these configurations will lead to a different chemical shift for that 13C, and furthermore will affect chemical shifts of neighboring 13C. The result is that a distribution of chemical shifts is actually measured for any given resonance. In principle, this effect should be more pronounced for Cβ resonances (compared to Cα), although we are not able to see this directly via separated resonances or clear shoulders on the measured resonances. For several residues, we are however able to measure both an Hα-Cα-Cβ correlation and an Hβ-Cβ-Cα correlation; for the Hα-Cα-Cβ correlation, the Cα is protonated and the Cβ is deuterated, whereas in the latter case, the Cβ is protonated and the Cα is deuterated. This results in the Cα shifting to lower ppm values and the Cβ shifting to higher ppm values, by |1ΔC(D)−2ΔC(D)| = 200 ppb. This is shown in Fig. 7 for several residues, where the average change in shift of 189 ppb is in good agreement with the expected shift. Although separated resonances or shoulders are not visible for the individual peaks, variability in the degree of deuteration between different HET-s(218–289) molecules may broaden the Cα and Cβ resonances in the Hα and HαHN samples. Also note that our fit of the one-, two-, and three- bond isotope effect (1ΔC(D), 2ΔC(D), 3ΔC(D)) is likely to be slightly biased because we have not considered that different molecules in the same sample can have some variability in the deuteration level.
Comparison of peak positions for Hα-Cα-Cβ (red) and Hβ-Cβ-Cα (blue) correlations for selected residues. Peaks that are detected at the Hα resonance frequency (see Fig. 1a for transfer sequence) do not have a one-bond isotope effect on the Cα (δ1, red), but have a one-bond isotope effect on Cβ (δ2, red), and likewise, peaks detected at the Hβ resonance frequency have a reduced one-bond isotope effect at the Cβ position (δ1, blue), but have a one-bond isotope effect at the Cα position (δ2, red). Therefore, when comparing the Cα–Cβ cross peaks at the Hα and Hβ positions from the same spectrum, one finds that the Cα has shifted to lower ppm values and the Cβ has shifted to higher ppm values. Note that the peaks appear in opposite quadrants of the C–C plane; here, the 13C axes of the Hβ-Cβ-Cα correlation have been switched to overlay the peaks. Additional broadening in the δ2 dimension is due to a shorter acquisition time (see Table 2)
We have not analyzed 15N data for the deuterium isotope effect, because differences in pH [5.0 in this study, 7.5 in (Van Melckebeke et al. 2010)], and also temperature differences convolute the chemical-shift effects. The differences in 15N chemical shift are nonetheless plotted in Fig. 5d, e, for the HαHN and HN samples, respectively, showing nearly the same average change in the chemical shift and usually giving similar trends between the two samples.
Conclusions
Hα and HN resonances have been assigned for partially protonated HET-s(218–289) samples. Additionally Cα, Cβ, and N resonances have been corrected for isotope effects as compared to previous assignments (Siemer et al. 2006; Van Melckebeke et al. 2010). Changes in the Cα and Cβ resonances could be attributed primarily to the deuterium isotope effect, and therefore it was possible to calculate the deuterium isotope effect and approximate the Cα and Cβ resonance frequencies, then use these for assignment of the partially deuterated samples. After reassignment, the deuterium isotope effect in HET-s(218–289) yielded values for the nΔC(D) that are in good agreement with values previously reported in the literature. Relative uniformity of the isotope effect for different proteins simplifies the choice of reference nΔC(D) used in estimating chemical shifts.
References
Agarwal V, Penzel S, Szekely K, Cadalbert R, Testori E, Oss A, Past J, Samoson A, Ernst M, Böckmann A, Meier BH (2014) De novo 3D structure determination from sub-milligram protein samples by solid-state 100 kHz MAS NMR spectroscopy. Angew Chem Int Ed 53:12253–12256
Albildgaard J, Hansen PE, Manalo ME, LiWang A (2009) Deuterium isotope effects on N-15 backbone chemical shifts in proteins. J Biomol NMR 44:119–126
Asami S, Schmieder P, Reif B (2010) High resolution 1H-detected Solid-state NMR spectroscopy of protein aliphatic resonances: access to tertiary structure information. J Am Chem Soc 132:15133–15135
Asami S, Szekely K, Schanda P, Meier BH, Reif B (2012) Optimal degree of protonation for 1H detection of aliphatic sites in randomly deuterated proteins as a function of the MAS frequency. J Biomol NMR 54:155–168
Balguerie A, Dos Reis S, Ritter C, Chaignepain S, Bénédicte C-S, Forge V, Bathany K, Lascu I, Schmitter J-M, Riek R, Saupe SJ (2003) Domain organization and structure–function relationship of the HET-s prion protein of Podospora anserina. EMBO J 22:2071–2081
Berthelot K, Lecomte S, Gean J, Immel F, Cullin C (2010) A yeast toxic mutant of HET-s(218–289) prion displays alternative intermediates of amyloidogenesis. Biophys J 99:1239–1246
Böckmann A, Gardiennet C, Verel R, Hunkeler A, Loquet A, Pintacuda G, Emsley L, Meier BH, Lesage A (2009) Characterization of different water pools in solid-state NMR protein samples. J Biomol NMR 45:319–327
Fowler DM, Koulov AV, Balch WE, Kelly JW (2007) Functional amyloid —from bacteria to humans. Trends Biochem Sci 32:217–224
Gardner KH, Kay LE (1998) The use of 2 H, 13 C, 15 N multidimensional NMR to study the structure and dynamics of proteins. Annu Rev Biophys Biomol Struct 27:357–406
Gardner KH, Rosen MK, Kay LE (1997) Global folds of highly deuterated, methyl-protonated proteins by multidimensional NMR. Biochemistry 36:1389–1401
Glass NL, Kaneko I (2003) Fatal attraction: nonself recognition and heterokaryon incompatibility in filamentous fungi. Eukaryot Cell 2:1–8
Hansen PE (1988) Isotope Effects in Nuclear Shielding. Prog Nucl Mag Res Sp 20:207–255
Hansen PE (2000) Isotope effects on chemical shifts of proteins and peptides. Magn Reson Chem 38:1–10
Hediger S, Meier BH, Ernst RR (1995) Adiabatic passage Hartmann-Hahn cross polarization in NMR under magic angle sample spinning. Chem Phys Lett 240:449–456
Huber M, Hiller S, Schanda P, Ernst M, Böckmann A, Verel R, Meier BH (2011) A proton-detected 4D solid-state nmr experiment for protein structure determination. Chem Phys Chem 12:915–918
Jameson CJ, Osten HJ (1984) The additivity of NMR isotope shifts. J Chem Phys 81:4293
Jameson CJ, Osten HJ (1986) Theoretical aspects of isotope effects on nuclear shielding. Ann R NMR S 17:1–78
Knight MJ, Webber AL, Pell AJ, Guerry P, Barbet-Massin E, Bertini I, Felli IC, Gonnelli L, Pierattelli R, Emsley L, Lesage A, Herrmann T, Pintacuda G (2011) Fast resonance assignment and fold determination of human superoxide dismutase by high-resolution proton-detected solid-state MAS NMR spectroscopy. Angew Chem Int Ed Engl 50:11697–11701
Knowles TPJ, Vendruscolo M, Dobson CM (2014) The amyloid state and its association with protein misfolding diseases. Nat Rev Mol Cell Bio 15:384–396
LeMaster DM, LaIuppa JC, Kushlan DM (1994) Differential deuterium isotope shifts and one-bond 1H-13C scalar couplings in the conformational analysis of protein glycine residues. J Biomol NMR 4:863–870
Lewandowski JR, Dumez J-N, Akbey Ü, Lange S, Emsley L, Oschkinat H (2011) Enhanced resolution and coherence lifetimes in the solid-state NMR spectroscopy of perdeuterated proteins under ultrafast magic-angle spinning. J Phys Chem Lett 2:2205–2211
Lundström P, Hansen DF, Vallurupalli P, Kay LE (2009) Accurate measurement of alpha proton chemical shifts of excited protein states by relaxation dispersion NMR spectroscopy. J Am Chem Soc 131:1915–1926
Maciel GE, Ellis PD, Hofer DC (1967) Carbon-13 chemical shifts of the carbonyl group. V. Observation of a deuterium isotope effect using carbon-13 field-frequency lock. J Phys Chem 71:2160–2164
Maji SK, Perrin MH, Sawaya MR, Jessberger S, Vadodaria K, Rissman RA, Singru PS, Nilsson KPR, Simon R, Schubert D, Eisenberg D, Rivier J, Sawchenko P, Wylie V, Riek R (2009) Functional amyloids as natural storage of peptide hormones in pituitary secretory granules. Science 325:328–332
Maltsev AS, Ying J, Bax A (2012) Deuterium isotope shifts for backbone 1 H, 15 N and 13 C nuclei in intrinsically disordered protein a-synuclein. J Biomol NMR 54:181–191
Meier BH, Böckmann A (2015) The structure of fibrils from ‘misfolded’ proteins. Curr Opin Struc Biol 30:43–49
Morris DG, Murray AM (1975) 13 C Chemical Shifts of I-Substituted Camphenes. Perkin Trans 2:539–541
Nietlistpach D, Clowes RT, Broadhurst RW, Ito Y, Keeler J, Kelly M, Ashurst J, Oschkinat H, Domaille PJ, Laue ED (1996) An approach to the structure determination of larger proteins using triple resonance NMR experiments in conjunction with random fractional deuteration. J Am Chem Soc 118:407–415
Ottiger M, Bax A (1997) An empirical correlation between amide deuterium isotope effects on 13Ca chemical shifts and protein backbone conformation. J Am Chem Soc 119:8070–8075
Penzel S, Smith AA, Agarwal V, Hunkeler A, Org M-L, Samoson A, Böckmann A, Ernst M, Meier BH (2015) Protein resonance assignment at MAS frequencies approaching 100 kHz: a quantitative comparison of J-coupling and dipolar—coupling-based transfer methods. J Biomol NMR 63:165–186
Saupe SJ (2000) Molecular genetics of heterokaryon incompatibility in filamentous ascomycetes. Microbiol Mol Biol Rev 64:489–502
Schubert M, Manolikas T, Rogowski M, Meier BH (2006) Solid-state NMR spectroscopy of 10 % 13 C labeled ubiquitin: spectral simplification and stereospecific assignment of isopropyl groups. J Biomol NMR 35:167–173
Siemer AB, Ritter C, Ernst M, Riek R, Meier BH (2005) High-resolution solid-state NMR spectroscopy of the prion protein HET-s in its amyloid conformation. Angew Chem Int Ed 44:2441–2444
Siemer AB, Ritter C, Steinmetz MO, Ernst M, Riek R, Meier BH (2006) 13 C, 15 N resonance assignment of parts of the HET-s prion protein in its amyloid form. J Biomol NMR 34:75–87
Smith AA, Testori E, Cadalbert R, Meier BH, Ernst M (2016) Characterization of fibril dynamics on three timescales by solid-state NMR. J Biomol NMR 65:171–191
Tang M, Comellas G, Mueller LJ, Rienstra CM (2010) High resolution 13C-detected solid-state NMR spectroscopy of a deuterated protein. J Biomol NMR 48:103–111
Van Melckebeke H, Wasmer C, Lange A, AB E, Loquet A, Böckmann A, Meier BH (2010) Atomic-resolution three-dimensional structure of HET-s(218–289) amyloid fibrils by solid-state NMR spectroscopy. J Am Chem Soc 132:13765–13775
Venters RA, Farmer BT II, Fierke CA, Spicer LD (1996) Characterizing the use of perdeuteration in NMR studies of large proteins: 13 C, 15N and 1H assignments of human carbonic anhydrase II. J Mol Biol 264:1101–1116
Vranken WF, Boucher W, Stevens TJ, Fogh RH, Pajon A, Llinas M, Ulrich EL, Markley JL, Ionides J, Laue ED (2005) The CCPN data model for NMR spectroscopy: development of a software pipeline. Proteins 59:687–696
Wasmer C, Lange A, Van Melckebeke H, Siemer AB, Riek R, Meier BH (2008) Amyloid fibrils of the HET-s(218–289) prion form a β solenoid with a triangular hydrophobic core. Science 319:1523–1526
Zhou DH, Rienstra CM (2008) High-performance solvent suppression for proton detected solid-state NMR. J Magn Res 192:167–172
Acknowledgements
This work has been supported by the Swiss National Science Foundation (Grants 200020_146757 and 200020_159707) and by the French ANR (ANR-14-CE09-0024B).
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Smith, A.A., Ravotti, F., Testori, E. et al. Partially-deuterated samples of HET-s(218–289) fibrils: assignment and deuterium isotope effect. J Biomol NMR 67, 109–119 (2017). https://doi.org/10.1007/s10858-016-0087-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10858-016-0087-0