Abstract
MYB transcription factors are encoded by a large family of highly conserved genes from plants to vertebrates. There are three members of the MYB gene family in human, namely, MYB, MYBL1, and MYBL2 that encode MYB/c-MYB, MYBL1/A-MYB, and MYBL2/B-MYB, respectively. MYB was the first member to be identified as a cellular homolog of the v-myb oncogene carried by the avian myeloblastosis virus (AMV) causing leukemia in chickens. Under the normal scenario, MYB is predominantly expressed in hematopoietic tissues, colonic crypts, and neural stem cells and plays a role in maintaining the undifferentiated state of the cells. Over the years, aberrant expression of MYB genes has been reported in several malignancies and recent years have witnessed tremendous progress in understanding of their roles in processes associated with cancer development. Here, we review various MYB alterations reported in cancer along with the roles of MYB family proteins in tumor cell plasticity, therapy resistance, and other hallmarks of cancer. We also discuss studies that provide mechanistic insights into the oncogenic functions of MYB transcription factors to identify potential therapeutic vulnerabilities.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
MYB transcription factors are encoded by a large family of highly conserved genes from plants to vertebrates, suggesting their significance in the fundamental biological processes [1, 2]. In humans and other vertebrates, there are three members in the MYB gene family, namely, MYB, MYBL1, and MYBL2 encoding MYB or c-MYB, A-MYB or MYBL1, and B-MYB or MYBL2, respectively. Invertebrates, however, harbor a single Myb gene only [1]. MYB was the first member to be identified in humans as a cellular homolog of the v-myb oncogene inserted in the genome of avian myeloblastosis virus (AMV) causing acute myeloblastic leukemia in chickens [3, 4]. Later, screening of T-cell cDNA libraries with low stringency hybridization with a c-myb probe identified A-MYB and B-MYB genes showing strong sequence homology [5, 6]. MYB is predominantly expressed in hematopoietic tissues, colonic crypts, and neural stem cells, whereas MYBL1 expression is restricted to gonadal tissue, germinal B lymphocytes, developing mammary gland, and central nervous system. In contrast, the expression of MYBL2 is ubiquitous in all proliferating cells [7, 8]. An aberrant expression of MYB genes has been reported in different cancers due to either gene amplification and/or transcriptional upregulation. Alternative splice variants and gene fusions of MYB, MYBL1, and MYBL2 have also been reported in some cancers [2, 9,10,11]. This review article sheds light on various MYB alterations reported in cancer along with the roles of MYB family proteins in tumor cell plasticity and therapy resistance sustaining the relentless growth and spread of cancer cells. We also discuss studies that provide insights into these oncogenic functions of MYB transcription factors to identify potential therapeutic vulnerabilities.
2 Genomic and proteomic organization of MYB transcription factors
According to UCSC genome browser (https://genome.ucsc.edu/), MYB gene is present at the chromosomal band 6q23.3 as a 37,865 bp long sequence containing 16 exons and 15 introns. MYBL1 is localized on chromosome 8 in the region q13.1 and is the largest in size (51,044 bp), also consisting of 16 exons and 15 introns. The chromosomal locus of MYBL2 is 20q13.12 and its size is 49,369 bp containing 14 exons and 13 introns (Fig. 1A). A large number of splice variants of MYBs are reported, of which some are translated, while others undergo non-sense-mediated decay or their fates remain unknown [12]. Full length MYB protein is composed of 642 amino acids having a molecular weight of ~ 75 kD. A-MYB/MYBL1 and B-MYB/MYBL2 are slightly heavier, weighing about 95 kD and 93 kD, and are composed of 752 and 742 amino acids, respectively [2]. Structurally, all MYB proteins have large similarities. They contain a highly conserved N-terminal DNA-binding domain (DBD), encompassing three tryptophan-rich tandemly repeated motifs of ~ 50 amino acids (R1, R2, and R3), a central trans-activated domain (TAD), and a C-terminal negative regulatory domain (NRD). The folding architecture is similar for each of the three repeats of DBD and each repeat contains a variation of helix-turn-helix (HTH) motif [13, 14] (Fig. 1B).
Genomic and proteomic organization of MYB transcription factors. A Genomic location of MYB transcription factors (MYB, MYBL1, and MYBL2) and depiction of their exons (E) and introns (I). B Comparative presentation of functional domains of MYBs. The N-terminal DNA binding domain (DBD) of MYB, MYBL1, and MYBL2 is composed of approximately ~ 150 amino acids and consists of three repeats, R1, R2, and R3. The central region contains transactivation domain (TAD), with different number of amino acids in MYB (~ 135 a.a.), MYBL1 (~ 65 a.a.) and MYBL2 (~ 33 a.a.). C-terminal negative regulatory domain (NRD) has variable number of amino acids in different MYB proteins and frequently undergoes posttranslational modifications, including phosphorylation (P), acetylation (Ac), and sumoylation (SUMO)
All MYB proteins, including v-myb, recognize and bind to the same DNA consensus sequence [5′-(YAACG/TG)-3′], known as the MYB recognition or MYB-binding site (MBS). “Y” in this sequence represents a pyrimidine base (C or T) [15, 16]. Structural analysis has revealed that R2 and R3 motifs are responsible for DNA recognition, while the R1 motif is dispensable [13, 17]. Both c-MYB and A-MYB share similarities in the nature of amino acid composition of TAD, having clusters of acidic amino acids, although there are some sequence-specific differences [18]. B-MYB TAD shows minimal homology to that of c-MYB despite containing the clusters of acidic residues indicating functional differences in gene activation [19]. Across different species, C-terminal regulatory domain shows conserved sequences, with most significant similarity observed in the central region [20, 21]. This domain contains a leucine zipper structure, which confers negative regulatory activity by forming a homodimer that interferes with the binding to the target DNA sequence [22,23,24]. Post-translational modifications in the regulatory domain inhibit its interaction with DBD, thus repressing the transcriptional activity [25,26,27]. Consequently, truncation or mutations in this domain are known to confer oncogenic ability to MYB resulting from its constitutive activation [28].
3 MYB alterations in cancers
While the expression of MYB family proteins is tightly regulated in healthy tissues, a number of alterations have been reported in human malignancies. These include gene amplification, mutations, and structural rearrangements due to chromosomal translocation or gene fusion resulting in their enhanced biological activity that promotes different aspects of tumorigenesis.
3.1 Gene copy number alterations
Analysis of candidate oncogenes in pancreatic cancer (PC) identified amplification at 6q24 chromosomal locus that houses MYB [29]. Similarly, a copy number gain of MYB was also detected in BRCA1-mutated breast tumors by fluorescence in situ hybridization analysis of 6q22-24 region [30]. An amplification of MYB has also been reported in pediatric low-grade gliomas (LGGs) [31]. High-density profiling of gastric adenocarcinomas revealed somatic copy number alterations in MYB oncogene associated with its overexpression [32]. Similarly, MYB amplification has also been reported in prostate cancer exhibiting enhanced amplification frequency as it progressed from hormone-sensitive to hormone-resistant state [33]. Amplification of MYB is reported to be of prognostic significance in esophageal carcinoma [34]. Pediatric low-grade gliomas (PLGGs) have the most significant gain in 8q13.1 chromosomal region resulting in the partial duplication of MYBL1 along with the deletion of its c-terminal negative-regulatory domain [35]. MYBL2 overexpression in breast cancer, malignant melanoma, and sporadic ovarian cancer is also shown to partly result from the amplification of 20q13 locus [36,37,38,39].
3.2 Gene mutations
Bioinformatics analysis of MYB (MYB, MYBL1, and MYBL2) genes predicted a total of 45 non-synonymous single-nucleotide polymorphisms (nsSNPs) associated with the high risk of cancer. Some of these mutations, which were located within the helix-turn-helix (HTH) domain, were predicted to be conserved and associated with a shift in DNA-binding specificity of the protein leading to altered protein function [40]. In another study, SNPs (rs619289, rs826943, and rs826944) in MYBL2 promoter regions were identified and associated with an increased susceptibility of breast cancer [39].
3.3 Chromosomal translocations
Another type of alteration in MYB genes results from chromosomal translocation. One among these is the translocation t(6;9) leading to the fusion of MYB and NFIB genes, which generates a chimeric transcripts consisting of exon 14 of MYB fused with the last coding exon of NFIB. This fusion product lacks MYB 3′UTR containing the binding sites for negative regulatory microRNAs leading to the overexpression of the chimeric MYB-NFIB transcript and protein [41, 42]. Such fusions have been detected in primary and metastatic Adenoid Cystic Carcinoma (AdCC) of salivary gland resulting in the overexpression of multiple chimeric variants [43]. Genomic analysis of pediatric low-grade gliomas (PLGG) identified MYB:QK1 fusion transcript that results in MYB activation due to the truncation of c-terminal negative regulatory domain and hemizygous loss of tumor suppressor QK1 expression [44]. More recently, MYB:QK1 fusion was also identified in pediatric high-grade glioma and adult angiocentric glioma (AG) [45]. Massively parallel sequencing analysis of breast adenoid cystic carcinoma lacking the MYB-NFIB fusion has identified the gene rearrangement in MYBL1, such as MYBL1-ACTN1 and MYBL1-NFIB, associated with its overexpression [46]. In addition, a novel MYBL1-NFIB gene fusion as a result of t(8;9) translocation and multiple other rearrangements in the MYBL1 gene has been reported [47]. A fusion transcript of MYBL1 and RAD51B of unknown functional significance is also reported in AdCC resulting from t(8;14) translocation. This fusion leads to antisense transcription of part of the RAD51B intron and truncation of the MYBL1/ A-MYB in the predicted fusion protein [10].
4 Multifaceted roles of MYB proteins in oncogenesis
Cancer development is a multistep process where the transformed cell gains unrestricted proliferation and survival abilities, becomes invasive, leaves the primary site, and establishes itself at secondary locations. This gradual process of evolution is facilitated by accumulation of a series of molecular alterations that work in concert influenced by the external microenvironment [48]. Among these, MYB proteins appear to play a central role in multiple malignancies by affecting the multiple aspects of cancer development as discussed below:
4.1 Cellular plasticity
The earliest work on MYB demonstrated its restricted expression in stem cells and later it was shown that it plays an essential role in the maintenance of the undifferentiated state [49,50,51]. Restored expression of MYB family genes in several malignancies suggests that it might play a similar role to support continued evolution of cancer. Indeed, cancer cells must exhibit adaptive capabilities to sustain their existence under constantly changing microenvironmental conditions. Thus, cellular plasticity is an important attribute that allows cancer cells to survive when they face stressful situations. Epithelial to mesenchymal transition (EMT), an evolutionarily conserved process involved in normal embryonic development and tissue regeneration, bestows such property to the cancer cells [52]. Almost a couple of decades ago, Dvorak et al. reported the role of MYB in the induction of EMT in trunk neural crest cells [53]. Later, Tanno et al.’s group demonstrated that MYB induced the mesenchymal phonotype in embryonic kidney and neuroblastoma cells via transcriptional upregulation of Slug (SLAI2) [54]. The same group later showed that TGFβ-induced EMT in ER( +) breast cancer cells was mediated through MYB, which enhanced the expression of Slug and Bcl-2 [55]. TGFβ/MYB axis is also shown to promote EMT phenotype in esophageal cancer cells [56]. Along with these observations, we also found a role of MYB in EMT in prostate cancer cells [57]. Moreover, in a very recent study, we have observed that MYB plays an essential role in metabolic plasticity of pancreatic cancer cells, especially when these cells are exposed to hypoxia. MYB induced the expression of several glycolytic genes through direct promoter binding and by enhancing the recruitment of HIF-1α on the shared target gene promoters [58].
MYB is crucial for the activation of discoidin domain receptor 2 (DDR2), a key player in matrix stiffness- induced EMT. It has been shown that increased cellular contractility on a stiff matrix recruits MYB and LEF1 to DDR2 promoter and promotes the expression of mesenchymal markers [59]. The aberrant expression of c-MYB has been reported in colorectal cancer (CRC), and its knockout inhibits EMT in CRC cells via a mechanism involving c-fos repression [60]. A negative correlation of MYB with E-cadherin and positive correlation with vimentin in salivary adenoid cystic carcinoma (SACC) also suggests its association with EMT [61]. Single-cell RNA sequencing of metastatic cells from the lungs of hepatoblastoma patients revealed distinct transcriptional signature and significant association of MYBL2 expression with poor prognosis of patients. Overexpression of MYBL2 in hepatoblastoma cancer cells (HCC) promoted the SNAI1 expression and Smad2/3 phosphorylation thereby promoting the EMT and tumorigenesis [62]. A higher expression of MYBL2 has also been reported in metastatic breast cancer cells and aggressive triple-negative subtype (TNBC) and shown to promote EMT [36, 63, 64]. MYBL1 also transcriptionally upregulates TWIST1, a promoter of EMT, in HCC cells [65]. These findings establish that MYB proteins afford cellular plasticity to cancer cells either directly and/or by altering the expression and transcriptional activity of other transcription factors to support their adaptive nature under harsh environments (Fig. 2).
Role of MYB family proteins in cancer cell plasticity. Aberrant expression/activation of MYB proteins promotes epithelial-to-mesenchymal transition either directly modulating the expression of relevant genes or by modulating the expression of known inducers of EMT such as, Slug, SNAI1, and TWIST1. In addition, MYB also regulates the expression of stem cell- associated proteins, including CD34, CXCR4, c-MYC, KLF4, and Nanog, to impart stemness properties. Under hypoxic conditions, MYB expression is induced and interacts with HIF1α to coordinately regulate gene expression associated with metabolic reprogramming
4.2 Cell proliferation and survival
EMT not only imparts aggressive behavioral properties to the cancer cells but also supports their survival. In addition, uncontrolled cell division is a crucial biological process that promotes cancer burden at the primary site and its establishment at the secondary metastatic sites. A very early report on MYB function demonstrated that MYB transcript levels are transiently increased via post-transcriptional mechanism during cell cycle progression in various cell types [66]. Later, this important function of MYB was confirmed by suppressing its expression that resulted in significantly decreased proliferation of myeloid-leukemia cells [67]. Gonda and colleagues demonstrated that MYB is regulated by estrogen/ER signaling and plays a role in the proliferation and survival of ER + breast cancer cells [68]. Transgenic knockout of MYB in murine models of breast cancer revealed its essential role in mammary tumorigenesis and cell survival. MYB promoted the expression of survival associated genes; Bcl-2 and GRP78/BiP in breast cancer cells compared to mammary epithelial cells [69]. In acute myeloid leukemia cells, MYB suppression promoted apoptosis and decreased cell survival due to enhanced expression of pro-apoptotic DRAK2 and increased caspase-9 activity [70]. Indeed, chromatin immunoprecipitation coupled with genome promoter tilling microarrays has demonstrated MYB binding to several gene promoters, including those involved in cell-cycle regulation and survival [71].
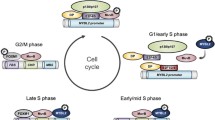
We have also found a role of MYB in cell cycle progression and survival of pancreatic and prostate cancer cells [57, 72, 73]. Recently, we have shown MYB expression is regulated by androgens in a bi-phasic manner mediating its growth-promoting and -suppressive effects [74]. At lower doses, androgens transcriptionally upregulated MYB whereas at high doses, androgens induced the expression of MYB-targeted miRNA miR-150 leading to its repression. The ubiquitous expression of B-MYB in proliferating cells and its regulation by E2F, a cell cycle-related transcription factor, suggest its important function in cell proliferation [75]. Indeed, the elevated MYBL2/B-MYB expression is shown to promote proliferation of bladder [76], liver [77], and lung [78] cancer cells through upregulation of cell-cycle-associated genes. EGFR signaling co-operates with E2F to enhance MYBL2 expression and promotes the proliferation of breast cancer cells [79]. MYBL2 silencing is also shown to inhibit the proliferation of myeloid or lymphoid cells [80]. There are, however, not many reports on the role of MYBL1/A-MYB. The expression of A-MYB is detected in proliferating B-cells, in the S and G2/M phases of the cell cycle, but not in the resting stage suggesting its role in cell cycle progression [81].
4.3 Invasion and metastasis
Most cancer deaths occur due to metastasis, which interferes with the vital organ functions. These attributes are also facilitated through EMT affording invasive capabilities to the cancer cells. A variety of reports have documented the role of MYB in behavioral properties that support the metastatic spread of the cancer cells (Fig. 3). MYB interacts with Wnt effector β-catenin and co-activates the downstream target genes involved in invasion and metastasis of breast cancer cells [82]. Ectopic expression of MYB in human and murine mammary cancer cells is also shown to enhance their potential to invade the Matrigel® by inducing the expression of cathepsin D and MMP9 but downregulating MMP1 [83]. By gain and loss of function studies and using an orthotopic mouse model, we have also demonstrated that MYB promotes invasiveness and metastatic spread of pancreatic cancer cells to the liver, lung, and spleen [72]. MYB knockout in colorectal cancer cells is also shown to inhibit the invasion and metastasis in vivo through a mechanism mediated through the repression of c-fos-induced EMT [60]. Interestingly, another report that measured MYB expression in CRC specimens showed its higher expression in primary lesions relative to the distant metastases [84]. This may suggest that likely tumor cells underwent a reversal of EMT facilitated through MYB downregulation as they established themselves at the secondary site.
Impact of MYB oncoproteins on various aspects of cancer cell growth and metastasis. MYB transcription factors play crucial roles in the regulation of proliferation, growth, invasion, and metastasis in various cancer types. MYB and MYBL2 are shown to promote cell proliferation by inducing the expression of cell cycle-related genes, c-MYC, and supporting the activation signaling cascades responsible for proliferation of the tumor cells at the primary and metastatic sites. MYB transcription factors also promote invasion and metastasis of cancer cells by increasing the expression of proteins involved in the degradation of extracellular matrix (MMP9, Cathepsin D), acquisition of migratory phenotype (DDR2, Wnt/β-catenin signaling, Smad2/3 SNAI1, TWIST, and ANGPT2) and likely anoikis resistance through upregulation of survival-associated genes (Bcl-2, MCL-1, Bcl-xL, Survivin, and Clusterin)
MYBL2 expression is upregulated in bladder cancer (BLCA), the most common malignancy associated with urinary tract system. Silencing of MYBL2 inhibited the migration and invasion of bladder cancer cells in vitro and reduced lung metastases in vivo. These processes involved the interaction of MYBL2 with FOXM1 and transactivation of CDCA3, a protein that could promote Wnt/β-catenin signaling, thus malignant phenotype of BLCA cells [76]. Overexpression of MYBL2 is also detected in non-small-cell lung cancer (NSCLC) and associated with advancing pathological grades and clinical stages. Using gene knockdown and overexpression approaches, it was shown that MYBL2 was involved in cell migration and invasion. RNA-seq analysis revealed an overexpression of various critical genes involved in cancer metastasis, likely through MYBL2-mediated activation of Erk and Akt signaling pathways [85]. In hepatocellular carcinoma, MYBL1/TWIST1 axis promoted aggressive behavior and metastasis in vitro and in vivo [65]. In another report, MYBL1 was shown to cause transcriptional upregulation of ANGPT2 to support neovascularization and metastasis in hepatocellular carcinoma [86].
4.4 Therapy resistance
Therapy resistance in cancer can develop through a variety of innate and acquired mechanisms, including the activation of drug efflux transporters, cell death inhibition (apoptosis suppression), altered drug metabolism, genetic and epigenetic modifications of drug targets, upregulation of DNA repair activity, and activation of bypass pathways [87]. A number of studies have shown the significant role of MYB proteins in supporting the cancer survival can be a major roadblock in the efficacy of anticancer drugs (Fig. 4). Indeed, a higher expression of MYB is reported in derived cisplatin-resistant colorectal cancer cells as compared to the parental cells and its silencing led to the increased sensitivity towards cisplatin-mediated toxicity [88]. Similarly, in another report, overexpression of MYB is shown to activate NF-κB and STAT3 signaling in ovarian cancer cells as a mechanism of cisplatin resistance [89]. In comparison to naïve parental MCF-7 cells, tamoxifen-resistant MCF-7 (TAM-MCF7) breast cancer cells show an upregulated expression of MYB. Repression of MYB in these cells re-sensitized them to the tamoxifen treatment [90]. MYB is also shown to regulate DNA damage and components of the homology-directed repair pathway in ER + ve breast cancer cells suggesting that MYB inhibition along with induction of DNA damage could yield improved therapeutic outcomes [91]. In nasopharyngeal cancer cells, overexpression of c-MYB promotes the resistance to apoptosis induced by ionizing radiation by regulating the PARP cleavage and cleaved caspase-3 [92]. A study from Pekarcikova et al. demonstrated the importance of c-MYB/NOX1/p38 signaling axis in chemoresistance of colorectal cancer cells. Ectopic expression of MYB protected these cells from oxaliplatin- and doxorubicin- induced apoptosis via activation of NOX-1 and p38 MAPK pathway [93]. In glioblastoma cells, ZEB1 is shown to promote MYB expression by downregulating miR200, a MYB-targeting microRNA, which in turn, promotes the expression of O-6-methylguanine-DNA methyltransferase (MGMT) to promote chemoresistance [94].
Promotion of therapeutic resistance by MYB family transcription factors. MYB proteins promote resistance against both targeted (shown in cyan) and non-targeted (shown in red) therapeutic drugs. MYB promotes cisplatin resistance by increasing the expression of NFκB and STAT3, carboplatin/etoposide resistance by increasing the expression of DNA damage response (DDR) pathway-related genes, radiation therapy resistance by preventing the PARP cleavage, and oxaliplatin/doxorubicin resistance by activation of NOX1/p38MAPK pathways in different cancer types. Similarly, MYBL2 contributes to resistance against doxorubicin and tamoxifen drugs by regulating the expression of Bcl-2 and PLK1, PRC1, BIRC5, and HMMR. Development of resistance against targeted therapies including radiation therapy, castration therapy, taxane, and sorafenib drug has also been shown to be associated with activation of MYB family transcription factors
Androgen deprivation therapy (ADT) or castration therapy (CT) has been the mainstay treatment for the advanced and metastatic prostate cancer [95]. Despite an initial response, prostate cancer relapses in most patients as a castration-resistant disease through aberrant activation of androgen receptor (AR) signaling [96]. In our studies, we found that MYB-overexpressing prostate cancer cells survived well under androgen-deprived condition and retained the expression of AR-responsive gene, KLK3/PSA [57]. Later, we demonstrated that MYB interacted with AR and retained it in the nucleus to sustain its transcriptional activity under androgen-reduced condition. Further, these findings were confirmed in an orthotopic model by castrating the mice. We observed that MYB-overexpressing cells sustained their growth following castration and quickly resumed the serum PSA levels [97]. In additional studies, we have observed racially disparate expression of MYB in prostate and ovarian malignancies associated with patient’s prognosis and disease recurrence (unpublished data; [98]). An upregulation of B-MYB is also reported in CRPC tissues and cell lines, where it supports the resistance to androgen-deprivation therapy and taxane drugs [99]. Another study suggested that B-MYB contributed to castration-resistance by activating the YAP1 transcription [100]. Overexpression of B-MYB in T-lymphoblastic cells enhanced the expression of Bcl-2 and resistance to killing by doxorubicin, ceramide, and dexamethasone [101]. MYBL2 transcriptionally upregulates Clusterin expression, which mediates at least in part, the antiapoptotic effects of B-MYB and confer doxorubicin resistance in neuroblastoma cells [102]. It is also shown to contribute to tamoxifen resistance in breast cancer cells by upregulating genes associated with survival [103]. A pan-cancer analysis of B-MYB function using various bioinformatics approaches also predicted its role in chemoresistance and immune escape via regulation of apoptosis and immune-checkpoint-associated genes [104]. Elevated expression of MYBL2 in lung adenocarcinoma drives the expression of a set of genes that mediate replication stress response and promote error-prone DNA repair which were also coupled with loss of cell cycle check-point regulators TP53 and RB1 [105]. Moreover, MYBL2 upregulates the expression of cell division cycle associated 8 (CDCA8) protein, a component of chromosomal passenger complex (CPC) and confers olaparib and cisplatin resistance in ovarian cancer cells by regulating the apoptosis and homologous recombination-mediated DNA damage repair [106]. The information on the role of A-MYB in therapy resistance is scarce. Hepatocellular carcinoma cells expressing higher A-MYB exhibit resistance to sorafenib and its inhibition abrogates this resistance [86]. Thus, it appears that MYB family proteins are good targets for achieving the therapeutic enhancement of existing targeted or non-targeted anticancer drugs. Moreover, expression of MYB proteins could also be used as a potential biomarker for therapeutic planning and predicting the response to chemo- and immune therapies.
5 Role of MYB in stromal remodeling and its impact on tumor cell plasticity, metastasis, and therapy resistance
Tumor cells continuously interact with other cells in the tumor microenvironment (TME), such as fibroblasts, endothelial cells, and immune cells, throughout the course of cancer evolution. These dynamic interactions create a tumor-supportive environment by modifying the phenotypes and makeup of the stromal cells as well as altering the composition of the extracellular matrix [107, 108]. We have shown that MYB-overexpression in pancreatic cancer cells promotes desmoplasia by increasing the secretion of sonic hedgehog (SHH) and adrenomedullin (ADM) [109]. MYB transcriptionally upregulated SHH and ADM, which activated pancreatic stellate cells (PSC) allowing their transition to myofibroblasts. A greater abundance of collagen-1, fibronectin, and α-Smooth muscle actin (α-SMA)-positive fibroblast cells was recorded in orthotopic xenografts derived from MYB-expressing pancreatic cancer (PC) cells than those from MYB-silenced cells [109]. This is interesting since extensive desmoplasia in pancreatic tumors has been reported to be a significant cause of chemoresistance. A seminal study by Olive et al. demonstrated that desmoplastic stroma restricted the delivery of gemcitabine to the tumor cells in a genetically engineered mouse model of pancreatic adenocarcinoma [110]. Increased desmoplasia also creates a more hypoxic environment that is known to cause EMT, promote metastasis, and reduce the efficacy of anticancer drugs in many cancers [111, 112].
High MYB expression in colorectal cancer has been associated with reduced infiltration of activated T-cells near the tumor and poor relapse-free survival. This reciprocal relationship indicatse that MYB could be useful as a marker to predict patient response to immunotherapy [113]. Further, MYB-promoted desmoplasia could also restrict immune cell infiltration and/or may be inhibitory to their proliferation and survival within the tumor microenvironment [114, 115]. In a recent study, loss of structural integrity of desmoplastic matrix promoted efficacy of tumor antigen (mesothelin)-targeted CAR-T cells and anti-PD-1 antibody therapies in solid tumors [115]. CXCL12, the ligand for chemokine receptor CXCR4 and abundantly expressed by activated tumor-associated fibroblasts, promotes fibrosis, and inhibition of CXCR4 is shown promote T-lymphocyte infiltration and induce an integrated immune response in breast, pancreatic and colorectal cancers [71, 116, 117]. Recent studies revealed that senescent fibroblasts near the tumor cells undergo changes in the expression of genes associated with cell cycle, metabolism, and secretory proteins leading to gain of a pro-inflammatory secretory phenotype [118]. The secretion of pro-tumorigenic SASP factors, such as osteopontin (OPN), IL-6, and IL-8, is regulated by MYB and promotes the growth and migration of cancer cells [119,120,121]. MYB expression is also upregulated in macrophages upon co-culture with the breast cancer cells. Further, it transcriptionally represses the 5-Lipoxygenase (5-LO), a key enzyme in leukotrienes biosynthesis, and leads to reduced T-cell recruitment favoring tumor progression [122]. Thus, MYB can not only affect tumor cell features through direct tumor-intrinsic actions but also by modulating the tumor microenvironment.
6 Conclusion
Identifying and characterizing the genes involved in tumorigenesis are crucial to develop novel molecular approaches for cancer management. From early reports demonstrating the expression of MYB in hematopoietic stem cells, the field has moved fast demonstrating the aberrant expression and/or activation of MYB family proteins in multiple malignancies. Further, we have learnt a great deal regarding the involvement of MYB proteins in multiple oncogenic processes, including proliferation, survival, stemness, invasion, and stromal remodeling. Also, these proteins appear to support the growth of cancer cells under harsh environmental conditions and fight therapeutic insults. More interestingly, racial differences in MYB expression are also reported suggesting its role in racially-disparate clinical outcomes. Thus, targeting MYB could be a useful strategy to effectively manage cancer and narrow the disparity gaps. Having said that, there is still a lot to learn about MYB functions in cancers. It is important to scan the complete spectrum of target genes of MYB proteins in different cancers and at different stages of cancer development. Most transcription factors work in concert with other proteins and the differential protein–protein interactions impact the transcriptional output. For example, we have found an interaction of MYB with HIF-1α, which is expressed at the protein level under an oxygen-reduced environment. Our initial data suggest that this interaction promotes metabolic reprogramming and helps the cells to switch from a proliferative state to a slow growing state, which is more invasive. This is a great example of the role of MYB in tumor cell plasticity and should be explored further in different tumor types and when the tumor cells are exposed to other environmental stressors, including therapeutic treatments. This new knowledge could be highly useful to develop approaches for therapeutic targeting of cancer-supporting MYB functions and therapeutic enhancement of existing treatment modalities. It is also important to delineate the molecular mechanisms involved in controlling the expression and activation of MYB proteins. Such an information can also provide additional therapeutic opportunities and even help in developing strategies for cancer prevention.
Data Availability
Data sharing is not applicable to this article.
References
Davidson, C. J., Tirouvanziam, R., Herzenberg, L. A., & Lipsick, J. S. (2005). Functional evolution of the vertebrate Myb gene family: B-Myb, but neither A-Myb nor c-Myb, complements Drosophila Myb in hemocytes. Genetics, 169, 215–229. https://doi.org/10.1534/genetics.104.034132
Ciciro, Y., & Sala, A. (2021). MYB oncoproteins: emerging players and potential therapeutic targets in human cancer. Oncogenesis, 10, 19. https://doi.org/10.1038/s41389-021-00309-y
Klempnauer, K. H., Gonda, T. J., & Bishop, J. M. (1982). Nucleotide sequence of the retroviral leukemia gene v-myb and its cellular progenitor c-myb: The architecture of a transduced oncogene. Cell, 31, 453–463. https://doi.org/10.1016/0092-8674(82)90138-6
Boyle, W. J., Lipsick, J. S., Reddy, E. P., & Baluda, M. A. (1983). Identification of the leukemogenic protein of avian myeloblastosis virus and of its normal cellular homologue. Proceedings of the National Academy of Science U S A, 80, 2834–2838. https://doi.org/10.1073/pnas.80.10.2834
Oh, I. H., & Reddy, E. P. (1999). The myb gene family in cell growth, differentiation and apoptosis. Oncogene, 18, 3017–33. https://doi.org/10.1038/sj.onc.1202839
Nomura, N., Takahashi, M., Matsui, M., Ishii, S., Date, T., Sasamoto, S., et al. (1988). Isolation of human cDNA clones of myb-related genes, A-myb and B-myb. Nucleic Acids Research, 16, 11075–11089. https://doi.org/10.1093/nar/16.23.11075
Ramsay, R. G., & Gonda, T. J. (2008). MYB function in normal and cancer cells. Nature Reviews Cancer, 8, 523–534. https://doi.org/10.1038/nrc2439
Trauth, K., Mutschler, B., Jenkins, N. A., Gilbert, D. J., Copeland, N. G., & Klempnauer, K. H. (1994). Mouse A-myb encodes a trans-activator and is expressed in mitotically active cells of the developing central nervous system, adult testis and B lymphocytes. The EMBO Journal, 13, 5994–6005. https://doi.org/10.1002/j.1460-2075.1994.tb06945.x
Kumar, A., Baker, S. J., Lee, C. M., & Reddy, E. P. (2003). Molecular mechanisms associated with the regulation of apoptosis by the two alternatively spliced products of c-Myb. Molecular and Cellular Biology, 23, 6631–45. https://doi.org/10.1128/MCB.23.18.6631-6645.2003
Brayer, K. J., Frerich, C. A., Kang, H., & Ness, S. A. (2016). Recurrent fusions in MYB and MYBL1 define a common, transcription factor-driven oncogenic pathway in salivary gland adenoid cystic carcinoma. Cancer Discovery, 6, 176–187. https://doi.org/10.1158/2159-8290.CD-15-0859
Chen, X., Feng, J., Zhang, Y., Liu, J., Zhang, L., Zeng, P., et al. (2023). MYBL2 alternative splicing-related genetic variants reduce the risk of triple-negative breast cancer in the Chinese population. Frontiers in Genetics, 14, 1150976. https://doi.org/10.3389/fgene.2023.1150976
O’Rourke, J. P., & Ness, S. A. (2008). Alternative RNA splicing produces multiple forms of c-Myb with unique transcriptional activities. Molecular and Cellular Biology, 28, 2091–2101. https://doi.org/10.1128/MCB.01870-07
Ogata, K., Kanei-Ishii, C., Sasaki, M., Hatanaka, H., Nagadoi, A., Enari, M., et al. (1996). The cavity in the hydrophobic core of Myb DNA-binding domain is reserved for DNA recognition and trans-activation. Natural Structural Biology, 3, 178–187. https://doi.org/10.1038/nsb0296-178
Ogata, K., Hojo, H., Aimoto, S., Nakai, T., Nakamura, H., Sarai, A., et al. (1992). Solution structure of a DNA-binding unit of Myb: A helix-turn-helix-related motif with conserved tryptophans forming a hydrophobic core. Proceedings of the National Academy of Sciences U S A, 89, 6428–6432. https://doi.org/10.1073/pnas.89.14.6428
Facchinetti, V., Loffarelli, L., Schreek, S., Oelgeschlager, M., Luscher, B., Introna, M., et al. (1997). Regulatory domains of the A-Myb transcription factor and its interaction with the CBP/p300 adaptor molecules. Biochemical Journal, 324(Pt 3), 729–36. https://doi.org/10.1042/bj3240729
Rushton, J. J., & Ness, S. A. (2001). The conserved DNA binding domain mediates similar regulatory interactions for A-Myb, B-Myb, and c-Myb transcription factors. Blood Cells, Molecules, and Diseases, 27, 459–63. https://doi.org/10.1006/bcmd.2001.0405
Ogata, K., Morikawa, S., Nakamura, H., Sekikawa, A., Inoue, T., Kanai, H., et al. (1994). Solution structure of a specific DNA complex of the Myb DNA-binding domain with cooperative recognition helices. Cell, 79, 639–648. https://doi.org/10.1016/0092-8674(94)90549-5
Bergholtz, S., Andersen, T. O., Andersson, K. B., Borrebaek, J., Luscher, B., & Gabrielsen, O. S. (2001). The highly conserved DNA-binding domains of A-, B- and c-Myb differ with respect to DNA-binding, phosphorylation and redox properties. Nucleic Acids Research, 29, 3546–3556. https://doi.org/10.1093/nar/29.17.3546
Nakagoshi, H., Takemoto, Y., & Ishii, S. (1993). Functional domains of the human B-myb gene product. Journal of Biological Chemistry, 268, 14161–14167. https://doi.org/10.1016/S0021-9258(19)85222-5
Sleeman, J. P. (1993). Xenopus A-myb is expressed during early spermatogenesis. Oncogene, 8, 1931–1941.
Katzen, A. L., Kornberg, T. B., & Bishop, J. M. (1985). Isolation of the proto-oncogene c-myb from D. melanogaster. Cell, 41, 449–456. https://doi.org/10.1016/S0092-8674(85)80018-0
Kanei-Ishii, C., MacMillan, E. M., Nomura, T., Sarai, A., Ramsay, R. G., Aimoto, S., et al. (1992). Transactivation and transformation by Myb are negatively regulated by a leucine-zipper structure. Proceedings National Academy Sciences U S A, 89, 3088–3092. https://doi.org/10.1073/pnas.89.7.3088
Nomura, T., Sakai, N., Sarai, A., Sudo, T., Kanei-Ishii, C., Ramsay, R. G., et al. (1993). Negative autoregulation of c-Myb activity by homodimer formation through the leucine zipper. Journal of Biological Chemistry, 268, 21914–21923. https://doi.org/10.1016/S0021-9258(20)80628-0
Takahashi, T., Nakagoshi, H., Sarai, A., Nomura, N., Yamamoto, T., & Ishii, S. (1995). Human A-myb gene encodes a transcriptional activator containing the negative regulatory domains. FEBS Letters, 358, 89–96. https://doi.org/10.1016/0014-5793(94)01402-M
Ziebold, U., & Klempnauer, K. H. (1997). Linking Myb to the cell cycle: Cyclin-dependent phosphorylation and regulation of A-Myb activity. Oncogene, 15, 1011–1019. https://doi.org/10.1038/sj.onc.1201282
Ramsay, R. G., Morrice, N., Van Eeden, P., Kanagasundaram, V., Nomura, T., De Blaquiere, J., et al. (1995). Regulation of c-Myb through protein phosphorylation and leucine zipper interactions. Oncogene, 11, 2113–2120.
Wijeratne, T. U., Guiley, K. Z., Lee, H. W., Muller, G. A., & Rubin, S. M. (2022). Cyclin-dependent kinase-mediated phosphorylation and the negative regulatory domain of transcription factor B-Myb modulate its DNA binding. Journal of Biological Chemistry, 298, 102319. https://doi.org/10.1016/j.jbc.2022.102319
Tomita, A., Watanabe, T., Kosugi, H., Ohashi, H., Uchida, T., Kinoshita, T., et al. (1998). Truncated c-Myb expression in the human leukemia cell line TK-6. Leukemia, 12, 1422–1429. https://doi.org/10.1038/sj.leu.2401113
Wallrapp, C., Muller-Pillasch, F., Solinas-Toldo, S., Lichter, P., Friess, H., Buchler, M., et al. (1997). Characterization of a high copy number amplification at 6q24 in pancreatic cancer identifies c-myb as a candidate oncogene. Cancer Research, 57, 3135–3139.
Kauraniemi, P., Hedenfalk, I., Persson, K., Duggan, D. J., Tanner, M., Johannsson, O., et al. (2000). MYB oncogene amplification in hereditary BRCA1 breast cancer. Cancer Research, 60, 5323–5328.
Tatevossian, R. G., Tang, B., Dalton, J., Forshew, T., Lawson, A. R., Ma, J., et al. (2010). MYB upregulation and genetic aberrations in a subset of pediatric low-grade gliomas. Acta Neuropathologica, 120, 731–743. https://doi.org/10.1007/s00401-010-0763-1
Schumacher, S. E., Shim, B. Y., Corso, G., Ryu, M. H., Kang, Y. K., Roviello, F., et al. (2017). Somatic copy number alterations in gastric adenocarcinomas among Asian and Western patients. PLoS ONE, 12, e0176045. https://doi.org/10.1371/journal.pone.0176045
Edwards, J., Krishna, N. S., Witton, C. J., & Bartlett, J. M. (2003). Gene amplifications associated with the development of hormone-resistant prostate cancer. Clinical Cancer Research, 9, 5271–5281.
Dong, G., Mao, Q., Yu, D., Zhang, Y., Qiu, M., Dong, G., et al. (2017). Integrative analysis of copy number and transcriptional expression profiles in esophageal cancer to identify a novel driver gene for therapy. Sciences Reports, 7, 42060. https://doi.org/10.1038/srep42060
Ramkissoon, L. A., Horowitz, P. M., Craig, J. M., Ramkissoon, S. H., Rich, B. E., Schumacher, S. E., et al. (2013). Genomic analysis of diffuse pediatric low-grade gliomas identifies recurrent oncogenic truncating rearrangements in the transcription factor MYBL1. Proceedings National Academy Sciences U S A, 110, 8188–8193. https://doi.org/10.1073/pnas.130025211
Bayley, R., Ward, C., & Garcia, P. (2020). MYBL2 amplification in breast cancer: Molecular mechanisms and therapeutic potential. Biochimica et Biophysica Acta - Reviews on Cancer, 1874, 188407. https://doi.org/10.1016/j.bbcan.2020.188407
Koynova, D. K., Jordanova, E. S., Milev, A. D., Dijkman, R., Kirov, K. S., Toncheva, D. I., et al. (2007). Gene-specific fluorescence in-situ hybridization analysis on tissue microarray to refine the region of chromosome 20q amplification in melanoma. Melanoma Research, 17, 37–41. https://doi.org/10.1097/CMR.0b013e3280141617
Tanner, M. M., Grenman, S., Koul, A., Johannsson, O., Meltzer, P., Pejovic, T., et al. (2000). Frequent amplification of chromosomal region 20q12-q13 in ovarian cancer. Clinical Cancer Research, 6, 1833–1839.
Shi, H., Bevier, M., Johansson, R., Grzybowska, E., Chen, B., Eyfjord, J. E., et al. (2011). Single nucleotide polymorphisms in the 20q13 amplicon genes in relation to breast cancer risk and clinical outcome. Breast Cancer Research and Treatment, 130, 905–916. https://doi.org/10.1007/s10549-011-1600-5
Lim, S. W., Tan, K. J., Azuraidi, O. M., Sathiya, M., Lim, E. C., Lai, K. S., et al. (2021). Functional and structural analysis of non-synonymous single nucleotide polymorphisms (nsSNPs) in the MYB oncoproteins associated with human cancer. Scientific Reports, 11, 24206. https://doi.org/10.1038/s41598-021-03624-x
Stenman, G., Andersson, M. K., & Andren, Y. (2010). New tricks from an old oncogene: Gene fusion and copy number alterations of MYB in human cancer. Cell Cycle, 9, 2986–2995. https://doi.org/10.4161/cc.9.15.12515
Persson, M., Andren, Y., Mark, J., Horlings, H. M., Persson, F., & Stenman, G. (2009). Recurrent fusion of MYB and NFIB transcription factor genes in carcinomas of the breast and head and neck. Proceedings National Academy Science U S A, 106, 18740–18744. https://doi.org/10.1073/pnas.0909114106
Mitani, Y., Li, J., Rao, P. H., Zhao, Y. J., Bell, D., Lippman, S. M., et al. (2010). Comprehensive analysis of the MYB-NFIB gene fusion in salivary adenoid cystic carcinoma: Incidence, variability, and clinicopathologic significance. Clinical Cancer Research, 16, 4722–4731. https://doi.org/10.1158/1078-0432.CCR-10-0463
Bandopadhayay, P., Ramkissoon, L. A., Jain, P., Bergthold, G., Wala, J., Zeid, R., et al. (2016). MYB-QKI rearrangements in angiocentric glioma drive tumorigenicity through a tripartite mechanism. Nature Genetics, 48, 273–282. https://doi.org/10.1038/ng.3500
Suh, Y. Y., Lee, K., Shim, Y. M., Phi, J. H., Park, C. K., Kim, S. K., et al. (2023). MYB/MYBL1::QKI fusion-positive diffuse glioma. Journal of Neuropathology and Experimental Neurology, 82, 250–260. https://doi.org/10.1093/jnen/nlac123
Kim, J., Geyer, F. C., Martelotto, L. G., Ng, C. K., Lim, R. S., Selenica, P., et al. (2018). MYBL1 rearrangements and MYB amplification in breast adenoid cystic carcinomas lacking the MYB-NFIB fusion gene. The Journal of Pathology, 244, 143–150. https://doi.org/10.1002/path.5006
Mitani, Y., Liu, B., Rao, P. H., Borra, V. J., Zafereo, M., Weber, R. S., et al. (2016). Novel MYBL1 gene rearrangements with recurrent MYBL1-NFIB Fusions in salivary adenoid cystic carcinomas lacking t(6;9) translocations. Clinical Cancer Research, 22, 725–733. https://doi.org/10.1158/1078-0432.CCR-15-2867-T
Hanahan, D., & Weinberg, R. A. (2011). Hallmarks of cancer: The next generation. Cell, 144, 646–674. https://doi.org/10.1016/j.cell.2011.02.013
Kuehl, W. M., Bender, T. P., Stafford, J., McClinton, D., Segal, S., & Dmitrovsky, E. (1988). Expression and function of the c-myb oncogene during hematopoietic differentiation. Current Topics in Microbiology and Immunology, 141, 318–323. https://doi.org/10.1007/978-3-642-74006-0_42
Dyson, P. J., Poirier, F., & Watson, R. J. (1989). Expression of c-myb in embryonal carcinoma cells and embryonal stem cells. Differentiation, 42, 24–27. https://doi.org/10.1111/j.1432-0436.1989.tb00603.x
Introna, M., Luchetti, M., Castellano, M., Arsura, M., & Golay, J. (1994). The myb oncogene family of transcription factors: Potent regulators of hematopoietic cell proliferation and differentiation. Seminars in Cancer Biology, 5, 113–124.
Huang, Y., Hong, W., & Wei, X. (2022). The molecular mechanisms and therapeutic strategies of EMT in tumor progression and metastasis. Journal of Hematology & Oncology, 15, 129. https://doi.org/10.1186/s13045-022-01347-8
Karafiat, V., Dvorakova, M., Krejci, E., Kralova, J., Pajer, P., Snajdr, P., et al. (2005). Transcription factor c-Myb is involved in the regulation of the epithelial-mesenchymal transition in the avian neural crest. Cellular and Molecular Life Sciences, 62, 2516–2525. https://doi.org/10.1007/s00018-005-5297-7
Tanno, B., Sesti, F., Cesi, V., Bossi, G., Ferrari-Amorotti, G., Bussolari, R., et al. (2010). Expression of Slug is regulated by c-Myb and is required for invasion and bone marrow homing of cancer cells of different origin. Journal of Biological Chemistry, 285, 29434–29445. https://doi.org/10.1074/jbc.M109.089045
Cesi, V., Casciati, A., Sesti, F., Tanno, B., Calabretta, B., & Raschella, G. (2011). TGFbeta-induced c-Myb affects the expression of EMT-associated genes and promotes invasion of ER+ breast cancer cells. Cell Cycle, 10, 4149–4161. https://doi.org/10.4161/cc.10.23.18346
Cheng, J., Wu, K., Yang, Q., Zhu, Z., & Zhao, H. (2023). RNF6 activates TGF-beta1/c-Myb pathway to promote EMT in esophageal squamous cell carcinoma. Frontiers in Oncology, 13, 1081333. https://doi.org/10.3389/fonc.2023.1081333
Srivastava, S. K., Bhardwaj, A., Singh, S., Arora, S., McClellan, S., Grizzle, W. E., et al. (2012). Myb overexpression overrides androgen depletion-induced cell cycle arrest and apoptosis in prostate cancer cells, and confers aggressive malignant traits: Potential role in castration resistance. Carcinogenesis, 33, 1149–1157. https://doi.org/10.1093/carcin/bgs134
Anand, S., Khan, M. A., Zubair, H., Sudan, S. K., Vikramdeo, K. S., Deshmukh, S. K., et al. (2023). MYB sustains hypoxic survival of pancreatic cancer cells by facilitating metabolic reprogramming. EMBO Reports, 24, e55643. https://doi.org/10.15252/embr.202255643
Kim, D., You, E., Jeong, J., Ko, P., Kim, J. W., & Rhee, S. (2017). DDR2 controls the epithelial-mesenchymal-transition-related gene expression via c-Myb acetylation upon matrix stiffening. Scientific Reports, 7, 6847. https://doi.org/10.1038/s41598-017-07126-7
Qu, X., Yan, X., Kong, C., Zhu, Y., Li, H., Pan, D., et al. (2019). c-Myb promotes growth and metastasis of colorectal cancer through c-fos-induced epithelial-mesenchymal transition. Cancer Science, 110, 3183–3196. https://doi.org/10.1111/cas.14141
Xu, L. H., Zhao, F., Yang, W. W., Chen, C. W., Du, Z. H., Fu, M., et al. (2019). MYB promotes the growth and metastasis of salivary adenoid cystic carcinoma. International Journal of Oncology, 54, 1579–1590. https://doi.org/10.3892/ijo.2019.4754
Wei, M., Yang, R., Ye, M., Zhan, Y., Liu, B., Meng, L., et al. (2022). MYBL2 accelerates epithelial-mesenchymal transition and hepatoblastoma metastasis via the Smad/SNAI1 pathway. American Journal of Cancer Research, 12, 1960–1981.
Tao, D., Pan, Y., Jiang, G., Lu, H., Zheng, S., Lin, H., et al. (2015). B-Myb regulates snail expression to promote epithelial-to-mesenchymal transition and invasion of breast cancer cell. Medical Oncology, 32, 412. https://doi.org/10.1007/s12032-014-0412-y
Fiscon, G., Pegoraro, S., Conte, F., Manfioletti, G., & Paci, P. (2021). Gene network analysis using SWIM reveals interplay between the transcription factor-encoding genes HMGA1, FOXM1, and MYBL2 in triple-negative breast cancer. FEBS Letters, 595, 1569–1586. https://doi.org/10.1002/1873-3468.14085
Xie, B., Liu, Y., Zhao, Z., Liu, Q., Wang, X., Xie, Y., et al. (2020). MYB proto-oncogene-like 1-TWIST1 axis promotes growth and metastasis of hepatocellular carcinoma cells. Molecular Therapy Oncolytics, 18, 58–69. https://doi.org/10.1016/j.omto.2020.05.016
Thompson, C. B., Challoner, P. B., Neiman, P. E., & Groudine, M. (1986). Expression of the c-myb proto-oncogene during cellular proliferation. Nature, 319, 374–80. https://doi.org/10.1038/319374a0
Anfossi, G., Gewirtz, A. M., & Calabretta, B. (1989). An oligomer complementary to c-myb-encoded mRNA inhibits proliferation of human myeloid leukemia cell lines. Proceedings of the Nationsl Academy of Sciences U S A, 86, 3379–3383. https://doi.org/10.1073/pnas.86.9.3379
Drabsch, Y., Hugo, H., Zhang, R., Dowhan, D. H., Miao, Y. R., Gewirtz, A. M., et al. (2007). Mechanism of and requirement for estrogen-regulated MYB expression in estrogen-receptor-positive breast cancer cells. Proceedings of the National Academy Sciences U S A, 104, 13762–13767. https://doi.org/10.1073/pnas.0700104104
Miao, R. Y., Drabsch, Y., Cross, R. S., Cheasley, D., Carpinteri, S., Pereira, L., et al. (2011). MYB is essential for mammary tumorigenesis. Cancer Research, 71, 7029–7037. https://doi.org/10.1158/0008-5472.CAN-11-1015
Ye, P., Zhao, L., & Gonda, T. J. (2013). The MYB oncogene can suppress apoptosis in acute myeloid leukemia cells by transcriptional repression of DRAK2 expression. Leukemia Research, 37, 595–601. https://doi.org/10.1016/j.leukres.2013.01.012
Quintana, A. M., Liu, F., O’Rourke, J. P., & Ness, S. A. (2011). Identification and regulation of c-Myb target genes in MCF-7 cells. BMC Cancer, 11, 30. https://doi.org/10.1186/1471-2407-11-30
Srivastava, S. K., Bhardwaj, A., Arora, S., Singh, S., Azim, S., Tyagi, N., et al. (2015). MYB is a novel regulator of pancreatic tumour growth and metastasis. British Journal of Cancer, 113, 1694–1703. https://doi.org/10.1038/bjc.2015.400
Azim, S., Zubair, H., Srivastava, S. K., Bhardwaj, A., Zubair, A., Ahmad, A., et al. (2016). Deep sequencing and in silico analyses identify MYB-regulated gene networks and signaling pathways in pancreatic cancer. Scientific Reports, 6, 28446. https://doi.org/10.1038/srep28446
Acharya, S., Anand, S., Khan, M. A., Zubair, H., Srivastava, S. K., Singh, S., et al. (2023). Biphasic transcriptional and posttranscriptional regulation of MYB by androgen signaling mediates its growth control in prostate cancer. Journal of Biological Chemistry, 299, 102725. https://doi.org/10.1016/j.jbc.2022.102725
Lyon, J., Robinson, C., & Watson, R. (1994). The role of Myb proteins in normal and neoplastic cell proliferation. Critical Reviews in Oncogenesis, 5, 373–388. https://doi.org/10.1615/critrevoncog.v5.i4.30
Liu, W., Shen, D., Ju, L., Zhang, R., Du, W., Jin, W., et al. (2022). MYBL2 promotes proliferation and metastasis of bladder cancer through transactivation of CDCA3. Oncogene, 41, 4606–4617. https://doi.org/10.1038/s41388-022-02456-x
Wei, T., Weiler, S. M. E., Toth, M., Sticht, C., Lutz, T., Thomann, S., et al. (2019). YAP-dependent induction of UHMK1 supports nuclear enrichment of the oncogene MYBL2 and proliferation in liver cancer cells. Oncogene, 38, 5541–5550. https://doi.org/10.1038/s41388-019-0801-y
Xiong, Y. C., Wang, J., Cheng, Y., Zhang, X. Y., & Ye, X. Q. (2020). Overexpression of MYBL2 promotes proliferation and migration of non-small-cell lung cancer via upregulating NCAPH. Molecular and Cellular Biochemistry, 468, 185–193. https://doi.org/10.1007/s11010-020-03721-x
Hanada, N., Lo, H. W., Day, C. P., Pan, Y., Nakajima, Y., & Hung, M. C. (2006). Co-regulation of B-Myb expression by E2F1 and EGF receptor. Molecular Carcinogenesis, 45, 10–17. https://doi.org/10.1002/mc.20147
Arsura, M., Introna, M., Passerini, F., Mantovani, A., & Golay, J. (1992). B-myb antisense oligonucleotides inhibit proliferation of human hematopoietic cell lines. Blood, 79, 2708–2716. https://doi.org/10.1182/blood.V79.10.2708.2708
Golay, J., Broccoli, V., Lamorte, G., Bifulco, C., Parravicini, C., Pizzey, A., et al. (1998). The A-Myb transcription factor is a marker of centroblasts in vivo. The Journal of Immunology, 160, 2786–2793. https://doi.org/10.4049/jimmunol.160.6.2786
Li, Y., Jin, K., van Pelt, G. W., van Dam, H., Yu, X., Mesker, W. E., et al. (2016). c-Myb enhances breast cancer invasion and metastasis through the Wnt/beta-catenin/axin2 pathway. Cancer Research, 76, 3364–3375. https://doi.org/10.1158/0008-5472.CAN-15-2302
Knopfova, L., Benes, P., Pekarcikova, L., Hermanova, M., Masarik, M., Pernicova, Z., et al. (2012). ”c-Myb regulates matrix metalloproteinases 1/9, and cathepsin D: implications for matrix-dependent breast cancer cell invasion and metastasis”. Molecular Cancer, 11, 15. https://doi.org/10.1186/1476-4598-11-15
Tichy, M., Knopfova, L., Jarkovsky, J., Pekarcikova, L., Veverkova, L., Vlcek, P., et al. (2016). Overexpression of c-Myb is associated with suppression of distant metastases in colorectal carcinoma. Tumour Biology, 37, 10723–10729. https://doi.org/10.1007/s13277-016-4956-7
Y. Jin, H. Zhu, W. Cai, X. Fan, Y. Wang, Y. Niu, et al. (2017). "B-Myb is up-regulated and promotes cell growth and motility in non-small cell lung cancer," Int J Mol Sci, 18 https://doi.org/10.3390/ijms18060860
Zhu, J., Wu, Y., Yu, Y., Li, Y., Shen, J., & Zhang, R. (2022). MYBL1 induces transcriptional activation of ANGPT2 to promote tumor angiogenesis and confer sorafenib resistance in human hepatocellular carcinoma. Cell Death Disease, 13, 727. https://doi.org/10.1038/s41419-022-05180-2
Mansoori, B., Mohammadi, A., Davudian, S., Shirjang, S., & Baradaran, B. (2017). The different mechanisms of cancer drug resistance: a brief review. Advanced Pharmaceutical Bulletin, 7, 339–348. https://doi.org/10.15171/apb.2017.041
Funato, T., Satou, J., Kozawa, K., Fujimaki, S., Miura, T., & Kaku, M. (2001). Use of c-myb antisense oligonucleotides to increase the sensitivity of human colon cancer cells to cisplatin. Oncology Reports, 8, 807–10. https://doi.org/10.3892/or.8.4.807
Tian, M., Tian, D., Qiao, X., Li, J., & Zhang, L. (2019). Modulation of Myb-induced NF-kB -STAT3 signaling and resulting cisplatin resistance in ovarian cancer by dietary factors. Journal of Cellular Physiology, 234, 21126–21134. https://doi.org/10.1002/jcp.28715
Gao, Y., Zhang, W., Liu, C., & Li, G. (2019). miR-200 affects tamoxifen resistance in breast cancer cells through regulation of MYB. Scientific Reports, 9, 18844. https://doi.org/10.1038/s41598-019-54289-6
Yang, R. M., Nanayakkara, D., Kalimutho, M., Mitra, P., Khanna, K. K., Dray, E., et al. (2019). MYB regulates the DNA damage response and components of the homology-directed repair pathway in human estrogen receptor-positive breast cancer cells. Oncogene, 38, 5239–5249. https://doi.org/10.1038/s41388-019-0789-3
Wang, W., Wu, S., Shi, Y., Miao, Y., Luo, X., Ji, M., et al. (2015). c-MYB regulates cell growth and DNA damage repair through modulating MiR-143. FEBS Letters, 589, 555–564. https://doi.org/10.1016/j.febslet.2015.01.012
Pekarcikova, L., Knopfova, L., Benes, P., & Smarda, J. (2016). c-Myb regulates NOX1/p38 to control survival of colorectal carcinoma cells. Cellular Signalling, 28, 924–936. https://doi.org/10.1016/j.cellsig.2016.04.007
Siebzehnrubl, F. A., Silver, D. J., Tugertimur, B., Deleyrolle, L. P., Siebzehnrubl, D., Sarkisian, M. R., et al. (2013). The ZEB1 pathway links glioblastoma initiation, invasion and chemoresistance. EMBO Molecular Medicine, 5, 1196–1212. https://doi.org/10.1002/emmm.201302827
P. Posdzich, C. Darr, T. Hilser, M. Wahl, K. Herrmann, B. Hadaschik, et al. (2023). "Metastatic Prostate cancer-a review of current treatment options and promising new approaches." Cancers (Basel), 15 https://doi.org/10.3390/cancers15020461.
Chandrasekar, T., Yang, J. C., Gao, A. C., & Evans, C. P. (2015). Mechanisms of resistance in castration-resistant prostate cancer (CRPC). Translational Andrology Urology, 4, 365–380. https://doi.org/10.3978/j.issn.2223-4683.2015.05.02
Srivastava, S. K., Khan, M. A., Anand, S., Zubair, H., Deshmukh, S. K., Patel, G. K., et al. (2022). MYB interacts with androgen receptor, sustains its ligand-independent activation and promotes castration resistance in prostate cancer. British Journal of Cancer, 126, 1205–1214. https://doi.org/10.1038/s41416-021-01641-1
Miree, O., Srivastava, S. K., Khan, M. A., Sameeta, F., Acharya, S., Ndetan, H., et al. (2021). Clinicopathologic significance and race-specific prognostic association of MYB overexpression in ovarian cancer. Scientific Reports, 11, 12901. https://doi.org/10.1038/s41598-021-92352-3
Yoshikawa, Y., Stopsack, K. H., Wang, X. V., Chen, Y. H., Mazzu, Y. Z., Burton, F., et al. (2022). Increased MYBL2 expression in aggressive hormone-sensitive prostate cancer. Molecular Oncology, 16, 3994–4010. https://doi.org/10.1002/1878-0261.13314
Li, Q., Wang, M., Hu, Y., Zhao, E., Li, J., Ren, L., et al. (2021). MYBL2 disrupts the Hippo-YAP pathway and confers castration resistance and metastatic potential in prostate cancer. Theranostics, 11, 5794–5812. https://doi.org/10.7150/thno.56604
Grassilli, E., Salomoni, P., Perrotti, D., Franceschi, C., & Calabretta, B. (1999). Resistance to apoptosis in CTLL-2 cells overexpressing B-Myb is associated with B-Myb-dependent bcl-2 induction. Cancer Research, 59, 2451–2456.
Sala, A., Bettuzzi, S., Pucci, S., Chayka, O., Dews, M., & Thomas-Tikhonenko, A. (2009). Regulation of CLU gene expression by oncogenes and epigenetic factors implications for tumorigenesis. Advances in Cancer Research, 105, 115–132. https://doi.org/10.1016/S0065-230X(09)05007-6
Li, X., Zhang, X., Wu, C. C., Li, P. P., Fu, Y. M., Xie, L. H., et al. (2021). The role of MYB proto-oncogene like 2 in tamoxifen resistance in breast cancer. Journal of Molecular Histology, 52, 21–30. https://doi.org/10.1007/s10735-020-09920-6
Chen, X., Lu, Y., Yu, H., Du, K., Zhang, Y., Nan, Y., et al. (2021). Pan-cancer analysis indicates that MYBL2 is associated with the prognosis and immunotherapy of multiple cancers as an oncogene. Cell Cycle, 20, 2291–2308. https://doi.org/10.1080/15384101.2021.1982494
Morris, B. B., Wages, N. A., Grant, P. A., Stukenberg, P. T., Gentzler, R. D., Hall, R. D., et al. (2020). MYBL2-driven transcriptional programs link replication stress and error-prone DNA repair with genomic instability in lung adenocarcinoma. Frontiers in Oncology, 10, 585551. https://doi.org/10.3389/fonc.2020.585551
Qi, G., Zhang, C., Ma, H., Li, Y., Peng, J., Chen, J., et al. (2021). CDCA8, targeted by MYBL2, promotes malignant progression and olaparib insensitivity in ovarian cancer. American Journal of Cancer Research, 11, 389–415.
Bussard, K. M., Mutkus, L., Stumpf, K., Gomez-Manzano, C., & Marini, F. C. (2016). Tumor-associated stromal cells as key contributors to the tumor microenvironment. Breast Cancer Research, 18, 84. https://doi.org/10.1186/s13058-016-0740-2
Sund, M., & Kalluri, R. (2009). Tumor stroma derived biomarkers in cancer. Cancer and Metastasis Reviews, 28, 177–183. https://doi.org/10.1007/s10555-008-9175-2
Bhardwaj, A., Srivastava, S. K., Singh, S., Tyagi, N., Arora, S., Carter, J. E., et al. (2016). MYB promotes desmoplasia in pancreatic cancer through direct transcriptional up-regulation and cooperative action of sonic hedgehog and adrenomedullin. Journal of Biological Chemistry, 291, 16263–16270. https://doi.org/10.1074/jbc.M116.732651
Olive, K. P., Jacobetz, M. A., Davidson, C. J., Gopinathan, A., McIntyre, D., Honess, D., et al. (2009). Inhibition of Hedgehog signaling enhances delivery of chemotherapy in a mouse model of pancreatic cancer. Science, 324, 1457–1461. https://doi.org/10.1126/science.1171362
Shah, V. M., Sheppard, B. C., Sears, R. C., & Alani, A. W. (2020). Hypoxia: Friend or foe for drug delivery in pancreatic cancer. Cancer Letters, 492, 63–70. https://doi.org/10.1016/j.canlet.2020.07.041
Yuen, A., & Diaz, B. (2014). The impact of hypoxia in pancreatic cancer invasion and metastasis. Hypoxia (Auckl), 2, 91–106. https://doi.org/10.2147/HP.S52636
Millen, R., Malaterre, J., Cross, R. S., Carpinteri, S., Desai, J., Tran, B., et al. (2016). Immunomodulation by MYB is associated with tumor relapse in patients with early stage colorectal cancer. Oncoimmunology, 5, e1149667. https://doi.org/10.1080/2162402X.2016.1149667
Watt, J., & Kocher, H. M. (2013). The desmoplastic stroma of pancreatic cancer is a barrier to immune cell infiltration. Oncoimmunology, 2, e26788. https://doi.org/10.4161/onci.26788
Xiao, Z., Todd, L., Huang, L., Noguera-Ortega, E., Lu, Z., Huang, L., et al. (2023). Desmoplastic stroma restricts T cell extravasation and mediates immune exclusion and immunosuppression in solid tumors. Nature Communications, 14, 5110. https://doi.org/10.1038/s41467-023-40850-5
Chen, I. X., Chauhan, V. P., Posada, J., Ng, M. R., Wu, M. W., Adstamongkonkul, P., et al. (2019). Blocking CXCR4 alleviates desmoplasia, increases T-lymphocyte infiltration, and improves immunotherapy in metastatic breast cancer. Proceeding of the National Academy Sciences U S A, 116, 4558–4566. https://doi.org/10.1073/pnas.1815515116
Biasci, D., Smoragiewicz, M., Connell, C. M., Wang, Z., Gao, Y., Thaventhiran, J. E. D., et al. (2020). CXCR4 inhibition in human pancreatic and colorectal cancers induces an integrated immune response. Proceedings of the National Academy Sciences U S A, 117, 28960–28970. https://doi.org/10.1073/pnas.2013644117
Coppe, J. P., Desprez, P. Y., Krtolica, A., & Campisi, J. (2010). The senescence-associated secretory phenotype: The dark side of tumor suppression. Annual Review of Pathology: Mechanisms of Disease, 5, 99–118. https://doi.org/10.1146/annurev-pathol-121808-102144
Flanagan, K. C., Alspach, E., Pazolli, E., Parajuli, S., Ren, Q., Arthur, L. L., et al. (2018). c-Myb and C/EBPbeta regulate OPN and other senescence-associated secretory phenotype factors. Oncotarget, 9, 21–36. https://doi.org/10.18632/oncotarget.22940
Wu, X., Tao, P., Zhou, Q., Li, J., Yu, Z., Wang, X., et al. (2017). IL-6 secreted by cancer-associated fibroblasts promotes epithelial-mesenchymal transition and metastasis of gastric cancer via JAK2/STAT3 signaling pathway. Oncotarget, 8, 20741–20750. https://doi.org/10.18632/oncotarget.15119
K. Jin, N. B. Pandey, and A. S. Popel 2017 "Crosstalk between stromal components and tumor cells of TNBC via secreted factors enhances tumor growth and metastasis." Oncotarget 8, 60210–60222 https://doi.org/10.18632/oncotarget.19417
Ringleb, J., Strack, E., Angioni, C., Geisslinger, G., Steinhilber, D., Weigert, A., et al. (2018). Apoptotic cancer cells suppress 5-lipoxygenase in tumor-associated macrophages. The Journal of Immunology, 200, 857–868. https://doi.org/10.4049/jimmunol.1700609
Acknowledgements
The authors would like to acknowledge the Department of Pathology and University of South Alabama Mitchell Cancer Institute for providing the necessary resources. The figures in this study have been created using biorender.
Funding
The authors are supported, in part, by funding from NIH/NCI (R01CA224306, R01CA231925) and Department of Defense (DOD)/US Army (HT94252310452 and W81XWH-22–1-0913).
Author information
Authors and Affiliations
Contributions
Concept, Planning, and Supervision: APS.
Literature search, review, and writing original draft: SA, KSV, SKS, AS, SA, and MAK.
Discussion of cited literature and feedback on the manuscript draft: SS.
Revision and final review of the manuscript: SA, KSV, SKS, AS, SA, MAK, SS, APS.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Conflict of interest
The authors declare ‘no conflict of interest’.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Anand, S., Vikramdeo, K.S., Sudan, S.K. et al. From modulation of cellular plasticity to potentiation of therapeutic resistance: new and emerging roles of MYB transcription factors in human malignancies. Cancer Metastasis Rev 43, 409–421 (2024). https://doi.org/10.1007/s10555-023-10153-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10555-023-10153-8