Abstract
The hyperspectral remote sensing datasets possess high capability in differentiating the spectrally similar features, and thus, they are immensely important in various forestry activities, especially vegetation classifications. But extracting endmembers for data training is a challenging task. The present study is focused on the use of automated endmember extraction technique for deriving endmembers during the unavailability of ground spectra. We used the Sequential Maximum Angle Convex Cone (SMACC) method on EO-1 Hyperion data for endmember extraction in the Barkot forest range of Dehradun district, Uttarakhand which were used for classification of the study area using support vector machine (SVM). Further, we estimated the vegetation health of the region by assigning the threshold weights for various derived environmental variables such as NDVI (Normalised Difference Vegetation Index), CRI (Carotenoid Reflectance Index), Anthocyanin Reflectance Index (ARI), Modified Simple Ratio (MSR), Modified Chlorophyll Absorption Ratio Index (MCARI) and WBI (Water Band Index). Then, to further validate the health of the forest types, we correlated it with the Land Surface Temperature (LST) from LANDSAT 5 ETM + data. The results showed a high classification accuracy of 89.13%. The healthy vegetation area coverage of the area was about 78.6% with most healthy class as Tectona grandis and Shorea robusta and its correlation with LST showed lower temperature range in healthy vegetation areas and vice versa. The study was useful in determining the superiority of SMACC automated endmember extraction and estimating the vegetation health.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
The pure spectral elements represent a quick synopsis of the vast hyperspectral image data (Thompson et al., 2010). An endmember is the unmixed representative of a sample and its extraction is an utmost important process carried out in a hyperspectral data analysis (Plaza & Chang, 2006). The spectral unmixing is an important step in hyperspectral data analysis to determine pure spectral elements for classification (Somers et al., 2012). According to Veganzones and Grana (2008), there are various methods of spectral unmixing such as geometric method, lattice computing method and heuristic method which further have subtypes. A number of methods of endmember extraction were compared by Plaza et al. (2004) to determine the better performing algorithm, and considered the linear mixture model to be more appropriate. Chen et al. (2018) used the Sequential Maximum Angle Convex Cone (SMACC) in combination with hyperspectral data for determining the metallic-rich zone in rocks. The work done by Aufaristama et al. (2018) involved the utilisation of SMACC for endmember selection for working on volcanic results considered SMACC to be quick in endmember selection. The classification performed using the endmembers could help in further analysis and monitoring of various parameters like vegetation.
Vegetation is an important component of the ecosystem owing to their contribution in atmospheric reciprocity and various climate change-related activities of the world (Pei et al., 2018). The impact of various natural and man-made phenomena, viz., urban sprawl and climate dynamics, highly determines floral behaviour (Pei et al., 2018; Ekwueme & Agunwamba, 2021). For example, the activity of vegetation is enhanced during carbon dioxide expansion (Piao et al., 2012) and diminuted in droughts (Ji & Peters, 2003). The dynamicity in biodiversity stability has been due to the climate change and land use patterns (Sala et al., 2000; Hansen et al., 2001). Global warming has been responsible for dragging down global biodiversity under the threat zone (Malcolm et al., 2006) and biological alterations (Parmesan & Yohe, 2003; Root et al., 2003). Studies have been carried out to determine the impact of global warming on biodiversity worldwide (Kappelle et al., 1999; Noss, 2001). Hence, it is of utmost importance to monitor the environment for detecting the changes in the biodiversity composition (Prasad et al., 2010) which includes the vegetation health.
The vegetation health monitoring is essential to determine stress conditions in vegetation and hyperspectral remote sensing plays an important role to serve the purpose (Kureel et al., 2021). Shafri and Hamdan (2009) considered the red edge-based technique better than the indices based while working on plant disease infection. However, the study by Dutta et al. (2009) determined the vegetation health using vegetation indices and random forest method and inferred the method to be reliable for vegetation health analysis if a threshold is provided for the vegetation indices to categorise to healthy or stressed class. Further, Kureel et al. (2021) derived the vegetation health of Lonar forest in Maharashtra using the hyperspectral remote sensing and a combination of several indices and concluded the method to be efficient in vegetation health analysis.
Remote sensing is a promising and more efficient technology which has gained weightage over the traditional methods of mapping (Kuenzer, 2011). The use of multispectral and hyperspectral data has been popular for the determination of the dynamics of the ecosystem (Shippert, 2003). But in recent years, the use of hyperspectral remote sensing has gained much popularity and has been considered a very useful technology (Navin & Agilandeeswari, 2020). But only a limited set of works have been carried out using SMACC for endmember extraction in combination with support vector machine (SVM). The present study highly contributes towards highlighting the advantages of the automated endmember extraction method SMACC for selecting the endmembers in hyperspectral data classification as we hypothesised that SMACC is highly beneficial for endmember selection and further the use of various environmental variables for the estimation of vegetation health. Hence, the objective of the study is (1) to determine the efficiency of SMACC for endmember extraction by deriving LULC of forest from it using SVM and (2) to further determine the vegetation health status from the same.
Materials and method
Study area
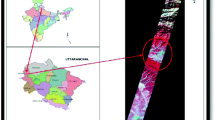
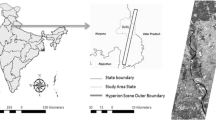
We took the Barkot forest range of Dehradun district in the state of Uttarakhand in India as the study area around 30° 06′ North longitude and 78° 18′ East latitude as in Fig. 1. This region is situated in the Himalayan foothills with an altitude range of 340 to 560 m above mean sea level (MSL) (Attri & Kushwaha, 2018). It covers the parts of Rajaji national park, and the range of Shivaliks. The main land use land cover (LULC) classes are forest, urban, water body, grassland and cropland. The rivers Ganga, Chandrabhaga and Song flow around the area.
Being bounded by the lush green forest ranges of Motichur (southern region) and Lachchhiwala (western region) and the urban settlements of Doiwala, Rishikesh and Bhaniawala, the region lies into the Sub-Group 3C North Indian Tropical Moist Deciduous Forests class of Champion and Seth’s (1968) forest classification of Indian forests (Attri & Kushwaha, 2018). Tectona grandis (Teak), Shorea robusta (Sal) and Mallotus phillipensis (Indian redwood) are the main tree species of forest (Bhattacharjee et al., 2019). Falling into the tropical to subtropical moist climate category, the temperature range varies from minimum of about 2 °C to maximum of about 41 °C with average annual precipitation of about 2300 mm (Nandy et al., 2017). The area consists of gentle slopes with fine and loamy thermic haplustalf type of soil (Shah & Subudhi, 2009).
Data used
For analysis, we used the cloud-free satellite data which was derived from the United State Geological Survey (USGS) (http://earthexplorer.usgs.gov/) open source geo-portal. The details of the data are given in Table 1.
Data pre-processing
We performed the pre-processing of the EO-1 Hyperion data to derive the desired results for analysis and it involved the bad band removal as the first step in which we selected the bands with non-zero data and we removed the bands with noise. We carried out de-stripping to remove noisy lines to obtain noise-free equivalent data. For this, we used the Tactical Hyperspectral Operations Resource (THOR) workflow. Further, we eliminated the bad column. In this, we selected the column and assigned them a value from the average values of the neighbouring pixels or columns.
After the elimination of the bad bands and the bad columns, then we performed the correction for atmospheric errors using the Fast Line of Sight Atmospheric Analysis of Spectral Hypercubes (FLAASH). It requires various parameters which vary according to a given set of conditions. The parameters used are depicted in Table 2.
Finally, we georeferenced the data using the Landsat 5 ETM + data.
Endmember selection
We performed the endmember selection for classification. The endmembers signify the pure classes of an image that have values not lower than zero. For this, the method used was the SMACC. SMACC finds the spectral endmembers and their abundances throughout an image with the help of the angle made with the current cone, of which the vector with maximum angle is selected for the endmember and the method is useful for hyperspectral dataset due to impairment of unmixing (Gruninger et al., 2004). The endmembers were selected for the ten classes S. robusta (Sal), T. grandis (Teak), mixed forest, scrub, grass, riverine forest, cropland, settlement, dry riverbed and water. Now, the selected endmembers were used to select region of interest (ROI). After that, we matched the spectra of the selected species to create its ROI. The classification was performed using SVM.
For the health assessment, we derived the spectral variables NDVI (Normalised Difference Vegetation Index), CRI (Carotenoid Reflectance Index), Anthocyanin Reflectance Index (ARI), Modified Simple Ratio (MSR), Modified Chlorophyll Absorption Ratio Index (MCARI) and WBI (Water Band Index), the details of which are mentioned in Table 3. We assigned the threshold range of values and accordingly weightage favouring the healthy vegetation for each of the variables based on various literatures (Rouse, 1974; Hati et al., 2020; Chen, 1996; Peñuelas et al., 1993) and is depicted in Fig. 2.
Further, we calculated the Land Surface Temperature (LST) where Pv was used to derive emissivity as \(\boldsymbol P\boldsymbol v\boldsymbol={\boldsymbol\lbrack\boldsymbol(\boldsymbol N\boldsymbol D\boldsymbol V\boldsymbol I\boldsymbol-\boldsymbol N\boldsymbol D\boldsymbol V\boldsymbol I\boldsymbol m\boldsymbol i\boldsymbol n\boldsymbol)\boldsymbol/\boldsymbol(\boldsymbol N\boldsymbol D\boldsymbol V\boldsymbol I\boldsymbol m\boldsymbol a\boldsymbol x\boldsymbol-\boldsymbol N\boldsymbol D\boldsymbol V\boldsymbol I\boldsymbol m\boldsymbol i\boldsymbol n\boldsymbol)\boldsymbol\rbrack}^{\mathbf2}\) (Chavez, 1996) where Pv = proportion vegetation, NDVImin = minimum value of NDVI, and NDVImax = maximum value of NDVI.
Now, ETM6 (emmissivity) was calculated as.
\({E}_{TM6}=0.004Pv+0.986\) (Moran et al., 1992).
Then, L (spectral radiance at sensor) calculated as \(\boldsymbol L=\lbrack({\boldsymbol L}_{\boldsymbol M\boldsymbol A\boldsymbol X}-{\boldsymbol L}_{\mathbf M\mathbf I\mathbf N})/(\boldsymbol Q\boldsymbol C\boldsymbol A{\boldsymbol L}_{\mathbf M\mathbf A\mathbf X}-\boldsymbol Q\boldsymbol C\boldsymbol A{\boldsymbol L}_{\mathbf M\mathbf I\mathbf N})\rbrack\ast\boldsymbol(\boldsymbol Q\boldsymbol C\boldsymbol A\boldsymbol L-\boldsymbol Q\boldsymbol C\boldsymbol A{\boldsymbol L}_{\mathbf M\mathbf I\mathbf N})+{\boldsymbol L}_{\mathbf M\mathbf I\mathbf N}\) (Brivio et al., 2006) where,
QCAL = quantized calibrated pixel value in DN, LMAX = spectral radiance scaled to QCALMIN (here LMAX = 15.303 W/(m2*sr*um)), LMIN = spectral radiance scaled to QCALMIN (here LMIN = 1.238 W/(m2*sr*um)), QCALMAX = maximum quantized pixed value in DN (here, QCALMAX = 255) and QCALMIN = minimum quantized pixed value in DN (here, QCALMIN = 1).
Then LST was calculated as \(\boldsymbol B\boldsymbol T=({\boldsymbol K}_{\mathbf2}/(\boldsymbol l\boldsymbol n({\boldsymbol K}_{\mathbf1}/\boldsymbol L)+\mathbf1))-\mathbf{273}\boldsymbol.\mathbf1\) where,
K1 = band-specific thermal conversion constant from the metadata (here, K1 = 607.76),
K2 = band-specific thermal conversion constant from the metadata (here, K2 = 1260.56) and L = spectral radiance at sensor. The methodology flowchart is depicted in Fig. 3.
Result and discussion
The LULC classification was derived along with the vegetation health and LST map.
The SVM classification shown in Fig. 4 resulted in an overall accuracy of 89.13% and fetched the kappa coefficient 0.87. The class S. robusta exhibited an accuracy of 93.40% which was the maximum among all the classes and it was followed by T. grandis (92.88%) and this explains the species level classification efficiency of SMACC in the hyperspectral data. The efficiency of SMACC has been earlier considered by Chen et al. (2018) and Aufaristama et al. (2018) for classification. Also, the high accuracy of the abovementioned classes might be due to the presence of more super pixels.
Further, the combination of the variables displayed the majority area (78.6%) falling under the most healthy to moderately healthy vegetation category (Fig. 5). The most healthy vegetation was found in the class T. grandis species followed by S. robusta. This might be due to lack of anthropogenic disturbances in the area and favourable environment, topographic and climatic factors. The least healthy categories included the riparian forest, croplands, river and settlement. The variations of the edaphic factors of riverine or riparian forests due to flood conditions such as the deposition of new sediments, duration of existence and number of times of occurrence of floods challenge the adaptability potential of the vegetation in those forests (Priyadarshana et al., 2009). Similarly, the impact of climate change on the hydrological dynamics (Oo et al., 2020; Faye et al., 2022) could contribute towards limiting the vegetation health. This justifies the riparian forests falling under the least healthy category. Similar work was done and the vegetation health was derived where the method was considered to serve the purpose efficiently (Kureel et al., 2021; Dutta et al., 2009). This may be due to the wide range of values that discriminate the vegetation classes according to various parameters determined by the variables, where the ones with most suitable values in all variables are considered to be healthy. This determines the effectiveness of environmental variables in vegetation analysis.
The minimum LST range was 13.13 to 18.40 °C which again was for the T. grandis and S. robusta class (Fig. 6). In the vegetation category, the highest LST was of the riparian forest. This showed the relation of vegetation health with the LST where LST is lowest in case of healthiest vegetation. Hernández-Clemente et al. (2019) considered the use of hyperspectral data and thermal data to be beneficial for signifying the vegetation health as they determine the biophysical changes.
The use of an efficient automated endmember extraction method like SMACC with one of the most systematic classification methods SVM into the hyperspectral data showcased the technological strength of the work. The vegetation health estimation and categorisation and correlation with LST could clear paths for numerous future research works based on the use of hyperspectral data. The hyperspectral data carries great potential in floral analysis on species level and the addition of the health estimation to it could further enhance the decision-making strength in various conservation plans. However, despite the perks of the number of bands and narrow band width, both the spatial and temporal limitations in the data availability of hyperspectral data are the major weakness. The multispectral data like Sentinel-2 cover the major regions on the earth and have the data repetivity which proves to be useful in current time data analysis but the same cannot be stated for the hyperspectral data as the coarse temporal resolution and limited region availability of data restricts the emergence of countless number of fruitful and great research works.
Conclusion
The study reflected the importance of automated endmember extraction method SMACC for the hyperspectral dataset which was useful for deriving pure endmembers for ROI according to our hypothesis. A good accuracy was observed in vegetation species level separability in SMACC-derived ROI. It was a quick method to derive the super pixels. The healthiest vegetation classes were T. grandis and D. sissoo whereas the riverine forests were categorised under the stressed category and expected to have deteriorating or poor health. Hence, some necessary measures are prescribed to drag it to the healthy vegetation category as it may harbour important, endemic and rare species which need conservation. The LST is inversely proportional to the vegetation health. Also the vegetation of poor health conditions lead to expansion in LST.
The global concern of climate change and global warming fetches the issue of biodiversity to be vulnerable to decline. This study contributes towards future studies as it is helpful for various forestry activities such as biodiversity conservation and contributes towards floral monitoring and mapping. However, the major limitation of the study lies in the lack of periodic data availability as the hyperspectral data are openly available for some limited region without regular repetivity. Also, the endmember selection process was approximate which might affect the precision value.
References
Attri, P., & Kushwaha, S. P. S. (2018). Estimation of biomass and carbon pool in Barkot forest range, UK using geospatial tools. ISPRS Annals of the Photogrammetry, Remote Sensing and Spatial Information Sciences, 4, 121–128. https://doi.org/10.5194/isprs-annals-IV-5-121-2018
Aufaristama, M., Ulfarsson, M. O., Höskuldsson, A., & Jónsdóttir, I. (2018, September). Application of airborne hyperspectral remote sensing for mapping surface mineral and volcanic products at 2014–2015 Holuhraun lava flow (Iceland) using Sequential Maximum Angle Convex Cone (SMACC) method. In 10th Cities on Volcanoes, CoV 2018. https://doi.org/10.13140/RG.2.2.32139.75042.
Bhattacharjee, R., Nandy, S., Sett, T., & Gupta, A. (2019). Tree parameters retrieval and volume estimation using terrestrial laser scanner: A case study on Barkot forest.
Brivio, P., Lechi-Lechi, G., & Zilioli, E. (2006). Principi e metodi di telerilevamento (pp. 1–525). CittaStudi.
Chavez, P. S. (1996). Image-based atmospheric corrections-revisited and improved. Photogrammetric Engineering and Remote Sensing, 62(9), 1025–1035.
Chen, J. M. (1996). Evaluation of vegetation indices and a modified simple ratio for boreal applications. Canadian Journal of Remote Sensing, 22(3), 229–242. https://doi.org/10.1080/07038992.1996.10855178
Chen, J., Bing, Z., Mao, Z., Zhang, C., Bi, Z., & Yang, Z. (2018). Using geochemical data for prospecting target areas by the sequential maximum angle convex cone method in the Manzhouli area,m China. Geochemical Journal, 52(1), 13–27. https://doi.org/10.2343/geochemj.2.0493
Daughtry, C. S., Walthall, C. L., Kim, M. S., De Colstoun, E. B., & McMurtrey Iii, J. E. (2000). Estimating corn leaf chlorophyll concentration from leaf and canopy reflectance. Remote Sensing of Environment, 74(2), 229–239. https://doi.org/10.1016/S0034-4257(00)00113-9
Dutta, D., Singh, R., Chouhan, S., Bhunia, U., Paul, A., Jeyaram, A., & Murthy, Y. K. (2009, September). Assessment of vegetation health quality parameters using hyperspectral indices and decision tree classification. In Proceedings of the ISRS Symposium, Nagpur, Maharashtra (pp. 17–19).
Ekwueme, B. N., & Agunwamba, J. C. (2021). Trend analysis and variability of air temperature and rainfall in regional river basins. Civil Engineering Journal, 7, 816–826. https://doi.org/10.28991/cej-2021-03091692
Faye, N., Diallo, A., Sagna, M. B., Peiry, J. L., Sarr, P. S., & Guisse, A. (2022). Influence of anthropic and eco-hydrological factors on the floristic diversity of the herbaceous vegetation around the temporary ponds in Ferlo, Northern Senegal. Journal of Plant Ecology, 15(1), 26–38.
Gitelson, A. A., Merzlyak, M. N., & Chivkunova, O. B. (2001). Optical properties and nondestructive estimation of anthocyanin content in plant leaves. Photochemistry and Photobiology, 74(1), 38–45. https://doi.org/10.1562/0031-8655(2001)0740038OPANEO2.0.CO2
Gitelson, A. A., Zur, Y., Chivkunova, O. B., & Merzlyak, M. N. (2002). Assessing carotenoid content in plant leaves with reflectance spectroscopy. Photochemistry and Photobiology, 75(3), 272–281. https://doi.org/10.1562/0031-8655(2002)0750272ACCIPL2.0.CO2
Gruninger, J. H., Ratkowski, A. J., & Hoke, M. L. (2004, August). The sequential maximum angle convex cone (SMACC) endmember model. In Algorithms and technologies for multispectral, hyperspectral, and ultraspectral imagery X (Vol. 5425, pp. 1–14). International Society for Optics and Photonics. https://doi.org/10.1117/12.543794
Hansen, A. J., Neilson, R. P., Dale, V. H., Flather, C. H., Iverson, L. R., Currie, D. J., & Bartlein, P. J. (2001). Global change in forests: Responses of species, communities, and biomes: Interactions between climate change and land use are projected to cause large shifts in biodiversity. BioScience, 51(9), 765–779. https://doi.org/10.1641/0006-3568(2001)051[0765:GCIFRO]2.0.CO;2
Hati, J. P., Goswami, S., Samanta, S., Pramanick, N., Majumdar, S. D., Chaube, N. R., & Hazra, S. (2020). Estimation of vegetation stress in the mangrove forest using AVIRIS-NG airborne hyperspectral data. Modeling Earth Systems and Environment. https://doi.org/10.1007/s40808-020-00916-5
Hernández-Clemente, R., Hornero, A., Mottus, M., Peñuelas, J., González-Dugo, V., Jiménez, J. C., & Zarco-Tejada, P. J. (2019). Early diagnosis of vegetation health from high-resolution hyperspectral and thermal imagery: Lessons learned from empirical relationships and radiative transfer modelling. Current Forestry Reports, 5(3), 169–183. https://doi.org/10.1007/s13593-014-0246-1
Ji, L., & Peters, A. J. (2003). Assessing vegetation response to drought in the northern Great Plains using vegetation and drought indices. Remote Sensing of Environment, 87(1), 85–98.
Kappelle, M., Van Vuuren, M. M., & Baas, P. (1999). Effects of climate change on biodiversity: A review and identification of key research issues. Biodiversity & Conservation, 8(10), 1383–1397. https://doi.org/10.1023/A:1008934324223
Kuenzer, C., Bluemel, A., Gebhardt, S., Quoc, T. V., & Dech, S. (2011). Remote sensing of mangrove ecosystems: A review. Remote Sensing, 3(5), 878–928. https://doi.org/10.3390/rs3050878
Kureel, N., Sarup, J., Matin, S., Goswami, S., & Kureel, K. (2021). Modelling vegetation health and stress using hypersepctral remote sensing data. Modeling Earth Systems and Environment. https://doi.org/10.1201/b11222-20
Level-III, L. I. L. I. Class description Champion and Seth (1968) class with.
Malcolm, J. R., Liu, C., Neilson, R. P., Hansen, L., & Hannah, L. E. E. (2006). Global warming and extinctions of endemic species from biodiversity hotspots. Conservation Biology, 20(2), 538–548. https://doi.org/10.1111/j.1523-1739.2006.00364.x
Moran, M. S., Jackson, R. D., Slater, P. N., & Teillet, P. M. (1992). Evaluation of simplified procedures for retrieval of land surface reflectance factors from satellite sensor output. Remote Sensing of Environment, 41(2–3), 169–184. https://doi.org/10.1016/0034-4257(92)90076-V
Nandy, S., Singh, R., Ghosh, S., Watham, T., Kushwaha, S. P. S., Kumar, A. S., & Dadhwal, V. K. (2017). Neural network-based modelling for forest biomass assessment. Carbon Management, 8(4), 305–317. https://doi.org/10.1080/17583004.2017.1357402
Navin, M. S., & Agilandeeswari, L. (2020). Multispectral and hyperspectral images based land use/land cover change prediction analysis: An extensive review. Multimedia Tools and Applications, 79(39), 29751–29774. https://doi.org/10.1007/s11042-020-09531-z
Noss, R. F. (2001). Beyond Kyoto: Forest management in a time of rapid climate change. Conservation Biology, 15(3), 578–590. https://doi.org/10.1046/j.1523-1739.2001.015003578.x
Oo, H. T., Zin, W. W., & Kyi, C. T. (2020). Analysis of streamflow response to changing climate conditions using SWAT model. Civil Engineering Journal, 6(2), 194–209.
Parmesan, C., & Yohe, G. (2003). A globally coherent fingerprint of climate change impacts across natural systems. Nature, 421(6918), 37–42. https://doi.org/10.1038/nature01286
Pei, F., Wu, C., Liu, X., Li, X., Yang, K., Zhou, Y., & Xia, G. (2018). Monitoring the vegetation activity in China using vegetation health indices. Agricultural and Forest Meteorology, 248, 215–227. https://doi.org/10.1016/j.agrformet.2017.10.001
Peñuelas, J., Filella, I., Biel, C., Serrano, L., & Save, R. (1993). The reflectance at the 950–970 nm region as an indicator of plant water status. International Journal of Remote Sensing, 14(10), 1887–1905. https://doi.org/10.1080/01431169308954010
Piao, S., Tan, K., Nan, H., Ciais, P., Fang, J., Wang, T., & Zhu, B. (2012). Impacts of climate and CO2 changes on the vegetation growth and carbon balance of Qinghai-Tibetan grasslands over the past five decades. Global and Planetary Change, 98, 73–80. https://doi.org/10.1016/j.gloplacha.2012.08.009
Plaza, A., & Chang, C. I. (2006). Impact of initialization on design of endmember extraction algorithms. IEEE Transactions on Geoscience and Remote Sensing, 44(11), 3397–3407. https://doi.org/10.1109/TGRS.2006.879538
Plaza, A., Martínez, P., Pérez, R., & Plaza, J. (2004). A quantitative and comparative analysis of endmember extraction algorithms from hyperspectral data. IEEE Transactions on Geoscience and Remote Sensing, 42(3), 650–663. https://doi.org/10.1109/TGRS.2003.820314
Prasad, P., Pattanaik, C., Prasad, S. N., & Dutt, C. B. S. (2010). Analysis of spatial and temporal changes in mangroves along Thane Creek of Mumbai (India) using geospatial tools. The IUP Journal of Environmental Sciences, 4(4), 52–59.
Priyadarshana, T., Asaeda, T., Manatunge, J., Fujino, T., & Gamage, N. P. (2009). Dynamics, threats, responses and recovery of riverine-riparian flora. Oceans and Aquatic Ecosytems, 1, 256–285.
Root, T. L., Price, J. T., Hall, K. R., Schneider, S. H., Rosenzweig, C., & Pounds, J. A. (2003). Fingerprints of global warming on wild animals and plants. Nature, 421(6918), 57–60. https://doi.org/10.1038/nature01333
Rouse, J. W. (1974). Monitoring the vernal advancement of retrogradation of natural vegetation. NASA/GSFC, type III, final report, greenbelt, MD, 371.
Sala, O. E., Chapin, F. S., Armesto, J. J., Berlow, E., Bloomfield, J., Dirzo, R., & Wall, D. H. (2000). Biodiversity-global biodiversity scenarios for the year 2100. Science, 287(5459), 1770–1774. https://doi.org/10.1126/science.287.5459.1770
Shafri, H. Z., & Hamdan, N. (2009). Hyperspectral imagery for mapping disease infection in oil palm plantation using vegetation indices and red edge techniques. American Journal of Applied Sciences, 6(6), 1031.
Shah, R. K., & Subudhi, S. P. (2009). Working plan of dehradun forest division. Dehradun (Shiwalik circle), Part-I, 1, 10.
Shippert, P. (2003). Introduction to hyperspectral image analysis. Online Journal of Space Communication, 2(3), 8.
Somers, B., Zortea, M., Plaza, A., & Asner, G. P. (2012). Automated extraction of image-based endmember bundles for improved spectral unmixing. IEEE Journal of Selected Topics in Applied Earth Observations and Remote Sensing, 5(2), 396–408. https://doi.org/10.1109/JSTARS.2011.2181340
Thompson, D. R., Mandrake, L., Gilmore, M. S., & Castano, R. (2010). Superpixel endmember detection. IEEE Transactions on Geoscience and Remote Sensing, 48(11), 4023–4033. https://doi.org/10.1109/TGRS.2010.2070802
Veganzones, M. A., & Grana, M. (2008, September). Endmember extraction methods: A short review. In International conference on knowledge-based and intelligent information and engineering systems (pp. 400–407). Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-540-85567-5_50.
Acknowledgements
We present our immense gratitude to USGS earth explorer for the provision of the EO-1 Hyperion data and Landsat 5 ETM+ data which were used in the analysis. Also, we are very thankful to ENVI and ESRI ARCGIS for providing the pre-processing, processing and result extraction platform.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Singh, R., Kumar, V. Evaluating automated endmember extraction for classifying hyperspectral data and deriving spectral parameters for monitoring forest vegetation health. Environ Monit Assess 195, 72 (2023). https://doi.org/10.1007/s10661-022-10576-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10661-022-10576-w