Abstract
Anthocyanins and proanthocyanidins are the primary pigments of red rice and are also important functional nutrients for human health. To identify novel quantitative trait loci (QTLs) underlying anthocyanins and proanthocyanidins (ANC and PAC) in rice, a recombinant inbred line (RIL) derived from a cross of red rice ‘Hong Xiang 1’ (‘HX1’) and white rice ‘Song 98-131’ (‘S98-131’) was cultivated in six environments. A genetic map containing 126 markers covering 1833.4 cM with an average of 14.55 cM between markers was constructed. A total of 21 additive QTLs (A-QTLs) for ANC and PAC were identified from six environments using the IciMapping v3.3 software. Two new QTLs, qANC3 and qPAC12-4, were detected in several environments, and explained significant phenotypic variance. Nine QTLs of ANC and PAC were detected with additive × environmental interaction effects (AE effects) by QTLNetwork 2.1 software, but no epistatic and epistatic × environmental interaction effects (AA and AAE effects) were detected. The information obtained in this study could be useful for fine mapping and molecular marker-assisted selection of ANC and PAC in rice.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Rice is the most important food for more than 50% of the world’s population, and approximately 90% of the world’s rice is produced and consumed in Asia (Liu et al. 2016; Ghosh et al. 2014). Rice, as one of the most important cereals, can provide human beings with energy, vitamins, minerals and amino acids (Cheng et al. 2016). Rice has been cultivated for nearly 10,000 years (Molina et al. 2011). Red rice, rice varieties having a red pericarp, is a kind of colored rice and has gained attention for its nutritional and medicinal value (Furukawa et al. 2006). Anthocyanins (ANC) and proanthocyanidins (PAC) are the primary pigments in colored rice. These pigments accumulate in the seed coat, pericarp, and aleurone layers of the rice grain that gives the rice different colors (Han et al. 2009). Anthocyanins are also important functional nutrients having huge health benefits for humans (Acquaviva et al. 2003). ANC have been suggested to have a positive effect on obesity, diabetes and cardiovascular disease (Williamson and Clifford 2010; Miller and Shukitt 2012; Norberto et al. 2013; Tsuda 2012). The hydrolysate of ANC can inhibit the growth of tumor cells significantly, and its antioxidant and anti-inflammatory activity can also reduce the physiological activity of serum cholesterol and cancer (Koide et al. 1996; Acquaviva et al. 2003; Lazze et al. 2003; Russo et al. 2005). Proanthocyanidins have important applications not only in agriculture but also in health care products, cosmetics and as functional polymers (Shi and Du 2006). PAC have inhibitory effects on cancer-causing peroxide and protect against DNA damage and apoptosis (Bagchi et al. 1999; Agarwal et al. 2000). Consumption of proanthocyanidin-rich foods, such as red wine and cocoa (chocolate), seems to decrease blood pressure and insulin resistance and reverse endothelial dysfunction (Grassi et al. 2005; Heiss et al. 2007, 2005; Taubert et al. 2007; Hayashi et al. 2010).
The breeding of rice varieties with high ANC or PAC is an effective method to increase the daily intake of these antioxidants for people. Using identified genes/QTLs and molecular markers associated with ANC and PAC for marker-assisted selection (MAS) is a promising method for breeding high ANC or PAC containing rice varieties. The ANC and PAC content were considered to be the quantitative traits. Quantitative trait loci (QTLs) of ANC and PAC have been identified in pepper, eggplant, grape, mimulus and carrot (Chaim et al. 2003; Barchi et al. 2012; Huang et al. 2012; Yuan et al. 2013; Cavagnaro et al. 2014), but no QTLs were detected in rice. Though certain genes necessary for ANC and PAC have been reported in rice (Furukawa et al. 2006; Kim et al. 2007), more gene loci should be identified to explain the complex ANC and PAC traits.
In the present study, a recombinant inbred line (RIL) population derived from a cross between rice cultivars, HX1 and S98-131, was used to determine ANC and PAC content in six environments. The objectives of this study were (1) to identify the stable QTLs controlling ANC and PAC in different environments and (2) to measure the additive and additive × environment interaction effects of QTLs controlling ANC and PAC. The results contribute to the fine mapping of ANC and PAC QTLs in rice and provide a foundation for breeding high ANC and PAC content rice varieties.
Materials and methods
Plant materials and growth conditions
The mapping populations of 182 F7, F8, and F9 recombinant inbred lines (RILs) that were advanced by single-seed-descent from the cross between two japonica varieties namely, Hongxiang No. 1 (HX1, developed by Jilin Academy of Agricultural Sciences, Changchun, China) and Song 98-131 (S98-131, developed by Heilongjiang Academy of Agricultural Sciences, Harbin, China). The unpolished rice of HX1 was red, and the ANC and PAC content was high. The unpolished rice of S98–131 was white, and the ANC and PAC content was low. Parents and RIL populations were planted in Harbin (N45°28′, E128°08′) in 2013, 2014 and 2015 (E1, E3, E6), Acheng (N44°29′, E126°42′) in 2013 and 2014 (E2, E4) and Suihua (N48°02′, E124°13′) in 2014 (E5). All of the test materials were grown in a randomized block design with three replications of single row plots with a 5 m row length. The space between each row was 30 cm, and each single plant per row was 10 cm apart. Field management followed standard agricultural practices. Only three middle plants per row were used for measuring the ANC and PAC content. After harvest, the seeds were dried naturally for 3 months to stabilize the physical and chemical characteristics.
Measurement of ANC and PAC
The ANC content was measured according to the methods used by (Pang et al. 2009). The unpolished rice samples from each parent and RIL was ground to a fine flour (100 mesh) using a grinding miller. Next, 0.1 g of ground sample was added to 2 mL of methanol-concentrated HCl (99/1 v/v) solution and incubated for 24 h at 4 °C. Next, the samples were centrifuged at 25,000 RPM for 10 min. An equal volume of water and chloroform was added to remove the chlorophyll, and the absorption of the aqueous phase was detected at 530 nm using an AAS-3500 atomic absorption spectrophotometer (Shanghai HP analysis instrument, China). The molar absorbance of cyanidin-3-glucoside was used to calculate the total ANC content, expressed as cyanidin glucoside equivalents in μg per g of dry matter.
The PAC content was measured according to the method described by (Przybylski et al. 1998). The unpolished rice flour of 0.1 g (100 mesh) was dissolved in a 70% acetone solution containing 0.5% acetic acid and was slowly shaken on a shaking Table (20 r/min) at 25 °C for 15 h. After centrifuging at 3500 RPM for 15 min, the supernatant was mixed with n-hexane and was centrifuged again at 3500 RPM for 10 min. The supernatant was removed, and the remaining sample was dried at 40 °C for 4 h. A solution of methanol containing 1.0% (w/v) of vanillin and 10.0 M H2SO4 was added to the sample solution dissolved in methanol. The mixture was allowed to stand for 20 min at 40 °C, and the absorbance was measured at 500 nm. To determine the content of PAC, catechin was used as a standard, and PAC content was expressed as catechin equivalents.
Construction of a genetic linkage map and QTL analysis
Total DNA of the parents and of each RIL were isolated from fresh leaf tissues by the CTAB method (Doyle 1990). A total of 1000 SSR markers, covering the whole genome of rice, were used to detect polymorphisms. The PCR reaction conditions were set at 94 °C for 5 min, followed by 35 cycles of 30 s at 94 °C, 30 s at 55 °C, and 30 s at 72 °C followed by 5 min at 72 °C after the last cycle. The PCR products were separated by 6% (w/v) polyacrylamide gel electrophoresis and visualized by silver staining (Trigiano and Caetano 1998). The 126 polymorphic markers were integrated into the genetic linkage map covering a total length of 1833.4 cM with an average distance of 14.55 cM between adjacent markers.
The identification of QTLs was performed using QTL IciMapping v3.3 (Li et al. 2007) software. For single environment analysis, additive effect QTLs (A-QTLs) were identified in the biparental populations (BIP) module using the Inclusive Composite Interval Mapping (ICIM) method. An LOD value corresponding to an experimental threshold of LOD > 2.5 was used to declare a QTL as significant. The estimate of the QTL position was the point of maximum LOD score in the region under consideration. QTLNetwork 2.1 (Yang et al. 2007), which is based on mixed-model-based composite interval mapping (MCIM), was used to conduct two-locus QTL analysis. This method allows identification of QTLs involved in additive effect (A-QTL), epistatic interactions (AA-QTL), additive × environmental interactions (AE-QTL), and epistatic × environmental interactions (AAE-QTL). A significance level of P < 0.05 was used to select associated markers and to declare a putative A-QTL or AA-QTL. This QTL nomenclature follows that of (McCouch et al. 1997). The ANC and PAC data and the correlation between ANC and PAC levels were analyzed by SPSS 20.0 software (SPSS Inc., Chicago, IL, USA). Values of P < 0.05 were considered to be statistically significant, and values of P < 0.01 were considered to be highly significant.
Results
Phenotypic variation and correlation among traits
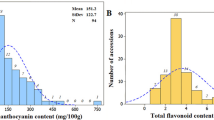
Table 1 shows the statistical analysis for ANC and PAC content of the parental lines and RIL populations from the six environments. ANC and PAC levels were significantly higher in ‘HX1’ compared to ‘S98-131’ for all six environments. For ANC and PAC content, the frequency of six environments in the RIL were normally distributed, and the absolute values of skewness and kurtosis for most environments were less than one (Fig. 1), which indicates that the data for ANC and PAC were suitable for QTL analysis.
The correlation analyses showed that ANC content was significantly and positively correlated with PAC content in the six environments, and the correlation coefficients were 0.271, 0.159, 0.348, 0.268, 0.163 and 0.247 for E1, E2, E3, E4, E5, and E6, respectively (Table 2).
A-QTLs for ANC and PAC content
A total of 21 A-QTLs for ANC and PAC content was detected using the ICIM method and QTL IciMapping v3.3 software (Table 3; Fig. 2). These QTLs were detected with LOD scores ranging from 2.5 to 4.7 and were distributed on chromosomes 1, 2, 3, 7, 8, 10 and 12. The explanation for the phenotypic variation by a single QTL varied from 3.8 to 34.8%.
Of the 21 A-QTLs, seven were detected for ANC content in all six different environments (Table 3). Two, three, three, two, two and one QTLs were detected in E1, E2, E3, E4, E5 and E6, respectively. The phenotypic variances of QTLs ranged from 7.1 to 34.8%. qANC7, qANC1-1 and qANC3 were detected in more than one environment. qANC3 was detected in E1, E3, E4, E5 and E6 and explained 17.6–34.8% of the phenotypic variance. The positive allele (an allele increasing the trait value) of qANC3 in five environments was carried by ‘HX1’. 14 A-QTLs were detected for PAC content in six different environments (Table 3). One, four, five, one, five and two QTLs were detected in E1, E2, E3, E4, E5 and E6, respectively. The phenotypic variances of the QTLs ranged from 3.8 to 23.3%. qPAC12-4, detected in E2, E3, E5 and E6 explained 14.0–23.3% of the phenotypic variance and was a major effect QTL for PAC content. The positive allele of qPAC12-4 in four environments was carried by ‘HX1’.
qANC2-2 and qPAC2-3 were in the SSR interval of RM530-RM166, and qANC10 and qPAC10 were in the SSR interval of RM24992–RM474. The phenotypic variances of the four QTLs were 10.4, 13.1, 10.1 and 11.2%, suggesting that common loci control the ANC and PAC content of rice.
AE-QTLs for ANC and PAC content
A total of nine QTLs were detected for ANC and PAC content by the combined analysis of phenotypic values using the MCIM method and QTLNetwork 2.1 software (Table 4; Fig. 3). All nine loci were detected with AE effects for the six environments but no AA and AAE effects were detected. Except for the locus RM1379–RM1347 on chromosome 2, other loci were detected by the ICIM method using QTL IciMapping v3.3 software including the two major effect QTLs, qANC3 and qPAC12-4 (Table 3).
For A-QTL, the effects of the QTLs ranged from − 0.1476 to 0.2205, and the explanation for the phenotypic variation by a single QTL varied from 0.25 to 2.01%. For AE-QTL, the effects of the QTLs ranged from − 0.441 to 0.1947, and the explanation for the phenotypic variation by a single QTL varied from 0.06 to 0.38%.
Discussion
Anthocyanins and proanthocyanidins are important bioactive substances in plants and constitute a major group of natural pigments in flowers, fruits and vegetables (Koponen et al. 2007; Finocchiaro et al. 2007). In this study, two parents of a RIL population, HX1, having red-colored rice, and S98-131, with white-colored grains, were crossed and studied. The ANC and PAC content were significantly higher in HX1 than in S98-131, indicating that the red rice has more anthocyanins and proanthocyanidins compared to common rice. In HX1, the PAC content was higher than the ANC content, suggesting that red rice has more proanthocyanidins compared to anthocyanins. This result was consistent with the research of (Oki et al. 2002) and (Finocchiaro et al. 2007, 2010).
Several genes necessary for anthocyanins and proanthocyanidins have been identified in rice seeds. Kim analyzed the expression levels of homologous genes from the anthocyanin biosynthetic pathway of Arabidopsis (Kim et al. 2007). Two genes, DFR and ANS, located on chromosome 1, had relatively high expression levels in the seeds from the black rice, Heugjinju. Some transcription factors associated with production of anthocyanin pigments were up-regulated or down-regulated in Heugjinju, but their functions in regulating anthocyanin biosynthesis need to be studied more deeply (Kim et al. 2010, 2011). Furukawa reported that the Rc and Rd genes are involved in proanthocyanidin synthesis in rice pericarp (Furukawa et al. 2006). The Rc gene encodes a bHLH protein and is located on 42.6–47.7 cM of chromosome 7; the Rd gene encodes a DFR protein and is located on 103.7–106.2 cM of chromosome 1. Rc, as a positive regulator of proanthocyanidin in rice, was reported by (Sweeney et al. 2006) around the same time. Dong reported four QTLs associated with the degree of red coloration (DRC) on rice chromosomes 1, 7, 9 and 11, and loci on chromosome 1 and 7 had the same location with previously characterized Rc and Rd genes (Dong et al. 2008). Therefore, the PAC content in rice was considered to be controlled by these genes on chromosome 1 and 7. In this study, 21 A-QTLs for ANC and PAC content were detected using the ICIM method. Among these QTLs, the major effect QTLs were qANC3 for ANC content and qPAC12-4 for PAC content, and each were detected in multiple environments. The two QTLs were located on different chromosomes from known genes, indicating that qANC3 and qPAC12-4 were new loci affecting ANC and PAC content. The accurate location of these loci on the chromosomes needs to be determined by QTL fine mapping.
qANC1-1 and qANC1-2 for ANC were located on 105.6–118.3 cM and 97.8–103.6 cM of chromosome 1, respectively. The two adjancent QTLs were most possibly the same QTL, but further fine mapping need to perform to verify it. qPAC2-1 and qPAC2-2 were also the possibility to be the same QTL. qANC1-1 and qANC1-2 locus had the same location on chromosome 1 with Rd gene controlling the PAC, and it was coincident with the positively correlated result of ANC and PAC. It is pleiotropism of Rd gene or close linkage of the two loci, which needs further verification. qPAC7 detected in E1 and E4 environments had the same location on chromosome 7 with Rc gene controlling the PAC. qPAC7 maybe the Rc gene detected in this study, but it has the very low LOD value and PVE. The key roles of Rc and Rd genes involve in PAC synthesis in red rice had been report (Furukawa et al. 2006; Sweeney et al. 2006), but in this study, major effect QTL in Rc gene locus were not detected. QTL mapping using different genetic populations or in different environments usually got various QTL loci. For example, thirteen of the 19 (68%) QTLs for rice grain appearance traits detected in Ha/Ne population did not appear in the Ha/IR population, including some large effect QTLs (Hosseini, et al. 2012). In addition, the measure method of PAC were obviously different from (Furukawa et al. 2006) and (Sweeney et al. 2006), so some new QTL loci for PAC were detected but the major effect Rc gene locus was not discovered.
qANC2-2 and qPAC2-3 on chromosome 2 and qANC10 and qPAC10 on chromosome 10 were detected within the same marker intervals, indicating a similar genetic background. The significantly positive correlation between ANC and PAC content also supports this conclusion. Several studies have demonstrated that anthocyanin and proanthocyanidin are produced in the early and middle steps of the flavonoid biosynthetic pathway and have a number of intersections (Winkel-Shirley 2001; Furukawa et al. 2006). The two pairs of QTLs detected in this study may play a common role to regulate the ANC and PAC content in rice.
Epistatic effects have been demonstrated to be prevalent for the inheritance of quantitative traits (Yu et al. 1997). When a QTL is involved in digenic epistatic interaction, its effects are mostly dependent on the genotypes at other QTLs (Hosseini et al. 2012). However, AA-QTLs were not detected in this study, suggesting that the interaction between QTLs affecting ANC or PAC content is feeble. QTL × environment interactions have been discussed in different plant breeding programs (Messmer et al. 2009; Ha et al. 2012; Bonneau et al. 2013). In the present study, eight of the total 21 A-QTLs were detected with AE effects by the combined analysis of phenotypic values in six environments using the MCIM method of QTLNetwork 2.1 software. From a breeding standpoint, a QTL with only additive effects that are independent of the environment are particularly appealing for MAS (Cao et al. 2001; Hosseini et al. 2012). However, QTLs dependent on the environment could also be useful in a specific environment, but be less important in other environments (Lark et al. 1995). Even so, the contribution rate of the explained phenotypic variation by A and AE-effects should also be considered. All of the AE-QTLs detected in this study have a tiny contribution rate of AE-effect compared to the A-effect; therefore, the AE effect of QTLs for ANC and PAC content could be negligible when using MAS.
Change history
14 December 2017
The author H.L. Chang is affiliated with “Innovation Center, Suihua Branch of Heilongjiang Academy of Agricultural Sciences, Suihua 152052, People’s Republic of China” in addition to the institution listed in the original publication.
References
Acquaviva R, Russo A, Galvano F, Galvano G, Barcellona ML, Volti GL, Vanella A (2003) Cyanidin and cyanidin 3-O-β-D-glucoside as DNA cleavage protectors and antioxidants. Cell Biol Toxicol 19(4):243–252
Agarwal C, Sharma Y, Zhao JF, Agarwal R (2000) A polyphenolic fraction from grape seeds causes irreversible growth inhibition of breast carcinoma MDA-MB468 cells by inhibiting mitogen-activated protein kinases activation and inducing G1 arrest and differentiation. Clin Cancer Res 6(7):2921–2930
Bagchi D, Krohn RL, Balmoori J, Bagchi M, Garg A, Stohs SJ (1999) Comparative in vitro and in vivo free radical scavenging abilities of a novel grape seed proanthocyanidin extract and selected antioxidants. Nat Antioxid Anticarcinog Nutr Health Dis 240:178–187
Barchi L, Lanteri S, Portis E, Valè G, Volante A, Pulcini L, Ciriaci T, Acciarri N, Barbierato V, Toppino L, Rotino GL (2012) A rad tag derived marker based eggplant linkage map and the location of QTLs determining anthocyanin pigmentation. PLoS ONE 7(8):e43740
Bonneau J, Taylor J, Parent B, Bennett D, Reynolds M, Feuillet C, Langride P, Mather D (2013) Multi-environment analysis and improved mapping of a yield-related QTL on chromosome 3B of wheat. Theore Appl Genet 126(3):747–761
Cao G, Zhu J, He C, Gao Y, Wu P (2001) impact of epistasis and QTL × environment interaction on the developmental behavior of plant height in rice (Oryza sativa L.). Theor Appl Genet 103(1):153–160
Cavagnaro PF, Iorizzo M, Yildiz M, Senalik D, Parsons J, Ellison S, Simon PW (2014) A gene-derived SNP-based high resolution linkage map of carrot including the location of QTL conditioning root and leaf anthocyanin pigmentation. BMC genom 15(1):1118
Chaim AB, Borovsky Y, Jong WD, Paran I (2003) Linkage of the A locus for the presence of anthocyanin and fs10.1, a major fruit-shape QTL in pepper. Theor Appl Genet 106(5):889–894
Cheng XD, Yan XJ, Hu MZ (2016) The effect of storage pressure on the mechanical properties of paddy grains. J Stored Prod Res 68:19–24
Dong YJ, Xu JL, Xiao K, Zhang YJ, Zhang JZ, Luo LJ, Mastuo M (2008) Genomic regions associated with the degree of red coloration in pericarp of rice (Oryza sativa L.). J Cereal Sci 48(2):556–560
Doyle J (1990) Isolation of plant dna from fresh tissue. Focus 12:13–15
Finocchiaro F, Ferrari B, Gianinetti A, Dall’Asta C, Galaverna G, Scazzina F, Pellegrini N (2007) Characterization of antioxidant compounds of red and white rice and changes in total antioxidant capacity during processing. Mol Nutr Food Res 51(8):1006–1019
Finocchiaro F, Ferrari B, Gianinetti A (2010) A study of biodiversity of flavonoid content in the rice caryopsis evidencing simultaneous accumulation of anthocyanins and proanthocyanidins in a black-grained genotype. J Cereal Sci 51(1):28–34
Furukawa T, Maekawa M, Oki T, Suda I, Iida S, Shimada H, Takamure I, Kadowaki K (2006) The Rc and Rd genes are involved in proanthocyanidin synthesis in rice pericarp. Plant J Cell Mol Biol 49(1):91–102
Ghosh M, Patra PK, Bhattacharyya C (2014) Effect of limited irrigation on growth and yield of rice varieties in a typic haplustalf soil of red and laterite zone of west bengal. J Crop Weed 10(634):42–47
Grassi D, Necozione S, Lippi C, Croce G, Valeri L, Pasqualetti P, Desideri G, Blumberg JB, Ferri C (2005) Cocoa reduces blood pressure and insulin resistance and improves endothelium-dependent vasodilation in hypertensives. Hypertension 46(2):398–405
Ha BK, Kim HK, Kang ST (2012) Mapping QTLs with epistatic effects and QTL-by-environment interactions for seed coat cracking in soybeans. Euphytica 186(3):933–942
Han L, Dong BC, Yang XJ, Huang CB, Wang XD, Wu XJ (2009) Study on flavonoids in the caryopsis of indica Rice Rdh. J Integr Agric 8(2):249–256
Hayashi K, Matsumoto S, Tsukazaki H, Kondo T, Kubo N, Hirai M (2010) Mapping of a novel locus regulating anthocyanin pigmentation in brassica rapa. Breed Sci 60(1):76–80
Heiss C, Kleinbongard P, Dejam A, Perré S, Schroeter H, Sies H, Kelm M (2005) Acute consumption of flavanol-rich cocoa and the reversal of endothelial dysfunction in smokers. J Am Coll Cardiol 46(7):1276–1283
Heiss C, Finis D, Kleinbongard P, Hoffmann A, Rassaf T, Kelm M, Sies H (2007) Sustained increase in flow-mediated dilation after daily intake of high-flavanol cocoa drink over 1 week. J Cardiovasc Pharmacol 49(2):74–80
Hosseini M, Houshmand S, Mohamadi S, Mohamadi S, Tarang A, Khodambashi M, Rahimsoroush H (2012) Detection of QTLs with main, epistatic and QTL × environment interaction effects for rice grain appearance quality traits using two populations of backcross inbred lines (BILs). Field Crops Res 135:97–106
Huang YF, Doligez A, Fournier-Level A, Cunff LL, Bertrand Y, Canaguier A, Morel C, Miralles V, Veran F, Souquet JM, Cheynier V, Terrier N, This P (2012) Dissecting genetic architecture of grape proanthocyanidin composition through quantitative trait locus mapping. BMC Plant Biol 12(1):30
Kim BG, Kim JH, Min SY, Shin KH, Kim JH, Kim HY, Ryu SN, Ahn JH (2007) Anthocyanin content in rice is related to expression levels of anthocyanin biosynthetic genes. J Plant Biol 50(2):156–160
Kim CK, Kikuchi S, Hahn JH, Park SC, Kim YH, Lee BW (2010) Computational identification of anthocyanin-specific transcription factors using a rice microarray and maximum boundary range algorithm. Evolut Bioinform 6(6):13
Kim CK, Cho MA, Choi YH, Kim JA, Kim YH, Kim YK, Park SH (2011) Identification and characterization of seed-specific transcription factors regulating anthocyanin biosynthesis in black rice. J appl genet 52(2):161–169
Koide T, Kamei H, Hashimoto Y, Kojima T, Hasegawa M (1996) Antitumor effect of hydrolyzed anthocyanin from grape rinds and red rice. Cancer Biother Radiopharm 11(4):273
Koponen JM, Happonen AM, Mattila PH, Törrönen AR (2007) Contents of anthocyanins and ellagitannins in selected foods consumed in Finland. J Agricu Food Chem 55(4):1612–1619
Lark KG, Chase K, Adler F, Mansur LM, Orf JH (1995) Interactions between quantitative trait loci in soybean in which trait variation at one locus is conditional upon a specific allele at another. Proc Natl Acad Sci USA 92(10):4656–4660
Lazze MC, Pizzala R, Savio M, Stivala LA, Prosperi E, Bianchi L (2003) Anthocyanins protect against DNA damage induced by tert-butyl-hydroperoxide in rat smooth muscle and hepatoma cells. Mutat Res 535(1):103–115
Li H, Ye G, Wang J (2007) A modified algorithm for the improvement of composite interval mapping. Genetics 175(1):361–374
Liu C, Ding NF, Fu QL, Brookes PC, Xu J, Guo B, Lin YC, Li H, Li NY (2016) The influence of soil properties on the size and structure of bacterial and fungal communities along a paddy soil chronosequence. Eur J Soil Biol 76:9–18
McCouch SR, Cho YG, Yano M, Paul E, Blinstrub M, Morishima H, Kinoshita T (1997) Report on QTL nomenclature. Rice Genet Newsl 14:11–14
Messmer R, Fracheboud Y, Bänziger M, Vargas M, Stamp P, Ribaut JM (2009) Drought stress and tropical maize: qTL-by-environment interactions and stability of QTLs across environments for yield components and secondary traits. Theor Appl Genet 119(5):913–930
Miller MG, Shukitt-Hale B (2012) Berry fruit enhances beneficial signaling in the brain. J Agric Food Chem 60(23):5709
Molina J, Sikora M, Garud N, Flowers JM, Rubinstein S, Reynolds A, Huang P, Jackson S, Schaal BA, Bustamante CD, Boyko AR, Purugganan MD (2011) Molecular evidence for a single evolutionary origin of domesticated rice. Proc Natl Acad Sci USA 108(20):8351
Norberto S, Silva S, Meireles M, Faria A, Pintado M, Calhau C (2013) Blueberry anthocyanins in health promotion: a metabolic overview. J Funct Foods 5(4):1518–1528
Oki T, Masuda M, Kobayashi M, Nishiba Y, Furuta S, Suda I, Sato T (2002) Polymeric procyanidins as radical-scavenging components in red-hulled rice. J Agric Food Chem 50(26):7524–7529
Pang YZ, Wenger JP, Saathoff K, Peel GJ, Wen JQ, Huhman D, Allen SN, Tang Y, Cheng XF, Tadege M, Ratet P, Mysore KS, Sumner LW, Marks MD, Dixon RA (2009) A WD40 repeat protein from Medicago truncatula is necessary for tissue-specific anthocyanin and proanthocyanidin biosynthesis but not for trichome development. Plant Physiol 151(3):1114–1129
Przybylski R, Lee YC, Eskin NAM (1998) Antioxidant and redicl-scavenging activities of buckwheat seed components. J American Oil Chemists’ Society 75(11):1595–1601
Russo A, Fauci LL, Acquavivi R, Campisi A, Raciti G, Scifo C, Renis M, Galvano G, Vanella A, Galvano F (2005) Ochratoxin A-induced DNA damage in human fibroblast: protective effect of cyanidin 3-O-β-D-glucoside. J Nutr Biochem 16(1):31–37
Shi B, Du X (2006) The progress on research and utilization of plant proanthocyanidins. J Sichuan Univ 38(5):16–24
Sweeney MT, Thomson MJ, Pfeil BE, Mccouch S (2006) Caught red-handed: Rc encodes a basic Helix-Loop-Helix protein conditioning red pricarp in rice. Plant Cell 18(2):283–294
Taubert D, Roesen R, Lehmann C, Jung N, Schömig E (2007) Effects of low habitual cocoa intake on blood pressure and bioactive nitric oxide: a randomized controlled trial. JAMA, J Am Med Assoc 298(1):49–60
Trigiano RN, Caetano-Anolles G (1998) Laboratory exercises on DNA amplification fingerprinting for evaluating the molecular diversity of horticultural species. Horttechnology 8(3):413–422
Tsuda T (2012) Dietary anthocyanin-rich plants: biochemical basis and recent progress in health benefits studies. Mol Nutr Food Res 56(1):159–170
Williamson G, Clifford MN (2010) Colonic metabolites of berry polyphenols: the missing link to biological activity? Br J Nutr 104(S3):S48–S66
Winkel-shirley B (2001) Flavonoid biosynthesis. a colorful model for genetics, biochemistry, cell biology, and biotechnology. Plant Physiol 126(2):485–493
Yang J, Zhu J, Williams RW (2007) Mapping the genetic architecture of complex traits in experimental populations. Bioinformatics 23(12):1527–1536
Yu SB, Li JX, Tan YF, Gao YJ, Li XH, Zhang QF, Maroof MAS (1997) Importance of epistasis as the genetic basis of heterosis in an elite rice hybrid. Proc Natl Acad Sci USA 94(17):9226–9231
Yuan YW, Sagawa JM, Young RC, Christensen BJ, Bradshaw HD Jr (2013) Genetic dissection of a major anthocyanin qtl contributing to pollinator-mediated reproductive isolation between sister species of mimulus. Genetics 194(1):255–263
Acknowledgements
This work was supported by the major science and technology fund bidding project in Heilongjiang province (GA14B102-02).
Author information
Authors and Affiliations
Corresponding author
Additional information
T. Y. Xu, J. Sun and H. L. Chang have contributed equally to this work.
A correction to this article is available online at https://doi.org/10.1007/s10681-017-2084-0.
Rights and permissions
About this article
Cite this article
Xu, T.Y., Sun, J., Chang, H.L. et al. QTL mapping for anthocyanin and proanthocyanidin content in red rice. Euphytica 213, 243 (2017). https://doi.org/10.1007/s10681-017-2035-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10681-017-2035-9