Abstract
Pseudocontact shift (PCS) effects induced by a paramagnetic lanthanide bound to a protein have become increasingly popular in NMR spectroscopy as they yield a complementary set of orientational and long-range structural restraints. PCS are a manifestation of the χ-tensor anisotropy, the Δχ-tensor, which in turn can be determined from the PCS. Once the Δχ-tensor has been determined, PCS become powerful long-range restraints for the study of protein structure and protein–ligand complexes. Here we present the newly developed package Numbat (New User-friendly Method Built for Automatic Δχ-Tensor determination). With a Graphical User Interface (GUI) that allows a high degree of interactivity, Numbat is specifically designed for the computation of the complete set of Δχ-tensor parameters (including shape, location and orientation with respect to the protein) from a set of experimentally measured PCS and the protein structure coordinates. Use of the program for Linux and Windows operating systems is illustrated by building a model of the complex between the E. coli DNA polymerase III subunits ε186 and θ using PCS.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Paramagnetic lanthanide ions bound to the natural metal-binding site of a metalloprotein or introduced via a lanthanide tag provide a number of paramagnetic effects that can be distance dependent (i.e. paramagnetic relaxation enhancement), orientation dependent (i.e. residual dipolar couplings, RDC), or a combination of both, like cross-correlated relaxation effects and pseudocontact shifts (PCS; Bertini et al. 2002; Pintacuda et al. 2004). PCS present particularly valuable structural restraints, as they are easy to measure and provide long-range information that would be difficult to obtain by other techniques. PCS originate from unpaired electron spins which lead to an anisotropic magnetic susceptibility tensor (χ-tensor). PCS restraints induced by lanthanide ions have been used to investigate structural and dynamical properties of proteins (Allegrozzi et al. 2000; Bertini et al. 2001, 2004; Gaponenko et al. 2004; Jensen et al. 2006; Eichmüller and Skrynnikov 2007; Wang et al. 2007) and protein–ligand complexes (John et al. 2006; Pintacuda et al. 2007).
In order to apply PCS restraints, eight variables have to be determined. These comprise the lanthanide position (three Cartesian coordinates), three angles (e.g. Euler angles) that relate the molecular frame to the χ-tensor frame, and the axial and rhombic anisotropy parameters of the χ-tensor. (Since PCS depend only on the χ-tensor anisotropy Δχ rather than the absolute magnitude of the χ-tensor, it is sufficient to determine the anisotropy parameters represented by the Δχ-tensor.) Several integrated software tools are available for the determination and study of the alignment tensor using RDCs (Dosset et al. 2000; Zweckstetter and Bax 2000; Valafar and Prestegard 2004; Wei and Werner 2006). For the situation where the 3D structure of the protein is known a priori, corresponding tools for the determination of the Δχ-tensor from PCS have been developed but are more limited in scope. The program Fantasia (Banci et al. 1996) and its extension Fantasian (Banci et al. 1997) can fit the magnitude and Euler angles of the Δχ-tensor using a set of experimental PCS but requires prior knowledge of the metal coordinates. The program Platypus (Pintacuda et al. 2004) can simultaneously fit the Δχ-tensor and assign the signals of 15N-HSQC spectra of samples containing diamagnetic and paramagnetic lanthanides, but assumes that the 15N-HSQC peaks are sufficiently well resolved such that the paramagnetic peaks can be unambiguously associated with their diamagnetic partners. The program Echidna (Schmitz et al. 2006) uses assigned diamagnetic 15N-HSQC cross-peaks of a uniformly 15N-labelled protein to determine the magnitude and Euler angles of the Δχ-tensor and, simultaneously, the assignment of the paramagnetic 15N-HSQC cross-peaks. It also requires prior knowledge of the approximate metal ion position. In principle, the structure refinement packages Xplor-NIH (Schwieters et al. 2003, 2006) with the module PARArestraint for Xplor-NIH (Banci et al. 2004), GROMACS (Van der Spoel et al. 2005) with an implementation of orientation restraints (Hess and Scheek 2003), or DYANA (Güntert et al. 1997) with the module PSEUDYANA (Banci et al. 1998) could be used for Δχ-tensor determination from PCS but the protocols would be cumbersome. Considering that simultaneous determination of the Δχ-tensor and metal ion position relative to a known protein structure is a commonly required task, we set out to design a tool to achieve this in an easier and user-friendly way.
While the metal coordinates of metalloproteins can be accurately determined by crystallography, the metal position must be fitted when no crystal structure is available, e.g., when the lanthanide is introduced via a lanthanide tag. None of the reported tools addresses this issue. Here we present the newly developed program Numbat (New User-friendly Method Built for Automatic Δχ-Tensor determination), which can simultaneously fit the Δχ-tensor and lanthanide coordinates using experimental PCS values and the coordinates of the protein. Furthermore, the program encompasses a number of useful tools for multiple data sets recorded with different paramagnetic lanthanides, for rigid-body docking using PCS, and for analysis and visualization of the results. Following a description of the algorithm on which the program builds and a presentation of the graphical user interface (GUI), we illustrate the use of Numbat for building the model of a complex in a rigid-body docking approach using PCS.
Algorithm
The Δχ-tensor can be determined and refined by the comparison between experimentally determined PCS values and PCS values back-calculated from the atomic coordinates of the molecular structure (Sherry and Pascual 1977; Lee and Sykes 1983; Emerson and La Mar 1990; Veitch et al. 1990; Banci et al. 1992; Capozzi et al. 1993). The pseudocontact shift of a nuclear spin i, PCS calc i , is given by (Bertini et al. 2002):
where \( \tilde x_i \), \( \tilde y_i \), \( \tilde z_i \) are the Cartesian coordinates of the nuclear spin i in the Δχ-tensor frame, r i is the distance between the spin i and the paramagnetic centre, and Δχax and Δχrh are the axial and rhombic components of the Δχ-tensor. The orientation of the Δχ-tensor frame with respect to the protein frame can be specified, e.g., by three Euler angles α, β and γ.
To quantify the difference between experimental and back-calculated PCS values we define a quadratic cost c:
where PCS i exp is the experimental PCS for the spin i, and tol i is its associated tolerance. The tolerance values can be used to reflect different uncertainties in the measurement of different PCS. When the lanthanide position is known, only five Δχ-tensor parameters have to be optimized. In this case, the least square fitting problem is linear, as can be seen from an alternate formulation of the PCS (Bertini et al. 2002):
where x i , y i , z i are the Cartesian coordinates of the spin i in an arbitrary frame f and Δχxx, Δχyy, Δχzz, Δχxy, Δχxz, Δχyz are the Δχ-tensor components in this frame. The Singular Value Decomposition (SVD) algorithm, which is commonly used to determine an alignment tensor from a set of experimental RDC (Valafar and Prestegard 2004; Wei and Werner 2006), would be a good candidate to minimize the cost c. The least square fitting, or the Simplex algorithm (Nelder and Mead 1965) has been applied in previous work (Emerson and La Mar 1990; Capozzi et al. 1993). However the most general problem one has to solve is non-linear since the metal ion position may be unknown. We consequently chose for the non-linear least square fitting procedure in Numbat the Levenberg–Marquardt algorithm (Marquardt 1963) as implemented in the GNU Scientific Library (Galassi et al. 2006).
Program features
GUI
The GUI of Numbat was built with the GTK + library (Krause 2007) that is commonly available on recent Linux systems. Figure 1 shows two screenshots of the main interface of Numbat illustrating the intuitive and flexible user interface.
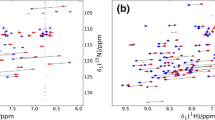
Screenshots of Numbat main windows. (a) Graphical User Interface for the tab Input Data. Four PCS data sets can be loaded simultaneously under the tabs PCS1 to PCS4. The list of all atoms is displayed in the main frame and can be filtered with the Display button to show only the atom or residue types of interest. The experimental PCS and the tolerance can be directly modified, and only atoms that are selected (see the column labelled “Use?”) are taken into account in the calculations. The distance between the respective atom to the metal ion, the calculated PCS and the deviation between experimental and predicted PCS are calculated and displayed after each fitting procedure. (b) Graphical User Interface for Tensor Fit. A Δχ-tensor can be fitted for each of the data sets PCS1 to PCS4. An additional tab (Multiple PCS) is for simultaneous fitting different data sets that share the same metal-ion centre. The frame Select conformers allows the choice of the model(s) to be used from a family of conformers loaded. The Tensor search restraints frame allows the individual selection of each of the eight variables to be free, fixed or constrained between two values. The computed Δχ-tensor values are displayed with error estimates from the GSL implementation of the Levenberg–Marquardt algorithm and the corresponding unique tensor representation (“UTR”) is reported
Input files
Numbat reads atomic coordinates from protein data bank (PDB; Berman et al. 2000) files. In the case of NMR structures, the entire ensemble of conformers is loaded and any subset can be selected for subsequent calculations. When optimizing the Δχ-tensor, PCS are back-calculated for each selected structure and averaged for the computation of the cost function c (Eq. 2). PCS data can be read either in the Xplor-NIH format or in a format specific to Numbat. For test purposes, Numbat also allows the generation of PCS data (optionally with addition of Gaussian noise) for a user-specified Δχ-tensor.
Methyl group definition
The 1H chemical shift of a rotating methyl group can be described as the average of the chemical shifts of the three 1H spins. The selection “methyl association” in the GUI allows definition of pseudoatom names for any methyl group for which the experimental PCS value is to be treated as the average of the PCS of the three 1H nuclei. The pseudoatom names can be used to identify the experimental PCS values of methyl groups in the input file. Alternatively, the PCS values of methyl groups can be interactively entered via the user-interface.
Optimization of the tensor parameters
In order to give the user a maximum of flexibility, any subset of the eight Δχ-tensor variables can be optimized with the remaining ones fixed to user-specified values. Such a situation occurs, for example, when a protein–ligand complex is studied where the protein is tagged with a lanthanide. First, the Δχ-tensor can be determined using the PCS measured for the protein. Fitting of the position and orientation of the Δχ-tensor with respect to the ligand can subsequently be performed with a minimal number of adjustable parameters by keeping the axial and rhombic components of the Δχ-tensor fixed at the values determined for the protein. The Δχ-tensors determined for the protein and the ligand can finally be superimposed to derive a model of the protein–ligand complex (Pintacuda et al. 2007).
Numbat also offers the option of restricting the Δχ-tensor variables within user-defined boundaries. This is useful if the magnitude, position and/or orientation of the Δχ-tensor is approximately known from previous studies (Su et al. 2008). Depending on the quality and quantity of PCS measurements available, the Δχ-tensor variables (especially the lanthanide coordinates) may only reach a local minimum during the optimization procedure. Therefore the starting values of all Δχ-tensor variables used to initialize the minimizer can be changed interactively within Numbat.
Residual Anisotropic Chemical Shifts (RACS)
Paramagnetic lanthanides bound to the protein weakly align the molecule in the magnetic field resulting in an incomplete averaging of the anisotropic chemical shifts. This can affect the PCS by a shift of up to 0.2 ppm for backbone 15N and 13C′ spins at a magnetic field of 18.8 T (John et al. 2005). The RACS correction term ΔδRACS for 1HN, backbone 15N and 13C′ spins can be calculated given the Δχ-tensor and the chemical shielding anisotropic tensor (CSA-tensor) using (John et al. 2005):
where B 0 is the magnetic field, μ 0 the induction constant, k the Boltzmann constant, T the temperature, σ CSA ii the principal components of the CSA-tensor, cosθ ij the nine direction cosines between pairs of the principal axis of the Δχ-tensor and the CSA-tensor, and Δχ jj the principal components of the Δχ-tensor. Numbat optionally uses the RACS correction term when generating PCS data and fitting Δχ-tensors. The orientations of the principal component axes of the nuclear CSA-tensors and the σ CSA ii values for 1HN, backone 15N and 13C′ spins are taken from Cornilescu and Bax (2000).
Multiple PCS data sets
A new PCS data set can be obtained by replacing one paramagnetic lanthanide with another paramagnetic lanthanide. Multiple PCS data sets obtained in this way share a conserved lanthanide position, but different orientations and magnitudes of the Δχ-tensors must be fitted to each individual PCS data set. Numbat can perform a simultaneous fit of the Δχ-tensors and the shared lanthanide position. This feature is of particular interest when only a limited number of PCS can be measured for each lanthanide ion, as fewer variables in the Δχ-tensor fit will facilitate the determination of accurate Δχ-tensor parameters. For example, a limited set of unambiguously measured PCS can be used to determine initial Δχ-tensor parameters from which the PCS of unassigned paramagnetic cross-peaks can be back-calculated, leading to assignments of additional paramagnetic cross-peaks and improved Δχ-tensor parameters. Similarly, applications to small ligand molecules with a small number of NMR signals are aided by limiting the number of adjustable variables to a minimum.
PCS modification
Once an initial Δχ-tensor has been fitted, Numbat computes and displays PCS values for all atoms. Doubtful assignments can easily be detected at this stage by inspection of the deviation between experimental and calculated values. Numbat allows interactive modification of PCS exp i and tol i as well as the input of additional PCS data.
PCS selection
The experimental PCS values to be used for the Δχ-tensor fit can be selected according to three criteria: A list of (i) residue types or (ii) atom types can be provided by the user. This is convenient in the case of selectively isotope-labelled proteins and allows a quick assessment of the amount of information necessary in order to retrieve a robust Δχ-tensor. (iii) Each individual PCS can be selected or deselected interactively via the GUI interface. This is particularly convenient if, after initial optimization of the Δχ-tensor, some of the back-calculated PCS consistently show large deviations with respect to the experimental values, which may be due to erroneous assignments or discrepancies between the atomic coordinates of the PDB file and the actual structure of the protein, as is often the case for flexible polypeptide segments. Deselecting the corresponding atoms is likely to improve the Δχ-tensor fit in the next iteration.
Conventions
Different conventions have been used in the literature to report Δχ-tensor parameters, including different definitions of Euler angles, choice of principal and secondary axis of the Δχ-tensor, and units of Δχ-tensor magnitudes. Numbat can report the Δχ-tensor parameters in many different conventions but uses as a default the following conventions: (i) The axes of the Δχ-tensor frame are labelled such that |Δχzz| ≥ |Δχyy| ≥ |Δχxx| in analogy to alignment tensor conventions (Clore et al. 1998). This ensures that axial and rhombic components are always of the same sign. (ii) The Euler angles α, β and γ are expressed in the “ZYZ” convention, i.e., the first rotation of angle α is around the z-axis of the protein frame, the second rotation of angle β is around the new y′ axis and the last rotation of angle γ is around the new z′′ axis (Fig. 2). (iii) While for an asymmetric object the Euler angles are uniquely defined if the angles α, β and γ are taken in the intervals [0, 2π[, [0, π[, [0, 2π[, respectively, ambiguities arise for symmetric objects. Therefore, we chose the interval [0, π[ for all three angles, eliminating the potential ambiguities arising from the four symmetry-related Δχ-tensors that generate the same PCS values. In the case of β = 0, an infinite number of combinations of α and γ would produce the same overall rotation. In this case, we set γ = 0. These two rules ensure that any Δχ-tensor is unambiguously reported as a single set of parameters which is referred to in the GUI as UTR (Unique Δχ-Tensor Representation).
Euler angle definitions used by Numbat. The relative orientation of the Δχ-tensor frame with respect to the protein frame is defined by Euler rotations of angle α, β and γ in the ZYZ convention: (a) A right-handed rotation of angle α around the z-axis is applied to the protein frame xyz to give the frame x′y′z′. (b) A second rotation of angle β around the new axis y′ is applied to the frame x′y′z′ to give x′′y′′z′′. (c) The last rotation of angle γ around the z′′-axis gives the Δχ-tensor frame
Error analysis
The Levenberg–Marquardt algorithm is used to minimize the cost c (Eq. 2), but the quality of the fit cannot be assessed without further error analysis. Therefore, in addition to the uncertainty values provided by the GSL implementation of the minimizer, Numbat embeds a Monte-Carlo protocol with random Gaussian noise added either to the atomic coordinates of the molecule or to the experimental PCS values. The robustness of the Δχ-tensor fit with respect to the PCS data set can also be tested by random subset selection of the PCS values used. Resulting Δχ-tensor orientations are displayed in a Sanson-Flamsteed projection (Bugayevskiy and Snyder 1995) using the plotting utility gnuplot.
Visualization
Graphical visualization of the Δχ-tensor frame and isosurfaces of PCS values in the structure of the molecule presents a convenient way to assess the similarity of the principal axes of multiple Δχ-tensors and the similarity of their respective isosurfaces. To this end Numbat interfaces with the molecular viewers MOLMOL (Koradi et al. 1996) and PyMOL (DeLano 2002) by generating suitable macro files and displaying the Δχ-tensor frame and corresponding PCS isosurfaces in superimposition with the protein studied, as illustrated in Fig. 3. The files of the macros, PCS potential and PDB file containing the coordinates of the protein together with coordinates of the metal ion and Δχ-tensor axes can also be saved for later use.
Visualisation of the Δχ-tensor in MOLMOL and PyMOL, and display of its orientational uncertainty in a Sanson-Flamsteed projection plot. Numbat can directly call MOLMOL and PyMOL to display the axes of the fitted Δχ-tensor and PCS isosurfaces at user-defined contour levels. The orientational uncertainty of the Δχ-tensor frame can be evaluated by a Monte-Carlo protocol with random additions of noise to the structure coordinates and/or PCS data, with optional random selection of subsets of data. Numbat calls gnuplot to display the results in a Sanson-Flamsteed projection plot
Output
The list of PCS can be saved in Xplor-NIH format and in a Numbat-specific format. The weak molecular alignment in the magnetic field resulting from a non-vanishing Δχ-tensor can be described by an alignment tensor with principal axes parallel to those of the Δχ-tensor and axial and rhombic components that are directly proportional to Δχax and Δχrh, respectively (Tolman et al. 1995). Numbat calculates the RDC between two spins A and B for the situation of a completely rigid molecule, using (Bertini et al. 2002)
where γ A and γ B are the magnetogyric ratios of spins A and B, respectively, ħ the Planck constant divided by 2π, S the order parameter, r AB the internuclear distance, and \( \tilde x_{AB} \), \( \tilde y_{AB} \), \( \tilde z_{AB} \) the coordinates of the vector AB expressed in the Δχ-tensor frame. The RDC values are reported in Xplor-NIH (Schwieters et al. 2003, 2006) and Pales (Zweckstetter and Bax 2000) format.
Finally, Numbat can generate PDB files where the Δχ-tensor is reported in a format ready for use with MOLMOL or PyMOL for rigid-body docking alignment, or for further refinement by Xplor-NIH.
Study case
The proteins ε and θ are subunits of the complex of proteins constituting E. coli DNA polymerase III. The complex between the N-terminal domain of ε (ε186) and θ has been extensively studied using PCS data (Pintacuda et al. 2006, 2007). In light of the recent crystal structure of the complex between ε186 and the θ homolog HOT (Kirby et al. 2006), we illustrate in the following the features of Numbat by revisiting the NMR structure of the complex between ε186 and θ which was derived from PCS induced by Dy3+ and Er3+ ions bound to the natural metal-binding site of ε186 (Pintacuda et al. 2006).
The coordinates of the A chain in the PDB deposition 2IDO (Kirby et al. 2006) was used as the structural model for ε186. The structural model of θ was conformer 10 of the NMR structure of θ in complex with ε186 (PDB accession code 2AXD; Keniry et al. 2006). This conformer was chosen because it has the lowest backbone RMSD to the HOT protein (2.1 Å) for residues 9–66 (the structurally defined region for which meaningful PCS could be measured). The experimentally determined PCS values of ε186 have been reported previously (Schmitz et al. 2006) and the PCS values of θ are provided in the Supporting Information. All Δχ-tensor optimizations were performed using Numbat including the RACS correction term and a tolerance value tol i of zero for all spins.
Subunit ε186
Table 1 presents the results of the Δχ-tensor fit to the PCS measured for ε186. Initially, individual eight-variable Δχ-tensor optimizations were performed using the PCS data of each lanthanide (Table 1, columns 1 and 2). Next, the Numbat GUI was updated to display the deviations between the experimental and back-calculated PCS for the Δχ-tensors found. Several atoms showed deviations >0.15 ppm between the experimental and back-calculated PCS (15 out of 199 and 8 out of 255 atoms in the case of Dy3+ and Er3+, respectively. Without the RACS correction, deviations >0.15 ppm where observed for 36 and 7 atoms, respectively). Assuming that these outliers were due to problematic measurements or inaccuracies of the 3D structure, these PCS were removed interactively using the GUI. Re-calculation of the Δχ-tensor was found not to change the fitted Δχ-tensor parameters significantly for any of the lanthanide ions (results not shown). This can be explained by the high quality and large number of experimental PCS data available for each lanthanide (backbone 13C′, 15N and 1HN spins), resulting in robust fits of the Δχ-tensors.
Since the coordinates of the Dy3+ and Er3+ found in the individual fits were very similar (Table 1, columns 1 and 2), we subsequently assumed that the Δχ-tensors induced by each lanthanide are centered at the same position relative to ε186. The results obtained by simultaneously fitting the distinct Δχ-tensors while restraining their metal coordinate to a common centre (Table 1, columns 3 and 4) show little difference to the Δχ-tensor parameters found when performing the individual optimizations.
For comprehensive error analysis, we introduced a random error into the structure coordinates of ε186, where the atomic coordinates were varied according to a Gaussian distribution with a standard deviation σ of 0.5 Å, resulting in a mean atom displacement of 0.8 Å. The resulting uncertainty in Δχ-tensor parameters was approximately equivalent to the uncertainty introduced by a random variation added to the measured PCS data sampled from a Gaussian distribution with a standard deviation σ of 0.15 ppm. The Δχ-tensor parameters of ε186 were well defined, as the values of all eight Δχ-tensor variables determined by 1,000 randomized pseudo-replicates of the structure were in good agreement with the Δχ-tensors fitted to the original structure (Table 2, column 1). To eliminate the possibility that the quality of the Δχ-tensor fit was significantly affected by the number of PCS measured, the error analysis for the Δχ-tensors fitted to ε186 was recalculated with random selection of only 20% of the measured PCS. The results (Table 2, column 2) show that the Δχ-tensor parameters of ε186 were still well defined.
Subunit θ
The results of the Δχ-tensor determination in the molecular frame of θ are presented in Table 1. There was only a small number of spins for which the back-calculated PCS deviated from the experimental PCS by more than 0.15 ppm (4 out of 50 in the case of Dy3+, 0 out of 41 for Er3+). Like for ε186, removal of these PCS from the optimization did not significantly change the parameters of the fitted Δχ-tensors. While the Δχax and Δχrh values of Er3+ determined from the PCS observed for θ and ε186 were very similar, the Δχrh value of the Dy3+ tensor found for θ was almost three times larger than that found for ε186. We subsequently performed an error analysis for θ as for the ε186 subunit, introducing either random variations into the atomic positions of θ according to a Gaussian distribution with a standard deviation σ of 0.5 Å or using a random selection of only 80% of the measured PCS. In either case, the Δχ-tensor parameters of θ proved to be less well defined than those of ε186 (Table 2). As θ samples a relatively small and remote volume of the Δχ-tensors due to its spatial separation from the metal ion, one would expect a less accurate determination of the Δχ-tensors from the θ data. The effect could be exacerbated by inaccuracies of the NMR structure.
In order to compensate for the smaller number of experimentally determined PCS available for θ (only 1HN spins) and the poorer quality of the Δχ-tensors fitted, we performed another fit with Δχax and Δχrh fixed to the values determined for ε186 (Table 1, columns 9 and 10). Analysis of the experimental versus back-calculated PCS, both for the eight- and six-variable fits of the Δχ-tensor to θ, showed that the PCS deviations were similar in magnitude and trends. Therefore, constraining Δχax and Δχrh did not significantly deteriorate the quality of the fit, despite considerable changes of the Δχ-tensor parameters (Table 1). A comparison of the Δχ-tensor parameters obtained by using a single model of θ and by averaging over all PDB models of θ is provided in the Supporting Information.
Modelling the complex between ε186 and θ
Numbat facilitates the modelling of protein–protein complexes by listing coordinates of the Δχ-tensor axes together with the protein coordinates in files in PDB format. Superimposition of the Δχ-tensors fitted to ε186 and θ for each lanthanide ion yields the three-dimensional structure of the ε186/θ complex by straightforward rigid-body docking. Standard PyMOL or MOLMOL commands can be used to align the Δχ-tensors. Numbat reports the coordinate system of the Δχ-tensor in such a way that all four degenerate solutions arising from the symmetry of the Δχ-tensor about the x, y and z-axes can easily be visualized. Identification of the correct solution requires additional information, such as proper steric interactions, chemical shift perturbation data or knowledge of the biological function of the complex. The most objective way, however, is by simultaneous evaluation of the Δχ-tensors of different lanthanides (Pintacuda et al. 2006).
In the case of the complex between ε186 and θ, the Δχ-tensor frames of Dy3+ and Er3+ share a common origin for both proteins. The lowest RMSD value resulting from all 16 possible 7-coordinate alignments between the two combined Δχ-tensors identified a single relative orientation of the two proteins as the best solution. The position of θ relative to ε186 derived from PCS data in this way was also the correct solution. It agreed with a model of the complex obtained by superimposition of θ onto HOT in the ε186/HOT complex, with a backbone RMSD of 4.4 Å. Similarly for the Δχ-tensor of θ calculated with fixed Δχax and Δχrh values, a backbone RMSD of 4.3 Å was calculated relative to HOT. When PCS data from only Dy3+ or Er3+ were used, the backbone RMSD values were, respectively, 4.2 Å and 4.4 Å for the best fit to the ε186/HOT complex. Figure 4 displays the model of the ε186/θ complex derived from the superimposition of the Δχ-tensors obtained by using the combined and fixed optimization protocols for ε186 and θ, respectively (Table 1).
The complex between ε186 and θ determined by superimposition of Δχ-tensors. The ε186/HOT complex (PDB accession code 2IDO) is shown for reference, with ε186 coloured in silver and HOT (residues 9–66) in orange. The isosurfaces correspond to the PCS induced by the Dy3+ ion (from individual optimization) contoured at ±1.5 ppm and ±0.5 ppm. Blue and red isosurfaces represent regions with positive and negative PCS, respectively. Residues 9–66 of θ are shown as a thin dark ribbon in the position defined by the Dy3+ and Er3+ tensors from the combined optimization for ε186 and the fixed optimization for θ shown in Table 1
Conclusion
The program Numbat is the first software package for fitting Δχ-tensors from PCS data with a user-friendly GUI. Numbat calculations are fast, as it was written with open-source Linux routines in C. While the main task of Numbat is the fit of the eight Δχ-tensor variables, the intuitive GUI combined with convenient data handling, including Monte-Carlo error analysis and links to the molecular viewers MOLMOL and PyMOL, offer high flexibility of use. The study case of the complex formed between the subunits ε186 and θ of E. coli DNA polymerase III illustrates the simplicity of use of Numbat.
The program for Linux and Windows operating systems is freely available under the GNU General Public License (GPL) upon request (see also http://compbio.chemistry.uq.edu.au/bmmg/christophe/numbat.html).
Abbreviations
- α:
-
Subunit α of the E. coli polymerase III
- ε186:
-
N-terminal 185 residues of the E. coli polymerase III subunit ε
- θ:
-
Subunit θ of the E. coli polymerase III
- CSA:
-
Chemical shielding anisotropy
- GUI:
-
Graphical user interface
- HOT:
-
The bacteriophage P1-encoded homolog of θ
- PCS:
-
Pseudocontact shift
- RACS:
-
Residual anisotropic chemical shift
- RDC:
-
Residual dipolar coupling
- UTR:
-
Unique Δχ-tensor representation
References
Allegrozzi M, Bertini I, Janik MBL, Lee Y-M, Liu G, Luchinat C (2000) Lanthanide-induced pseudocontact shifts for solution structure refinements of macromolecules in shells up to 40 Å from the metal ion. J Am Chem Soc 122:4154–4161
Banci L, Dugad LB, La Mar GN, Keating KA, Luchinat C, Pierattelli R (1992) 1H nuclear magnetic resonance investigation of cobalt(II) substituted carbonic anhydrase. Biophys J 63:530–543
Banci L, Bertini I, Bren KL, Cremonini MA, Gray HB, Luchinat C, Turano P (1996) The use of pseudocontact shifts to refine solution structures of paramagnetic metalloproteins: Met80Ala cyano-cytochrome c as an example. J Biol Inorg Chem 1:117–126
Banci L, Bertini I, Savellini GG, Romagnoli A, Turano P, Cremonini MA, Luchinat C, Gray HB (1997) Pseudocontact shifts as constraints for energy minimization and molecular dynamics calculations on solution structures of paramagnetic metalloproteins. Proteins Struct Funct Genet 29:68–76
Banci L, Bertini I, Cremonini MA, Savellini GG, Luchinat C, Wüthrich K, Güntert P (1998) PSEUDYANA for NMR structure calculation of paramagnetic metalloproteins using torsion angle molecular dynamics. J Biomol NMR 12:553–557
Banci L, Bertini I, Cavallaro G, Giachetti A, Luchinat C, Parigi G (2004) Paramagnetism-based restraints for Xplor-NIH. J Biomol NMR 28:249–261
Berman HM, Westbrook J, Feng Z, Gilliland G, Bhat TN, Weissig H, Shindyalov IN, Bourne PE (2000) The protein data bank. Nucleic Acids Res 28:235–242
Bertini I, Donaire A, Jiménez B, Luchinat C, Parigi G, Piccioli M, Poggi L (2001) Paramagnetism-based versus classical constraints: an analysis of the solution structure of Ca Ln calbindin D9k. J Biomol NMR 21:85–98
Bertini I, Luchinat C, Parigi G (2002) Magnetic susceptibility in paramagnetic NMR. Prog NMR Spectrosc 40:249–273
Bertini I, Del Bianco C, Gelis I, Katsaros N, Luchinat C, Parigi G, Peana M, Provenzani A, Zoroddu MA (2004) Experimentally exploring the conformational space sampled by domain reorientation in calmodulin. Proc Natl Acad Sci U S A 101:6841–6846
Bugayevskiy LM, Snyder JP (1995) Map projections: a reference manual. Taylor & Francis, London
Capozzi F, Cremonini MA, Luchinat C, Sola M (1993) Assignment of pseudo-contact-shifted 1H NMR resonances in the EF site of Yb3+-substituted rabbit parvalbumin through a combination of 2D techniques and magnetic susceptibility tensor determination. Magn Reson Chem 31:S118–S127
Clore GM, Gronenborn AM, Bax A (1998) A robust method for determining the magnitude of the fully asymmetric alignment tensor of oriented macromolecules in the absence of structural information. J Magn Reson 133:216–221
Cornilescu G, Bax A (2000) Measurement of proton, nitrogen, and carbonyl chemical shielding anisotropies in a protein dissolved in a dilute liquid crystalline phase. J Am Chem Soc 122:10143–10154
DeLano WL (2002) The PyMOL molecular graphics system. Palo Alto, CA, USA
Dosset P, Hus J-C, Blackledge M, Marion D (2000) Efficient analysis of macromolecular rotational diffusion from heteronuclear relaxation data. J Biomol NMR 16:23–28
Eichmüller C, Skrynnikov NR (2007) Observation of μs time-scale protein dynamics in the presence of Ln3+ ions: application to the N-terminal domain of cardiac troponin C. J Biomol NMR 37:79–95
Emerson SD, La Mar GN (1990) NMR determination of the orientation of the magnetic susceptibility tensor in cyanometmyoglobin: a new probe of steric tilt of bound ligand. Biochemistry 29:1556–1566
Galassi M, Davies J, Theiler J, Gough B, Jungman G, Booth M, Rossi F (2006) GNU scientific library reference manual. Network Theory Ltd, Bristol
Gaponenko V, Sarma SP, Altieri AS, Horita DA, Li J, Byrd RA (2004) Improving the accuracy of NMR structures of large proteins using pseudocontact shifts as long-range restraints. J Biomol NMR 28:205–212
Güntert P, Mumenthaler C, Wüthrich K (1997) Torsion angle dynamics for NMR structure calculation with the new program DYANA. J Mol Biol 273:283–298
Hess B, Scheek RM (2003) Orientation restraints in molecular dynamics simulations using time and ensemble averaging. J Magn Reson 164:19–27
Jensen MR, Hansen DF, Ayna U, Dagil R, Hass MAS, Christensen HEM, Led JJ (2006) On the use of pseudocontact shifts in the structure determination of metalloproteins. Magn Reson Chem 44:294–301
John M, Park AY, Pintacuda G, Dixon NE, Otting G (2005) Weak alignment of paramagnetic proteins warrants correction for residual CSA effects in measurements of pseudocontact shifts. J Am Chem Soc 127:17190–17191
John M, Pintacuda G, Park AY, Dixon NE, Otting G (2006) Structure determination of protein–ligand complexes by transferred paramagnetic shifts. J Am Chem Soc 128:12910–12916
Keniry MA, Park AY, Owen EA, Hamdan SM, Pintacuda G, Otting G, Dixon NE (2006) Structure of the θ subunit of Escherichia coli DNA polymerase III in complex with the ε subunit. J Bacteriol 188:4464–4473
Kirby TW, Harvey S, DeRose EF, Chalov S, Chikova AK, Perrino FW, Schaaper RM, London RE, Pedersen LC (2006) Structure of the Escherichia coli DNA polymerase III ε-HOT proofreading complex. J Biol Chem 281:38466–38471
Koradi R, Billeter M, Wüthrich K (1996) MOLMOL: a program for display and analysis of macromolecular structures. J Mol Graphics 14:51–55
Krause A (2007) Foundations of GTK + development. Apress. Berkeley, CA, USA
Lee L, Sykes BD (1983) Use of lanthanide-induced nuclear magnetic resonance shifts for determination of protein structure in solution: EF calcium binding site of carp parvalbumin. Biochemistry 22:4366–4373
Marquardt DW (1963) An algorithm for least-squares estimation of nonlinear parameters. J Soc Ind Appl Math 11:431–441
Nelder JA, Mead R (1965) A simplex method for function minimization. Comput J 7:308–313
Pintacuda G, John M, Su X-C, Otting G (2007) NMR structure determination of protein–ligand complexes by lanthanide labeling. Acc Chem Res 40:206–212
Pintacuda G, Keniry MA, Huber T, Park AY, Dixon NE, Otting G (2004) Fast structure-based assignment of 15N HSQC spectra of selectively 15N-labeled paramagnetic proteins. J Am Chem Soc 126:2963–2970
Pintacuda G, Park AY, Keniry MA, Dixon NE, Otting G (2006) Lanthanide labeling offers fast NMR approach to 3D structure determinations of protein–protein complexes. J Am Chem Soc 128:3696–3702
Schmitz C, John M, Park AY, Dixon NE, Otting G, Pintacuda G, Huber T (2006) Efficient χ-tensor determination and NH assignment of paramagnetic proteins. J Biomol NMR 35:79–87
Schwieters CD, Kuszewski JJ, Tjandra N, Clore GM (2003) The Xplor-NIH NMR molecular structure determination package. J Magn Reson 160:65–73
Schwieters CD, Kuszewski JJ, Clore GM (2006) Using Xplor-NIH for NMR molecular structure determination. Prog NMR Spectrosc 48:47–62
Sherry AD, Pascual E (1977) Proton and carbon lanthanide-induced shifts in aqueous alanine. Evidence for structural changes along the lanthanide series. J Am Chem Soc 99:5871–5876
Su X-C, McAndrew K, Huber T, Otting G (2008) Lanthanide-binding peptides for NMR measurements of residual dipolar couplings and paramagnetic effects from multiple angles. J Am Chem Soc 130:1681–1687
Tolman JR, Flanagan JM, Kennedy MA, Prestegard JH (1995) Nuclear magnetic dipole interactions in field-oriented proteins: Information for structure determination in solution. Proc Natl Acad Sci U S A 92:9279–9283
Valafar H, Prestegard JH (2004) REDCAT: a residual dipolar coupling analysis tool. J Magn Reson 167:228–241
Van der Spoel D, Lindahl E, Hess B, Groenhof G, Mark AE, Berendsen HJC (2005) GROMACS: fast, flexible, and free. J Comput Chem 26:1701–1718
Veitch NC, Whitford D, Williams RJP (1990) An analysis of pseudocontact shifts and their relationship to structural features of the redox states of cytochrome b5. FEBS Lett 269:297–304
Wang X, Srisailam S, Yee AA, Lemak A, Arrowsmith C, Prestegard JH, Tian F (2007) Domain-domain motions in proteins from time-modulated pseudocontact shifts. J Biomol NMR 39:53–61
Wei Y, Werner MH (2006) iDC: a comprehensive toolkit for the analysis of residual dipolar couplings for macromolecular structure determination. J Biomol NMR 35:17–25
Zweckstetter M, Bax A (2000) Prediction of sterically induced alignment in a dilute liquid crystalline phase: aid to protein structure determination by NMR. J Am Chem Soc 122:3791–3792
Acknowledgment
Financial support from the Australian Research Council for project grants to G.O and T.H. is gratefully acknowledged.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Schmitz, C., Stanton-Cook, M.J., Su, XC. et al. Numbat: an interactive software tool for fitting Δχ-tensors to molecular coordinates using pseudocontact shifts. J Biomol NMR 41, 179–189 (2008). https://doi.org/10.1007/s10858-008-9249-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10858-008-9249-z