Abstract
Bacterial spot caused by several Xanthomonas spp. is an economically important disease of tomato (Solanum lycopersicum L.). Host resistance to the disease is partially dominant or incomplete, which requires accurate assessment of disease severity for genetic studies of resistance. In the present study, three independent experiments were conducted to investigate the feasibility of using image analysis to estimate foliar disease severity of bacterial spot in tomato. The resistant line PI 114490 and the susceptible line OH 88119 were used in the first experiment, five tomato lines (PI 114490, PI 128216, Hawaii 7981, Hawaii 7998, and Fla. 7600) with a range of resistance and OH 88119 were used in the second experiment, and 439 F2 individuals from a cross between OH 88119 and PI 114490 were used in the third experiment. Tomato plants were spray-inoculated with bacterial spot race T3. Five diseased leaves from each plant were randomly collected and scanned to obtain digital images 21 days after inoculation. The disease severity (% leaf area) was measured using image analysis. The susceptible line OH 88119 showed the most severe disease. The resistant line PI 114490 showed the least severe disease, and was not significantly different to PI 128216 or Hawaii 7981. These results indicated that image analysis could be used to distinguish tomato lines with different resistance to bacterial spot. Marker-trait association analysis identified four quantitative trait loci conferring resistance to race T3 in PI 114490 using data obtained from image analysis, the Horsfall-Barratt (HB) category scale data, and HB midpoint converted values. However, the disease severity was slightly underestimated using the HB category scale and the phenotypic variation explained by each marker was overestimated using the HB category data compared to using the image analysis-measured disease severity data. Therefore, image analysis could provide a consistent, accurate and reliable method compared to the HB scale to estimate disease severity for genetic studies of foliar bacterial spot in tomato.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Bacterial spot, caused by four species of Xanthomonas (X. euvesicatoria, X. vesicatoria, X. perforans, and X. gardneri) is a serious disease of tomato (Solanum lycopersicum L.) particularly in wet and humid environments (Jones et al. 1991). The disease occurs in open field production causing severe losses in total and marketable yields (Yang et al. 2007; Stall et al. 2009). More recently, the disease has become widespread in protected tomato production areas including greenhouses in China (Wang et al. 2005; Liu et al. 2008; Guo et al. 2008; Zhang et al. 2010). The existence of multiple species and races of these pathogens, as well as lack of efficacy of commonly applied bactericides makes it difficult to effectively control the disease. The use of host resistance is believed to be an inexpensive, effective, and environmentally friendly approach to manage the disease.
Accurate assessment of disease severity is critical for several purposes including monitoring epidemics, evaluation of germplasm, genetic studies of resistance, and breeding. Although the bacterial spot pathogen can infect all above ground parts of tomato plants, most studies have been based on assessment of foliar disease (Yang et al. 2007). Several disease assessment variables including visual estimates of disease severity (Yang 2013), percentage defoliation (Scott and Jones 1989; Somodi et al. 1994), and number of lesions on leaves (Sotirova and Bogatsevska 2000) have been used to rate the severity of bacterial spot on tomato. The Horsfall-Barratt (HB) category scale has been used for the visual assessment of plant disease severity as an effort to compensate for human error in estimating the percentage of foliage infected (Horsfall and Barratt 1945). The HB scale is a 12-category, non-linear scale with logarithmically increasing interval sizes symmetrical around 50 % (Horsfall and Barratt 1945). Thus the interval size up to 12 % severity is 3 to 6 % percentage points, but from 12 to 50 % is 12 to 25 % percentage points. It is the most frequently used method in visual estimates of disease severity of tomato bacterial spot (Somodi et al. 1994; Scott et al. 1995, 2003; Zhang et al. 2009; Hutton et al. 2010a, b; Sun et al. 2011). However, the accurancy and reliability of visual assessment using the HB scale has been questioned (Nutter and Esker 2006; Bock et al. 2009b, 2010a). Even without using a category scale, visual assessment of disease severity can be inaccurate and unreliable (Price et al. 1993; Nita et al. 2003; Nutter et al. 1993; Parker et al. 1995; Moya et al. 2005; Bade and Carmona 2011; Bock et al. 2013).
Various image analysis systems have been developed and applied to measure disease severity on foliage (Bock et al. 2010b). They can provide a more accurate estimation of the disease severity than visual assessment (Price et al. 1993; Moya et al. 2005; Bock et al. 2010b; Bade and Carmona 2011). The software Assess (Lamari 2008) is a popular image analysis software program that has been widely used. The measurements of diseased areas obtained using Assess can be accurate and reliable, and image analysis is often used to gather “actual” values (Bock et al. 2008, 2009a, 2010b). Assess V2.2, the most recent version of the software, incorporates an automatic threshold feature so that each image can be treated and measured separately. It has been successfully used to measure severity of rice blast (Campos-Soriano et al. 2012) and to study quantitative disease resistance to common bacterial blight in common bean (Xie et al. 2012) . Image analysis has not been used to study the genetics of quantitative resistance in tomato.
Resistance to bacterial spot in tomato is partially dominant or incomplete (Yang et al. 2007). Genetic analyses of resistance to the pathogen has suggested multigene models for field resistance (Yang et al. 2007; Sun et al. 2011; Hutton et al. 2010a, b; Sharma et al. 2011). Although visual estimates of disease severity have been widely used to obtain data for genetic studies of resistance to bacterial spot in tomato, it is critical to have accurate phenotypic data for fine-mapping of these quantitative trait loci (QTL). The objective of these experiments was to assess severity of bacterial spot on tomato leaves using image analysis, compare the measurements with visual estimates using the HB category scale, and compare the methods for providing accurate phenotypic data for QTL mapping.
Materials and methods
Plant material and experimental design
Three experiments were conducted to evaluate the feasibility of using image analysis to measure disease severity for genetic analysis of resistance in tomato to bacterial spot. Two tomato lines were used in the first experiment. S. lycopersicum var. cerasiforme accession PI 114490 is an indeterminate cherry tomato with resistance to tomato bacterial spot races T1-T4 (Scott et al. 2003; Hutton et al. 2010b; Sun et al. 2011), while OH 88119 is an elite processing tomato breeding line susceptible to bacterial spot (Yang et al. 2005). In addition to PI 114490 and OH 88119, four additional tomato lines were used in the second experiment to verify the potential of image analysis in distinguishing lines with different levels of resistance. The unimproved breeding line Hawaii 7981 and S. pimpinellifolium accession PI 128216 have partial resistance, and cultivar Fla. 7600 and unimproved breeding line Hawaii 7998 have tolerance to race T3 of tomato bacterial spot (Scott et al. 1995). For the first and second experiments, measurements of disease severity were made on three replicates with 30 plants of each line in each replicate. Since both PI 114490 and OH 88119 were included in the second experiment, only the second experiment was repeated. The third experiment included an F2 population consisting of 439 individuals derived from a cross between OH 88119 and PI 114490 as well as 10 plants of each parent.
Seeds of all lines were germinated in 288 Square Plug Trays (Taizhou Longji Gardening Materials Co., Ltd, Zhejiang, China) in a mixture of peat and vermiculite (3:1). For the first and second experiments, 1-month old seedlings were transplanted into 10 cm (diameter) × 8 cm (height) pots filled with the same peat : vermiculite mixture and placed in a growth chamber (22/30 °C night/day cycle with a 14 h photoperiod). Water and fertilizer were provided as needed. The third experiment was conducted in a protected field at the Shangzhuang Research Station of the China Agricultural University (Beijing, China) following conventional field management practices (Zhang 2005). Seedlings were transplanted 45 days after sowing in trays.
Inoculum preparation and inoculation
X. perforans race T3 strain Xv829 was obtained from the University of Florida (Dr. J. B. Jones). The bacteria were grown in Petri plates on yeast, dextrose, and calcium carbonate (YDC) agar medium (Lelliot and Stead 1987) at 28 °C for 48 to 72 h. Bacteria were washed from the agar with sterile double-distilled water (ddH2O) and the suspension was adjusted to approximately 3 × 108 colony forming units (CFU) per ml confirmed using a spectrophotometer (A600 = 0.15). Plants were spray-inoculated with the bacterial suspension using a handheld sprayer approximately 2 weeks after transplanting for the first two experiments or 1 week before transplanting in the case of the third experiment. The plants were misted with water twice a day (9:00 am and 5:00 pm) from 1 day before inoculation to 21 days after inoculation to increase humidity and prolong leaf wetness.
Image acquisition and lesion area analysis
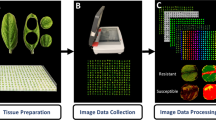
Twenty-one days after inoculation, five diseased leaves were randomly collected from each plant and wiped with a moist cloth to remove dust. The five leaves were scanned as a single TIFF image using a BenQ 5560 Color Scanner (BenQ, Shanghai, China) at 600 × 600 dpi.
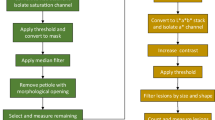
The disease severity (percentage of leaf area covered by lesions) was measured using the image analysis software Assess V2.2 using hue, saturation and intensity colour space and the median filter in the Manual Panel. To determine a suitable approach to measure the severity for the five leaves in one image, two measurements were taken in the first experiment. First, the overall percentage area diseased for all five leaves in the single image was measured. Second, the diseased area was measured separately for each leaf and the mean severity was calculated. The association between the data obtained using these two measurements was analyzed by correlation (SAS V9.4, SAS Institute, Cary, NC, USA). Based on the results, the disease severity in the second and third experiments was measured using the most accurate approach. A general linear model (GLM) was used to analyze disease severity data for the first two experiments. Mean separations were based on least significant difference (LSD) (P = 0.05).
The resistance to tomato bacterial spot race T3 in PI 114490 has been investigated using the HB category scale in an inbred backcross population (Sun et al. 2011). To determine whether the disease severity data obtained by image analysis was comparable to the HB scale for quantitative trait loci (QTL) mapping, all leaf samples in the third experiment were also visually assessed using the HB scale by a trained plant pathologist. The raw HB scale of each leaf was converted to the HB midpoint value (Bock et al. 2010a), and the HB midpoint values of the five leaves for each individual in the F2 population were averaged for further analysis. The relationships between the image analysis measurements and the raw HB scale estimates or the HB midpoint converted values were determined using Microsoft Excel 2007 (Microsoft Corporation, Redmond, WA, USA). The agreement in the HB midpoint values and the image analysis measurements was determined with Lin’s concordance correlation coefficient (ρ c ) (Lin 1989; Nita et al. 2003).
Determining the bacterial population
For the first two experiments, the bacterial population in inoculated tissue of the tomato lines (OH88119 and PI114490 in experiment 1, and OH88119, Ha7998, Fla7600, PI114490, PI128216 and Ha7981 in experiment 2) was determined by a dilution plate method (Yang et al. 2005). Approximately 0.1 g of leaf tissue from each plant was harvested, surface-sterilized with 70 % ethanol, homogenized in 1 ml ddH2O, and subjected to 10-fold serial dilutions. One hundred μl of each serial dilution was plated onto YDC medium, and incubated at 28 °C for 2–3 days prior to counting colonies. Bacterial population sizes were log-transformed to stabilize sample variances prior to statistical analysis. GLM was used to analyze the data of log-transformed colony forming units (log10CFU) per gram of tissue. Mean separations were based on least significant difference (LSD) (P = 0.05).
Genetic analysis of resistance in tomato line PI 114490
Genomic DNA was isolated from young leaves of each plant of the two parents and each individual in the F2 population of the OH 88119 × PI 114490 cross using the modified CTAB method (Kabelka et al. 2002). We identified four QTLs for resistance to race T3 of bacterial spot using an inbred backcross population derived from PI 114490 in a previous study (Sun et al. 2011). However, phenotypic variation explained by each QTL (6.5–11.7 %) was low. To detect whether there were additional QTLs contributing resistance to race T3 in PI 114490, a total of 44 previously defined simple sequence repeat (SSR), insertion/deletion (InDel) polymorphism, single nucleotide polymorphism (SNP) and conserved ortholog set (COSI) markers (Frary et al. 2005; Van Deynze et al. 2007; Sim et al. 2009; Wang et al. 2010; Sun et al. 2011) from 12 chromosomes were used in this study (Table 1), of which 20 were novel for characterizing genetics of resistance to race T3 in PI 114490. The InDel marker ‘DHS12’ (forward primer: 5′-TTATGTGGACGGAAATTCTGC-3′, reverse primer: 5′-CACAAATCGGACATCAGTCG-3′) was newly identified by comparing the genomic DNA sequences between OH 88119 and PI 114490 obtained in our lab.
Genotypic data was obtained by PCR amplification of the genomic DNA of each individual plant using the 44 markers listed in Table 1. PCRs were conducted in a 10-μl reaction volume containing 10 mM Tris–HCl (pH 9.0 at room temperature), 50 mM KCl, 1.5 mM MgCl2, 100 μM each of dNTPs, 0.1 μM each primer, 10 ng of genomic DNA template, and 1 unit of Taq DNA polymerase. Initial denaturing was at 94 °C for 3 min followed by 36 cycles of 30 s at 94 °C, 30 s at 50 °C, and 30 s at 72 °C, and a final extension stage at 72 °C for 5 min. Amplification was performed in a programmable thermal controller (PTC-100; MJ Research, Inc., Watertown, MA). Following the amplification reactions, the PCR products were separated on 7 % polyacrylamide gel. The gel was stained with silver staining approach following the protocol described in SILVER SEQUENCETM DNA Sequencing System (Promega, USA). Genotypic data from molecular markers were scored as homozygous for PI 114490 (band from PI 114490), homozygous for OH 88119 (band from OH 88119), or as heterozygous (bands from both PI 114490 and OH 88119).
Association between disease severity and marker genotype in the F2 population of OH 88119 × PI 114490 was determined by a Kruskal-Wallis test (Kruskal and Wallis 1952) (PROC NPAR1WAY in SAS). The non-parametric analysis was chosen due to non-normality of the data. The significance of association between a marker locus and resistance was indicated by an H-value > 5.99 (degree of freedom = 2) at P ≤ 0.05.
Results
Symptoms of bacterial spot on leaves of tomato lines PI 114490 and OH 88119
In the first experiment, the development of lesions was monitored each day for 1 month after inoculation. The occurrence of lesions on leaves of both PI 114490 and OH 88119 was observed 4 days after inoculation (DAI). The lesions on leaves of PI 114490 plants were small and did not expand (Fig. 1a), and the plants stayed healthy although blighting of the leaf edges occasionally occurred by 24 DAI (Fig. 1b). In contrast, on leaves of OH 88119 plants, the number of lesions increased, they expanded, and coalesced (Fig. 1c). Some diseased leaves wilted and defoliation was observed 24 DAI (Fig. 1d). Therefore, a sampling time of 21 DAI was chosen for all three experiments.
Foliar lesions and symptoms on tomato plants after inoculation with bacterial spot race T3 (Xanthomonas perforans) strain Xv829. a Lesions on a leaf of the resistant line PI 114490 at 21 days after inoculation. b Symptoms on a plant of the resistant line PI 114490 at 24 days after inoculation. c Lesions on a leaf of the susceptible line OH 88119 at 21 days after inoculation. d Symptoms on a plant of the susceptible line OH 88119 at 24 days after inoculation
Comparison of diseased area from group or leaf-by-leaf measurements
The percentage diseased area obtained using image analysis to measure severity on a group of 5 leaves at the same time was correlated to the mean obtained using the leaf-by-leaf measurements (r = 0.94, P < 0.0001). When the colours of the five leaves in the image were similar, the percentage diseased areas measured using the two approaches were almost the same. However, when the colours of the five leaves in the image varied, the percentage diseased area measured by the overall approach was either higher or lower than that on the basis of the leaf-by-leaf approach. Leaves from approximately 12.8 % of the plants in the first experiment showed discrepancy between the two approaches. Due to the variation in leaf colour, only the leaf-by-leaf approach was used for measuring the disease severity in the second and third experiments.
Comparisons of diseased areas and bacterial populations between tomato lines PI 114490 and OH 88119
In the first experiment, the severity on leaves was significantly higher (F = 88.11, P < 0.0001) on OH 88119 plants compared to the PI 114490 plants. The mean disease severity for all sampled leaves of PI 114490 was 1.5 % with a range of 0.8 to 2.2 %, while the mean disease severity for OH 88119 was 32.7 % with a range of 6.5 to 69.6 %. There was no significant difference (F = 2.84, P = 0.1) for log10CFU between OH 88119 (7.84) and PI 114490 (7.46).
Comparisons of diseased areas and bacterial populations among six tomato lines
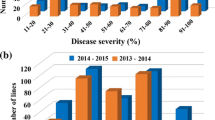
There was a range in the number of lesions on leaves of plants of different lines, with the most on OH 88119 and fewest on PI 114490. The number of lesions on leaves of PI 128216 and Hawaii 7981 was numerically greater than that on leaves of PI 114490, but lower when compared to Hawaii 7998 and Fla. 7600 (Fig. 2).
The disease severities were significantly different among the six tomato lines (Table 2). The disease severity was the least on leaves of PI 114490, but not significantly different from that on leaves of PI 128216 and Hawaii 7981. The susceptible line OH 88119 had the most severe disease. The disease severity on leaves of Hawaii 7998 and Fla. 7600 was less than that on leaves of OH 88119 but greater compared to that on leaves of the other lines. The bacterial populations were not significantly different between lines PI 114490, Hawaii 7998, Fla. 7600 and OH 88119. However, the bacterial populations were significantly lower in leaves of PI 128216 and Hawaii 7981 compared to other lines, and Hawaii 7981 had the lowest population of all lines (Table 2).
Genetics of resistance to bacterial spot race T3 in PI 114490
The mean disease severity measured by image analysis on leaves was 2.3 % on PI 114490 and 63.2 % on OH 88119. In the F2 population of the OH 88119 × PI 114490 cross, the severity of bacterial spot for all sampled leaves ranged from 0.6 to 92.2 %, and the mean severity for individual plants (a mean of estimates from five leaves) ranged from 1.2 to 59.4 % with a mean of 12.0 %. The distribution of the severities in the F2 population was skewed towards low disease severity (Fig. 3a). The HB category scale estimates for all individuals ranged from 2 to 7, and had a normal distribution (Fig. 3b). When the HB category scale was converted to the HB midpoint values, the resulting distribution had a bias towards low disease severity (Fig. 3c). Visual rating (HB category scale estimates) had a positive, logarithmic association with image analysis measurement of disease severity based on mean estimates from five leaves (Fig. 4). More severe disease was measured by image analysis, but was not estimated as severe using the HB scale, such that the HB estimates reached a plateau when disease severity was ~20 %. This indicated that visual estimates using the HB category scale did not differentiate well more severe disease (where the HB scale has wide intervals). A logarithmic regression function provided the best fit to these data (coefficient of determination, R 2 = 0.87). A linear relationship was observed when comparing the HB midpoint converted values with the disease severity obtained using image analysis based on mean estimates from five leaves (Fig. 5). The correlation coefficient (r = 0.91) and concordance (representing perfect agreement between actual and estimated severity) correlation coefficient (ρ c = 0.91) were high. However, there was a slight underestimation of disease severity because the u value (constant bias, height differences between the concordance and the best fitting lines) was less than 0. The υ value (systematic bias, the difference in disease severity between the HB midpoint and image analysis measurements with increasing actual disease) was greater than 1, suggesting that the slope for the estimated severity was slightly greater than the concordance line at low disease severity (Fig. 5).
Distributions of disease severities (percentage diseased area) obtained by image analysis measurements (a), disease severity obtained by visual estimates using the Horsfall and Barratt (1945) category scale (b), and the midpoint converted from the HB scale (c) in the F2 population of OH 88119 × PI 114490 cross 21 days after inoculation of bacterial spot race T3 (Xanthomonas perforans) strain Xv829. Both the actual disease severity and HB values are averages from five leaf samples per individual plant
The relationship between the actual disease severity (percentage diseased area measured using image analysis) and the visual estimates using the 12-point Horsfall and Barratt (1945) category scale in the F2 population of the cross OH 88119 × PI 114490. Both the actual disease severity and HB values are averages from five leaf samples per individual plant
The relationship between the actual disease severity (percentage diseased area measured using image analysis) and the visual estimates using midpoint (%) converted from the 12-point Horsfall and Barratt (1945) category scale in the F2 population of the cross OH 88119 × PI 114490. Both the actual disease severity and HB midpoint values are averages from five leaf samples per individual. The solid line is the concordance line, and the dash line is the best fitting line. Accuracy is determined with Lin’s (1989) concordance correlation coefficient (ρ c ), calculated as the product of the correlation coefficient (r) and the bias correction factor (Cb). Cb = 2/(υ + 1/υ + u 2), where u is height differences between the concordance and the best fitting lines and υ is the difference between measured and actual values (Nita et al. 2003)
The Kruskal and Wallis (1952) test was used to determine marker-trait association due to the skewed distribution of the disease severity data in the F2 population. Among the 439 individuals in the F2 population of the OH 88119 × PI 114490 cross, seven (1.59 %) had the same resistance as the parent PI 114490 based on the disease severities obtained using image analysis measurement, while six (1.37 %) were similar to PI 114490 based on the HB category scale or the HB midpoint converted values. The proportion of resistant plants in the F2 population was very close to the expected value of 1.56 % assuming a three-gene model requiring all six alleles for maximum resistance. Based on marker-trait association analysis using the diseases severity data obtained through image analysis measurement, six markers (SSR134, SL10480i, LEOH185, SL10690i, SL10737i, and SSR20) on four chromosomal regions were significantly (H > 5.99, degree of freedom =2, P ≤ 0.05) associated with disease resistance, suggesting that four QTLs confer resistance to race T3 in PI 114490 (Table 1). Locations of markers on chromosomes 1, 3, and 11 were consistent with previous QTL positions (Sun et al. 2011). Marker SSR20 was linked to a novel QTL on chromosome 12. In addition to the above six markers, the marker SSR601 on the same region of chromosome 3 was also associated with resistance when the HB category scale data and the HB midpoint converted values were used for marker-trait analysis.
Discussion
The leaf-by-leaf approach measures disease severity using image analysis software based on a single leaf. The approach has advantages in simplicity but there is a need to consider leaf colour variation in the image and possibly adjust the parameters to maintain accuracy between leaves. It has been widely used in many plant disease severity assessments (Bock et al. 2010b). Due to the need to recalibrate for each leaf, it can take time to acquire and analyze images for a large number of leaf samples in a segregating population. In this study, we tested measuring disease on five leaves from the same individual plant scanned in a single image. The disease severity was measured once for all five leaves and separately for each leaf. This allowed us to determine the most suitable approach to measure disease severity for large number of samples. The results showed close correlation between measurement of five-leaf samples at once and leaf-by-leaf measurements, suggesting that the grouped leaf measurements could be used to assess disease severity for all leaves from one plant at once, despite colour variation among the sampled leaves. Minimizing colour variation among leaves in one image or a leaf-by-leaf approach would increase the accuracy of measurement.
Bacterial populations in plants have been used as an index of disease severity for QTL analysis of resistance to race T1 of bacterial spot in tomato line Hawaii 7998 (Yang et al. 2005) because the number of bacteria in diseased tomato leaves is highly correlated with field severity ratings (Somodi et al. 1991). However, the numbers of bacteria in diseased leaves of tomato lines Hawaii 7981 and PI 128216 were consistent with their field resistance, while the bacterial population in the resistance tomato line PI 114490 was not significantly different from that of the susceptible line OH 88119 in this study. As is established, the hypersensitive response (HR) is a mechanism used by plants to limit the growth of microbial pathogen. Both PI 128216 and Hawaii 7981 show HR to race T3 while PI 114490 does not (Scott et al. 1995; Wang et al. 2011; Pei et al. 2012), which might explain the difference in bacterial population in vivo. Therefore, genotypic differences should be considered when using the bacterial population as an index of disease severity.
The relationship between actual disease (0 to 100 %), visually estimated disease (0 to 100 %) and that estimated by the HB category scale could be misunderstood. The HB category scale is one of the most widely used quantitative scales for the visual assessment of plant disease. One assumption for developing the HB scale was that the human eye reads in logarithms, requiring a scale that has logarithmic categories over the 0 to 100 % disease severity ratio scale (Horsfall and Barratt 1945). Comparing either actual disease or visually estimated disease with disease estimated using the HB scale results in a logarithmic relationship unless a midpoint conversion is made prior to taking averages (Bock et al. 2010a). Thus a logarithmic relationship was observed between the HB category scale and percentage disease severity obtained by image analysis in this study. However, it has been demonstrated that the psychophysical basis for the HB scale is flawed, and there is a linear increase in visual stimulus with increased disease severity (Nita et al. 2003; Nutter and Esker 2006; Bock et al. 2010b). Linear relationships between visual estimates using percent disease severity and the actual disease severity obtained by image analysis has been confirmed in a number of studies (Price et al. 1993; Nita et al. 2003; Nutter and Esker 2006; Bock et al. 2008, 2010a). The relationship between the disease severity estimated using a linear scale with equal 5 % intervals and the actual disease is also linear (Nita et al. 2003). These suggest that direct estimation of disease severity is more reliable and accurate than the HB scale (Bock et al. 2010b). The relationship between the visually estimated HB scale disease severity and that estimated or measured as actual disease severity using the 0–100 % ratio scale should be carefully interpreted.
It has been suggested that the HB category scale is less useful for hypothesis testing than nearest percent estimates (Bock et al. 2010a). Inaccuracy of visual assessment using the HB category scale can lead to a greater risk of type II errors when the population size sampled is inadequate (Bock et al. 2009b, 2010a; Nita et al. 2003; Nutter and Esker 2006). However, visual assessment using category scales like the HB scale are useful tools for selection in breeding programs, especially for screening large breeding populations, and some previous research suggests certain category scales have little or no effect on QTL mapping (Poland and Nelson 2011). Our previous work suggests that the resistance in PI 114490 to bacterial spot race T3 is conditioned by at least four QTLs having small effects (6.5–11.7 %). To establish an approach for fine-mapping of these QTLs, the disease severities were measured using image analysis and the data used for QTL detection in this study. Although using data from both the actual disease severity measured by image analysis and the raw HB category scale resulted in indentifying the same QTLs in the F2 population of the OH 88119 × PI 114490 cross, the phenotypic variation explained by each marker was higher using the raw HB category data compared to using the image analysis-measured disease severity data. This suggested that the disease severity obtained using the HB category scale might result in overestimation of the actual effect of QTL. A similar phenomenon was observed in a comparative analysis of quantitative resistance to common bacterial blight in common bean using an ordinal scale and image analysis (Xie et al. 2012). The overestimation of phenotypic variation might be due to the lack of additivity when disease severity was visually assessed using a category scale (Xie et al. 2012). In addition, the use of both the raw HB scale data and the HB midpoint converted values identified one additional marker SSR601 associated with resistance in this study. This might be due to existence of missing genotypic data for approximately 48.0 % of individuals in the F2 population. The individuals with missing genotypic data were automatically removed from statistical analysis in the SAS program, which might cause false association or lack of ability to detect QTL in different data. Poland and Nelson (2011) also reported that additional QTLs for resistance to northern leaf blight in maize could be detected using a 0–9 category rating scale (11 QTLs) compared to using percentage estimates on the 0–100 % scale (9 QTLs). They proposed that the detection of smaller-effect QTL might be sensitive to the type of scale and the resulting distribution. However, care should be taken when estimating QTL allele effects based on visual disease assessments because the magnitudes of estimated allele effects by different raters for identified QTLs varied drastically (Poland and Nelson 2011). Conversion of the raw HB scale data to midpoint values changed the relationship between the disease severity obtained using image analysis and visual estimates, but did not increase the precision and accuracy in this study. All these suggest that image analysis is a more reliable and suitable method than visual estimates using a category scale for phenotypic evaluations in quantitative genetic studies.
Visual assessment based on the HB scale has been widely applied to investigate genetics of resistance to bacterial spot in tomato due to its simplicity and flexibility (Yang 2013). Two strategies were adapted to increase the accuracy and precision of visual estimates using the HB scale for QTL identification in previous studies (Scott et al. 2003; Yang et al. 2005; Hutton et al. 2010a, b; Sun et al. 2011). First, disease severity assessments were conducted on replicated field trials with specific populations (e.g. advanced backcross population or inbred backcross population) derived from resistant lines. This allowed estimating the environmental effect on disease severity and identifying the real QTLs. Second, the raw HB scale estimates were made based on the average disease severity of at least 10 plants of the same genotype in a plot to minimize any potential artifacts. Experienced raters are needed to obtain a relatively accurate and precise estimate using the HB scale, as with the 0 to 100 % scale. Image analysis is a useful approach for fine-mapping of QTLs for quantitative disease resistance due to its sensitivity and the truly quantitative nature of the measurements that lend themselves to direct parametric analysis (unlike HB measurements that must be mid-point converted prior to analysis).
In conclusion, image analysis is an accurate method to distinguish tomato lines with different susceptibility to bacterial spot, and to identify QTLs for resistance in PI 114490. These results suggest that image analysis can be used for measuring disease for genetic analysis of quantitative resistance to plant disease and fine-mapping of QTLs in tomato. Although the same QTLs can be identified using both the HB scale data and the actual disease severity measured by image analysis, underestimation of disease severity and overestimation of phenotypic variation using the HB scale data suggest that the 0–100 % ratio scale is better than the HB category scale for disease severity assessment and QTL identification.
References
Bade, C. I. A., & Carmona, M. A. (2011). Comparison of methods to assess severity of common rust caused by Puccinia sorghi in maize. Tropical Plant Pathology, 36(4), 264–266.
Bock, C. H., Parker, P. E., Cook, A. Z., & Gottwald, T. R. (2008). Visual rating and the use of image analysis for assessing different symptoms of citrus canker on grapefruit leaves. Plant Disease, 92, 530–541.
Bock, C. H., Cook, A. Z., Parker, P. E., & Gottwald, T. R. (2009a). Automated image analysis of the severity of foliar citrus canker symptoms. Plant Disease, 93, 660–665.
Bock, C. H., Gottwal, T. R., Parker, P. E., Cook, A. Z., Ferrandino, F., Parnel, S., & van den Bosch, F. (2009b). The Horsfall-Barratt scale and severity estimates of citrus canker. European Journal of Plant Pathology, 125, 23–38.
Bock, C. H., Gottwald, T. R., Parker, P. E., Ferrandino, F., Welham, S., van den Bosch, F., & Parnell, S. (2010a). Some consequences of using the Horsfall-Barratt scale for hypothesis testing. Phytopathology, 100, 1031–1041.
Bock, C. H., Poole, G. H., Parker, P. E., & Gottwald, T. R. (2010b). Plant disease severity estimated visually, by digital photography and image analysis, and by hyperspectral imaging. Critical Reviews in Plant Sciences, 29, 59–107.
Bock, C. H., Wood, B. W., van den Bosch, F., Parnell, S., & Gottwald, T. R. (2013). The effect of Horsfall-Barratt category size on the accuracy and reliability of estimates of pecan scab severity. Plant Disease, 97, 797–806.
Campos-Soriano, L., García-Martínez, J., & San Segundo, B. (2012). The arbuscular mycorrhizal symbiosis promotes the systemic induction of regulatory defence-related genes in rice leaves and confers resistance to pathogen infection. Molecular Plant Pathology, 13, 579–592.
Frary, A., Xu, Y., Liu, J., Mitchell, S., Tedeschi, E., & Tanksley, S. D. (2005). Development of a set of PCR-based anchor markers encompassing the tomato genome and evaluation of their usefulness for genetics and breeding experiments. Theoretical and Applied Genetics, 111, 291–312.
Guo, S. C., Zhuang, J. R., Li, Q., & Dong, Q. C. (2008). Investigation and integrative management of bacterial spot in Fei County. China Vegetables, 5, 60–61.
Horsfall, J. G., & Barratt, R. W. (1945). An improved grading system for measuring plant diseases. Phytopathology, 35, 655.
Hutton, S. F., Scott, J. W., & Jones, J. B. (2010a). Inheritance of resistance to bacterial spot race T4 from three tomato breeding lines with differing resistance backgrounds. Journal of the American Society for Horticultural Science, 135, 150–158.
Hutton, S. F., Scott, J. W., Yang, W. C., Sim, S. C., Francis, D. M., & Jones, J. B. (2010b). Identification of QTL associated with resistance to bacterial spot race T4 in tomato. Theoretical and Applied Genetics, 121, 1275–1287.
Jones, J. B., Jones, J. P., Stall, R. E., & Zitter, T. A. (Eds.). (1991). Compendium of tomato diseases. Minnesota, USA: APS Press.
Kabelka, E., Franchino, B., & Francis, D. M. (2002). Two loci from Lycopersicon hirsutum confer resistance to strains of Clavibacter michiganensis subsp. michiganensis. Phytopathology, 92, 504–510.
Kruskal, W. H., & Wallis, W. A. (1952). Use of ranks in one-criterion variance analysis. Journal of the American Statistical Association, 47, 583–621.
Lamari, L. (2008). ASSESS2.0: Image analysis software for plant disease quantification. Minnesota, USA: APS Press.
Lelliot, R. A., & Stead, D. E. (1987). Methods for the diagnosis of bacterial diseases of plants. Oxford: Blackwell Scientific Publication Ltd.
Lin, L. I. K. (1989). A concordance correlation coefficient to evaluate reproducibility. Biometrics, 45, 255–268.
Liu, Y. Y., Cui, Y. Q., & Bao, L. L. (2008). New features of occurrence and approaches for no pollution management for tomato disease in greenhouse. Jinlin Vegetables, 2, 47.
Moya, E. A., Barrales, L. R., & Apablaza, G. E. (2005). Assessment of the disease severity of squash powdery mildew through visual analysis, digital image analysis and validation of these methodologies. Crop Protection, 24, 785–789.
Nita, M., Ellis, M. A., & Madden, L. V. (2003). Reliability and accuracy of visual estimation of Phomopsis leaf blight of strawberry. Phytopathology, 93, 995–1005.
Nutter, F. W., Jr., & Esker, P. D. (2006). The role of psychophysics in phytopathology: the Weber-Fechner law revisited. European Journal of Plant Pathology, 114, 199–213.
Nutter, F. W., Jr., Gleason, M. L., Jenco, J. H., & Christians, N. C. (1993). Assessing the accuracy, intra-rater repeatability, and inter-rater reliability of disease assessment systems. Phytopathology, 83, 806–812.
Parker, S. R., Whelan, M. J., & Royle, D. J. (1995). Reliable measurement of disease severity. Aspects of Applied Biology, 43, 205–214.
Pei, C. C., Wang, H., Zhang, J. Y., Wang, Y. Y., Francis, D. M., & Yang, W. C. (2012). Fine mapping and analysis of a candidate gene in tomato accession PI128216 conferring hypersensitive resistance to bacterial spot race T3. Theoretical and Applied Genetics, 124, 533–542.
Poland, J. A., & Nelson, R. J. (2011). In the eye of the beholder: the effect of rater variability and different rating scales on QTL mapping. Phytopathology, 101, 290–298.
Price, T. V., Gross, R., Ho Wey, J., & Osborne, C. F. (1993). A comparison of visual and digital image-processing methods in quantifying the severity of coffee leaf rust (Hemileia vastatrix). Australian Journal of Experimental Agriculture, 33, 97–101.
Scott, J. W., & Jones, J. B. (1989). Inheritance of resistance to foliar bacterial spot of tomato incited by Xanthomonas campestris pv. vesicatoria. Journal of the American Society for Horticultural Science, 114, 111–114.
Scott, J. W., Jones, J. B., & Somodi, G. C. (1995). Screening tomato accessions for resistance to Xanthomonas campestris pv. vesicatoria, race T3. HortScience, 30, 579–581.
Scott, J. W., Francis, D. M., Miller, S. A., Somodi, G. C., & Jones, J. B. (2003). Tomato bacterial spot resistance derived from PI 114490; inheritance of resistance to race T2 and relationship across three pathogen races. Journal of the American Society for Horticultural Science, 128, 698–703.
Sharma, D., Gupta, S. K., & Kumud, J. (2011). Studies on the inheritance pattern of bacterial spot (Xanthomonas vesicatoria) in tomato. Indian Phytopathology, 64(2), 128–130.
Sim, S. C., Robbins, M. D., Chilcott, C., Zhu, T., & Francis, D. M. (2009). Oligonucleotide array discovery of polymorphisms in cultivated tomato (Solanum lycopersicum L.) reveals patterns of SNP variation associated with breeding. BMC Genomics, 10, 466.
Somodi, G. C., Jones, J. B., & Scott, J. W. (1991). Populations of Xanthomonas campestris pv. vesicatoria in lesions of susceptible and resistant tomato genotypes. Plant Disease, 75, 357–360.
Somodi, G. C., Jones, J. B., Scott, J. W., & Jones, J. P. (1994). Screening tomato seedlings for resistance to bacterial spot. HortScience, 29, 680–682.
Sotirova, V., & Bogatsevska, N. (2000). Evaluation of tomato wild species for resistance to race T1 nad T3 of Xanthomonas vesicatoria. Acta Physiologiae Plantarum, 22, 256–259.
Stall, R. E., Jones, J. B., & Minsavage, G. V. (2009). Durability of resistance in tomato and pepper to Xanthomonads causing bacterial spot. Annual Review of Phytopathology, 47, 265–284.
Sun, H. J., Zhang, J. Y., Wang, Y. Y., Scott, J. W., Francis, D. M., & Yang, W. C. (2011). QTL analysis of resistance to bacterial spot race T3 in tomato. Acta Horticulturae Sinica, 38, 2297–2308.
van Deynze, A., Stoffel, K., Buell, C. R., Kozik, A., Liu, J., van der Knaap, E., & Francis, D. (2007). Diversity in conserved genes in tomato. BMC Genomics, 8, 465.
Wang, Z. X., Wang, Z. Q., & Wang, G. F. (2005). Integrative technologies for controlling bacterial spot in tomato. Northwest Horticulture, 7, 30.
Wang, Y. Y., Chen, J., Francis, D. M., Shen, H. L., Wu, T. T., & Yang, W. C. (2010). Discovery of intron polymorphisms in cultivated tomato using both tomato and Arabidopsis genomic information. Theoretical and Applied Genetics, 121, 1199–1207.
Wang, H., Hutton, S. F., Robbins, M. D., Sim, S. C., Scott, J. W., Yang, W. C., Jones, J. B., & Francis, D. M. (2011). Molecular mapping of hypersensitive resistance from tomato cv. Hawaii 7981 to Xanthomonas perforans race T3. Phytopathology, 101, 1217–1223.
Xie, W., Yu, K., Pauls, K. P., & Navabi, A. (2012). Application of image analysis in studies of quantitative disease resistance, exemplified using common bacterial blight-common bean pathosystem. Phytopathology, 102, 434–442.
Yang, W. (2013). Recent advances on genetics and mapping of resistance to bacterial spot in tomato. Acta Horticulturae Sinica, 40, 1731–1740.
Yang, W., Sacks, E. J., Ivey, M. L., Miller, S. A., & Francis, D. M. (2005). Resistance in Lycopersicon esculentum intraspeciflc crosses to race T1 strains of Xanthomonas campestris pv. vesicatoria causing bacterial spot of tomato. Phytopathology, 95, 519–527.
Yang, W. C., Chen, J., Zhang, X. M., & Francis, D. M. (2007). Recent advances in classification of tomato bacterial spot pathogen, genetics of resistance, and marker-assisted selection. Scientia Agricultura Sinica, 40, 283–290.
Zhang, Z. X. (Ed.). (2005). Vegetable cultivation. Beijing, China: China Agricultural University Press.
Zhang, X. M., Francis, D. M., & Yang, W. C. (2009). Evaluation of resistance to bacterial spot in varieties growing in China and marker-assisted selection. Acta Agriculturae Boreali-Sinica, 24(4), 183–187.
Zhang, W. L., Wang, G. Y., Li, Q. S., & Yang, S. B. (2010). Recognization and prevention of tomato bactrerial diseases in greenhouse. China Agricltural Technology Extension, 26(2), 40–41.
Acknowledgments
The authors would like to thank Dr. David M. Francis in the Department of Horticulture and Crop Science at the Ohio State University for providing the seeds of tomato lines and Dr. Jeffrey B. Jones in the Department of Plant Pathology at the University of Florida for providing race T3 of the bacterial spot pathogen. The authors also thank Dr. C.H. Bock for his critical comments and suggestions on manuscript revision. The work was partly supported by the National Natural Science Foundation of China (30972003), the Specialized Research Fund for the Doctoral Program of Higher Education (20110008110019) and the Beijing Modern Agro-industry Technology Research System.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Sun, H., Wei, J., Zhang, J. et al. A comparison of disease severity measurements using image analysis and visual estimates using a category scale for genetic analysis of resistance to bacterial spot in tomato. Eur J Plant Pathol 139, 125–136 (2014). https://doi.org/10.1007/s10658-013-0371-8
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10658-013-0371-8