Abstract
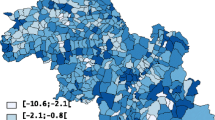
Boundary analysis of cancer maps may highlight areas where causative exposures change through geographic space, the presence of local populations with distinct cancer incidences, or the impact of different cancer control methods. Too often, such analysis ignores the spatial pattern of incidence or mortality rates and overlooks the fact that rates computed from sparsely populated geographic entities can be very unreliable. This paper proposes a new methodology that accounts for the uncertainty and spatial correlation of rate data in the detection of significant edges between adjacent entities or polygons. Poisson kriging is first used to estimate the risk value and the associated standard error within each polygon, accounting for the population size and the risk semivariogram computed from raw rates. The boundary statistic is then defined as half the absolute difference between kriged risks. Its reference distribution, under the null hypothesis of no boundary, is derived through the generation of multiple realizations of the spatial distribution of cancer risk values. This paper presents three types of neutral models generated using methods of increasing complexity: the common random shuffle of estimated risk values, a spatial re-ordering of these risks, or p-field simulation that accounts for the population size within each polygon. The approach is illustrated using age-adjusted pancreatic cancer mortality rates for white females in 295 US counties of the Northeast (1970–1994). Simulation studies demonstrate that Poisson kriging yields more accurate estimates of the cancer risk and how its value changes between polygons (i.e., boundary statistic), relatively to the use of raw rates or local empirical Bayes smoother. When used in conjunction with spatial neutral models generated by p-field simulation, the boundary analysis based on Poisson kriging estimates minimizes the proportion of type I errors (i.e., edges wrongly declared significant) while the frequency of these errors is predicted well by the p-value of the statistical test.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Best N, Richardson S and Thomson A (2005). A comparison of Bayesian spatial models for disease mapping. Stat Methods Med Res 14: 35–59
Clayton DG and Kaldor J (1987). Empirical Bayes estimates of age-standardized relative risks for use in disease mapping. Biometrics 43: 671–681
Goovaerts P (2001). Geostatistical modelling of uncertainty in soil science. Geoderma 103: 3–26
Goovaerts P (2005). Geostatistical analysis of disease data: estimation of cancer mortality risk from empirical frequencies using Poisson kriging. Int J Health Geogr 4: 31
Goovaerts P (2006). Geostatistical analysis of disease data: visualization and propagation of spatial uncertainty in cancer mortality risk using Poisson kriging and p-field simulation. Int J Health Geogr 5: 7
Goovaerts P and Jacquez GM (2004). Accounting for regional background and population size in the detection of spatial clusters and outliers using geostatistical filtering and spatial neutral models: the case of lung cancer in Long Island, New York. Int J Health Geogr 3: 14
Jacquez G (1998). GIS as an enabling technology. In: Gatrell, A and Loytonen, M (eds) GIS and health, pp 17–28. Taylor and Francis, London
Jacquez GM and Greiling DA (2003). Geographic boundaries in breast, lung and colorectal cancers in relation to exposure to air toxics in Long Island, New York. Int J Health Geogr 2: 4
Lu H and Carlin BP (2005). Bayesian areal wombling for geographical boundary analysis. Geogr Anal 37: 265–285
Monestiez P, Dubroca L, Bonnin E, Durbec JP and Guinet C (2006). Geostatistical modelling of spatial distribution of Balenoptera physalus in the Northwestern Mediterranean Sea from sparse count data and heterogeneous observation efforts. Ecol Model 193: 615–628
Mungiole M, Pickle LW and Hansen Simonson K (1999). Application of a weighted head-banging algorithm to mortality data maps. Stat Med 18: 3201–3209
Oden NL, Sokal RR, Fortin M-J and Goebl H (1993). Categorical wombling: detecting regions of significant change in spatially located categorical variables. Geogr Anal 25: 315–336
Pardo-Iguzquiza E (1999). VARFIT: a Fortran-77 program for fitting variogram models by weighted least squares. Comput Geosci 25: 251–261
Pickle LW, Mungiole M, Jones GK and White AA (1999). Exploring spatial patterns of mortality: the new Atlas of United States mortality. Stat Med 18: 3211–3220
Rushton G, Elmes G and McMaster R (2000). Considerations for improving geographic information system research in public health. J Urban Regional Inform Syst Assoc 12: 31–49
Swets JA (1988). Measuring the accuracy of diagnostic systems. Science 240: 1285–1293
Waller LA and Gotway CA (2004). Applied spatial statistics for public health data. Wiley, New Jersey
Womble W (1951). Differential systematics. Science 114: 315–322
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Goovaerts, P. Accounting for rate instability and spatial patterns in the boundary analysis of cancer mortality maps. Environ Ecol Stat 15, 421–446 (2008). https://doi.org/10.1007/s10651-007-0064-6
Received:
Revised:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10651-007-0064-6