Abstract
The common bean (Phaseolus vulgaris L., Fabaceae) is the most important legume of the genus Phaseolus, and the most cultivated in family farming. This study aimed to characterize landrace beans by SSR markers and morphoagronomic variables, comparing them to commercial cultivars. A total of 185 accessions (158 landrace and 27 commercial) were evaluated using SSR, and later 50 accessions (37 landrace and 13 commercial cultivars) were evaluated by 12 morphoagronomic variables. The estimators of genetic diversity, expected heterozygosity, allelic number, private, and rare alleles showed greater magnitude in landrace accessions, indicating the potential of these accessions as a reservoir of rare and private alleles compared to commercial stocks. Ten groups were observed in the cluster analysis, of which one group had 89% of accessions (138 landrace and all commercial ones). For morphoagronomic characterization, three field experiments were carried out in randomized blocks with three replications, one in Alegre, ES and two in Mimoso do Sul, ES. The accessions Verde-111, Branco-110, Pérola-102, Preto-13-69, Feijão Rainha-18, and Branco-05 showed high productivity levels (above 3000 kg ha−1) and six groups were formed. Most landrace accessions and commercial cultivars were allocated into the same group, suggesting that landrace beans have characteristics of interest compared to the commercial ones. The variability detected in landrace beans is of strategic importance, given the selection carried out by farmers for materials adapted to their local conditions. Thus, the development of policies to support the conservation of bean trees on rural properties is indicated as a measure to preserve diversity.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The common bean (Phaseolus vulgaris: Fabaceae) is a legume of great socio-economic importance in several countries in Latin America, Africa, and Asia, where it is consumed as a staple food by a large part of the population (Akibode and Maredia 2011; Beebe 2012). Brazil stands out as the third-largest producer, with ~ 2.9 million tons (FAO 2019). The common bean is cultivated in most Brazilian states, under the most varied edaphoclimatic conditions and in different times and cultivation systems. It constitutes an essential source of protein, dietary fiber, and complex carbohydrates, especially for low-income groups (Castro-Rosas et al., 2016; Santiago-Ramos et al. 2018).
The species is predominantly autogamous, with cross-fertilization rates lower than 5% (Graham and Ranalli 1997). It has a diploid genome with 11 chromosomes (2n = 2x = 22) and estimated size of ~ 587 Mbp (Schmutz et al. 2014). It originated in Mesoamerica (Bitocchi et al. 2012) and then went through two independent domestication events giving rise to the main gene pools of the species, the Mesoamerican and Andean gene pool (Gepts et al. 1986; Bitocchi et al. 2013). Varieties are well defined within each gene pool due to a divergence estimated to occur ~ 165,000 years ago (Schmutz et al. 2014). Genotypes from the Andean gene pool generally have large grains, while the Mesoamerican ones have small or medium-sized grains (Evans 1980).
For millennia, the improvement of plants was carried out empirically by farmers, which led to the formation of local seed stocks, i.e., landrace beans. Brazilian landrace beans present high diversity due to different environments, multiple production systems, and farmer preferences (Bertoldo et al. 2014). Brazil is considered a secondary center of diversification due to the success of bean cultivation in the country (Burle et al. 2010). In the national scenario, beans are characterized by family farming and are considered a commercial alternative with potential for expansion. Approximately 99.08% of farms present beans planted in 50 hectares, 0.77% planted between 50 and 200 hectares, and 0.15% planted in more than 200 hectares (Silva and Wander 2013).
Landrace beans can show tolerance to biotic and abiotic stresses and carry genes related to several quality attributes. These features represent a precious genetic value to maintain food safety and biodiversity, and their preservation and evaluation are essential in programs of crop genetic improvement (Sifres et al. 2010; Savić et al. 2021). Also, some accessions have specific attributes (size, color, and grain shape) desired by consumers who sometimes pay more for these products (Negri and Tosti 2002; Murekezi et al. 2017; Choudhary, et al. 2022).
Studies have identified greater diversity within the landrace accessions group than commercial cultivars, emphasizing the importance of such varieties as sources of genetic variation for the crop (Maciel et al. 2003; Perseguini et al. 2016; Carias et al. 2018; Jannat et al. 2022). The genetic diversity of landrace beans makes them a valuable resource, as they are potential parents in cross-breedings aimed at developing new cultivars. Several characteristics can be used to identify and estimate genetic diversity, including molecular markers and morphoagronomic variables. The combination of molecular and morphological data can be more objective and accurate to assess genetic variation and reported in different studies (Delfino et al. 2017).
DNA-based molecular markers provide a reasonable estimate of genetic diversity as they are almost unlimited in number and are not influenced by the environment (Naghavi et al. 2009; Ambadeet al. 2015). The discovery and use of molecular markers offer an excellent opportunity to understand and identify diverse genetic material in agricultural species (Kapoor and Choudhary 2017). Among the molecular markers, microsatellites have a high degree of polymorphism, stability, ease of handling, speed, and reproducibility (Ellegren 2004; Kumar et al. 2009; Vieira 2016). They are also successfully used in beans to determine genetic diversity and population structure, revealing wide variability and confirmed varieties within each gene pool (Blair et al. 2009; Kwak et al. 2009; Gioia et al. 2019).
Morphoagronomic markers are crucial during pre- and post-improvement studies of genetic variation in common beans (Cabral et al. 2011; Okii et al. 2017; Carias et al. 2018; Sinkovii et al. 2019). They are used in the indirect analysis of genetic diversity, reflecting the real productive potential of the plant and enabling its direct use in improvement.
In this perspective, the objective of this work was to characterize a work collection using microsatellite and morphoagronomic molecular markers of common bean landrace accessions cultivated in Espírito Santo and compare them to commercial genotypes from different improvement programs.
Material and methods
Plant material
One hundred and eighty-five accessions were evaluated in this work, of which: 27 commercial cultivars that included 23 cultivars from Embrapa Rice and Beans (Empresa Brasileira de Pesquisa Agropecuária—Arroz e Feijão); one from IAPAR (Instituto Agronômico do Paraná); and three from Incaper. The remaining 158 accessions were landrace, with 91 of them having information on the place of cultivation. The others were from previous collections in the state of Espírito Santo with no identification of the municipality of origin (Table S1, Fig. 1).
Municipalities in the state of Espírito Santo where 91 accessions of landrace beans were harvested. Alegre (42 accessions); Cachoeiro de Itapemirim (1), Domingos Martins (7), Guaçuí (16), Ibitirama (2), Irupi (5), Muniz Freire (4), Santa Angélica (3), Santa Tereza (4), Venda Nova (2), Vila Velha (2), Vitória (3)
The genetic diversity of the 185 accessions was initially analyzed using microsatellite markers. Afterward, 50 accessions were selected for morphoagronomic characterization, according to (1) representativeness of a subset of highly variable common beans according to the genetic dissimilarity obtained by the analysis of microsatellite markers; (2) organoleptic qualities in the regional market; (3) be widely cultivated by farmers in the main cultivation areas, and (4) quantity of seeds sufficient to be submitted to a field experiment (Table S2).
Genetic diversity by microsatellites
Seeds of each genotype were germinated in the sand to obtain leaf tissue for genomic DNA extraction. Extraction was performed from individual plant tissue of each accession by the CTAB (cetyl-trimethyl ammonium bromide) method (Doyle and Doyle 1990). DNA concentrations and integrity were estimated using a Nanodrop™ 2000 (Thermo Scientific) spectrophotometer. A set of 39 microsatellite loci was selected due to high polymorphism information content (PIC) and genome distribution (Benchimol et al. 2007; Gaitán-Solís et al. 2002; Hanai et al. 2007; Garcia et al. 2011), and tested in 10 bean accessions. Thirteen out of the 39 loci were selected for genotyping of the other accessions due to the consistency of amplification and polymorphism (Gaitán-Solís et al. 2002; Benchimol et al. 2007). Seven of these loci were derived from expressed regions of EST (Hanai et al. 2007; Garcia et al. 2011) (Table S3).
PCR reactions were done with a final volume of 15 µL containing 50 ng of genomic DNA, 1X buffer (I0 Phoneutria®—500 mM KCl, 100 mM Tris-HCl pH 8.4, 1% Triton X-100), 0.15 µM Primer (Invitrogem®), 1.5 mM MgCl2 (Phoneutria®), 0.15 U Taq DNA polymerase (Phoneutria®), and 0.25 mM DNTP (Invitrogem®). Amplifications occurred in a Veriti™ 96-Well Thermal Cycler (Thermo Scientific®) under the following conditions: 94 °C for 4 min; 30 cycles of 94 °C for 0, 45 min, annealing temperature (Table S3) for 1 min, 72 °C for 1 min, and 72 °C for 8 min. The amplification products were separated by electrophoresis on 10% polyacrylamide gels. As standard, the 100 bp molecular weight marker (Kasvi, K9-100 L) was used. After electrophoresis, the gels were stained with GelRed® Nucleic Acid Gel Stain—Biotium (1.33X) and visualized by a photo documentation system (Bio-Rad Gel Doc™ EZ Imager). The images were processed, and the bands were analyzed with ImageLab 6.0 software (Bio-Rad Laboratories Inc.) to estimate the size of the fragments of the PCR products.
Morphoagronomic characterization
An experiment was carried out in the Área Experimental do Centro de Ciências Agrárias e Engenharias of the Universidade Federal do Espírito Santo in the municipality of Alegre, ES (latitude 20°45′S, longitude 41°31′W, and 138 m of altitude) (sowing in March of 2017 and harvest in June 2017). Another two were performed in a private area in the municipality of Mimoso do Sul, ES (latitude 21°01′S, longitude 41°17′W, and 250 m of altitude) (sowing in April and September 2017, and harvest in July 2017 and January 2018, respectively). The months of work duration were characterized by a season with rainfall below the expected normal and poor spatial distribution. The spatiotemporal distribution of rainfall was irregular, resulting in a low-quality rainfall regime (INCAPER 2017a, 2017b, 2018).
The design used was a randomized block one with three replications and plots composed of four rows of 1.2 m in length, 0.5 m between them, and ten seeds per linear meter, with a useful area of 2.4 m2. The first and last lines and the first and last plants of each central line per plot were considered borders. The useful plot consisted of 10 random plants. The cultural practices were carried out per the recommendations for the crop (Carneiro et al. 2015).
The accessions were evaluated with 12 post-harvest morphoagronomic descriptors, selected according to the list of minimum descriptors for the bean crop in the Registration in the national registry of cultivars (RNC) recommended in Decree No. 2366 of November 5, 1997 (BRASIL 1997), also based on the literature (Collicchio et al. 1997). The focus is on technological quality, management and production, and information obtained through questionnaires answered by producers at the collection of accessions. The characteristics evaluated were: stem diameter (DIAM) in mm from an evaluation carried out at 2 cm above the ground with a digital caliper; plant height (ALT), determined in cm by the distance between the collar and the last fully expanded leaf, number of nodes at harvest (NN); number of racemes (NR), and number of productive racemes (NRP), evaluated by the average of 10 plants from the base to the first inflorescence; first pod height (APV), measured in cm based on the distance between the collar and the insertion of the first pod; number of pods per plant (NVP), number of locules per plant (NLP) and number of seeds per plant (NSP), average measured in each of the 10 plants at crop density; grain mass per plant (MG), in g plant−1 determined by the mean seed weight of 10 plants using a precision scale; weight in grams of 100 grains (P100), evaluated in g of 100 seeds to the second decimal place at 12–14% moisture; grain yield (RG) estimated in kg ha−1 using the total seed weight of 10 plants × 10 (number of plants evaluated)/useful area, transforming to kg ha−1.
Analysis of molecular data
The parameters to measure the genetic diversity were the mean number of alleles per loci (A/loci), expected heterozygosity (He), observed heterozygosity (Ho), private alleles (AP), rare alleles (AR), mean inbreeding coefficient (F), and polymorphism information content (PIC) using the GENES software (Cruz et al. 2013).
The simple correspondence dissimilarity index estimated the Dissimilarity matrix with 1,000 bootstrap interactions, implemented in the DARwin 5.0 program (Perrier and Jacquemoud-Collet 2006). The accessions were then grouped hierarchically using the Unweighted Pair-Group Method with Arithmetic Means (UPGMA). Also, a principal coordinate analysis (PCoA) was performed using GenAlEx 6.5 software (Peakall and Smouse 2006).
A Bayesian cluster analysis was performed using STRUCTURE 2.3.4. software (PRITCHARD et al. 2000), with no a priori information on groups. The set of the number of runs was analyzed with a burn-in of 500,000 iterations, followed by 1,000,000 MCMC (Monte Carlo Markov Chain) iterations, with ten independent simulations. The tested K values ranged from 1 to 10. The ΔK method by Evanno et al. (2005) was used to find the best K, implemented in STRUCTURE HARVESTER (Earl and Vonholdt 2012).
Analysis of morphoagronomic data
The data obtained were subjected to analysis of variance (ANOVA). Subsequently, the Pearson linear correlation (r) coefficient matrix between the analyzed characteristics was estimated, and Student's t-test verified the significance at a 5% probability level. A multicollinearity analysis was also performed, and the relative importance of quantitative variables (S.j) was analyzed by Singh’s method (1981). These analyses were performed using the GENES software (Cruz et al. 2013).
The morphoagronomic data were submitted to multivariate analyses, considering the average of the cultivars in the three environments. The Mahalanobis generalized distance (D2) was obtained from the residual variance and covariance matrices. Then, a UPGMA hierarchical cluster analysis was performed, and the groups were defined by the non-hierarchical k-means method (Macqueen 1967). These analyses were performed using the R software (R Core Team 2020).
A principal component analysis (PCA) was performed to verify the relationship between variables and samples and group them. The main components whose sum of their variances represented at least 80% of the accumulated variance were considered. More attention was paid to the component with the most significant influence of the sensory note. A biplot graph was constructed from the two main components to observe the grouping of samples and variables and their performance concerning each other, mainly to the sensory note.
Results
Genetic diversity by microsatellite markers
The markers used were informative in the studied samples. The mean number of alleles per genomic SSR locus was 4.5, ranging from 3 to 6 alleles, and for EST-SSRs, it was 5.71 ranging from 5 to 8 alleles. The highest number of alleles per loci (8) and PIC (0.77) were observed in the loci PvM30. The lowest in the genomic SSR-IAC54 (A/loci = 3 and PIC 0,11). Mean He was higher for SSR-EST (0.54), compared to genomic SSR (0.46); however, not for Ho (EST-SSR = 0.01; SSR genomic = 0.03). The low value of Ho indicates an excess of homozygotes, which is confirmed by f, positive and greater than 0.8 (Table 1).
According to the diversity indices evaluated, He and Ho in commercial cultivars were 0.42 and 0.02 and in landrace accessions 0.50 and 0.02. Two private alleles were found in commercial beans and 29 in landrace. Concerning rare alleles, i.e., with a frequency below 0.05%, eight were found in commercial beans and 30 in landrace (Table 2).
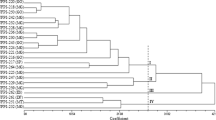
The clustering revealed the formation of 24 groups. Groups G11, G12, G13, G14, G15, G18, G23, G24 were formed with landrace and commercial accessions (Fig. 2A). The value of the cophenetic correlation coefficient was 0.86. The SSRs used did not detect dissimilarity between BRS Campeiro-28 and BRS Esplendor-30 accessions, black beans. Genetic distances ranged in magnitude from 0 to 1, suggesting extensive genetic variability within the set of accessions studied.
Evaluation of genetic diversity among 185 bean accessions using 13 microsatellite markers. A UPGMA dendrogram constructed from measurements of the simple coincidence distance of 185 bean accessions. Names in black are landrace accessions, and in red are commercial cultivars. B Cluster analysis based on 13 microsatellite loci evaluated in 185 common, landrace, and commercial bean accessions. Individual assignment to each group for K = 2 and K = 3. A line represents each individual, and the proportion of each color indicates each group's ancestry. C Analysis of principal coordinates obtained through 13 microsatellite loci evaluated in 185 common bean accessions, representing the 27 commercial beans (red), the 158 landrace accessions (green), and of these, the 50 selected for field evaluation (black border)
According to STRUCTURE, the adequate number of groups (K) identified was equal to two. The second-highest peak was equal to three, thus dividing the accessions into groups G, G2, and G3 (Figure S1). G1 comprised 56 accessions, all landrace, of which 37 are large seeds, and the remaining 16 are small and medium-sized seeds. G2 comprised 61 landrace accessions and 13 commercial cultivars. G3 comprised 40 landrace and 15 commercial ones (Fig. 2B).
The graphical representation by PCoA allowed splitting the diversity between landrace and commercial accessions, with the first two components accounting for 20.34% of the accumulated variation (Fig. 2C). Landrace accessions showed more significant variability, consistent with the greater number of alleles found in this group (Table 2).
AMOVA split the variation in 9.88% between landrace accessions and commercial cultivars and 90.11% within populations. Although the most significant genetic variation is found in the intrapopulation component, this was not significant, and the differentiation between populations was significant (p < 0.01) (Table 3). FST value according to Wright’s method was 0.074, which shows a moderate genetic variability between commercial cultivars and landrace accessions and high differentiation within groups according to the FIS of 0,967.
The genetic diversity observed in Figs. 2A, B, and C was evaluated in 185 accessions, and 50 of them were selected for morphoagronomic characterization (Table S1, S2).
Morphoagronomic characterization
ANOVA revealed significant differences among accessions for almost all variables (Table S4), showing variability between accessions. A perfect correlation (1.0) was observed between yield and grain mass, which was expected since the yield is calculated using the grain mass. Thus, the grain mass was excluded from further analyses due to the high correlation and a lower agronomic value than the grain yield. The correlation analysis showed that yield is positively and significantly correlated with number of nodes (p < 0.05), number of pods (p < 0.001), number of locules per plant (p < 0.001), diameter (p < 0.01), and weight of 100 grains (p < 0.01) (Fig. 3B). The number of seeds per plant showed multicollinearity of 439.82, considered moderate to strong, negatively influencing the multivariate analysis. Its removal reduced the value to 63.22, considered low using the Classification of Montgomery and Peck (1981) criteria, thus remaining 10 of the 12 variables.
Evaluation of genetic diversity among 50 bean accessions using ten morphoagronomic variables (ALT: plant height, APV: first pod height, DIAM: stem diameter, NN: number of nodes at harvest, NR: number of racemes, NRP: number of productive racemes, NV: number of pods per plant, NLV: number of locules per pod, P100: weight of 100 grains; RG: grain yield). A Dendrogram with Mahalanobis distance and UPGMA clustering method. The arrows indicate accessions with the weight of 100 seeds above 30 g. The vector loads of the first four main components are also presented. The vector loads are the weights given to each variable for each component through which they allow the interpretation of the components. B Pearson’s correlation matrix, the asterisks indicate the confidence levels of p < 0.1, p < 0.01, p < 0.001 for *, **, and ***, respectively. C Biplot of the first two axes of principal component analysis (PC1 and PC2)
The variables with the most significant relative contributions were height (29.20%), followed by the number of nodes (15.0%), number of locules per plant (10.7%), number of racemes (10.1%), number of pods per plant (8.5%), grain yield (8.1%), and weight of 100 seeds (7.8%). Conversely, the variables with less significant contributions in the separation of genotypes were the number of productive racemes (0.3%), stem diameter (4.6%), and first pod height (5.5%) (Table 4).
The mean value of the plant height ranged from 30.89 (Vermelho-132) to 102.29 cm (BRS Pérola-22). The height of the first pod ranged from 8.37 (Vermelho-19) to 22.98 cm (Vagem riscada-10). The diameter ranged from 4.15 (Carioca vermelho-168) to 15.87 (Bege-135). The largest number of nodes was detected in accession No. 10 Quadrado-156 (13.12), the number of racemes in Branco-5 (6.16), and the number of productive racemes in Branco-5 (5.09). The highest number of pods, number of locules, and seeds per plant were respectively 15.00, 80.99, and 87.18, all detected in Preto 13–69 accession. The highest weight of 100 grains and yield was recorded for Branco-5 accession (18.03 g and 4062.30 kg ha−1, respectively) (Table S5).
UPGMA enabled the distribution of the 50 accessions into six groups defined by the non-hierarchical k-means method (Macqueen 1967) (Fig. 3A). The smallest dissimilarity was between Verde Cristo rei-81 and Feijão verde-26 accessions (0,266), and the largest was between Branco-5 and No. 10 Quadrado-156 (30.274) (Fig. 3A).
Groups G1 (Branco-5) and G2 (No. 10 Quadrado-156) were formed by a landrace accession. G3 was formed by 16 landrace beans (Verde cristo rei 01-81, Feijão verde-26, Amarelo-133, Preto-104, Palhacinho-1, Amendoim-14, Verde-114, Verde 02-68, Verde-111, Verde-109, Vermelho-132, Beje-135, Manteiga beje claro-3, Vermelho-9, No. 10 Quadrado-156, Branco-5) and two commercial cultivars (CNFC 15,625-20, IPR Colibri-25). G4 was formed by two landraces (Carioca-105 and Amendoim-42). G5 was formed by two accessions, one landrace (Feijão rainha-18) and one commercial (BRS Pérola-22). G6 comprised 19 landrace (Vagem riscada-172, Preto-4, Bico de ouro-128, Enxofre-13, Vagem riscada-11, Feijão 04-149, Preto-13-69, Preto-118, Vermelho-174, Carioca vermelho-168, Bico de ouro-120, Vagem riscada-10, Vagem riscada-8, Vagem riscada-92, Ouro vermelho-117, Marrom-112, Vermelho-19, Bico de ouro-12, Branco-110) and nine commercial cultivars (CNFC 15,462-21, BRS Supreme-181, CNFP 15,304-32, BRS Notável-23, BRS Esplendor-30, IPR Uirapuru-89, Emcapa 404, Serrano-180, BRS Estilo-24, BRS Notável-150) (Fig. 3A).
In the biplot (Fig. 3C), variables are represented by vectors and samples by numbers. The larger the vector, the greater the variable's influence on the grouping; the smaller the angle between vectors, the more significant the correlation between variables. The first three components of the PCA accounted for about 81.49% of the variation: first—46.57%, second—22.23%, and third—12.69%. The first two components accounted for 68.8% of the variation. Even with the low variance values, the analysis identified six groups according to quality by the biplot of the two components (Fig. 3C), with an agreement between the groups formed by the UPGMA clustering (Fig. 3A).
Ten principal components were obtained, of which three were selected because they accumulated 81.49% of the total variance (Table 5). The first principal component (PC1) showed the most significant influence of the variable number of pods per plant, followed by the number of productive racemes and number of nodes.
Discussion
This study demonstrates common bean accessions' genetic diversity and structure in a representative work collection from the state of Espírito Santo using microsatellite markers and morphoagronomic variables. Bean production in the state is carried out through subsistence agriculture with commercialization in local markets and fairs, which favors the development of landrace accessions with vernacular names attributed by the producers themselves. They collect a bulk of seeds from several plants with no guarantee of homogeneity and plant them in the next harvest, repeating the procedure generation after generation.
The present study analyzed 185 common bean accessions cultivated in the state of Espírito Santo using 13 microsatellite loci. Of these, 50 accessions were selected for evaluation with 12 morphoagronomic variables. Of the 50 accessions, 41 were collected in 12 municipalities, and three regions of the state expanded the scope of studies on beans in the state compared to previous studies. Cabral et al. (2010, 2011) evaluated 57 accessions of landrace beans and commercial cultivars in the municipality of Muqui-ES, in which wide genetic diversity was detected using SSR and morphoagronomic variables. Studies with the characterization of landrace accessions compared to commercial ones using molecular markers and morphoagronomic variables are reliable for the evaluation of genetic diversity, aiming to discriminate promising accessions for bean crop improvement programs and genetic resource conservation (Sharma et al. 2013; Scaranoa et al. 2014; Rana et al. 2015).
The collections of landrace beans carried out in the state of Espírito Santo allowed to discriminate commercial types. The predominant seeds were of small and medium-size (characteristics of the Mesoamerican gene pool), mainly of the red (16 accessions), black (15), and carioca (10). While in the national scene, black beans are the most traditional and widely cultivated type in Brazil, carioca beans characterized by beans with black stripes on a light cream background became the distinct commercial type with 70% of the consumer market (Peloso and Melo, 2005). Beans with large grains were also found (characteristic of the Andean gene pool), among them Jalo (5 accessions), striped (3), white (5), large red (5), in addition to beans designated by the producers with the vernacular name of “Palhacinho” (4 accessions). These types are not very representative in the national market, but their organoleptic attributes are desired by consumers who sometimes pay more for these products (Negri and Tosti 2002; Galvan et al. 2006; Murekezi et al. 2017). Furthermore, they are alternatives to serve the export market (Pereira et al. 2010).
Genetic diversity by microsatellite markers
The study of the genetic diversity of 185 accessions (158 landrace and 27 commercial) using microsatellites revealed Ho and mean inbreeding coefficient with values equal to or close to one, showing generally homozygous loci. This is expected for bean plants, as the species is preferentially autogamous, showing self-fertilization equal to or greater than 95% (Graham and Ranalli 1997).
According to the classification presented by Botstein et al. (1980), PIC values below 0.25 are not very informative, from 0.25 to 0.5 moderately informative and above 0.5 very informative. In this study, PIC ranged from little (0.11-SSR-IAC54) to very informative (0.77-PvM30), with an overall mean of 0.46, indicating that the markers together presented medium quality. However, these values were higher than those detected by Cabral et al. (2011), who used 16 SSR in 57 bean accessions and found PIC ranging from 0.11 to 0.51. The values are similar to those obtained by Perseguini et al. (2015), who studied 58 SSR in 500 bean accessions and detected PIC ranging from 0.17 to 0.86. Therefore, the SRRs used in this work were efficient, indicating their potential to access genetic diversity in common beans.
Landrace accessions had a more significant number of AP and AR (29 and 30, respectively) than commercial accessions (2 and 8, respectively) (Table 2). Private alleles result from mutation, migration, adaptation, and selection to specific environmental conditions. An allele may appear occasionally and may have increased its frequency in the population through hybridization or genetic bottleneck, giving rise to a private allele. In beans, due to the predominance of autogamy, this event is not commonly expected. However, for the studied landrace accessions, they were found on a large scale. This fact is interesting, as the presence of private alleles in landrace accessions may be related to the adaptation to the different environments in which beans are cultivated in the state of Espírito Santo. Delfini et al. (2017) identified 24 private alleles in 16 of the loci analyzed in the carioca group and 22 in the black group. Cardoso et al. (2014) analyzed 114 bean genotypes, including 50 commercial cultivars from public institutions in Brazil and other countries, and 64 lines used in improvement programs. They found 35 private alleles at 15 analyzed microsatellite loci and observed that newer cultivars have more private alleles.
Molecular cluster analysis with 13 SSR markers was efficient to assess genetic diversity among common bean accessions. The commercial cultivars were distributed among the landrace beans, which allows us to conclude that the landrace beans cultivated in the state of Espírito Santo retain both diversity and similarity with commercial beans. It is an essential aspect from the point of view of improvement since landrace beans may already have characteristics of interest. The SSRs used did not detect dissimilarity between BRS Campeiro-28 e o BRS Esplendor-30 accessions, both black grains and developed by the same improvement program.
Morphoagronomic characterization
Multivariate analysis methods are considered ideal to describe genetic diversity. They treat variables together and unify multiple information on a set of characters of interest in genetic improvement (Cruz and Regazzi 2001) to detect a possible group structure by cluster analysis.
The MANOVA of quantitative data showed significance for the analyzed variables, indicating genetic variability among accessions. Positive and significant correlations between grain yield and number of nodes, number of pods, and number of locules per pod corroborate similar results found by Singh et al. (1995), Assady et al. (2005), and Salehi et al. (2008) in common bean. Such results show that these variables are helpful to select high yield in common bean improvement programs.
The knowledge of the correlation informs the improver associations that occur in a given trait due to selection in another correlated trait (Ramalho et al. 1993). Thus, a balance is sought between the variables so that the plant has an ideotype that maximizes the grain yield, considering the other variables, for example, a significant increase in the number of pods may not be supported by a plant that does not present an increase in the stem diameter. Positive and significant correlations were observed for grain yield with the weight of 100 grains, number of racemes, and number of productive racemes, which indicates that selecting one feature would increase another one's expression.
Among the morphoagronomic variables, plant height showed a greater discriminatory power by the Singh test (1981). The higher the value of this variable, the lower the losses in cultural practices and mechanized harvest. Avoiding the direct contact of the legume with the ground provides a better phytosanitary state for the seed. In the present study, plant height ranged from 30.89 (Vermelho-132) to 102.29 cm (BRS Pérola-22). The range of variation for this variable was larger, compared to that observed by Salgado et al. (2011), who observed values of 67 to 108 cm, demonstrating the diversity of the studied collection, which serves as a basis for future improvement works that seek different plant heights. According to Simone et al. (1992), ideal plants for mechanized harvest present height above 50 cm. In the present study, 68% of the total accessions fit this pattern. However, it is noteworthy that plants with very high heights, especially compared to smaller ones, provide a greater occurrence of lodging and falling caused by the action of wind, making mechanized harvest impossible and increasing grain losses. On the other hand, the short stature presented by 32% of the remaining accessions is not a limitation for bean cultivation, as these accessions can be used by small farmers who use family labor.
The stem diameter and first pod height had low discriminatory importance (4.6 and 5.5%, respectively) by the Singh test (1981). However, these variables are included in common bean improvement programs to represent advantages for common bean producers. The stem diameter is critical because it promotes better plant support and avoids lodging, aiming to reduce losses and favor mechanized harvest, particularly Preto 13-69 accession, with 6.6 cm (Table S5). Most of the existing cultivars in Brazil have the height of insertion of the first pod between 5 and 10 cm (Silva and Fonseca 1996; Salgado et al. 2011). According to Silva et al. (2009), plants with insertion of the first pod greater than 9.3 cm favor the direct harvest of beans with a combine harvester. In the present study, one of the accessions had a first pod height smaller than 9.3 cm (Vermelho-19, with 8,37 cm), and the others had a height ranging from 9.91 (Verde-109) to 22.98 (Vagem riscada-10). Thus, it is possible to obtain a plant pattern that allows mechanized harvest among the studied accessions (Table S5). Afférri et al. (2008) evaluated 17 lineages and 12 varieties, including BRS Pérola, finding values between 10 and 27 cm. Plants whose first pod does not directly contact the soil do not suffer from decay due to excess moisture and fungal attack, thus promoting a better phytosanitary state for the seeds (Salgado et al. 2012).
Production components such as grain yield, number of pods per plant, number of seeds per pod, and grain mass per plant are related to the productivity of the bean crop (Zilio et al. 2011). The mean productivity found in landrace accessions of the Andean gene pool, Mesoamerican, and commercial cultivars was 1868.65 kg ha−1, 1.639,73 kg ha−1, and 2.017,83 kg ha−1, respectively. The highest value was observed for the landrace Branco-5 accession (4.062,30 kg ha−1), with this superiority probably due to the greater grain mass (20.31), the greater number of seeds (40.43), and pods per plant (13.34) in the landrace accession (Table S5).
Thus, even with unfavorable climatic conditions in the experimental environments due to irregular rainfall causing low-quality rainfall regime (INCAPER 2017a, 2017b, 2018), the productivity had a high magnitude compared to the 1st Brazilian crop of 2017/2018, which averaged 1728.00 kg ha−1 (Companhia Nacional de Abastecimento 2018). The present study results support those of Ramos Junior et al. (2005), Afférri et al. (2008), who obtained yields ranging from 316 to 3587 kg ha−1.
Regarding the mass of 100 grains, superiority is observed in the CNFC 10,470 lineage with 39.13 g and the cultivars Pérola and BRSMG Majestoso with 38.77 g and 37.36 g, respectively (Table S5). These results agree with those obtained by Ramos Junior et al. (2005) and Hoffmann Júnior et al. (2007), who obtained values ranging from 16.8 to 37.9 g. The discriminated accessions provided indications of genotypes from different centers of origin. According to Coelho et al. (2007), genotypes with 100 seeds weighing less than 25 g belong to the Mesoamerican center and, when greater than 33 g, to the Andean center. The mass of 100 grains verified for landrace varieties was generally similar to the results obtained by other authors (Coelho et al. 2010; Lima et al. 2012). According to Singh et al. (1989), the size of cultivated bean seeds can vary from less than 15 to 90 g per 100 seeds and are grouped into small ones. The identity of the common bean gene pool is substantiated by seed characteristics, with size as an essential identification criterion. In this study, groups G1, G4, G5, and G6 presented the grains with a weight of 100 seeds ranging from 20.85 to 58.28 g, showing the possible presence of representatives of the Andean gene pool (Table S5). The accession that presented the largest grain weight was Branco-5, with 58.28 g, the only accession in group 1 (Table S5). Ramos Junior et al. (2005) and Hoffmann Júnior et al. (2007) obtained values ranging from 16.8 to 37.9 g. In groups G2 and G3, the weight of 100 seeds ranged from 16.37 (Vagem riscada-8) to 29.49 g (Feijão Rainha-18), suggesting there are representatives of the Mesoamerican gene pool (Table S5). Based on this classification, 38% of accessions belong to the Andean gene pool, including beans with beige, brown, red, yellow, green, white, and mixed colors seeds. A total of 62% are part of the Mesoamerican gene pool, mainly black, red, and carioca beans. Thus, while the Mesoamerican gene pools have more representatives, the Andean one has a greater diversity in seed morphology. This result was similar to those found by Blair et al. (2013) and Cardoso et al. (2014) analyzing collections of Brazilian beans.
According to Falconer (1987), variability is critical for forming populations with selection potential. The distribution of accessions on the dendrogram shows the formation of six groups. The combination with greater dissimilarity occurred between accessions Branco-5 (from the Andean group) and No. 10 Quadrado-156 (from the Mesoamerican gene group), with a distance of 30.274, revealing a tremendous genetic variability existing between these accessions. The most appropriate would be to recommend cross-breeding divergent parents that present superior performance for the desirable or potential variables, such as grain yield, making it possible to obtain superior accessions in segregating generations. The combination with the lowest magnitude of dissimilarity (distance = 0.266) was between the accessions Verde Cristo rei-81 and Feijão verde-26 (both from the Andean gene group), indicating proximity between these accessions. Despite the extensive genetic variability in Brazil, low variability was observed among current cultivars, mainly due to market pressure for specific types of grain and the expressive planting of improved cultivars. Such variability makes them more vulnerable to pests, diseases, and abiotic stresses, such as drought, resulting in lower grain yields (Abdurakhmonov and Abdukarimov 2008).
By grouping, G1 to G4 were found to differ from the others, formed only with landrace accessions. The other two groups (G5 and G6) comprised landrace and commercial together, demonstrating their similarity considering the variables under study. Landrace accessions can possess many valuable genes, essential resources for improving biotic and abiotic stress resistance/tolerance (Hanson et al. 2007; Sifres et al. 2010).
The principal component analysis showed that the ten morphoagronomic variables participated in the common bean grouping, with the number of pods per plant and weight of 100 seeds having the largest participation. These variables could be explored as useful morphoagronomic markers to discriminate bean genotypes (Gana et al. 2013).
Landrace genotypes Verde-111, Branco-110, Preto13-69, Feijão Rainha-18, Branco-05, and the commercial cultivar Pérola-102 showed high levels of productivity (above 3000 kg ha−1) and can be incorporated into crop improvement programs or suggested for farmers. Of these, the weight of 100 seeds of accessions Preto 13-69 and Pérola-102 are smaller than 30 g. Therefore, they are considered of Mesoamerican origin.
The most divergent accessions by morphoagronomic variables were Branco-5 and No. 10 Quadrado-156 (distance = 30.27). The variables with the most significant discriminatory power were plant height, number of nodes, and number of locules per plant. These variables contributed to the separation of accessions and can be used in genetic divergence studies to identify duplicates in work collections. Both SSR and morphoagronomic variables allowed a detailed description of the diversity present in the collection. Landrace and commercial accessions were allocated in the same group, evidencing that landrace accessions have characteristics of interest since the commercial ones have already been submitted to some process of genetic improvement.
The studied landrace beans are a promising genetic resource due to their performance equal to or superior to the yield components of the commercial cultivars analyzed. They also showed greater diversity at the molecular and morphoagronomic level, requiring the planning of conservation strategies and use in improvement programs.
References
Abdurakhmonov IY, Abdukarimov A (2008) Application of association mapping to understanding the genetic diversity of plant germplasm resources. Int J Plant Genomics 2008:574927. https://doi.org/10.1155/2008/574927
Afférri FS, Martins EP, Peluzio JM, Fidelis RR, Rodrigues VM (2008) Espaçamento e densidade de semeadura para a cultura do milho, em plantio tardio, no Estado do Tocantins. Pesquisa Agropecuária Tropical 128–133
Akibode CS, Maredia M (2011) Global and regional trends in production, trade and consumption of food legume crops. Report submitted to the Standing Panel on Impact Assessment (SPIA) of the CGIAR Science Council, FAO, Rome
Ambade RL, Verma SK, Nanda HC, Nair SK, Verulkar SB (2015) Genetic diversity based on molecular markers in Grasspea (Lathyrus sativus L.). Legume Res 38:43–46. https://doi.org/10.5958/0976-0571.2015.00007.7
Assady B, Dorri HR, Vaezi S (2005) Study of genetic diversity of bean (Phaseolus vulgaris L.) genotypes by multivariate analysis methods. In: The First Iranian Pulse Symposium, Research Center for Plant Sciences, Ferdowi University of Mashhad, Iran, p 650
Beebe SE (2012) Common bean breeding in the tropics. In: Janick J (ed) Plant breeding reviews, vol 36. Wiley, New York, pp 357–426
Benchimol LL, Campos T, De Carbonell SAM, Colombo CA, Chioratto AF, Formighieri EFL, Gouve RM, Souza APD (2007) Structure of genetic diversity among common bean (Phaseolus vulgaris L.) varieties of Mesoamerican and Andean origins using new developed microsatellite markers. Genet Resour Crop Evol 54:1747–1762. https://doi.org/10.1007/s10722-006-9184-3
Bertoldo JG, Nodari RO, Coimbra JLM, Guidolin AF, Toaldo D, Pinho de Morais PP, Elias HT (2014) Genetic progress of black bean (Phaseolus vulgaris L.) over seven years. Interciencia 39:24–31
Bitocchi E, Bellucci E, Giardini A, Rau D, Rodriguez M, Biagetti E, Santilocchi R, Spagnoletti Zeuli P, Gioia T, Logozzo G, Attene G, Nanni L, Papa R (2013) Molecular analysis of the parallel domestication of the common bean (Phaseolus vulgaris) in Mesoamerica and the Andes. New Phytol 197:300–313. https://doi.org/10.1111/j.1469-8137.2012.04377.x
Bitocchi E, Nanni L, Bellucci E, Rossi M, Giardini A, Zeuli P, Logozzo G, Stougaard J, McClean P, Attene G, Papa R (2012) Mesoamerican origin of the common bean (Phaseolus vulgaris L.) is revealed by sequence data. Proc Natl Acad Sci USA 109:788–796. https://doi.org/10.1073/pnas.1108973109
Blair MW, Brondani RVP, Díaz LM, Del Peloso MJ (2013) Diversity and population structure of common bean from Brazil. Crop Sci 53:1983–1993. https://doi.org/10.2135/cropsci2012.09.0516
Blair MW, Díaz LM, Buendía HF, Duque MC. (2009) Genetic diversity, seed size associations and population structure of a core collection of common beans (Phaseolus vulgaris L.). Theoretical and Applied Genetics. 119:955–972. https://doi.org/10.1007/s00122-009-1064-8
Botstein D, White R, Skolnick M, Davis RW (1980) Construction of a genetic linkage map in man using restriction fragment length polymorphisms. Am J Hum Genet 32:314–331
BRASIL. Presidência da República. Decreto no 2.366, de 05 de novembro de (1997) Regulamenta a Lei no 9.456, de 25 de abril de 1997, que institui a Proteção de Cultivares, dispões sobre o Serviço Nacional de Proteção de Cultivares - SNPC, e dá outras providências. Diário Oficial [da] República Federativa do Brasil, Brasília, no 216, 7 de nov. 1997. Seção 1, pp 25333–25354.
Burle ML, Fonseca JR, Kami JA, Gepts P (2010) Microsatellite diversity and genetic structure among common bean (Phaseolus vulgaris L.) landraces in Brazil, a secondary center of diversity. Theor Appl Genet 121:801–813. https://doi.org/10.1007/s00122-010-1350-5
Cabral PDS, Soares TCB, Lima ABP, de Miranda FD, Souza FB, Gonçalves LSA (2011) Genetic diversity in local and commercial dry bean (Phaseolus vulgaris) accessions based on microsatellite markers. Genet Mol Res 10(1):140–149. https://doi.org/10.4238/vol10-1gmr993
Cabral PDS, Taís CBS, Leandro SAG, Antonio TAJ, Andreia BPL, Rosana R, de Frederico PM (2010) Quantification of the diversity among common bean accessions using Ward-MLM strategy. Pesq Agropecu Brasília 45(10):1124–1132. https://doi.org/10.1590/S0100-204X2010001000011
Cardoso PCB, Brondani C, Menezes IPP, Valdisser PAMR, Borba TCO, Del Peloso MJ, Vianello RP (2014) Discrimination of common bean cultivars using multiplexed microsatellite markers. Genet Mol Res 13(1):1964–1978. https://doi.org/10.4238/2014.March.24.1
Carias CMDOM, Guilhen JHS, de Souza MT, Ferreira A, Ferreira MFDS (2018) Genetic divergence toward the selection of promising bean progenitors via mixed multivariate models. Cien Inv Agr 45:251–262. https://doi.org/10.7764/rcia.v45i3.1902
Carneiro JE de S, de Paula Júnior TJ, Borém A (2015) Feijão: do plantio à colheita. UFV, Viçosa, p 384
Castro-Rosas J, Aguirre-Santos EA, Gómez-Aldapa CA, Valle-Cervantes S, Ochoa-Martínez LA, Hernández-Santos B, Rodríguez-Miranda J (2016) Effect of harvest year on the physical properties, chemical composition and cooking time of three common bean varieties that are grown in Mexico. Qual Assur Saf Crop 8:339–348. https://doi.org/10.3920/QAS2015.0693
Choudhary N, Gupta M, Shafi S, Jan S, Mir AH, Singh B, Varshney R (2022) Molecular diversity and nutriment studies of common bean (Phaseolus vulgaris L.) from the two hot-spots of Western Himalayas of Jammu and Kashmir. Crop and Pasture Science
Coelho GMM, Coimbra JLM, Souza CA, Bogo A, Guidolim AF (2007) Diversidade genética em acessos de feijão (Phaseolus vulgaris L.). Cienc Rural 37:1241–1247. https://doi.org/10.1590/S0103-84782007000500004
Coelho CMM, Zilio M, Souza CA, Guidolin AF, Miquelluti DJ (2010) Características morfo-agronômicas de cultivares crioulas de feijão comum em dois anos de cultivo. Semina: Ciências Agrárias 31(1):1177–1186
Collicchio E, Ramalho MAP, Abreu ÂFB (1997) Associação entre o porte da planta do feijoeiro e o tamanho dos grãos. Pesq Agropec Bras 32:297–304
Companhia Nacional de Abastecimento (2018) Perspectivas para a agropecuária. Conab, Brasília
Cruz CD (2013) GENES—a software package for analysis in experimental statistics and quantitative genetics. Acta Sci 35:271–276. https://doi.org/10.4025/actasciagron.v35i3.21251
Cruz CD (2001) Programa Genes: aplicativo computacional em genética e estatística. UFV, Viçosa, 648p
Cruz CD, Regazzi AJ (2001) Modelos biométricos aplicados ao melhoramento genético. UFV. 390p
Del Peloso MJ, Melo LC (2005) Potencial de rendimento da cultura do feijoeiro comum. Santo Antônio de Goiás: Embrapa Arroz e Feijão, p 131
Delfini J, Moda-Cirino V, de Fátima RC, dos Santos NJ, Ruas PM, Buratto JS, Ruas EA, Gonçalves LSA (2017) Distinctness of Brazilian common bean cultivars with carioca and black grain by means of morphoagronomic and molecular descriptors. PLoS ONE 12:e0188798. https://doi.org/10.1371/journal.pone.0188798
Doyle JJ, Doyle JL (1990) Isolation of plant DNA from fresh tissue. Focus 12:13–15
Earl DA, VonHoldt BM (2012) Structure harvester: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Resour 4(2):359–361. https://doi.org/10.1007/s12686-011-9548-7
Ellegren H (2004) Microsatellites: simple sequences with complex evolution. Nat Rev Genet 5:435–445. https://doi.org/10.1038/nrg1348
Evans AM (1980) Structure, variation, evolution, and classification in Phaseolus. In: Summerfield RJ, Bunting AH (eds) Advances in legume science. Royal Botanic Gardens, Kew, pp 337–347
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14(8):2611–2620. https://doi.org/10.1111/j.1365-294X.2005.02553.x
Falconer DS (1987) Introdução à genética quantitativa. UFV, Viçosa
FAO (2019) New standards to curb the global spread of plant pests and diseases. http://www.fao.org/news/story/en/item/1187738/icode/. Accessed May 2020
Gaitán-Solís E, Duque MC, Edwards KJ, Tohme J (2002) Microsatellite repeats in common bean (Phaseolus vulgaris): isolation, characterization, and cross-species amplification in Phaseolus ssp. Crop Sci 42:2128–2136. https://doi.org/10.2135/cropsci2002.2128
Galvan MZ, Menendez-Sevillano MC, De Ron AM, Santalla M, Balatti PA (2006) Genetic diversity among wild common beans from north-western Argentina based on morpho-agronomic and RAPD data. Genet Resour Crop Evol 53:891–900. https://doi.org/10.1007/s10722-004-0981-2
Gana AS, Shaba SZ, Tsado EK (2013) Principal component analysis of morphological traits in thirty-nine accessions of rice (Oryza sativa L.) grown in a rainfed lowland ecology of Nigeria. J Plant Breed Crop Sci 5:120–126. https://doi.org/10.5897/JPBCS12.065
Garcia RAV, Rangel PN, Brondani C, Martins WS, Melo LC, Carneiro MS, Borba TCO, Brondani RPV (2011) The characterization of a new set of EST-derived simple sequence repeat (SSR) markers as a resource for the genetic analysis of Phaseolus vulgaris. BMC Genet 12:41. https://doi.org/10.1186/1471-2156-12-41
Gepts P, Osborn TC, Rashka K, Bliss FA (1986) Phaseolin-protein variability in wild forms and landraces of the common bean (Phaseolus vulgaris): evidence for multiple centers of domestication. Econ Bot 40:451–468. https://doi.org/10.1007/BF02859659
Gioia T, Logozzo G, Marzario S, Spagnoletti Zeuli P, Gepts P (2019) Evolution of SSR diversity from wild types to U.S. advanced cultivars in the Andean and Mesoamerican domestications of common bean (Phaseolus vulgaris). PLoS One 14:e0211342. https://doi.org/10.1371/journal.pone.0211342
Graham PH, Ranalli P (1997) Common bean (Phaseolus vulgaris L.). Field Crops Res 53:131–146. https://doi.org/10.1016/S0378-4290(97)00112-3
Hanai LR, de Campos T, Camargo LEA, Benchimol LL, de Souza AP, Melotto M, Carbonell SAM, Chioratto AF, Consoli L, Formighieri EF, Siqueira M, Tsai SM, Vieira MLC (2007) Development, characterization, and comparative analysis of polymorphism at common bean SSR loci isolated from genic and genomic sources. Genome 50:266–277. https://doi.org/10.1139/G07-007
Hanson PM, Sitathani K, Sadashiva AT, Yang RY, Graham E, Ledesma D (2007) Performance of Solanum habrochaites LA 1777 introgression line hybrids for marketable tomato fruit yield in Asia. Euphytica 158:167–178. https://doi.org/10.1007/s10681-007-9440-4
Hoffmann Júnior L, Ribeiro ND, Rosa SSD, Jost E, Poersch NL, Medeiros SLP (2007) Resposta de cultivares de feijão à alta temperatura do ar no período reprodutivo. Cienc Rural 37:1543–1548. https://doi.org/10.1590/S0103-84782007000600006
INCAPER (2018) Instituto Capixaba de Pesquisa e Assistência Têcnica Rural. Boletim Climatológico Trimestral do Espírito Santo Jan–Mar 2018. Vitória
INCAPER a (2017) Instituto Capixaba de Pesquisa e Assistência Têcnica Rural. Boletim Climatológico Trimestral do Espírito Santo Out–Dez 2017. Vitória
INCAPER b (2017) Instituto Capixaba de Pesquisa e Assistência Têcnica Rural. Boletim Climatológico Trimestral do Espírito Santo Abr – Jun 2017. Vitória
Jannat S, Shah AH, UlHassan M, Sher A, Fiaz S, Elesawy BH, Ismailf KA, Askaryf A, Qayyum A (2022) Genetic diversity of common bean (Phaseolus vulgaris L) ecotypes from Pakistan using simple sequence repeats. Saudi J Biol Sci. https://doi.org/10.1016/j.sjbs.2022.103300
Kapoor R, Choudhary K (2017) Genetic diversity analysis of fodder oats (Avena sativa L.) germplasm by microsatellite markers. J Agr Sci Tech 19: 1369–1379. http://hdl.handle.net/123456789/3577
Kumar P, Gupta VK, Misra AK, Modi DR, Pandey BK (2009) Potential of molecular markers in plant biotechnology. Plant Omics J 2(4):141–162
Kwak M, Gepts P (2009) Structure of genetic diversity in the two major gene pools of common bean (Phaseolus vulgaris L., Fabaceae). Theor Appl Genet 118:979–992. https://doi.org/10.1007/s00122-008-0955-4
Lima MS, Carneiro JES, Carneiro PCS, Pereira CS, Vieira RF, Cecon PR (2012) Characterization of genetic variability among common bean genotypes by morphological descriptors. Crop Breed Appl Biotechnol 12:76–84. https://doi.org/10.1590/S1984-70332012000100010
Maciel F, Echeverrigaray S, Gerald L, Grazziotin F (2003) Genetic relationship and diversity among Brazilian cultivar and of common beans (Phaseolus vulgaris L.) revealed by AFLP markers. Genet Resour Crop Evol 50:887–893. https://doi.org/10.1023/A:1025994513043
MacQueen J (1967) Some methods for classification and analysis of multivariate observations. In: Proceedings of the 5th Berkeley symposium on mathematical statistics and probability, University of California Press, City, pp 281–297
Montgomery DC, Peck EA (1981) Introduction to linear regression analysis. John Wiley, New York. 504p
Murekezi A, Oparinde A, Birol E (2017) Consumer market segments for biofortified iron beans in Rwanda: evidence from a hedonic testing study. Food Policy 66:35–49. https://doi.org/10.1016/j.foodpol.2016.11.005
Naghavi MR, Malaki M, Alizadeh H, Pirseiedi M, Mardi M (2009) An assessment of genetic diversity in wild diploid wheat Triticum boeoticum from west of Iran using RAPD, AFLP and SSR markers. J Agric Sci Tech 11:585–598
Negri V, Tosti N (2002) Phaseolus genetic diversity maintained on farm in Central Italy. Gen Res Crop Evol 49:511–520. https://doi.org/10.1023/A:1020902508166
Okii D, Tukamuhabwa P, Tusiime G, Talwana H, Odong T, Mukankusi C, Male A, Amongi W, Sebuliba S, Paparu P, Nkalubo S, Ugen M, Buah S, Gepts P (2017) Agronomic qualities of genetic pyramids of common bean developed for multiple-disease-resistance. Afr Crop Sci J 25:457–472. https://doi.org/10.4314/acsj.v25i4.5
Peakall R, Smouse PE (2006) GENALEX 6: genetic analysis in Excel. Population genetic software for teaching and research. Mol Ecol Notes 6:288–295. https://doi.org/10.1111/j.1471-8286.2005.01155.x
Pereira HS, Melo LC, Del Peloso MJ, Diaz JLC, Magaldi MCS, Faria LC, Abreu AFB, Pereira Filho IA, Moreira JAA, Martins M, Wendland A, Costa JGC (2010) Evaluation of export common bean genotypes in Brazil. Annu Rep Bean Improv Coop 53:276–277
Perrier X, Jacquemoud-Collet J (2006) DARwin software: dissimilarity analysis and representation for windows. http://darwin.cirad.fr/darwin. Accessed 1 Feb 2018
Perseguini JMKC, Oblessuc PR, Rosa JRBF, Gomes KA, Chiorato AF, Carbonell SA, Garcia AA, Vianello RP, Benchimol-Reis LL (2016) Genome-wide association studies of anthracnose and angular leaf spot resistance in common bean (Phaseolus vulgaris L). PLoS ONE 11(3):e0150506. https://doi.org/10.1371/journal.pone.0150506
Perseguini JMKC, Silva GMB, Rosa JRBF, Gazaffi R, Marçal JF, Carbonell SAM, Chiorato AF, Zucchi MI, Garcia AAF, Benchimol-Reis LL (2015) Developing a common bean core collection suitable for association mapping studies. Genet Mol Biol 38:67–78. https://doi.org/10.1590/S1415-475738120140126
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959. https://doi.org/10.1111/j.1471-8286.2007.01758.x
R Core Team (2020) R: a language and environment for statistical computing. Version 3.6.3. https://www.R-project.org
Ramalho MAP, Santos JB, Zimmermann MJO (1993) Genética quantitativa em plantas autógamas; aplicações ao melhoramento do feijoeiro. UFG, Goiânia
Ramos Junior EU, Lemos LB, Silva TRB (2005) Componentes da produção, produtividade de grãos e características tecnológicas de cultivares de feijão. Bragantia 64:75–82. https://doi.org/10.1590/S0006-87052005000100008
Rana JC, Sharma TR, Tyagi RK, Chahota RK, Gautam NK, Singh M, Sharma PN, Ojha SN (2015) Characterisation of 4274 accessions of common bean (Phaseolus vulgaris L.) germplasm conserved in the Indian gene bank for phenological, morphological and agricultural traits. Euphytica 205:441–457. https://doi.org/10.1007/s10681-015-1406-3
Salehi M, Tajick M, Ebadi AG (2008) The study of relationship between different traits in common bean with multivariate statistical methods. Am-Eurasian J Agric Environ Sci 3:806–809
Salgado FHM, Fidelis RR, Carvalho GL, dos Santos RS, Canceillier EL, Silva GF (2011) Comportamento de genótipos de feijão, no período da entressafra, no sul do estado de Tocantins. Biosci J 27:52–58
Salgado FHM, SilvaJ OliveiraTC, Tonello LP, Passos NG, Fidelis RR (2012) Efeito do nitrogênio em feijão cultivado em terras altas no sul do estado de Tocantins. Ambiência 8:125–136
Santiago-Ramos D, Juan de Dios Figueroa-Cárdenas JD, José Juan Véles-Medina JJ, Salazar (2018) Physicochemical properties of nixtamalized black bean (Phaseolus vulgaris L.) flours. Food Chem 240:456–462. https://doi.org/10.1016/j.foodchem.2017.07.156
Scarano D, Rubio F, José Ruiz J, Raoa R, Corradoa G (2014) Morphological and genetic diversity among and within common bean (Phaseolus vulgaris L.) landraces from the Campania region (Southern Italy). Sci Hortic 180:72–78. https://doi.org/10.1016/j.scienta.2014.10.013
Schmutz J, McClean P, Mamidi S, Wu GA, Cannon SB, Grimwood J, Jenkins J, Shu S, Song Q, Chavarro C et al (2014) A reference genome for common bean and genome-wide analysis of dual domestications. Nat Genet 46:707–713. https://doi.org/10.1038/ng.3008
Sharma PN, Diaz LM, Blair MW (2013) Genetic diversity of two Indian common bean germplasm collections based on morphological and microsatellite markers. Plant Genet Resour 11:121–130. https://doi.org/10.1017/S1479262112000469
Savić A, Pipan B, Vasić M, Meglič V (2021) Genetic diversity of common bean (Phaseolus vulgaris L) germplasm from Serbia, as revealed by single sequence repeats (SSR). Sci Hortic 288:110405
Sifres A, Blanca J, Nuez F (2010) Pattern of genetic variability of Solanum habrochaites in its natural area of distribution. Genet Resour Crop Evol. https://doi.org/10.1007/s10722-010-9578-0
Silva JG, Aidar H, Kluthcouski J (2009) Colheita direta de feijão com colhedora automotriz axial. Pesq Agropec Trop 39:371–379
Silva JG, Fonseca JR da (1996) Colheita. In: Araujo RS, Rava CA, Stone LF, Zimmermann MJ de O. Cultura do feijoeiro comum no Brasil. POTAFOS, Piracicaba pp 523–541.
Silva OF, Wander AE (2013) O feijão-comum no Brasil: Passado, Presente e Futuro. Embrapa-CNPAF, Santo Antônio de Goiás
Simone M de, Failde V, Garcia S, Panadero PC (1992) Adaptación de variedades y líneas de judías secas (Phaseolus vulgaris L.) a la recolección mecanica directa. INTA, Salta
Singh D (1981) The relative importance of characters affecting genetic divergence. Indian J Gene Plant Breed 41:237–245
Singh B, Makkar HPS, Negi SS (1989) Rate and extent of digestion and potentially digestible dry matter and cell wall of various tree leaves. J Dairy Sci 72(12):3233–3239. https://doi.org/10.3168/jds.S0022-0302(89)79482-0
Singh SP, Molina A, Gepts P (1995) Potential of wild common bean for seed yield improvement of cultivars in the tropics. Can J Plant Sci 75:807–812. https://doi.org/10.4141/cjps95-136
Sinkovič L, Pipan B, Sinkovič E, Meglič V (2019) Morphological seed characterization of common (Phaseolus vulgaris L.) and runner (Phaseolus coccineus L.) bean germplasm: a Slovenian gene bank example. BioMed Res 2019:1–13. https://doi.org/10.1155/2019/6376948
Vieira ML, Santini L, Diniz AL et al (2016) Microsatellite markers: what they mean and why they are so useful. Genet Mol Biol 39:312–328. https://doi.org/10.1590/1678-4685-GMB-2016-0027
Zilio M, Coelho CMM, Souza CA, Santos JCP, Miquelluti DJ (2011) Contribuição dos componentes de rendimento na produtividade de genótipos crioulos de feijão (Phaseolus vulgaris L.). Rev Ciênc Agron 42:429–438. https://doi.org/10.1590/S1806-66902011000200024
Acknowledgements
The authors thank the Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq, Brasília, DF, Brazil), Fundação de Amparo à Pesquisa e Inovação do Espírito Santo (FAPES, Vitória, ES, Brazil), and Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES, Brasília, DF, Brazil) for financial support.
Funding
The research leading to these results received funding from Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq, Brasília, DF, Brasil), Fundação de Amparo à Pesquisa do Espírito Santo (FAPES, Vitória, ES, Brasil) e Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES), Brasília, DF, Brasil) provided financial support of this study.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Carvalho, M.S., de Oliveira Moulin Carias, C.M., Silva, M.A. et al. Microsatellites and agronomic approaches reveal the diversity of beans (Phaseolus vulgaris L.) cultivated in Espírito Santo—Brazil, by family farms. Genet Resour Crop Evol 70, 731–747 (2023). https://doi.org/10.1007/s10722-022-01457-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-022-01457-9