Abstract
To understand the CO2 concentration mechanism in the phaeophyte kelp Saccharina japonica, a full-length complementary DNA (cDNA) encoding carbonic anhydrase (CA) was cloned from the gametophytes based on the two screened clones from a suppressive subtracted cDNA library. The cDNA sequence was composed of 2,804 bp in length, including a 166-bp 5′-untranslated region (UTR), a 1,765-bp 3′-UTR, and an 873-bp open read frame. No intron separating this gene was found after comparing its cDNA and DNA sequences. The deduced precursor protein of S. japonica CA consisted of 290 amino acids with a typical signal peptide cleavage site between Gly 20 and Val 21 from the N terminus. The mature protein contained three conserved His residues chelated with a zinc ion constituting a catalytic active site. This cloned CA gene from S. japonica could be grouped into the α-type as shown by a constructed phylogenetic tree and most identity within the conserved domains characteristic of the α-CA. Quantitative real-time PCR results demonstrated that the diurnal transcription of this CA gene in the gametophytes cultured under the addition of CO2 or HCO3 − were not significantly different from those cultured only with filtered air supply at any sampling time, suggesting that this CA might not be periplasmic. After preparation of polyclonal antibody with this recombinant CA in Escherichia coli, the gold immunolocalization showed that this α-CA was associated with the chloroplast envelopes and thylakoid membranes suggesting that this α-CA could provide the chloroplasts with sufficient CO2 for carbon assimilation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Over the past 250 years, atmospheric carbon dioxide (CO2) levels have increased by nearly 40 % (Solomon et al. 2007), while the ocean has absorbed about one quarter of anthropogenic carbon emissions (Sabine et al. 2004; Sabine and Feely 2007). Accompanying the ocean CO2 uptake, seawater pH decreases and the fundamental chemical balances of seawater are gradually altered, both of which are commonly referred to as ocean acidification (Turley 2005; Henderson 2006). As photoautotrophs, marine plants need to take up CO2 from seawater for photosynthesis. For example, the light-saturated net photosynthetic rates in the brown algae range from 100 to 1670 μmol CO2 g (dry wt)−1 h−1 (Gao and McKinley 1994). Therefore, the biological uptake CO2 from the ocean is one of the basic mechanisms controlling carbon distribution in the ocean (Siegenthaler and Sarmiento 1993). The cultivation of seaweeds such as Saccharina japonica (Aresch.) C. E. Lane, C. Mayes, Druehl et G. W. Saunders (=Laminaria japonica Aresch.) is thought to slow down ocean acidification.
Saccharina japonica is an economically important brown seaweed, especially in East Asian countries such as China, Japan, and South Korea (Tseng 2001). In the high pH environment where this kelp lives, the dominant species of inorganic C in seawater is HCO3 − (Axelsson et al. 2000). Since there is a fairly high photosynthetic rate in these kelps (Axelsson et al. 1991), an efficient HCO3 − utilization mechanism is expected to exist. A common strategy evolved in algae is extracellular dehydration of the HCO3 −, which is catalyzed by carbonic anhydrase (CA) (Haglund et al. 1992). CA is a zinc metalloenzyme that catalyses the interconversion of CO2 and HCO3 − (Khalifah 1971). As a result, much attention has been paid to CA in algae, especially from microalgae such as Chlamydomonas reinhardtii (Spalding 2008; Moroney et al. 2011), but there are fewer reports regarding CA in S. japonica and other brown seaweeds (Dring et al. 1994; Schmid et al. 1996; Moulin et al. 1999; Axelsson et al. 2000). There is only one CA gene from Laminaria digitata and six CA genes from Ectocarpus siliculosus in brown algae.
CAs are generally divided into three distinct classes (α, β, and γ) that have no sequence homology and which have evolved independently (Hewett-Emmett and Tashian 1996), and which are an example of convergent evolution of catalytic function (Moroney et al. 2001). In this study, a full-length CA encoding cDNA was cloned and characterized from S. japonica gametophytes. The subcellular location of the gene was studied by gold immunoelectron microscopy using a CA polyclonal antibody and the diurnal transcription of this α-CA gene as detected by quantitative real-time PCR (Q-RT-PCR).
Materials and methods
The Rongfu strain of Saccharina japonica was used in the present research. Both female and male gametophyte clones germinated from the released zoospores were isolated according to cell sizes under a microscope and cultured separately under vegetative growth conditions of 30 μmol photons m−2 s−1 s at 17 ± 1 °C with a photoperiod of 12:12 light/dark, as described previously (Zhou and Wu 1998). The PES medium (Starr and Zeikus 1993) was replaced once every 2 weeks.
DNA and RNA extraction
Genomic DNA was extracted separately from the freshly harvested female and male gametophytes according to the modified cetyltrimethyl ammonium bromide method as described by Hu and Zhou (2001). Female and male gametophyte filaments were individually harvested and frozen in liquid nitrogen for total RNA extraction using TRIzol reagent (Invitrogen, USA). The complementary DNA was synthesized using the Reverse Transcribed Kit II (TaKaRa, China).
Cloning of α-CA gene from S. japonica gametophytes
Two 376-bp expressed sequence tags (ESTs) were screened from a suppression subtractive hybridization cDNA library of S. japonica male gametophytes subtracted from females (Shi et al. 2005) since they had a 70 % identity to a CA gene (GenBank accession no. AJ130777) from L. digitata. Based on the ESTs and the conserved sequence of L. digitata CA gene, partial sequence of open reading frame (ORF) of S. japonica CA gene was amplified with primers J1-U and J1-D (Table 1). Based on the ORF partial sequence obtained, a full-length cDNA of the S. japonica CA gene, designated as SjCA, was cloned from the kelp gametophytes (Yu et al. 2011) using the rapid amplification of cDNA ends technique. Finally, the DNA sequence of SjCA was also cloned on the basis of its corresponding cDNA sequence.
Southern blot and bioinformatics analysis of SjCA
Southern blot analysis was made using the genomes of the gametophytes to gather more information about this CA gene. Aliquots of isolated DNA (approximately 2 μg per sample) were digested to completion at 37 °C for 4-6 h, independently, with EcoRI and XbaI. The digested DNA samples were fractionated on a 0.8 % agarose gel, blotted onto a positively charged nylon membrane (Pall, USA), and hybridized with the probe. The probe was synthesized with the SBL and SBR primers (Table 1) and labeled with biotin-dUTP using a North2South Biotin Random Prime Labeling Kit (Thermo, USA). Hybridized bands were detected using a North2South Chemiluminescent Hybridization and Detection Kit (Thermo, USA) and signals were visualized by exposure to XBT-1 film (Kodak, USA) (Liu et al. 2012).
The amino acid composition, molecular weight, and isoelectric point of this putative SjCA were analyzed using ProtParam (http://www.expasy.org/tools/protparam.html). The structure of this CA was predicted using Phyre 2 Server (http://www.sbg.bio.ic.ac.uk/phyre2/html/page.cgi?id=index). The signal peptide and transmembrane region of SjCA were predicted using the SignalP 4.1 Server (http://genome.cbs.dtu.dk/services/SignalP/) and the TMHMM (http://genome.cbs.dtu.dk/services/TMHMM/). A Blastp analysis (http://blast.ncbi.nlm.nih.gov/Blast.cgi) was used to obtain homologous sequences of the CAs. The sequences were inferred from the corresponding deduced amino acid sequences and aligned using the ClustalX Program (Thompson et al. 1997). A phylogenetic tree was constructed using the neighbor-joining (NJ) and maximum likelihood (ML) methods in the MEGA 4.0 program (Tamura et al. 2007).
SjCA transcription levels under different carbon resources
The gametophytes were cultivated in medium with filtered air, filtered air + 3 % CO2 and 0.018 M NaHCO3 independently, and were collected at 00:00, 08:00, 16:00, and 24:00 hours, and the number of replicates was 3. Total RNA was extracted using TRIzol reagent (Invitrogen, USA) with these male and female gametophytes individually, and was used to synthesize the oligo(dT) primed first-strand cDNA as described above. S. japonica actin (GenBank accession no. EU293553) served as the housekeeping gene. The primers used in Q-RT-PCR for both the SjCA (DL and DR) and actin (SL and SR) genes are also shown in Table 1. The Q-RT-PCR amplification was performed in a Bio-Rad iQ™5 multicolor real-time PCR detection system (USA) with the SYBR® RT-PCR Kit (TaKaRa, China). After predenaturing at 95 °C for 15 s, the Q-RT-PCR amplification was programmed for 40 denaturing cycles at 95 °C for 15 s and annealing and extension at 59.6 °C for 1 min, in a 20 μL reaction mixture containing 1 μL of each primer (10 μM) and 2 μL first-strand cDNA as a template and 10.5 μL SYBR Script™ (TaKaRa, China). The relative gene transcription data from triplicate reactions performed for each incubation time are expressed as the means ± standard error using the 2−ΔΔCt method (Livak and Schmittgen 2001). The significance was estimated by the Student's t-test.
Heterologous expression and preparative of SjCA polyclonal antibody
The CA gene without signal peptide was cloned using the LFT and NRG primers (Table 1), and ligated into the NcoI and XhoI digested pET-28a prokaryotic expression vector. The resulting construct pET28a-CA was transformed into Escherichia coli BL21 competent cells. Colony PCR and nucleotide sequencing were performed to confirm the orientation and reading frame of this ligated insert.
Expression of the recombinant protein in transformed E. coli cultures was induced by the addition of 1 mM isopropyl-β-d-thiogalactoside (IPTG) as described in Chen et al. (2010). Crude cell extracts were prepared by sonicating cells suspended in phosphate-buffered saline (PBS) (0.137 M NaCl, 2.7 mM KCl, 10 mM Na2HPO4, and 2 mM KH2PO4) at 4 °C. Denaturing sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE) was performed according to Laemmli (1970), and protein was quantified using the Bradford (1976) method. Antiserum was obtained from rabbits that had been immunized with the recombinant protein, which was purified from the transformed E. coli pET28a-CA/BL21. The antibody was purified according to the method as described by Olmsted (1981) and Ritter (1991).
Western blot
This SjCA antibody was used in Western analysis for the detection of the CA protein. Western blot procedures were carried out essentially as described by Chen et al. (2010). The total protein of the transformed E. coli, with and without the insertion of the CA gene was separated by 12 % SDS-PAGE. After electrophoresis, the proteins were electronically transferred onto nitrocellulose membranes. The protein blot was blocked with 5 % skim milk powder in Tris-buffered saline Tween-20 buffer (TBST) (0.137 M NaCl, 2.7 mM KCl, 0.025 M Tris, and 0.05 % Tween 20 at pH 7.4) and incubated with the purified antibodies in a suitable dilution for 1 h at room temperature. Next, washed in TBST for several times and incubated with the a secondary antibody, antirabbit IgG labeled by horseradish peroxidase (Shanghai Youke Biotechnology Co., Ltd.) diluted 1:1,200 in TBST at room temperature for 1 h and washed again. The color reaction was visualized with diaminobenzidine following the manufacturer's instructions (Tiangen Biotech Co., Ltd.). The Western blot was performed with gradient dilutions of antibody until the hybridization signals were strong but less background. Then, the Western blot in female and male gametophytes was performed as above.
Ultrastructure observation and immunoelectron microscopy
Since CAs play different roles in the different places and their functions are usually associated with their locations (Karlsson et al. 1998; Sinetova et al. 2012), we performed immunoelectron microscopy to precisely define the subcellular localization of this CA in S. japonica. Actively growing S. japonica male and female gametophytes, cultured in complete PES medium, were collected separately. The samples were fixed with 4 % (w/v) paraformaldehyde and 0.5 % glutaraldehyde in distilled seawater for 3-12 h at 4 °C (He et al. 2002), and rinsed with 0.1 M PBS at pH 7.4. The fixed samples were then subjected to post-fixation (3 h) with 1 % osmium tetroxide in 0.1 M PBS. The postfixed samples were dehydrated in ethanol/acetone series, and then were infiltrated and embedded in epoxy resin E51 as described by Ouyang et al. (2012).
The ultrathin sections used for immunoelectron microscopy were collected on 200-mesh nickel grids with a noncarbon-coated Formvar-supporting film. Nickel grids carrying ultrathin sections were treated with 1 % (w/v) sodium metaperiodate in PBS (Bendayan and Zollingerm 1983) and preincubated the sections with 50 mM glycine for 30 min (Miller and Howell 2006) and 5 % (w/v) bovine serum albumin (BSA) in PBS for 20 min (Andreu et al. 2007), and then incubated with the purified anti-CA antibody at 4 °C for 48 h or with the primary antibody absent as a control. The optimal working titer of the primary antibody was based on serial dilutions of the antibody from 1:1,000 to 1:3,600 as described by Miller and Howell (2006) and Shi et al. (2011). After washes in PBS and another 20 min preincubation with 1 % (w/v) BSA, the grids were incubated with the secondary antibody, antirabbit IgG conjugated to 10 nm gold particles (Sigma, USA) at room temperature for 1 h. Following sequential washes in PBS and water and dehydration in the air, the sections were stained with 3 % uranyl acetate-lead citrate, and observed in a JEOL-1230 transmission electron microscope at 80 kV.
The labeling density was defined as the number of gold particles per area unit (μm2) as described by Bernal (2007), and the area was estimated using Adobe Photoshop software (ver. 3.0). Following counting, the gold particles on chloroplasts and the other area calculated by subtracting the total area of chloroplasts in each micrograph, the percentage of particles versus the total number of particles was calculated. The statistical analysis of subcellular distribution of the CA protein was carried out using the independent sample test in SPSS Statistics 17.0 program.
Results
Characterization of the CA in Saccharina japonica
The complete coding sequence of SjCA contained 2,804 bp including a 166-bp 5′-untranslated region (UTR), a 1,765-bp 3′-UTR, and an 873-bp ORF. This gene has been submitted to GenBank under the accession no. JF827608. DNA cloning of this gene showed that there was no difference between the males and females, not only in the cDNA but also the DNA sequences. No intron separating this gene was found after comparing its cDNA and DNA sequences. Southern blot analysis (Fig. 1) indicated that this CA was a single gene copy in the gametophyte genomes as digested by EcoRI and XbaI independently. The deduced precursor protein of S. japonica CA consisted of 290 amino acids that possessed a typical signal peptide cleavage site between Gly 20 and Val 21. The mature protein, after digestion, was composed of 270 amino acids. The molecular weight of the mature protein was 30.34 kDa and its pI was at 5.06. There was no transmembrane domain in the mature CA protein though the predicted structure consisted of five α-helices since all these helices were too short. Therefore, this CA protein could not be an intrinsic membrane protein.
Electrophoresis and Southern blot profiles of SjCA from the female and male gametophytes of Saccharina japonica. Genomic DNA from S. japonica gametophytes was digested separately with EcoRI and XbaI, electrophoresed on 0.8 % agarose, blotted and hybridized with a labeled 569-bp PCR-amplified fragment of SjCA. M D-2000 marker of standard DNA (Tiangen Biotech Co., Ltd.)
The phylogenetic tree (Fig. 2) inferred from the selected 41 CA proteins illustrated that they could be grouped into three clades, α-, β- and γ-CA, with high support values (all 100 at the root of each clade by the NJ and 84, 77, and 92 by the ML). SjCA was clustered within the α-CA clade (Fig. 2), and it had 88 and 53 % similarity with α-CAs of L. digitata (CAB61337.1) and E. siliculosus (CBN76519.1), respectively, according to the Blastp.
Neighbor-joining phylogenetic tree inferred from the amino acid sequences deduced from the nucleotide sequences of CA genes from the selected species using the neighbor-joining and maximum likelihood methods. Bootstrap values were computed independently for 1,000 resamplings of both NJ analysis (before the slash) and ML analysis (after the slash). Values lower than 50 are not shown. The accession numbers of the CA proteins are indicated in the parentheses after the Latin names of each species. Asterisk denotes the position of Saccharina japonica
SjCA possessed 24 of the 36 amino acid residues (Fig. 3) with side chains that project into or border the active site cavity as reported in mammalian α-CA research by Hewett-Emmett and Tashian (1996). It has the characteristics of α-CA with the identical conserved domain, which was thought to be the active sites of α-CA specific inhibitors (Huang et al. 1998), especially containing Cys 211 (Fig. 3) that could form a disulfide bond with Cys 46 to keep the structure of SjCA stable. The three histidine residues, i.e., His 118, His 120, and His 137 (Fig. 3), which function as Zn ligands within the active site of all α-CAs (Liljas and Laurberg 2000) are also present in SjCA.
Alignment of α-CA proteins from various species containing Zn-binding sites. The accession numbers of the proteins corresponding to Latin-names of each species are referred to in Fig. 2, but the accession number for Chlamydomonas reinhardtii is AAC49983. Amino acids of high identity more than 70 % are indicated by asterisks. The solid arrows show the Zn-binding sites, and the box and the empty arrow indicate the conservative domain of α-CA proteins and the conserved Cys residue, respectively
In general, taking the cluster with other α-CAs at high bootstrap value as illustrated by the phylogenetic tree and the identity within the conserved domains that are characteristics of α-CA into consideration, this cloned SjCA coding for a S. japonica CA is shown to be a α-type CA gene. This SjCA could be imported from the outside of gametophyte cells under the guidance of the signal peptide.
Effect of different carbon sources on the transcription of SjCA
The transcription or expression of most periplasmic CAs in C. reinhardtii has been reported to be strongly regulated by changed levels of CO2 in the medium (Fukuzawa et al. 1990). To understand whether this SjCA was periplasmic, we investigated the relative transcription level in gametophytes cultivated in PES medium with different carbon sources by Q-RT-PCR. The results (Fig. 4) showed that there is no significant difference (P > 0.05) of SjCA transcription between the control and the gametophytes treated with CO2 or NaHCO3 at any time of the day. These Q-RT-PCR results also demonstrated that the transcription of SjCA in the female gametophytes (Fig. 4a) differed insignificantly (P > 0.05) from that in the males (Fig. 4b). It indicated that the SjCA transcription had a similar profile over the day in both female and male gametophytes and that there was no significant effect of carbon sources on the SjCA transcription. Therefore, it is suggested that this SjCA does not function as a periplasmic CA.
Heterologous expression and preparative antibody of SjCA
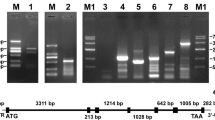
To reveal the precise location of this SjCA in the gametophyte filament cells by using immunoelectron microscopy, the polyclonal antibody of this protein was prepared. Upon cloning of the mature SjCA protein coding sequence, construction of pET28a-CA/BL21 and transformation into E. coli BL21, a transformant was screened out. Electrophoresis (Fig. 5a) of the colony PCR amplified products of the transformant E. coli harboring pET28a-CA/BL21 illustrated that only one specific band. Sequencing analysis of this product showed that it was 821 bp in length containing 813 bp of the mature SjCA protein coding sequence and eight nucleotides added suitable for endonulease restriction digestion indicating that the SjCA gene was successfully transformed into E. coli.
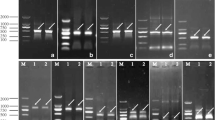
Electrophoresis profiles of colony PCR amplified product (a) and overexpression (b) of recombinant SjCA in Escherichia coli and a Western blot using the SjCA-specific polyclonal antibody in Escherichia coli (c) and in Saccharina japonica female and male gametophytes (d). Lane 1 Colony PCR amplified product with 813-bp target plus 8 nucleotides added for endonuclease digestion; lane 2 empty vector as a control without insertion of SjCA; lane 3 overexpression of SjCA induced by IPTG for 3 h; lane 4 the recombinant protein of SjCA; lane 5 empty vector as a control without insertion of SjCA; lane 6 the total soluble proteins from the female gametophytes; lane 7 the total soluble proteins from the male gametophytes; M1 D-2000 marker of standard DNA (Tiangen Biotech Co., Ltd.); M2 prestained protein ladder of standard protein (Fermentas)
A recombinant protein of SjCA was induced to be expressed in E. coli by the addition of ITPG, and its molecular weight was about 32.5 kDa (Fig. 5b) as expected to contain the 30.34 kDa mature SjCA moiety and the fused His-tag sequence at the C terminus. The polyclonal antibody of this SjCA was obtained from the immunized rabbits with this purified recombinant protein. The specificity of this SjCA antibody was verified by Western blot (Fig. 5c), and only one hybridization signal corresponding to the expected size of SjCA was present in the transformant harboring pET28a-CA/BL21 and female and male gametophytes (Fig. 5d), but absent in the transformed E. coli without insertion of SjCA (Fig. 5c). Therefore, the SjCA was proved to be present in both the male and female S. japonica gametophytes, and the purified anti-CA antibody could be used in immunoelectron microscopy for subcellular localization of this SjCA in the kelp gametophyte cells.
Subcellular localization of SjCA using immunoelectron microscopy
The ultrathin sections of S. japonica gametophytes incubated sequentially with the purified antibody of SjCA and the secondary antibody, antirabbit IgG conjugated gold particles, showed that more than 92 % of immunogold particles labeled with the anti-SjCA antibody were found decorating the kelp chloroplasts, and the labeling appeared either as isolated or clustered particles (Fig. 6). The immunogold control experiment with the primary antibody absent did not show any significant labeling. The labeling density in the chloroplasts was 7.3 particles μm−2 (n = 10), which was much higher than that in the other regions of the gametophyte cells (1.4 particles μm−2). Of the labeled gold particles in the chloroplasts, 20.6 % were found on the chloroplast envelopes, whereas 70.1 % were on the thylakoid membranes to face to the stroma, but only 9.3 % were in the stroma of chloroplasts. This subcellular distribution of the SjCA protein was significantly different (P < 0.05), but there was no difference of labeled gold particles present between the male and female gametophytes.
Transmission electron micrograph showing the immunogold labeling in the gametophyte cells of Saccharina japonica probed with the SjCA antibody. a Ultrastructure of a whole cell of S. japonica gametophytes. b–d Enlarged images corresponding to the marked areas 1, 2, and 3 in a showing the immunogold labeling distribution. e Shows the immunogold labeling distribution especially in a pyrenoid. Gold particles were localized on the envelopes and thylakoid membranes of chloroplasts as indicated by white arrows. Ch chloroplast, Nu nucleus, Py pyrenoid
This immunoelectron microscopic result indicated the precise subcellular location of this SjCA in the gametophyte cells in addition to the prediction based on both the presence of a plastid transit peptide at the N terminus of the SjCA and the insignificantly changed transcription levels of this SjCA under different carbon sources. Because the SjCA was not a transmembrane protein as mentioned above, this SjCA protein should be an extrinsic membrane protein associated with the chloroplast envelopes and the thylakoid membranes.
Discussion
In this study, we have documented the cloning and characterization of a gene, which encodes an intracellular α-type CA from S. japonica gametophytes. Using specific antibodies raised against the recombinant SjCA protein, immunoelectron micrographs illustrate that this SjCA is located on the chloroplast envelopes and thylakoid membranes.
CAs are zinc-metalloenzymes that catalyze the reversible interconversion of CO2 and HCO3 − (Badger and Price 1992). The known CAs in eukaryotes are generally divided into six families (Moroney et al. 2011), of which α-, β-, and γ-CA have been reported to be distributed widely in microalgae and higher plants, whereas δ-, ε-, and ζ-CA are found only in some diatoms (Roberts et al. 1997), bacteria (So et al. 2004), and marine protists (Lane and Morel 2000; Park et al. 2007). Most α-CAs are active as monomers of about 30 kDa with three histidines coordinating the zinc atom (Moroney et al. 2001). In contrast, the β-CA family is characterized as the histidine and two cysteine residues coordinating with the zinc atom, and all the γ-CAs that are reported to be localized only in mitochondria of C. reinhardtii (Ynalvez et al. 2008). The putative molecular mass of 30.34 kDa (Fig. 5), conserved His 118, His 120 and His 137 (Fig. 3) acting as Zn ligands, phylogenetic tree (Fig. 2), and localization on chloroplasts (Fig. 6) indicate that the cloned SjCA from S. japonica gametophytes belongs to α-CA rather than the β- or γ-CA families.
Chlamydomonas reinhardtii, a model organism, has 12 genes coding for CA (Moroney et al. 2011). It has been reported that two CAs, i.e., CAH3 and CAH6, are situated in the chloroplasts of this model organism. The N-terminal sequence of CAH3, an α-CA gene product, is consistent with a lumenal localization as it has a two-part leader sequence, one part required to enter the chloroplast and a second part directing the protein to the thylakoid lumen. In the second part of this leader sequence, there is a hydrophobic region often found in proteins directed to the thylakoid lumen, which contains a twin arginine motif followed by a conserved lumen peptidase cleavage site, Ala–X–Ala (Karlsson et al. 1998; Sinetova et al. 2012). However, the conserved lumen peptidase cleavage site was not found in the SjCA. CAH6 encoded by a β-CA gene is present in the chloroplast stroma of C. reinhardtii strain D66 instead of the thylakoid lumen like CAH3, since it has only one leader sequence that is consistent with a chloroplast location (Mitra et al. 2004). Regarding the similar subcellular localization, it is expected that this SjCA functions in the kelp chloroplasts as same as C. reinhardtii CAH6 even though they does not belong to a same family or have homologous sequences.
It has been reported that Immunogold density of CAH6 is fourfold higher in the area around the pyrenoids compared to that in the other areas in the chloroplast stroma of C. reinhardtii (Mitra et al. 2004). However, this distribution of CAH6 was not found in S. japonica. The SjCA was scattered in instead of surrounding the pyrenoids of S. japonica even though there was a slightly more density of labeling (Fig. 6e). Thus, what the CAH6 reduced CO2 leakage out of the C. reinhardtii pyrenoids was not proposed to be a role that this SjCA played. Accordingly, with the help of SjCA, the supply of CO2, usually diffusing very slowly in the cytosol or stroma, is expectedly speeded up to the RuBisCo as same as CAH6 does in C. reinhardtii (Mitra et al. 2004). The colocalization of SjCA and RuBisCo in this kelp will surely provide direct evidence for this suggested function.
Both the absence of a hydrophobic transmembrane helix and the majority of immunogold particles very close to the chloroplast envelopes and thylakoid membranes (Fig. 6) suggest that the SjCA might be a peripheral protein. However, we do not know how this SjCA is associated with the chloroplast envelopes and thylakoid membranes. In C. reinhardtii CW92, CAH3 is reported to be associated with PSII (Park et al. 1999). That is to say, CAH3 is associated with thylakoid membranes via the PSII complex. Accordingly, the association of the SjCA with the chloroplast envelopes and thylakoid membranes would likely be through some protein–protein interaction, as the mature SjCA has some predicted binding sites, such as the Arg- and Lys-enriched domain RKLAPKK at its N terminus. In Nicotiana tabacum, Jebanathirajah and Coleman (1998) found the chloroplast α-CA was associated with RuBisCo at the stromal periphery of a Calvin cycle enzyme complex in which phosphoribulokinase was more centrally located and associated with thylakoid membranes. A number of Lys and Arg residues were suggested to be involved in the specific protein–protein interactions (Süss et al. 1993; Jebanathirajah and Coleman 1998). This association in higher plants directs us to the future studies on this SjCA using subcellular fractions, immunoprecipitation, yeast two-hybrid, and Western blot analysis.
It needs to be pointed out that the present study only focuses on the microscopic gametophytes of S. japonica. In comparison to the gametophytes, the macroscopic sporophytes of this kelp are multicellular in morphology. Supply of CO2 to the RuBisCo for photosynthetic fixation in sporophytes could be different from that in gametophytes since α-CA messenger RNA was shown by both real-time PCR and Northern blot analyses to be present only in the gametophytes of L. digitata (Moulin et al. 1999). Recently, the transcriptomic analysis of this kelp (Deng et al. 2012) and of another species, S. latissima (Heinrich et al. 2012), in the same genus, at least reveals that all three CA genes, i.e., α-, β-, and γ-CAs, are present and expressed in this kelp. Upon cloning and subcellular localization of these CA genes like this SjCA, the mechanism of CO2 supply for photosynthetic fixation in this marine macroalga will be unveiled.
References
Andreu V, Collados R, Testillano PS, Risueno MC, Picorel M, Alfonso M (2999) In situ molecular identification of the plastid omega-3 fatty acid desaturase FAD7 from soybean: evidence of thylakoid membrane localization. Plant Physiol 145:1336–1344
Axelsson J, Mercado JM, Figueroa FL (2000) Utilization of HCO3 − at high pH by the brown macroalga Laminaria saccharina. Eur J Phycol 35:53–59
Axelsson L, Uusitalo J, Ryberg H (1991) Mechanisms for concentrating and storage of inorganic carbon in marine macroalgae. In: García Reina G, Pedersén M (eds) Seaweed cellular biotechnology, physiology and intensive cultivation, COST -48. Universidad de las Palmas de Gran Canaria, Las Palmas, pp 185–198
Badger MR, Price GD (1992) The CO2 concentrating mechanism in cyanobacteria and microalgae. Physiol Plant 84:606–615
Bendayan M, Zollinger M (1983) Ultrastructural localization of antigenic sites on osmium-fixed tissues applying the protein A-gold technique. J Histochem Cytochem 31:101–109
Bernal M, Testillano PS, Alfonso M, Del Carmen Risueño M, Picorel R, Yruela I (2007) Identification and subcellular localization of the soybean copper P1B-ATPase Gm HMA8 transporter. J Struct Biol 158:46–58
Bradford MM (1976) A rapid and sensitive method for quantification of microgram quantities of protein utilizing the principles of protein dye binding. Anal Biochem 72:245–248
Chen J, Wang L-L, Shi W-W, Zhou Z-G (2010) Cloning of srp gene from the gametophytes of Laminaria japonica and its expression in Escherichia coli. J Fish China 34:1165–1173
Deng Y, Yao Y, Wang X, Guo H, Duan D (2012) Transcriptome sequencing and comparative analysis of Saccharina japonica (Laminariales, Phaeophyceae) under blue light induction. PLoS ONE 7:e39704
Dring MJ, Forster RM, Schmid R (1994) Ecological significance of blue light stimulation of photosynthetic capacity in Laminaria spp. and other brown algae. Mar Ecol Prog Ser 113:271–277
Fukuzawa H, Fujiwara S, Yamamoto Y, Dionisio-Sese ML, Miyachi S (1990) cDNA cloning, sequence, and expression of carbonic anhydrase in Chlamydomonas reinhardtii: regulation by environmental CO2 concentration. Proc Natl Acad Sci U S A 87:4383–4387
Gao K, McKinley KR (1994) Use of macroalgae for marine biomass production and CO2 remediation: a review. J Appl Phycol 6:45–60
Haglund K, Ramazanov Z, Mtolera M, Pedersen M (1992) Role of external carbonic anhydrase in light-dependent alkalization by Fucus serratus L. and Laminaria saccharina (L.) Lamour. (Phaeophyta). Planta 188:1–6
He P, Wu W, Zhao J, Chen G, Zhang D (2002) Studies on ultrastructure of pyrenoid from several algae. Acta Hydrobiol Sinica 26:327–334
Heinrich S, Valentin KU, Frickenhaus S, John U, Wiencke C (2012) Transcriptomic analysis of acclimation to temperature and light stress in Saccharina latissima (Phaeophyceae). PLoS ONE 7:e44342
Henderson C (2006) Ocean acidification: the other CO2 problem. New Scientist. Retrieved from http://environment. newscientist.com/article/mg19125631.200.
Hewett-Emmett D, Tashian RE (1996) Functional diversity, conservation, and convergence in the evolution of the α-, β-, and γ-carbonic anhydrase gene families. Mol Phylogenet Evol 5:50–77
Hu Y-J, Zhou Z-G (2001) Extraction of RAPD-friendly DNA from Laminaria japonica (Phaeophyta) after enzymatic dissociation of the frozen sporophyte tissue. J Appl Phycol 13:415–422
Huang S, Xue Y, Sauer-Eriksson E, Chirica L, Lindskog S, Jonsson BH (1998) Crystal structure of carbonic anhydrase from Neisseria gonorrhoeae and its complex with the inhibitor acetazolamide. J Mol Biol 283:301–310
Jebanathirajah JA, Coleman JR (1998) Association of carbonic anhydrase with a Calvin cycle enzyme complex in Nicotiana tabacum. Planta 204:177–182
Karlsson J, Clarke AK, Chen Z-Y, Hugghins SY, Park Y-I, Husic HD, Moroney JV, Samuelsson G (1998) A novel α-type carbonic anhydrase associated with the thylakoid membrane in Chlamydomonas reinhardtii is required for growth at ambient CO2. EMBO J 17:1208–1216
Khalifah RG (1971) The carbon hydroxide hydration activity of carbonic anhydrase. I. Stop-flow kinetic studies on the native human isozymes B and C. J Biol Chem 246:2561–2573
Laemmli UR (1970) Cleavage of structural proteins during the assembly of the head of Bacteriophage T4. Nature 227:680
Lane TW, Morel FMM (2000) Regulation of carbonic anhydrase expression by zinc, cobalt, and carbon dioxide in the marine diatom Thalassiosira weissflogii. Plant Physiol 123:345–352
Liljas A, Laurberg M (2000) A wheel invented three times: the molecular structures of three carbonic anhydrases. EMBO Rep 1:16–17
Liu Y, Bi YH, Gu JG, Zhou ZG (2012) Localization of a female-specific marker on the chromosomes of the brown seaweed Saccharina japonica using fluorescence in situ hybridization. PLoS ONE 7:e48784
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔCT method. Methods 25:402–408
Miller SE, Howell DN (2006) Immunoelectron Microscopy. In: Howard GC, Kaser MR (eds) Making and using antibodies: a practical handbook. CRC, Boca Raton, pp 315–337
Mitra M, Lato SM, Ynalvez RA, Xiao Y, Moroney JV (2004) Identification of a new chloroplast carbonic anhydrase in Chlamydomonas reinhardtii. Plant Physiol 135:173–182
Moroney JV, Bartlett SG, Samuelsson G (2001) Carbonic anhydrases in plants and algae. Plant Cell Environ 24:141–153
Moroney JV, Ma Y, Frey WD, Fusilie KA, Pham TT, Simms TA, DiMario RJ, Yang J, Mukherjee B (2011) The carbonic anhydrase isoforms of Chlamydomonas reinhardtii: intracellular location, expression, and physiological roles. Photosynth Res 109:133–149
Moulin P, Crépineau F, Kloareg B, Boyen C (1999) Isolation and characterization of six cDNAs involved in carbon metabolism in Laminaria digitata (Phaeophyceae). J Phycol 35:1237–1245
Olmsted JB (1981) Affinity purification of antibodies from diazotized paper blots of heterogeneous protein samples. J Biol Chem 256:11955–11957
Ouyang LL, Du DH, Yu SY, Li CY, Zhang CW, Gao HJ, Zhou ZG (2012) Expressed sequence tags analysis revealing the taxonomic position and fatty acid biosynthesis in an oleaginous green microalga, Myrmecia incisa Reisigl (Trebouxiophyceae, Chlorophyta). Chin Sci Bull 57:3342–3352
Park YI, Karlsson J, Rojdestvenski I, Pronina N, Klimov V, Öquist G, Samuelsson G (1999) Role of a novel photosystem II-associated carbonic anhydrase in photosynthetic carbon assimilation in Chlamydomonas reinhardtii. FEBS Lett 444:102–105
Park H, Song B, Morel FMM (2007) Diversity of the cadmiumcontaining carbonic anhydrase in marine diatoms and natural waters. Environ Microbiol 9:403–413
Ritter K (1991) Affinity purification of antibodies from sera using polyvinylidenedifluoride (PVDF) membranes as coupling matrices for antigens presented by autoantibodies to triosephosphate isomerase. J Immunol Methods 137:209–215
Roberts SB, Lane TW, Morel FMM (1997) Carbonic anhydrase in the marine diatom Thalassiosira weissflogii (Bacillariophyceae). J Phycol 33:845–850
Sabine CL, Feely RA (2007) The oceanic sink for carbon dioxide. In: Reay D, Hewitt N, Grace J, Smith K (eds) Greenhouse gas sinks. CABI, Oxfordshire, pp 31–49
Sabine CL, Feely RA, Gruber N, Key RM, Lee K, Bullister JL, Wanninkhof R, Wong CS, Wallace DWR, Tilbrook B, Millero FJ, Peng TH, Kozyr A, Ono T, Rios AF (2004) The oceanic sink for anthropogenic CO2. Science 305:367–371
Schmid R, Mills JA, Dring MJ (1996) Influence of carbon supply on the stimulation of light-saturated photosynthesis by blue light in Laminaria saccharina: implications for the mechanism of carbon acquisition in higher brown algae. Plant Cell Environ 19:383–391
Shi XZ, Bi YH, Zhou ZG (2005) Cloning and screening of differentially expressed genes from the male gametophytes of Laminaria japonica Aresch. J Fish China 29:666–669
Shi J, Tan H, Yu LY, Liang W, Ranathunge K, Benni Franke R, Schreiber L, Wang Y, Kai G, Shanklin J, Ma H, Zhang D (2011) Defective Pollen Wall is required for anther and microspore development in rice and encodes a fatty acyl carrier protein reductase. Plant Cell 23:2225–2246
Siegenthaler U, Sarmiento JI (1993) Atmospheric carbon dioxide and the ocean. Nature 365:119–125
Sinetova MA, Kupriyanova EV, Markelova AG, Allakhverdiev SI, Pronina NA (2012) Identification and functional role of the carbonic anhydrase Cah3 in thylakoid membranes of pyrenoid of Chlamydomonas reinhardtii. Biochim Biophys Acta 1817:1248–1255
So AK, Espie GS, Williams EB, Shively JM, Heinhorst S, Cannon GC (2004) A novel evolutionary lineage of carbonic anhydrase (epsilon class) is a component of the carboxysome shell. J Bacteriol 186:623–630
Solomon S, Qin D, Manning M, Marquis M, Averyt KB, Tignor M, Mille HL, Chen Z (2007) Climate change 2007: the physical science basis. Cambridge University Press, Cambridge, 996 pp
Spalding MH (2008) Microalgal carbon-dioxide-concentrating mechanisms: Chlamydomonas inorganic carbon transporters. J Exp Bot 59:1463–1473
Starr RC, Zeikus JA (1993) UTEX-The culture collection of algae at the University of Texas at Austin. J Phycol 29/Suppl:1–106
Süss K-H, Arkona C, Manteuffel R, Adler K (1993) Calvin cycle multienzyme complexes are bound to chloroplast thylakoid membranes of higher plants in situ. Proc Natl Acad Sci U S A 90:5514–5518
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: Molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucl Acids Res 25:4876–4882
Tseng CK (2001) Algal biotechnology industries and research activities in China. J Appl Phycol 13:375–380
Turley C (2005) The other CO2 problem. openDemocracy. http://www.acamedia.info/sciences/sciliterature/globalw/reference/carol turley.html
Ynalvez RA, Xiao Y, Ward AS, Cunnusamy K, Moroney JV (2008) Identification and characterization of two closely related β-carbonic anhydrases from Chlamydomonas reinhardtii. Physiol Plant 133:15–26
Yu Z, Bi Y-H, Zhou Z-G (2011) Cloning and characterization of carbonic anhydrase (CA) gene from Laminaria japonica gametophytes. J Fish China 35:1343–1353
Zhou Z-G, Wu C-Y (1998) Clone culture of Laminaria japonica and induction of its sporophytes. Chin J Biotech 14:109–111
Acknowledgments
This research was supported by the National High Technology Research and Development Program of China (2012AA10A406) and the National Natural Science Foundation of China (31201992 and 30471328).
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Ye, RX., Yu, Z., Shi, WW. et al. Characterization of α-type carbonic anhydrase (CA) gene and subcellular localization of α-CA in the gametophytes of Saccharina japonica . J Appl Phycol 26, 881–890 (2014). https://doi.org/10.1007/s10811-013-0221-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10811-013-0221-2