Abstract
Common bean (Phaseolus vulgaris) is an important export crop in Kyrgyzstan since the end of the twentieth century. Genetic diversity analysis of common bean populations is useful for breeding programs, as it helps to select genetic material to be used for further crossing. Twenty-seven common bean accessions were analyzed using 13 qualitative morphological traits. In some cases, obtained morphological data were combined with previously published results based on microsatellite markers. The similarity matrices generated from the molecular and morphological data were significantly correlated (r = 0.49, P < 0.01). Cluster analyses based on Dice’s similarity coefficient were constructed based on morphological data and the combined data set of morphology and microsatellite, and both grouped the 27 accessions according to their origin: 15 belonged to the Andean and 12 to the Mesoamerican gene pool. On average, the Andean accessions were less diverse than the Mesoamerican accessions. The average diversity based on the Shannon diversity index for the 13 qualitative morphological traits was 0.05. Overall, this study revealed that qualitative morphological markers are efficient in assigning modern cultivars to their gene pools of origin.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Common bean (Phaseolus vulgaris L.) is an annual, self-pollinated crop which is grown on more than 12 million ha in the world. Common bean provides a cheap source of protein in the developing world, especially in Latin America and Africa (CIAT 1989; Graham and Ranalli 1997). The common bean was domesticated independently 8,000 to 10,000 years ago in South America (The Andes) and 7,000 years ago in Mesoamerica (Kaplan 1981). Bean type classification into two gene pools was made based on morphological traits and phaseolin seed proteins by Gepts et al. (1986) and Gepts (1988), and based on morphological characters and allozymes by Singh et al. (1991c). The two gene pools were further differentiated into races using agro-morphological traits (Singh et al. 1991a) and later confirmed by different types of DNA markers such as random amplified polymorphic DNA (RAPD; Beebe et al. 2000), amplified fragment length polymorphism (AFLP; Beebe et al. 2001), and microsatellites (SSR; Blair et al. 2003, 2006, 2009; Diaz et al. 2011).
In 1916, the Russian geneticist Nicolai Vavilov organized a plant collecting expedition to Central Asia (Tajikistan, Kyrgyzstan and Uzbekistan) and brought two cultivated P. vulgaris samples to the VIR institute from the Pamir Mountains (Tajikistan). Common bean cultivars in the former Soviet Union were initially developed from cultivars collected in the country and foreign breeding material and cultivars introduced from 1921 to 1923. The introduced material originated from American and Canadian breeding stations and seed companies: Hidatsa red, North Dakota, Refugee, Valley Seed Co, Sacramento, Cal and others (Buravtseva and Egorova 2012).
Common bean cultivars were most likely introduced to Central Asia (including Kyrgyzstan) by the Soviets during the last century (Hegay et al. 2012). When Kyrgyzstan achieved its independence in 1991, the agricultural land belonged to the State but this changed after the privatization process. From 1991 to 1996 collective farms (kolkhozes and sovhozes) were transformed into private farms. About 344,500 small-scale farms are registered today and they own together 1.28 million ha, (6.4 %) of the agricultural land in Kyrgyzstan (STATCOM 2011). The majority of the population depends on agriculture. Farmers grow cereals crops like wheat and barley, but small-scale farmers from the Talas and Chui oblasts are increasingly switching to common beans. Consequently, in these two oblasts the population meets the food calorie and protein requirements when compared to other oblasts (Asanaliev and Nurgaziev 2012).
The Kyrgyz common bean market started to develop in the end of the twentieth century. In 2010, 71,400 t of beans were produced (FAOSTAT 2010), and 90 % of the harvest was exported mainly to Turkey, Bulgaria and Russia (STATCOM 2011). Kyrgyzstan has a moderate bean production compared with other grain-bean producing countries (Beebe et al. 2011), but ranks however among the top 20 bean grain exporters worldwide (Akibode and Maredia 2011). The income from selling common beans (grains) is 1 billion Kyrgyz soms (approx. US$ 20 million; FAOSTAT 2009). Kyrgyz farmers grow different types of market bean classes, and sometimes use cultivar mixtures because they believe these will give a higher yield. Furthermore, the market price for different types of seeds is not stable from year to year, which also supports these cultivation practices.
The objective of the present study was to assess the diversity of Kyrgyz cultivars and a reference set of foreign common bean accessions using morphological qualitative traits and compare the results with previously published microsatellite marker data. The ultimate goal is to identify genetic variation useful for the Kyrgyz bean breeding program.
Materials and methods
Plant material
Five Kyrgyz cultivars were selected since they are widely grown in Kyrgyzstan. Seeds of foreign accessions were kindly provided by Michigan State University (East Lansing), and the United State Department of Agriculture (Pullman). Altogether, 27 accessions (Table 1) were characterized using morphological trait descriptors described by Singh et al. (1991a) (Table 2).
Data analysis
Data for 13 qualitative morphological traits were recorded on 10 randomly chosen individual plants per accession (Table 2). Qualitative morphological traits were binary-coded as 1 for presence or 0 for absence for each individual plant (e.g., pod beak position placental: presence (1) or absence (0); pod beak position central: presence (1) or absence (0) since common bean is a self-pollinated crop and we did not expect to find any heterozygotes. The Shannon diversity index (I) and percent polymorphism (%P) were calculated for each accession using POPGENE version 1.31 (Yeh and Boyle 1997).
Previously published microsatellite data (Hegay et al. 2012) for the accessions used in this study were also binary-coded and combined with the morphological data for cluster, principal coordinate analysis (PCoA) and STRUCTURE analyses in order to obtain a better genetic information about the common bean accessions. Cluster analyses were performed based on Dice’s similarity coefficient (Dice 1945) according to the unweighted pair group method with arithmetic average (UPGMA) using the sequential agglomerative hierarchical nested clustering (SAHN). The analyses were done using NTSYS-pc and FreeTree software (Rohlf 2000; Pavlicek et al. 1999). Principal coordinate analyses (PCoA) were performed based on the simple matching coefficient since this coefficient takes into account both the shared presence and absence of a particular character when estimating the similarity between two individuals. A two-way Mantel (1967) test was used to test the hypothesis of an equal precision for genotypic and phenotypic data to classify the bean accessions into a gene pool. The goodness of fit for the UPGMA trees and PCoA (using Dcenter and Eigenvectors) matrices were performed using NTSYS-pc with 10,000 random permutations. The bootstrap values for the UPGMA dendrograms were obtained via a 1,000 resampling procedure using the FreeTree program (Pavlicek et al. 1999). The TreeView program (Page 1996) was used to display the trees. The software STRUCTURE (Pritchard et al. 2000) was used for structure analysis based on the combination of morphological and microsatellite data. The admix model with 5,000 burning periods and 50,000 replicates was used to estimate each K value, with ten independent runs from K = 1 to 10. Delta K (population number) was estimated as described by Evanno et al. (2005), and population clusters were produced using the DISTRUCT software (Rosenberg 2004).
Discriminant analysis (DA), principal component analysis (PCA) and the best subset regression were used to estimate the diversity among accessions and their grouping, and were performed using the Minitab 15 statistical software (Minitab Inc 2008). DA was used to distinguish between accessions and divide them into groups based on morphological traits. DA grouped accessions with typical characters, and estimated the correct and incorrect percentage of classifications (Dytham 2011). DA maximizes differences between classes while minimizing those within classes, which is different from the PCA. PCA was used to analyze the diversity and to identify the optimum number of morphological traits which explain a high proportion of the variability. The Scree plot was used to display Eigenvalues and number of morphological traits in PCA. The best subset regression statistical method was used to determine a model for the grouping of accessions based on morphological traits.
Results
Diversity within and among common bean accessions
The average Shannon diversity index within common bean accessions estimated based on qualitative morphological traits was 0.05 (Table 1). Among the 27 accessions, only four were polymorphic for the thirteen traits. Accession PI527537 from Burundi had the highest diversity and the highest percent polymorphism. No polymorphism in all traits was observed in the Kyrgyz bean cultivars. On average, a higher genetic diversity was observed in the 12 Mesoamerican accessions (Shannon index of 0.08) as compared to the 15 Andean accessions (Shannon index of 0.02). The pair-wise comparisons of Dice’s coefficient of similarity between accessions estimated based on the qualitative morphological traits ranged from 0.154 (between PI543043 and PI451886) to 0.923 (between Kytayanka and Lopatka).
Grouping of accessions based on the qualitative morphological traits
The principal component analysis (PCA) showed that three of the thirteen morphological traits were the most important components for explaining the grouping of accessions (Fig. 1). To identify these three most important traits, a subset regression analysis was performed. The analysis determined seed size, pod beak position, size and shape of the bract as the most important with a correlation of R 2 = 79 % (Table 3). Morphological traits as plant growth habit and pod string may be included as secondary predictors, while the other eight morphological traits were not identified as important by this analysis. A discriminant analysis based on morphological traits was used to differentiate between Andean and Mesoamerican accessions and to assign them to the correct group (Table 4). Seed size and pod beak position were effective for the grouping of the accessions. Andean accessions had medium, large or very large seed sizes and a central beak position, while Mesoamerican accessions had small and medium seed sizes and a placental beak position. Overall, the morphological traits included in this study were able to properly assign 99 % of individuals into their respective gene pools.
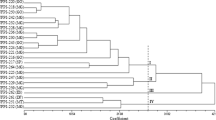
Two main groups were observed in the UPGMA cluster analysis with 100 % bootstrap support (Fig. 2). The first group (cluster I) included the 12 Mesoamerican accessions and the second group (cluster II) consisted the 15 Andean accessions. The principal coordinate analysis based on a combination of microsatellite and morphological data grouped the accessions into two main clusters (Fig. 3). The goodness of fit of matrix comparisons (phenotypic versus genotypic data) was r = 0.49 (P < 0.01). The first and second co-ordinate explained 52 % of the total variation. Cluster Ia and Ib included accessions belonging to the Mesoamerican gene pool, and cluster II comprised accessions that belong to the Andean gene pool (Fig. 3). A UPGMA dendrogram generated based on the combination of microsatellite and morphological data (Fig. 4) was similar with that constructed based only on morphological data (Fig. 2). In both dendrograms, the accessions were grouped into two main groups with high bootstrap support. A two-way Mantel test showed a highly significant correlation of the cophenetic values from the two independent UPGMA cluster analyses (r = 0.95, P = 0.01). The cluster analysis with K = 2 using Evanno’s methods of STRUCTURE defined two groups of accessions corresponding to the Mesoamerican and the Andean gene pools, and showed that accession PI527537 was a mixture according to both morphological and molecular data (Fig. 5).
Dice’s similarity coefficient based unweighted pair group method with arithmetic average (UPGMA) cluster analysis that clustered 27 common bean accessions into Mesoamerican (cluster I) and Andean (cluster II) groups based on qualitative morphological data. The bootstrap value from 1,000 resampling is indicated in between two branches
Discussion
The average genetic diversity within common bean accessions estimated using morphological qualitative traits in the present study (0.05) was lower than that estimated with microsatellite markers (0.07) (Hegay et al. 2012. In the present study, common bean accessions were clearly separated into two groups corresponding to the Mesoamerican and Andean gene pools. There was, however, no large variation in a majority of the morphological traits among the accessions from the two gene pools. This is probably due to the fact that many of the accessions were received from gene banks and may suffer from a reduction of polymorphism in the past; i.e., accessions with traits not amenable to industrial bean production have been eliminated.
Qualitative morphological traits and appropriate statistical methods such as PCA, DA and STRUCTURE were used to assign accessions into gene pools, which agreed with those made by Burle et al. (2011). DA, best subset regression analysis and PCA identified seed size, pod beak positions and size and shape of the bract as important for grouping of accessions into gene pools, which agree with the results by Singh et al. (1991a). Variation in pod string and growth habit can also be suitable traits for gene pool classification (Table 3). As a whole, qualitative morphological traits were useful when differentiating accessions and assigning them into respective gene pools (Table 5).
The results from the present study were in line with previous study by Singh et al. (1991b), who reported that variation of morphological characters in common beans could be independent variables and that the same morphological pattern can be found in different gene pools. The high correlation of the cophenetic values from the UPGMA analyses (r = 0.95) agrees with Kumar et al. (2008), who found a high correlation value (r = 0.934) for the clustering pattern when AFLP markers were used for diversity analysis in Indian common bean accessions.
The similarity coefficients of Jaccard and Dice were highly correlated with the simple matching coefficient, which was visually demonstrated in the analysis of P. vulgaris (Beharav et al. 2010; Duarte et al. 1999), and the use of either of these coefficients did not affect the grouping of common bean accessions into their genetic origin. In the present study, cluster analysis (UPGMA) and PCoA grouped bean accessions based on their genetic relationships or morphology rather than by country of origin, which agrees with previous research (Islam et al. 2002; Sharma et al. 2013). Principal coordinate analysis based on microsatellite data separated most accessions into two groups and only one accession (PI527537 from Burundi) was intermediate. The UPGMA dendrogram supported the PCoA grouping with an intermediate placement of accession PI527537. Seed mixtures, which are preferred by local consumers in Burundi (Wortmann et al. 1998) or the result of natural crosses between individuals from the two gene pools, could explain the placement in between two gene pools, which was suggested also by Hegay et al. (2012). In addition, we found that accession PI337090 from Brazil showed variability for both morphological and molecular data (Table 1); but, unlike accession PI527537, it did not include genotypes from the Andean gene pool (Fig. 5). This result was not surprising, because the small-scale farmers represent the Brazilian common bean industry and they usually grow commercial varieties together without purity control of the varieties (Burle et al. 2011).
The STRUCTURE analysis verified the two gene pools and grouped 12 accessions into the Mesoamerican gene pool and 15 accessions into the Andean gene pool. This clear clustering suggests that the recombination between the two gene pools is limited. Hybridizations between gene pools depend on the presence of the complementary dominant Dl 1 and Dl 2 genes which control traits that provide barriers between common beans of different geographic origins (Singh and Gutierrez 1984).
There was a moderate significant correlation between matrices derived from qualitative morphological traits and microsatellite data (r = 0.49, Fig. 3), which agrees with results from other matrix comparisons (r = 0.50) by Asfaw et al. (2009). However, the presence of two gene pools was more strongly supported by microsatellite data than by data from morphological traits. Microsatellites occur in coding and non-coding regions that are not always linked to the genes expressing morphological traits. Hybrid phenotypes after introgressing genes from one gene pool to another are difficult to differentiate based on morphology (Paredes and Gepts 1995).
Whether to use only morphological traits (both qualitative and quantitative) or to combine them with DNA markers is a matter of research aim and practical applications. For example, gene bank curators and plant breeders often use a combination of (usually more than 10) morphological traits and DNA markers for germplasm characterization, evaluation and utilization. In the present study, the use of only qualitative morphological traits was sufficient for separation of common bean accessions and for assigning them into two main gene pools, but the pattern was even more distinct when molecular markers were added. The number of morphological traits (13) agrees with the number of traits used for common bean in ex situ preservation in gene banks (Chiorato et al. 2006), or in situ preservation on farm management, and for diversity research (Gomez et al. 2005).
In conclusion, common beans characterized both with morphological traits and microsatellites were grouped into clusters corresponding to their gene pools of origin. Kyrgyz cultivars belonged to both Andean and Mesoamerican gene pools as previously shown by Hegay et al. (2012). Classification and divergence between common bean accessions analyzed in this study may help to preserve plant material both in situ and ex situ. Furthermore, our study provides important information to the Kyrgyz breeders that helps to optimize the selection of plant material to be used in breeding programs of this very important grain legume crop.
References
Akibode S, Maredia M (2011) Global and regional trends in production, trade and consumption of food legume crops. Department of Agricultural, Food and Resource Economics, USA
Alves-Santos FM, Cordeiro-Rodrigues L, Sayagues JM, Martin-Dominguez R, Garcia-Benavides P, Crespo MC, Diaz-Minguez JM, Eslava AP (2002) Pathogenicity and race characterization of Fusarium oxysporum f. sp phaseoli isolates from Spain and Greece. Plant Pathol 51:605–611
Asanaliev AJ, Nurgaziev RZ (2012) Food chain of agriculture of Kyrgyzstan—gained experience, learned lessons and development perspectives. In: Alpas H, Smith M, Kulmyrzaev A (eds) Strategies for achieving food security in Central Asia. Springer, The Netherlands, pp 21–30
Asfaw A, Blair MW, Almekinders C (2009) Genetic diversity and population structure of common bean (Phaseolus vulgaris L.) landraces from the East African highlands. Theor Appl Genet 120:1–12
Balardin RS, Jarosz AM, Kelly JD (1997) Virulence and molecular diversity in Colletotrichum lindemuthianum from South, Central, and North America. Phytopathology 87:1184–1191
Beebe S, Skroch PW, Tohme J, Duque MC, Pedraza F, Nienhuis J (2000) Structure of genetic diversity among common bean landraces of Middle American origin based on correspondence analysis of RAPD. Crop Sci 40:264–273
Beebe S, Rengifo J, Gaitan E, Duque MC, Tohme J (2001) Diversity and origin of Andean landraces of common bean. Crop Sci 41:854–862
Beebe S, Ramirez J, Jarvis A, Rao IM, Mosquera G, Bueno JM, Blair MW (2011) Genetic improvement of common beans and the challenges of climate change. In: Shyam RJR, Yadav S, Hatfield JL, Lotze-Campen H, Hall AE (eds) Crop adaptation to climate change. Wiley-Blackwell, UK, pp 356–369
Beharav A, Maras M, Kitner M, Šuštar-Vozlič J, Sun GL, Doležalová I, Lebeda A, Meglič V (2010) Comparison of three genetic similarity coefficients based on dominant markers from predominantly self-pollinating species. Biol Plantarum 54:54–60
Blair MW, Pedraza F, Buendia HF, Gaitan-Solis E, Beebe SE, Gepts P, Tohme J (2003) Development of a genome-wide anchored microsatellite map for common bean (Phaseolus vulgaris L.). Theor Appl Genet 107:1362–1374
Blair MW, Giraldo MC, Buendia HF, Tovar E, Duque MC, Beebe SE (2006) Microsatellite marker diversity in common bean (Phaseolus vulgaris L.). Theor Appl Genet 113:100–109
Blair MW, Diaz LM, Buendia HF, Duque MC (2009) Genetic diversity, seed size associations and population structure of a core collection of common beans (Phaseolus vulgaris L.). Theor Appl Genet 119:955–972
Buravtseva TV, Egorova GP (2012) Common bean collection—100 years. J Legum Groat Crops 4:46–52
Burle ML, Fonseca JR, Jose del Peloso M, Melo LC, Temple SR, Gepts P (2011) Integrating phenotypic evaluations with a molecular diversity assessment of a Brazilian collection of common bean landraces. Crop Sci 51:2668–2680
Chiorato AF, Carbonell SAM, Dias LADS, Moura RR, Chiavegato MB, Colombo CA (2006) Identification of common bean (Phaseolus vulgaris) duplicates using agromorphological and molecular data. Genet Mol Biol 29:105–111
CIAT (1989) Centro Internacional de Agricultura Tropical. In: Schwartz HF, Pastor-Corrales MA (eds) Bean production problem in the tropics, 2nd edn. Cali, Colombia
Diaz LM, Buendia HF, Duque MC, Blair MW (2011) Genetic diversity of Colombian landraces of common bean as detected through the use of silver-stained and fluorescently labelled microsatellites. Plant Genet Resour 9:86–96
Dice LR (1945) Measures of the amount of ecologic association between species. Ecology 26:297–302
Duarte JM, dos Santos JB, Melo LC (1999) Comparison of similarity coefficients based on RAPD markers in the common bean. Genet Mol Biol 22:427–432
Dytham C (2011) Choosing and using statistics: a biologist’s guide. Blackwell, London
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620
FAOSTAT (2009) United Nations Food and Agriculture Organization. Statistical database. http://faostat.fao.org/site/567/default.aspx#ancor. Accessed 10 Aug 2012
FAOSTAT (2010) United Nations Food and Agriculture Organization. Statistical database. http://faostat.fao.org/site/567/default.aspx#ancor. Accessed 12 Jan 2013
Gepts P (1988) Phaseolin as an evolutionary marker. In: Gepts P (ed) Genetic resources of Phaseolus beans. Kluwer, The Netherland, pp 215–241
Gepts P, Osborn TC, Rashka K, Bliss FA (1986) Phaseolin protein variability in wild forms and landraces of the common bean (Phaseolus vulgaris): evidence for multiple centers of domestication. Econ Bot 40:451–468
Gomez OJ, Blair MW, Frankow-Lindberg BE, Gullberg U (2005) Comparative study of common bean (Phaseolus vulgaris L.) landraces conserved ex situ in genebanks and in situ by farmers. Genet Resour Crop Evol 52:371–380
Graham PH, Ranalli P (1997) Common bean (Phaseolus vulgaris L.). Field Crop Res 53:131–146
Hegay S, Geleta M, Bryngelsson T, Gustavsson L, Persson-Hovmalm H, Ortiz R (2012) Comparing genetic diversity and population structure of common beans grown in Kyrgyzstan using microsatellites. Sci J Crop Sci 1:63–75
Islam FMA, Basford KE, Redden RJ, Gonzalez AV, Kroonenberg PM, Beebe S (2002) Genetic variability in cultivated common bean beyond the two major gene pools. Genet Resour Crop Evol 49:271–283
Johnson WC, Gepts P (1994) Two new molecular markers linked to bc-3. Annu Rep Bean Improv Coop 37:206–207
Kaplan L (1981) What is the origin of the common bean? Econ Bot 35:240–254
Kumar V, Sharma S, Kero S, Sharma S, Sharma AK, Kumar M, Bhat V (2008) Assessment of genetic diversity in common bean (Phaseolus vulgaris L.) germplasm using amplified fragment length polymorphism (AFLP). Sci Hortic 116:138–143
Kwak M, Kami JA, Gepts P (2009) The putative Mesoamerican domestication center of Phaseolus vulgaris is located in the Lerma-Santiago basin of Mexico. Crop Sci 49:554–563
Mantel N (1967) The detection of disease clustering and a generalized regression approach. Cancer Res 27:209–220
Miklas PN, Hang AN, Kelly JD, Strausbaugh CA, Forster RL (2002) Registration of three kidney bean germplasm lines resistant to bean common mosaic and necrosis potyviruses: USLK-2 light red kidney, USDK-4 dark red kidney, and USWK-6 white kidney. Crop Sci 42:674–675
Minitab Inc (2008) Minitab. Minitab Inc, State College, PA
Page RDM (1996) TreeView: an application to display phylogenetic trees on personal computers. Comput Appl Biosci 12:357–358
Paredes OM, Gepts P (1995) Extensive introgression of Middle American germplasm into Chilean common bean cultivars. Genet Resour Crop Evol 42:29–41
Pastor-Corrales MA, Otoya MM, Molina A, Singh SP (1995) Resistance to Colletotrichum lindemuthianum isolates from Middle America and Andean South America in different common bean races. Plant Dis 79:63–67
Pavlicek A, Hrda S, Flegr J (1999) Free-Tree: freeware program for construction of phylogenetic trees on the basis of distance data and bootstrap/jackknife analysis of the tree robustness. Application in the RAPD analysis of genus Frenkelia. Folia Biol 45:97–99
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Rohlf FJ (2000) NTSYS-pc: numerical taxonomy and multivariate analysis system. Setanket, New York
Rosenberg NA (2004) DISTRUCT: a program for the graphical display of population structure. Mol Ecol Notes 4:137–138
Sharma PN, Diaz LM, Blair MW (2013) Genetic diversity of two Indian common bean germplasm collections based on morphological and microsatellite markers. Plant Genet Resour. doi:10.1007/s10722-013-9959-2
Singh SP, Gutierrez JA (1984) Geographical distribution of the Dl1 and Dl2 genes causing hybrid dwarfism in Phaseolus vulgaris L., their association with seed size, and their significance to breeding. Euphytica 33:337–345
Singh S, Teran H (1995) Evaluating sources of water-stress tolerance in common bean. Annu Rep Bean Improv Coop 38:42–43
Singh SP, Gepts P, Debouck DG (1991a) Races of common bean (Phaseolus vulgaris, Fabaceae). Econ Bot 45:379–396
Singh SP, Gutierrez JA, Molina A, Urrea C, Gepts P (1991b) Genetic diversity in cultivated common bean: II. Marker-based analysis of morphological and agronomic traits. Crop Sci 31:23–29
Singh SP, Nodari R, Gepts P (1991c) Genetic diversity in cultivated common bean: I. Allozymes. Crop Sci 31:19–23
STATCOM (2011) National Statistical Committee of the Kyrgyz Republic. http://stat.kg/images/stories/docs/KZ/Agri/agri1.pdf. Accessed 5 Feb 2013
Strausbaugh CA, Myers JR, Forster RL, McClean PE (1999) Bc-1 and Bc-u—two loci controlling bean common mosaic virus resistance in common bean are linked. J Am Soc Hortic Sci 124:644–648
Wortmann CS, Kirkby RA, Eledu CA, Allen DJ (1998) Atlas of common bean (Phaseolus vulgaris L.) production in Africa. CIAT, Cali, Colombia
Yeh FC, Boyle TJB (1997) Population genetic analysis of co-dominant and dominant markers and quantitative traits. Belg J Bot 129:157
Acknowledgments
This research was undertaken as part of the PhD study of the first author. We are thankful for the funding provided by the Swedish International Development Agency (Sida) and and Einar and Inga Nilsson fund. We thank Michigan State University (East Lansing) and United States Department of Agriculture (Pullman) for providing the common bean germplasm used in this study. Special thanks to Dr. Elie Marx (INRA, France) for guidance regarding phenotyping of accessions. We are grateful to Anna Zborowska (SLU) for her laboratory assistance.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Hegay, S., Geleta, M., Bryngelsson, T. et al. Genetic diversity analysis in Phaseolus vulgaris L. using morphological traits. Genet Resour Crop Evol 61, 555–566 (2014). https://doi.org/10.1007/s10722-013-0056-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-013-0056-3