Abstract
Dissimilatory arsenate reduction from arsenic (As)-bearing minerals into highly mobile arsenite is one of the key mechanisms of As release into groundwater. To detect the microbial diversity and As-metabolizing gene clusters of indigenous arsenate-reducing bacteria in high As groundwater in the Hetao Plain of Inner Mongolia, China, three anaerobic arsenate-reducing bacteria were isolated and arrA and arsC gene-based clone libraries of four in situ groundwater samples were constructed. The strains IMARCUG-11(G-11), IMARCUG-C1(G-C1) and IMARCUG-12(G-12) were phylogenetically belonged to genera Paraclostridium, Citrobacter and Klebsiella, respectively. They could reduce >99% of 1 mM arsenate under anoxic conditions with lactate as a carbon source in 60 h, 72 h and 84 h, respectively. As far as we know, this was the first report of arsenate reduction by genus Paraclostridium. Compared with strain G-11 (arsC) and G-C1 (arsRBC), strain G-12 contained two incomplete ars operons (operon1: arsABC, operon2: arsBC), indicating that these strains might present different strategies to resist As toxicity. Phylogenetic analysis illuminating by the arrA genes showed that in situ arsenate-reducing bacterial communities were diverse and mainly composed of Desulfobacterales (53%, dominated by Geobacter), Betaproteobacteria (12%), and unidentified groups (35%). Based on the arsC gene analysis, the indigenous arsenate-reducing bacterial communities were mainly affiliated with Omnitrophica (88%) and Deltaproteobacteria (11%, dominated by Geobacter and Syntrophobacterales). Results of this study expanded our understanding of indigenous arsenic-reducing bacteria in high As groundwater aquifers.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
As a worldwide environmental issue, exposure to high arsenic (As) groundwater has affected more than 140 million people from countries such as Bangladesh, China, Vietnam and the USA (Barringer et al. 2011; Harvey et al. 2002; Li et al. 2014; Winkel et al. 2011). Groundwater arsenic can originate from geogenic As-laden iron (Fe) oxide minerals (Fendorf et al. 2010) or anthropogenic utilization of As-containing fertilizers as well as mining of As-bearing minerals (Neidhardt et al. 2012). Consumption of As containing groundwater as drinking water is harmful to human health and can cause many serious diseases including vascular and respiratory disease, melanosis, and even skin and heart cancer (Ferreccio and Sancha 2006; Maleki et al. 2015).
Numerous studies have been devoted to investigate the release and transformation mechanisms of As in groundwater aquifers (Fan et al. 2008; Jiang et al. 2014; Winkel et al. 2011) and results have shown that several processes may lead to the mobilization of As into groundwater, including reductive dissolution of iron (Fe) oxide minerals, oxidation of sulfide minerals, anion competition for desorption sites and microbial reduction of arsenate (As (V)). Among these processes, the microbially mediated As (V) reduction is mainly caused by two kinds of microorganisms including dissimilatory As (V)-respiring prokaryotes (DARPs) and As (V)-resistant microbes (ARMs) (Tsai et al. 2009; Liao et al. 2011). The DARPs can use arsenate (As (V)) as terminal electron acceptor and gain energy for metabolic growth under anaerobic conditions. The arrA gene, encoding the respiratory arsenate reductase subunit ArrA, is a credible biomarker for detection of respiratory As-reducing bacteria. The resistance of ARMs is encoded by the ars operon which usually contains the As (V) reductase ArsC and the regulatory protein ArsR (Branco et al. 2008). Different from the DARPs, the ARMs can survive in both aerobic and anaerobic conditions without energy conservation (Macy et al. 2000; Saltikov et al. 2005). To date, numerous phylogenetically diverse prokaryotes capable of As (V) reduction have been isolated from terrestrial and aquatic habitats (Chang et al. 2012; Handley et al. 2009; Niggemyer et al. 2001). ARMs are widespread almost in all bacterial phyla, while DARPs are affiliated mainly in Proteobacteria, Firmicutes, Aquificae, Deferribacteres, Chryosiogenetes and Archaea (Cavalca et al. 2013).
Hetao plain of Inner Mongolia is one of China’s representative high As affected area with the highest As concentration in groundwater up to 1.74 mg/L and more than 250,000 residents under As threat (Deng et al. 2009; Deng et al. 2011). Previous studies have showed that As release in Hetao Plain is the consequence of several biogeochemical processes such as reductive dissolution of As-rich Fe(III) oxyhydroxides and dissimilatory reduction of As (Cavalca et al. 2013; Deng 2008; Deng et al. 2009; Guo et al. 2011). Results of our previous studies have demonstrated that bacterial communities (16S rRNA genes based) in high As groundwater of Hetao Plain are dominated by Acinetobacter, Pseudomonas, Brevundimonas, Geobacter, Thiobacillus and large quantity of unidentified bacterial groups (Li et al. 2013; Wang et al. 2016). However, until now, only several As (V)-resistant strains have been isolated from the Hetao Plain, including two aerobic bacteria affiliated with Pseudomonas sp. and Bacillus sp. and three anaerobic bacteria belonged to Bacillus sp. Desulfitobacterium sp. and Exiguobacterium (Cai et al. 2016; Guo et al. 2015). Besides, studies on the community composition and diversity of indigenous As-reducing bacteria are still limited in the high As aquifers.
Therefore, the main objectives of the present study were to (1) isolate functional bacteria capable of As reduction and detect their arsenic-metabolizing gene clusters and (2) investigate the diversity and composition of indigenous As(V)-reducing bacterial communities via arrA and arsC genes from high As groundwater of the Hetao Plain, Inner Mongolia, China.
Material and methods
Groundwater sampling and geochemical analysis
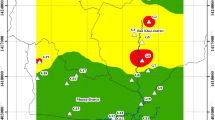
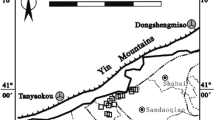
Four representative shallow groundwater samples IMCUGGW05 (N40.908°, E106.829°), IMCUGGW07 (N40.865°, E106.884°), IMCUGGW13 (N41.202°, E107.362°) and IMCUGGW17 (N41.070°, E107.159°) with differentiated geochemical characteristics were collected from Hangjinhouqi County, an As threatened area located in the Hetao Plain of western Inner Mongolia. Groundwater samples were pumped out from local domestic wells (depth: 20-25 m) for about 10 min to get a stabilized Eh values. Then, about 10 L fresh groundwater samples were filtered through 0.2 μm filters (Millipore, Germany). The filter membranes were collected into sterilized tubes and frozen on dry ice for further analysis. For strain isolation, the prepared serum bottles with sterile and deoxygenized chemically defined medium (CDM) were injected with approximately 1 mL groundwater samples and kept at room temperature during transportation.
Field and laboratory geochemical analyses including temperature, pH and oxidation-reduction potential (ORP) of groundwater samples as well as As and Fe speciation were performed with same procedures as in our previous studies (Wang et al. 2015; Li et al. 2017). Details were shown in Supporting Information (SI).
Strain isolation and identification
For enrichment and isolation, CDM was used (Weeger et al. 1999). It contained the following (g/L): NaCl 0.46, MgSO4•7H2O 0.117, KH2PO4 0.225, K2HPO4 0.225 and (NH4)2SO4 0.225. Additional 20 mM lactate, 0.1% yeast extract and 0.1% resazurin solution (1%) were added into the medium as carbon source, nutrient supplement and dissolved oxygen indicator, respectively. The medium was divided into 100 mL serum bottles, deoxygenated with pure N2, and then autoclaved. Arsenate (1 mM), L-cysteine (1 mM) and NaHCO3 (5 mM) were filtered through 0.22 μm filters and then added into the medium. After three times of transfer, diluted suspension was inoculated into anaerobic tubes containing CDM-As and purified agar (1.8%, w/v) following the Hungate technique (Hungate 1969). Single clones were picked out and inoculated into new medium. After several times of isolation, three pure cultures were obtained (IMARCUG-11, IMARCUG-C1 and IMARCUG-12).
Growth experiments of the isolated strains were conducted with 40 mL CDM in 100 mL serum bottles in triplicate. Gradient temperature and pH were set to detect the most suitable growth conditions. Different carbon sources with the same carbon concentrations of 20 mM were added into the medium to test the potential utilization of carbon sources. 1 mM As (V) was added into the pre-cultured stain suspensions to confirm the ability of As (V) reduction. Mediums without strain suspensions were cultured as controls. All experimental sets were performed in triplicate at 30 °C under anaerobic conditions. At different time intervals, 1 mL of the mixture were taken out from serum bottles and equally divided into two aliquots. One aliquot was filtered (0.22 μm) for the detection of As speciation, while the other aliquot was analyzed for protein concentration to monitor the bacterial growth (Lv and Yu 2013). All the experiments taken above were accomplished in the anaerobic glove box.
Genomic DNA of the isolates was extracted using the FastDNA spin kit for soil (MP Bio Laboratories, USA) following the manufacturer’s procedures. The 16 S rRNA gene fragment of the isolates were amplified using universal primer pair 27 F and 1492 R (Table 1). The PCR programs were the same as in our previous studies (Dai et al. 2016). Successful PCR products were purified and then commercially sequenced. The obtained sequences were blasted in NCBI to search sequences with high similarities and construct neighbor-joining tree with MEGA 7.0 software. The 16 S rRNA gene sequences of strain G-11, G-C1 and G-12 were submitted to the GenBank under accession numbers MK027325, MG198846 and MK027326, respectively.
Amplification of functional genes of isolates
Based on conservative sites from several arsC gene-related sequences in the genus Paraclostridium, a primer pair PB2-F and PB2-R (Table 1) was designed to amplify the arsC gene of strain G-11 with the target length of 202 bp. The arsB gene of strains G-C1 and G-12 were amplified with primer pair darsB1F and darsB1R (Achour et al. 2007) (Table 1). Amplification details of arsC and arsB gene were showed in the SI. Based on acquired arsB gene sequences, three primer pairs (Z540F/Z155R, K551F/K38R, 53BF/53BR, Table 1) were designed and two kinds of restriction endonucleases (EcoR I and Hind III, TaKaRa) were selected according to the inverse-PCR method (Ochman et al. 1988). Different from the common way, the forward primers here were designed at the end of the obtained arsB gene sequences while the reverse primers were designed at the head. Besides, it must be sure that the arsB gene could not be cut off by selected endonucleases. Restriction digests were carried out using 2 μg of extracted DNAs of strain G-C1 and G-12, which were treated with 15 U EcoR I and 15 U Hind III in a 50 μL mixture at 37 °C for 5 h followed by an inactivation of endonucleases at 65 °C for 20 min. For circularization, digested DNAs were purified using method mentioned above and the ligation reaction was performed by the addition of T4 DNA ligase (TaKaRa) to a concentration of 1 U/μL and the reaction was proceeded at 25 °C for 5 h and the circularized DNAs were purified again for PCR amplification. The PCR was performed manually in reactions containing 5 x PrimeSTAR GXL Buffer, 200 μM dNTP mixture, 0.2 μM designed primers, 50 ng circularized DNA and 1 U of PrimeSTAR GXL DNA Polymerase (TaKaRa, Japan). We used 30 cycles of 98 °C for 10 s, 55 °C for 15 s, and 68 °C for 1 min according to manufacturer’s guide. The PCR products were fractionated at 1% agarose gel, purified and then formed sticky ends with a 20 μL mixture containing 0.5 U of Taq DNA polymerase and 200 μM dNTP mixture at 72 °C for 20 min. Finally, the extended DNA products were purified again for ligating, transforming as described above and several single clones were picked out and sequenced.
PCR amplification and clone library analyses of arsenate reductase genes in groundwater samples
Total genomic DNA was extracted and purified from about one-quarter of each filter membrane of water samples using FastDNA spin kit for soil (MP Bio Laboratories, USA). The concentration and quality of extracted DNA were checked using Nanodrop 2000 (Thermo Fisher Scientific, USA). The arrA gene and arsC gene of groundwater samples were amplified with primer sets arrA-CVF1/arrA-CVR1(Mirza et al. 2016) and P52F/P323R (Bachate et al. 2009) (Table 1), respectively. Detailed information of PCR mixture and protocol was shown in SI. After clone screening procedure, several clones were picked out and sequenced. Data from the obtained raw sequences were processed following our previous methods (Li et al. 2014; Wang et al. 2018). The GenBank accession number of the arrA and arsC gene sequences were MK038511-MK038604.
Results and discussion
Isolation, identification and As (V) reduction
Three As (V)-reducing strains were obtained from this study. Phenotypic analysis found strain G-11 to be a gram-positive, strictly anaerobic bacterium, while strain G-C1 and G-12 were gram-negative, facultatively anaerobic bacteria. Strain G-11 and G-C1 had a pH range of 4.0–9.0 with the optimum pH of 8.0 and the optimum growth temperature of 37 °C, while strain G-12 could survive at the wide pH range of 3.0-10.0 (optimum at 8.0) and at the optimal growth temperature of 35 °C. All these strains could utilize different carbon sources including maltose, fructose, glucose, sucrose, galactose, sodium citrate, sodium acetate and sodium lactate.
BLAST analysis of the 16 s rRNA gene sequences indicated that these three strains belonged to genera Paraclostridium, Citrobacter and Klebsiella, respectively. Strain G-11 presented highest similarities with two microorganisms referred to as P. benzoelyticum strain JC272 (100%) isolated from marine sediment (Jyothsna et al. 2016) and P. bifermentans strain HYN0063 (99.93%) isolated in South Korea (NCBI database, accession number MF988694). Strain G-C1 showed high similarity (99.93%) with C. freundii strain S1 which was isolated from soil (Kary and Alizadeh 2016), while G-12 presented high similarity (99.57%) with K. variicola strain XF4 (Zhang and Kong 2014) (Fig. 1). Several previous studies reported that Klebsiella and Citrobacter were capable of As (V) reduction (Maeda et al. 1992; Shakoori et al. 2010; Wang et al. 2017), with the maximum tolerated concentration (MTC) of As for Klebsiella oxytoca was 3.2 mM (Shakoori et al. 2010). The three strains isolated in the present study could tolerate As (V) up to 30 mM (Fig. S1), indicating their strong As resistance. To date, there have been rare reports on reduction of As (V) by Paraclostridium while it was reported to be involved mainly in hydrogen or alcohol production under strictly anaerobic conditions (Liu et al. 2014; Yang et al. 2019).
Neighbor-joining tree of 16 S rRNA gene sequences of strains G-11, G-C1 and G-12. The sequence of Bacteroides fragilis strain ATCC 25285 was used as the outgroup. Only bootstrap values above 50% (for 1000 iterations) are shown. The GenBank accession number for each reference strain is shown before the strain name
In CDM containing 1 mM As (V), strain G-11, G-C1 and G-12 could reduce > 99% of As (V) to As (III) in 60 h, 72 h and 84 h, respectively, suggesting strain G-11 possessed the strongest ability with least amount of biomass (Fig. 2). The As (V) reduction abilities of the strains in the present study were comparable with the results of a previous study (Saleh and Ekram 2013) where Klebsiella oxytoca and Rahnella aquatilis could reduce 1.33 mM As (V) by percentages of 75 and 69%, respectively.
Arsenic-reducing clusters of As(V) reduction
In order to investigate the arsenic-metabolizing gene clusters of the isolated strains, putative arsC gene sequence (Accession No. MK038613) was amplified from strain G-11 using the primer set PB2-F/PB2-R. However, no arsC gene sequence could be amplified from strain G-C1 and G-12 with any reported primer sets (Table S1) (Bachate et al. 2009; Macur et al. 2004; Sun et al. 2004). Fortunately, the arsB gene sequences (750 bp) were obtained from genomic DNA of strain G-C1 and G-12. Using the reverse PCR technique with designed reverse PCR primers (Table 1, Z540/Z155R for strain G-C1, K551F/K38R and 53BF/53BR for strain G-12), the flanked sequences by the arsB gene from strain G-C1 and G-12 were successfully amplified. Finally, the arsC family genes including arsR gene (188 bp, MK038605), arsB gene (1290 bp, MK038606), arsC gene (426 bp, MK038607) were obtained from genomic DNA of strain G-C1.
For strain G-12, two incomplete ars operons consisting of arsA gene (888 bp, MK038608), arsB gene (1290 bp, MK038609) and arsC gene (426 bp, MK038610) for operon 1, and arsB gene (1119 bp, MK038611) and arsC gene (426 bp, MK038612) for operon 2 were successfully amplified (Fig. 3). It was worth mentioning that the genome of G-12 presented two arsB and arsC genes. Previous studies showed that more than one ars operon might be involved in the microbe genome such as strain Citrobacter freundii UMH16 (Anderson et al. 2018) and Pseudomonas putida BIRD-1 (Matilla et al. 2011). The transcription of ars operon might be regulated by environmental factors such as temperature (Páez‐Espino et al. 2015; Wang et al. 2016). Further investigations are needed to find the possible key environmental factors regulating ars systems in strain G-12.
Several universal primer sets from previous studies (Fisher et al. 2008; Kulp et al. 2006; Malasarn et al. 2004; Mirza et al. 2016; Song et al. 2009) and specific primer sets (designed in the present study from arrA genes of the same genera as the isolated strains) (Table S1) were chosen but failed to amplify any arrA related genes, implying that even the same genus could present different genomic compositions (Escudero et al. 2013).
Biodiversity of in situ As(V)-reducing bacterial communities
A total of 160 arrA and 302 arsC gene sequences were obtained to construct phylogenetic trees. The coverage values of arrA and arsC gene clone libraries were 0.80-0.89 and 0.90-0.91, respectively. With a 97% sequence similarity, the arrA gene sequences were classified into 8, 14, 12 and 10 OTUs, while the arsC gene sequences were classified into 11, 13, 13 and 11 OTUs, respectively (Table S2). Shannon and Chao1 indices of arrA and arsC gene clone libraries ranged from 1.00 to 2.35, 1.45 to 1.84 and from 13.00 to 20.00, 16.75 to 41.00, respectively (Table S2). The diversity indices of the functional genes and geochemical characteristics (Table S3) did not present significant correlations according to statistical correlations analysis (Table S4), indicating that diversity of As-reducing bacteria in groundwater were affected by various geochemical variables.
The amino acid sequences derived from the arrA gene sequences were phylogenetically clustered into Betaproteobacteria, Deltaproteobacteria and an unidentified group, with Deltaproteobacteria being dominant by the percentage of 53.13% (Fig. 4a). In the class Deltaproteobacteria, 48.75% and 4.37% of sequences were related to arrA amino acid sequences of Desulfuromonadales and Desulfobacterales, respectively. The cluster associated with Desulfuromonadales were mainly composed of Geobacter and showed high similarities to the arsenate respiratory reductase of Geobacter sp. OR-1, Geobacter thiogenes and Geobacter uraniireducens, which were previously reported isolated from sediments or paddy soils (Ohtsuka et al. 2013; Qiao et al. 2017; Upadhyaya et al. 2012). Previous studies showed that Geobacter sp. was also the dominant Fe(III)-reducing bacteria in many anaerobic environments and relatively high percentages of Geobacter sp. in microbial communities might be correlated with Fe(III) reduction and As(III) release in aquifers (Li et al. 2013; Rowland et al. 2007). Furthermore, G. uraniireducens was capable of reducing sorbed As(V) on Fe(III) oxyhydroxides to more mobile As (III) (Héry et al. 2010). These results suggested that Geobacter sp. could play an important role in As mobilization in groundwater via either direct or indirect mechanisms. Seven clones belonged to Desulfobacterales were similar with sequences of Desulfopila aestuarii DSM 18488, Desulfofustis glycolicus DSM 9705 and Desulfobulbaceae sp. A2 in sulfidic groundwater-fed fountain (Sharrar et al. 2017). The second cluster associated with Betaproteobacteria accounted for 11.88% of clones and was highly similar to ArrA like protein of Betaproteobacteria and Sulfuritalea hydrogenivorans sk43H from Rifle groundwater or freshwater (Anantharaman et al. 2016; Watanabe et al. 2014). Besides, 35% of sequenced clones were affiliated with putative ArrA proteins from New Jersey groundwater (Mumford et al. 2012), paddy soil (Zhang et al. 2015) or Cache Valley Basin (Mirza et al. 2014) and showed extremely low similarity with previously identified arrA gene sequences from cultured isolates, suggesting the possible existence of unique dissimilatory As(V)-reducing bacteria in groundwater of the Hetao Plain.
The phylogenetic tree of deduced amino acid sequences based on the arrA and arsC gene sequences of four samples from the in situ high As groundwater. One representative type within each OTU was shown. The GenBank accession number for each reference strain is shown before the strain name. Numbers in parentheses represent sequences numbers obtained for each OTU. Only bootstrap values above 50% (for 1000 iterations) are shown. □: IMCUGGW05; ■: IMCUGGW07; ☆: IMCUGGW13; ★: IMCUGGW17. (a) arrA gene based; (b) arsC gene based
Unlike the results of arrA genes, sequences derived from arsC genes were widely distributed among the microbial phyla, indicating that the detoxification was the most common mechanism of As reduction, which was consistent with the result of our previous study (Li et al. 2017). The translated arsC amino acid sequences were divided into three clusters: Omnitrophica, Deltaproteobacteria and Firmicutes (Fig. 4b). The largest cluster which belonged to Omnitrophica (88.08%) was similar to those bacteria of Candidatus Omnitrophica with arsenate reductase ArsC in groundwater (Anantharaman et al. 2016). Candidatus Omnitrophica which was previously known as OP3 belonged to the PVC superphylum and had not get any pure cultures yet (Devos and Ward 2014). What differed from the result of a previous study which indicated that OP3 was more abundant in low As groundwater samples (Wang et al. 2016), results of the present study showed that OP3 endowed with cytosolic arsenate reductase gene might be widely distributed in groundwater samples with different As concentrations. Twenty-one sequenced clones belonged to Syntrophobacterales which showed similarity with ArsC protein of Syntrophus aciditrophicus and Syntrophobacter sp. SbD1 (NCBI database, accession numbers WP011417387 and WP106821703). Two clones belonged to Bacillaceae showed similarities with Oceanobacillus chungangensis which was isolated from a sand dune (Lee et al. 2013). Besides, several sequences which belonged to Proteobacteria and Firmicutes have been proved to be involved in the As reduction processes (Cavalca et al. 2013). The cluster associated with Geobacter accounted for 4.3% of arsC amino acid sequences and was related to arsenate reductase ArsC of Geobacter such as Geobacter. sp. metabat 2.715 which was isolated from wetland sediments (Martins et al. 2018). These results once again proved the important role of Geobacter sp. in As cycling of the high As groundwater.
Collectively, results of the indigenous As (V) reducing communities showed that Geobacter and Omnitrophica were the major populations of DARPs and ARMs, respectively. Meanwhile, Geobacter was ever reported to be the dominant genus of microbial communities in high As groundwater of the Hetao Plain (Li et al. 2013). Previous studies found that Geobacter species were one kind of typical dissimilatory iron-reducing bacteria with the ability of extracellular electron transfer (Liu et al. 2018; Shi et al. 2009). In high arsenic groundwater, arrA-harboring Geobacter species could transfer electrons through cytomembrane using their specific extracellular electron transport system. The electrons might be sequestrated by ArrA reductase during the electrons being transferred through periplasmic space, leading to arsenate reduction. Besides, consistent with the failed amplification of arrA genes from isolated strains of the present study, no Gammaproteobacteria and Firmicutes were detected in the analysis of arrA based As (V) reducing bacterial communities, suggesting that the isolates in this study might be ARMs.
Conclusion
Three As (V)-reducing bacterial strains with strong As (V)-reducing abilities were isolated from high As groundwater of the Hetao Plain and identified as Paraclostridium, Citrobacter and Klebsiella, respectively. This was the first report of As (V) reduction by genus Paraclostridium. These strains might present different detoxifying strategies of As. The predominant As (V)-reducing populations in indigenous high As groundwater were diverse, which were dominated with Geobacter by targeting the functional arrA genes and dominated with Omnitrophica as well as Geobacter by targeting the arsC genes, respectively. The study might further expand our understanding of microbially-mediated As reduction and release in high As groundwater aquifers.
References
Achour AR, Bauda P, Billard P (2007) Diversity of arsenite transporter genes from arsenic-resistant soil bacteria. Res Microbiol 158:128–137
Anantharaman K, Brown CT, Hug LA, Sharon I, Castelle CJ, Probst AJ, Thomas BC, Singh A, Wilkins MJ, Karaoz U (2016) Thousands of microbial genomes shed light on interconnected biogeochemical processes in an aquifer system. Nat Commun 7:13219
Anderson MT, Mitchell LA, Zhao L, Mobley HL (2018) Citrobacter freundii fitness during bloodstream infection. Sci Rep 8:11792
Bachate S, Cavalca L, Andreoni V (2009) Arsenic‐resistant bacteria isolated from agricultural soils of Bangladesh and characterization of arsenate‐reducing strains. J Appl Microbiol 107:145–156
Barringer JL, Reilly PA, Eberl DD, Blum AE, Bonin JL, Rosman R, Hirst B, Alebus M, Cenno K, Gorska M (2011) Arsenic in sediments, groundwater, and streamwater of a glauconitic coastal plain terrain, New Jersey, USA—chemical “fingerprints” for geogenic and anthropogenic sources. Appl Geochem 26:763–776
Branco R, Chung A-P, Morais PV (2008) Sequencing and expression of two arsenic resistance operons with different functions in the highly arsenic-resistant strain Ochrobactrum tritici SCII24 T. BMC Microbiol 8:95
Cai X, Zhang Z, Yi N, Du H, Li Z, Cui Y (2016) Comparison of arsenate reduction and release by three As (V)-reducing bacteria isolated from arsenic-contaminated soil of Inner Mongolia, China. Chemosphere 161:200–207
Cavalca L, Corsini A, Zaccheo P, Andreoni V, Muyzer G (2013) Microbial transformations of arsenic: perspectives for biological removal of arsenic from water. Fut Microbiol 8:753–768
Chang YC, Nawata A, Jung K, Kikuchi S (2012) Isolation and characterization of an arsenate-reducing bacterium and its application for arsenic extraction from contaminated soil. J Ind Microbiol Biotechnol 39:37–44
Dai X, Li P, Tu J, Zhang R, Wei D, Li B, Wang Y, Jiang Z (2016) Evidence of arsenic mobilization mediated by an indigenous iron reducing bacterium from high arsenic groundwater aquifer in Hetao Basin of Inner Mongolia, China. Int Biodeterior Biodegrad 128:22–27
Deng Y (2008) Geochemical processes of high arsenic groundwater system at western Hetao Basin. China University of Geosciences (Wuhan)
Deng Y, Wang Y, Ma T (2009) Isotope and minor element geochemistry of high arsenic groundwater from Hangjinhouqi, the Hetao Plain, Inner Mongolia. Appl Geochem 24:587–599
Deng Y, Wang Y, Ma T, Yang H, He J (2011) Arsenic associations in sediments from shallow aquifers of northwestern Hetao Basin, Inner Mongolia. Environ Earth Sci 64:2001–2011
Devos DP, Ward NL (2014) Mind the PVCs. Environ Microbiol 16:1217–1221
Escudero LV, Casamayor EO, Chong G, Pedrós-Alió C, Demergasso C (2013) Distribution of microbial arsenic reduction, oxidation and extrusion genes along a wide range of environmental arsenic concentrations. PLoS ONE 8:e78890
Fan H, Su C, Wang Y, Yao J, Zhao K, Wang G (2008) Sedimentary arsenite‐oxidizing and arsenate‐reducing bacteria associated with high arsenic groundwater from Shanyin, Northwestern China. J Appl Microbiol 105:529–539
Fendorf S, Michael HA, van Geen A (2010) Spatial and temporal variations of groundwater arsenic in south and southease Asia. Science 328:1123–1127
Ferreccio C, Sancha AM (2006) Arsenic exposure and its impact on health in Chile. J Health Popul Nutr 24:164–175
Fisher E, Dawson AM, Polshyna G, Lisak J, Crable B, Perera E, Ranganathan M, Thangavelu M, Basu P, Stolz JF (2008) Transformation of inorganic and organic arsenic by Alkaliphilus oremlandii sp. nov. strain OhILAs. Ann NY Acad Sci 1125:230–241
Guo H, Liu Z, Ding S, Hao C, Xiu W, Hou W (2015) Arsenate reduction and mobilization in the presence of indigenous aerobic bacteria obtained from high arsenic aquifers of the Hetao basin, Inner Mongolia. Environ Pollut 203:50–59
Guo H, Zhang B, Li Y, Berner Z, Tang X, Norra S, Stüben D (2011) Hydrogeological and biogeochemical constrains of arsenic mobilization in shallow aquifers from the Hetao basin, Inner Mongolia. Environ Pollut 159:876–883
Héry M, Van Dongen B, Gill F, Mondal D, Vaughan D, Pancost R, Polya D, Lloyd J (2010) Arsenic release and attenuation in low organic carbon aquifer sediments from West Bengal. Geobiology 8:155–168
Handley KM, Héry M, Lloyd JR (2009) Redox cycling of arsenic by the hydrothermal marine bacterium Marinobacter santoriniensis. Environ Microbiol 11:1601–1611
Harvey CF, Swartz CH, Badruzzaman A, Keon-Blute N, Yu W, Ali MA, Jay J, Beckie R, Niedan V, Brabander D (2002) Arsenic mobility and groundwater extraction in Bangladesh. Science 298:1602–1606
Hungate R (1969) A roll tube method for cultivation of strict anaerobes. Methods Microbiol 3B:117–132
Jiang Z, Li P, Wang Y, Li B, Deng Y, Wang Y (2014) Vertical distribution of bacterial populations associated with arsenic mobilization in aquifer sediments from the Hetao plain, Inner Mongolia. Environmental Earth Sciences 71:311–318
Jyothsna TS, Tushar L, Sasikala C, Ramana CV (2016) Paraclostridium benzoelyticum gen. nov., sp. nov., isolated from marine sediment and reclassification of Clostridium bifermentans as Paraclostridium bifermentans comb. nov. Proposal of a new genus Paeniclostridium gen. nov. to accommodate Clostridium sordellii and Clostridium ghonii. Int J Syst Evolut Microbiol 66:1268–1274
Kary NE, Alizadeh Z (2016) Non-symbiotic association of Citrobacter freundii and Staphylococcus succinus with the entomopathogenic nematode Steinernema feltiae. J Entomological Soc Iran 36:111–119
Kulp T, Hoeft S, Miller L, Saltikov C, Murphy J, Han S, Lanoil B, Oremland R (2006) Dissimilatory arsenate and sulfate reduction in sediments of two hypersaline, arsenic-rich soda lakes: Mono and Searles Lakes, California. Appl Environ Microbiol 72:6514–6526
Lee DC, Kang H, Weerawongwiwat V, Kim B, Choi Y-W, Kim W (2013) Oceanobacillus chungangensis sp. nov., isolated from a sand dune. Int J Syst Evolut Microbiol 63:3666–3671
Li P, Jiang D, Li B, Dai X, Wang Y, Jiang Z, Wang Y (2014) Comparative survey of bacterial and archaeal communities in high arsenic shallow aquifers using 454 pyrosequencing and traditional methods. Ecotoxicology 23:1878–1889
Li P, Jiang Z, Wang Y, Deng Y, Van Nostrand JD, Yuan T, Liu H, Wei D, Zhou J (2017) Analysis of the functional gene structure and metabolic potential of microbial community in high arsenic groundwater. Water Res 123:268–276
Li P, Wang Y, Jiang Z, Jiang H, Li B, Dong H, Wang Y (2013) Microbial diversity in high arsenic groundwater in Hetao Basin of Inner Mongolia, China. Geomicrobiol J 30:897–909
Liao VHC, Chu YJ, Su YC, Hsiao SY, Wei CC, Liu C-W, Liao C-M, Shen W-C, Chang F-J (2011) Arsenite-oxidizing and arsenate-reducing bacteria associated with arsenic-rich groundwater in Taiwan. J Contaminant Hydrology 123:20–29
Liu X, Gu Q, Liao C, Yu X (2014) Ehancing butanol tolerance and preventing degeneration in Clostridium acetobutylicum by 1-butanol-glycerol storage during long-term preservation. Biomass Biogenergy 69:192–197
Liu X, Shi L, Gu J-D (2018) Microbial electrocatalysis: Redox mediators responsible for extracellular electron transfer. Biotechnol Adv 36:1815–1827
Lv W, Yu Z (2013) Isolation and characterization of two thermophilic cellulolytic strains of Clostridium thermocellum from a compost sample. J Appl Microbiol 114:1001–1007
Macur RE, Jackson CR, Botero LM, Mcdermott TR, Inskeep WP (2004) Bacterial populations associated with the oxidation and reduction of arsenic in an unsaturated soil. Environ Sci Technol 38:104–111
Macy JM, Santini JM, Pauling BV, O’Neill AH, Sly LI (2000) Two new arsenate/sulfate-reducing bacteria: mechanisms of arsenate reduction. Arch Microbiol 173:49–57
Maeda S, Ohki A, Miyahara K, Naka K, Higashi S (1992) Metabolism of methylated arsenic compounds by arsenic-resistane bacteria (Klebsiella oxytoca and Xanthomonas sp.). Appl Organometallic Chem 6:415–420
Malasarn D, Saltikov C, Campbell K, Santini J, Hering J, Newman D (2004) arrA is a reliable marker for As (V) respiration. Science 306:455–455
Maleki A, Pajootan E, Hayati B (2015) Ethyl acrylate grafted chitosan for heavy metal removal from wastewater: equilibrium, kinetic and thermodynamic studies. J Taiwan Inst Chem Eng 51:127–134
Martins PD, Danczak RE, Roux S, Frank J, Borton MA, Wolfe RA, Burris MN, Wilkins MJ (2018) Viral and metabolic controls on high rates of microbial sulfur and carbon cycling in wetland ecosystems. Microbiome 6:138
Matilla MA, Pizarro-Tobias P, Roca A, Fernández M, Duque E, Molina L, Wu X, van der Lelie D, Gómez MJ, Segura A (2011) Complete genome of the plant growth-promoting rhizobacterium Pseudomonas putida BIRD-1. J Bacteriol 193:1290–1290
Mirza BS, Muruganandam S, Meng X, Sorensen DL, Dupont RR, McLean JE (2014) Arsenic (V) reduction in relation to iron (III) transformation and molecular characterization of the structural and functional microbial community in sediments of a basin-fill aquifer in Northern Utah. Appl Environ Microbiol 80:3198–3208
Mirza BS, Sorensen DL, Dupont RR, McLean JE (2016) New arsenate reductase gene (arrA) PCR primers for diversity assessment and quantification in environmental samples. Appl Environ Microbiol 83:e02725–16
Mumford AC, Barringer JL, Benzel WM, Reilly PA, Young L (2012) Microbial transformations of arsenic: mobilization from glauconitic sediments to water. Water Res 46:2859–2868
Neidhardt H, Norra S, Tang X, Guo H, Stuben D (2012) Impact of irrigation with high arsenic burdened groundwater on the soil-plant system: Results from a case study in the Inner Mongolia, China. Environ Pollut 163:8–13
Niggemyer A, Spring S, Stackebrandt E, Rosenzweig RF (2001) Isolation and characterization of a novel As (V)-reducing bacterium: implications for arsenic mobilization and the genus Desulfitobacterium. Appl Environ Microbiol 67:5568–5580
Ochman H, Gerber AS, Hartl DL (1988) Genetic applications of an inverse polymerase chain reaction. Genetics 120:621–623
Ohtsuka T, Yamaguchi N, Makino T, Sakurai K, Kimura K, Kudo K, Homma E, Dong DT, Amachi S (2013) Arsenic dissolution from Japanese paddy soil by a dissimilatory arsenate-reducing bacterium Geobacter sp. OR-1. Environ Sci Technol 47:6263–6271
Páez‐Espino AD, Durante‐Rodríguez G, de Lorenzo V (2015) Functional coexistence of twin arsenic resistance systems in Pseudomonas putida KT 2440. Environ Microbiol 17:229–238
Qiao JT, Li XM, Hu M, Li FB, Young LY, Sun WM, Huang W, Cui JH (2017) Transcriptional activity of arsenic-reducing bacteria and genes regulated by lactate and biochar during arsenic transformation in flooded paddy soil. Environ Sci Technol 52:61–70
Rowland H, Pederick R, Polya D, Pancost R, Van Dongen B, Gault A, Vaughan D, Bryant C, Anderson B, Lloyd J (2007) The control of organic matter on microbially mediated iron reduction and arsenic release in shallow alluvial aquifers, Cambodia. Geobiology 5:281–292
Saleh MA, Ekram AE (2013) Arsenate reduction by some bacteria isolated from industrial effluents of rajshahi, Bangladesh. Int J Appl Nat Sci 2:23–30
Saltikov CW, Wildman RA, Newman DK (2005) Expression dynamics of arsenic respiration and detoxification in Shewanella sp. strain ANA-3. J Bacteriology 187:7390–7396
Shakoori FR, Aziz I, Rehman A, Shakoori A (2010) Isolation and characterization of arsenic reducing bacteria from industrial effluents and their potential use in bioremediation of wastewater. Pak J Zool 42:331–338
Sharrar AM, Flood BE, Bailey JV, Jones DS, Biddanda BA, Ruberg SA, Marcus DN, Dick GJ (2017) Novel large sulfur bacteria in the metagenomes of groundwater-fed chemosynthetic microbial mats in the Lake Huron basin. Front Microbiol 8:791
Shi L, Richardson DJ, Wang Z, Kerisit SN, Rosso KM, Zachara JM, Fredrickson JK (2009) The roles of outer membrane cytochromes of Shewanella and Geobacter in extracellular electron transfer. Environ Microbiol Rep 1:220–227
Song B, Chyun E, Jaffé PR, Ward BB (2009) Molecular methods to detect and monitor dissimilatory arsenate-respiring bacteria (DARB) in sediments. FEMS Microbiol Ecol 68:108–117
Sun Y, Polishchuk EA, Radoja U, Cullen WR (2004) Identification and quantification of arsC genes in environmental samples by using real-time PCR. J Microbiol Methods 58:335–349
Tsai S, Singh S, Chen W (2009) Arsenic metabolism by microbes in nature and the impact on arsenic remediation. Curr Opin Biotechnol 20:659–667
Upadhyaya G, Clancy TM, Brown J, Hayes KF, Raskin L (2012) Optimization of arsenic removal water treatment system through characterization of terminal electron accepting processes. Environ Sci Technol 46:11702–11709
Wang J, Zeng X, Zhu X, Chen X, Zeng X, Mu Y, Yang Y, Wang Y (2017) Sulfate enhances the dissimilatory arsenate-respiring prokaryotes-mediated mobilization, reduction and release of insoluble arsenic and iron from the arsenic-rich sediments into groundwater. J Hazard Mater 339:409–417
Wang L, Zhuang X, Zhuang G, Jing C (2016) Arsenic resistance strategy in Pantoea sp. IMH: organization, function and evolution of ars genes. Sci Rep 6:39195
Wang Y, Li P, Dai X, Zhang R, Jiang Z, Jiang D, Wang Y (2015) Abundance and diversity of methanogens: Potential role in high arsenic groundwater in Hetao Plain of Inner Mongolia, China. Sci Total Environ 515:153–161
Wang Y, Li P, Jiang Z, Liu H, Wei D, Wang H, Wang Y (2018) Diversity and abundance of arsenic methylating microorganisms in high arsenic groundwater from Hetao Plain of Inner Mongolia, China. Ecotoxicology 27:1047–1057
Wang Y, Li P, Jiang Z, Sinkkonen A, Wang S, Tu J, Wei D, Dong H, Wang Y (2016) Microbial community of high arsenic groundwater in agricultural irrigation area of Hetao Plain, Inner Mongolia. Front Microbiol 7:1917
Watanabe T, Kojima H, Fukui M (2014) Complete genomes of freshwater sulfur oxidizers Sulfuricella denitrificans skB26 and Sulfuritalea hydrogenivorans sk43H: genetic insights into the sulfur oxidation pathway of betaproteobacteria. Syst Appl Microbiol 37:387–395
Weeger W, Lievremont D, Perret M, Lagarde F, Hubert J-C, Leroy M, Lett M-C (1999) Oxidation of arsenite to arsenate by a bacterium isolated from an aquatic environment. Biometals 12:141–149
Winkel LH, Trang PTK, Lan VM, Stengel C, Amini M, Ha NT, Viet PH, Berg M (2011) Arsenic pollution of groundwater in Vietnam exacerbated by deep aquifer exploitation for more than a century. Proc Natl Acad Sci 108:1246–1251
Yang G, Yin Y, Wang J (2019) Microbial community diversity during fermentative hydrogen production inoculating various pretreated cultures. Int J Hydrogen Energy 44:13147–13156
Zhang C, Kong F (2014) Isolation and identification of potassium-solubilizing bacteria from tobacco rhizospheric soil and their effect on tobacco plants. Appl Soil Ecology 82:18–25
Zhang SY, Zhao FJ, Sun GX, Su JQ, Yang XR, Li H, Zhu YG (2015) Diversity and abundance of arsenic biotransformation genes in paddy soils from southern China. Environ Sci Technol 49:4138–4146
Webster G, Newberry CJ, Fry JC, Weightman AJ (2003) Assessment of bacterial community structure in the deep sub-seafloor biosphere by 16S rDNA-based techniques: a cautionary tale. J Microbiol Methods 55(1):155–164
Acknowledgements
This research was financially supported by National Natural Science Foundation of China (Grant Nos. 91851115, 41702365 and 41702260).
Funding
This study was funded by National Natural Science Foundation of China (grant numbers 91851115, 41702365, and 41702260)
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Wang, Y., Wei, D., Li, P. et al. Diversity and arsenic-metabolizing gene clusters of indigenous arsenate-reducing bacteria in high arsenic groundwater of the Hetao Plain, Inner Mongolia. Ecotoxicology 30, 1680–1688 (2021). https://doi.org/10.1007/s10646-020-02305-1
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10646-020-02305-1