Abstract
Purpose
Breast cancer stem cells (CSCs) are a small subpopulation of cancer cells that have high capability for self-renewal, differentiation, and tumor initiation. CSCs are resistant to chemotherapy and radiotherapy, and are responsible for cancer recurrence and metastasis.
Methods
By utilizing a panel of breast cancer cells and mammospheres culture as cell-based screening platforms, we performed high-throughput chemical library screens to identify agents that are effective against breast CSCs and non-CSCs. The hit molecules were paired with conventional chemotherapy to evaluate the combinatorial treatment effects on breast CSCs and non-CSCs.
Results
We identified a total of 193 inhibitors that effectively targeting both breast CSCs and non-CSCs. We observed that histone deacetylase inhibitors (HDACi) synergized conventional chemotherapeutic agents (i.e., doxorubicin and cisplatin) in targeting breast CSCs and non-CSCs simultaneously. Further analyses revealed that quisinostat, a potent inhibitor for class I and II HDACs, potentiated doxorubicin-induced cytotoxicity in both breast CSCs and non-CSCs derived from the basal-like (MDA-MB-468 and HCC38), mesenchymal-like (MDA-MB-231), and luminal-like breast cancer (MCF-7). It was also observed that the basal-like breast CSCs and non-CSCs were more sensitive to the co-treatment of quisinostat with doxorubicin compared to that of the luminal-like breast cancer subtype.
Conclusion
In conclusion, this study demonstrates the potential of HDACi as therapeutic options, either as monotherapy or in combination with chemotherapeutics against refractory breast cancer.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Increasing evidence indicates that many solid cancers, including breast cancer, contain a small subpopulation of cancer stem cells (CSCs) capable of self-renewal and differentiation into various cell types, contributing to cellular heterogeneity in tumors [1]. Breast CSCs are inherently resistant to chemotherapy and radiotherapy, and are a major factor contributing to treatment resistance, relapse, and metastasis [2]. The elucidation of pathways that regulate these cells has led to the identification of several potential therapeutic targets, including Wnt [1, 2], Notch [1, 2], Hedgehog (Hh) [1, 2], mTOR [3, 4], CDK [5, 6], and IGF-1R [7,8,9] signaling.
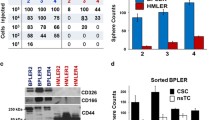
The initial description of human breast CSCs involved the prospective isolation of the CSC populations based on the positive expression of epithelial-specific antigen (ESA) and CD44 cell surface markers and the absence of CD24 expression [10]. The isolated breast CSCs (ESA+/CD44+/CD24−) were able to generate tumors in immunosuppressed non-obese diabetic/severe combined immunodeficient (NOD/SCID) mice with as little as 100 cells. In contrast, the non-CSCs isolated from the same tumors were non-tumorigenic and required 100-fold more cells to generate a tumor in the NOD/SCID mice. Importantly, the tumors generated from the isolated breast CSCs recapitulated the heterogeneity of the original tumor upon transplantation in mice, demonstrating the plasticity of the breast CSCs [10].
Recently, it has been demonstrated that established breast cancer cell lines contain cell hierarchies driven by a population that expresses cancer stem cell markers [11, 12]. Indeed, breast CSCs isolated from primary cultures of hormone-dependent and hormone-independent breast tumors as well as the MCF7 cell line could be cultured under anchorage-independent conditions to form clonal mammospheres [13, 14]. The mammosphere model system has been established in several breast cancer cell lines and represents a robust in vitro model for studying breast cancer initiation and screening for CSC-targeting agents [14]. Importantly, the mammospheres in vitro assays have been validated using xenotransplantation models, which are considered to be the gold standard assay for cancer stem cells. Using the mammosphere culture, our group and others have previously identified metformin as a selective breast CSC inhibitor [14,15,16,17,18].
Although development of CSC-targeted agents are promising, CSC-specific agents (e.g., salinomycin and abamectin) alone might not be effective in reducing the tumor bulk (non-CSCs) because these inhibitors are less potent compared to conventional chemotherapeutic agents [19,20,21]. In this case, dual targeting agents or combination therapy consisting of CSC inhibitors and conventional cytotoxic agents are expected to better eradicate both CSCs and non-CSCs simultaneously, and hence improve the clinical outcomes.
In the current study, we conducted a high-throughput screens for small chemical inhibitors that kill breast CSCs and non-CSCs simultaneously. We observed that histone deacetylase inhibitors (HDACi) alone or in combination with conventional chemotherapy were able to inhibit both breast CSCs and non-CSCs simultaneously. Thus, this combination could be considered as an effective therapeutic strategy for breast cancer treatment.
Materials and methods
Cell lines and cell culture
MDA-MB-468, MDA-MB-231, HCC38, and MCF-7 breast cancer cell lines were obtained from the American Type Culture Collection (ATCC; Manassas, VA, USA). Cells were maintained in RPMI 1640 (Corning Incorporated, New York, USA) containing 10% fetal bovine serum (Sigma-Aldrich, St. Louis, MO, USA), 100 IU/mL penicillin and 100 μg/mL streptomycin (Biowest, Nuaillé, France). All breast cancer cells were kept in culture for less than 6 months and maintained in logarithmic growth in a humidified 37 °C, 5% CO2 incubator.
Mammosphere culture
Mammosphere culture was performed as recommended by Stem Cell Technologies. Briefly, all the cells were grown in MammoCult™ Basal Medium (Stem Cell Technologies, Vancouver, BC, Canada) supplemented with MammoCult™ Proliferation Supplement (Stem Cell Technologies, Vancouver, BC, Canada), 4 µg/mL heparin (Stem Cell Technologies, Vancouver, BC, Canada), 0.48 µg/mL hydrocortisone (Sigma-Aldrich, St. Louis, MO, USA), 100 IU/mL penicillin, and 100 µg/mL streptomycin (Biowest, Nuaillé, France). The single cell suspensions of breast cancer cells were cultured in clear 6-well ultra-low attachment multiple well plates (Corning Incorporated, New York, USA) at humidified 37 °C, 5% CO2 for 5 days. Mammospheres were collected by gentle centrifugation and the pellets were gently triturated into single sphere suspensions with trypsin–EDTA (Sigma-Aldrich, St. Louis, MO, USA). Enrichment of greater than 80% of CD44+/CD24−/low CSC mammospheres from existing breast cancer cell lines is observed within 5 days of mammosphere culture [13, 14].
Chemical library screening
A chemical library consisting of 1672 diverse bioactive small molecules was obtained from Selleckchem (Houston, TX, USA) to screen for candidate molecules targeting the non-CSCs and/or CSCs in breast cancer. Both MDA-MB-468 breast CSCs and non-CSCs in the logarithmic growth phase were seeded overnight at a density of 5000 cells/well, respectively, and treated with 10 µM of each compound. The cells were then incubated at 37 °C in a humidified 5% CO2 incubator for 72 h. Cell proliferation was examined using CellTiter-Glo® Luminescent Cell Viability Assay (Promega Corporation, Madison, WI, USA) according to the manufacturer protocol [22]. The luminescent signal was measured by SpectraMax® M3 Multi-Mode Microplate Reader (Molecular Devices, Sunnyvale, CA, USA). Compounds that induced growth inhibition of more than 50% in CSCs and non-CSCs were considered as “hits”. The Redundant siRNA Activity (RSA) analysis method was employed to examine the rank distribution of the collective activities based on the known target(s) of the compounds, and p values were calculated to indicate the statistical significance of hit compounds with the same targets being remarkably distributed toward the top ranking slots [23, 24].
Cell proliferation assays
The effects of drug combination treatment on breast CSCs and non-CSCs cell proliferation were determined by CellTiter 96® AQueous One Solution Cell Proliferation Assay (also known as MTS assay; Promega Corporation, Madison, WI, USA) and methyl thiazolyl tetrazolium (MTT) assay (Sigma-Aldrich, St Louis, MI, USA), respectively [25, 26]. Briefly, breast CSCs and non-CSCs were seeded overnight in 96-well plates at a density of 5000 cells/well and treated with either HDACi alone (quisinostat, trichostatin A, givinostat, entinostat, belinostat, and vorinostat), chemotherapeutic agents alone (doxorubicin, cisplatin, and paclitaxel), or in combination for 72 h. The absorbance of the formazan solution as a result of cell-mediated reduction of MTT/MTS by the viable cells was determined using a SpectraMax® M3 Multi-Mode Microplate Reader (Molecular Devices, Sunnyvale, CA, USA) or Tecan Infinite® F200 Microplate Reader (Tecan Group, Ltd., CH, Männedorf, Switzerland) at 490 nm or 570/630 nm, respectively.
CD44 and CD24 flowcytometry
Analysis of breast CSCs populations were performed on single cell suspensions using flow cytometry as described previously [14]. Briefly, cells were stained with CD44-APC and CD24-PE (BD Biosciences, San Jose, CA, USA) for 30 min, washed, and re-suspended in PBS supplemented with 1% FBS. CSCs populations in breast cancer cell lines were identified as CD44+/CD24−. All cells were analyzed using a FACSCalibur flow cytometer and the CellQuest Pro software (version 5.1.1; BD Biosciences, USA) for acquisition and Flowing Software (Version 2.5.0; University of Turku, Turku, Finland) for data analysis.
Drug combination analyses
The combinatory effects of HDACi and chemotherapeutic agents on breast CSCs and non-CSCs were evaluated using the Chou–Talalay method and Highest Single Agent (HSA) models. Multiple drug dose–effect calculations, combination index (CI), and drug reduction index (DRI) were generated using CalcuSyn version 2.1 software (Biosoft, Cambridge, UK) according to the Chou–Talalay method, in which CI values of < 1, = 1, and > 1 indicate synergism, additive effect, and antagonism respectively as previously described [27,28,29]. DRI values were used to describe the dose reduction potential of the agents when combined. In principle, dose reduction potential with DRI > 1 can be clinically valuable in reducing the risk of developing drug toxicity towards the host while retaining the therapeutic efficacy in a synergistic drug combination [27, 30, 31]. Drug interaction was further analyzed using the HSA model (Combenefit software, Cancer Research UK Cambridge Institute) [32].
Results
Identification of chemical inhibitors targeting breast CSCs and non-CSCs through high-throughput phenotypic screens
A chemical library consisting of 1672 diverse bioactive small molecules was used for rapid identification of candidate molecules that could target both breast CSCs and non-CSCs. To determine the inhibitory effects of small molecules against breast CSCs and non-CSCs, a cell-based high-throughput screen was performed using breast CSC-enriched mammmospheres and parental breast cancer cells of MDA-MB-468 (Fig. 1a). Of note, unlike the MDA-MB-231 and SUM159 basal mesenchymal-like cell line (also known as Basal B cell line) which mainly showed CD44+/CD24− feature, the triple-negative (ER, PR and HER2 negative) MDA-MB-468 basal epithelial cells (also known as Basal A cell line) mainly showed CD44+/CD24+ feature with EGFR amplification and p53 mutation, closely resembling the refractory basal-like tumors in patients [14, 33,34,35,36,37].
High-throughput phenotypic screens identifying 193 bioactive small molecules targeting both MDA-MB-468 breast CSCs and non-CSCs. a By combining the screening data from MDA-MB-468 breast CSCs and non-CSCs, compounds that exerted selective inhibitory effects against both breast CSCs and non-CSCs were identified. Green circles, molecules targeting breast CSCs only; blue circles, molecules targeting breast non-CSCs; red circles, molecules targeting both breast CSCs and non-CSCs; gray circles, molecules lacking anti-proliferative activities. b Compounds which inhibited both breast CSCs and non-CSCs (viability < 50%) were considered as “hits”. c Compound similarity-based clustering of hits. The dendrogram of chemical structure similarities among the hits was constructed using extended-connectivity fingerprint 4 (ECFP 4) module of the C-SPADE [72]. Note that the hits are structurally diverse and do not share chemotype similarity to compounds within the same target class, with the exception of the EGFR inhibitors
As expected, the most malignant basal mesenchymal cell line MDA-MB-231 mainly showed CD44 +/CD24− feature (Fig. 1a,b), while the other three cell lines did not, in accordance with the previous findings showing that CD44 +/CD24−/low is a stem-like marker highly related to the malignance of breast cancer [19, 38, 39]. We also found that the luminal A cell line MCF-7 and the HER2-OE cell line SK-BR-3 were mainly composed of cells bearing the CD44−/CD24+ phenotype, while the basal epithelial cell line MDA-MB-468 mainly showed CD44+/CD24+ (Fig. 1a, b).
Out of the 1672 compounds tested, a total of 193 (11.5%) compounds were found to target both CSCs and non-CSCs and were identified as hits (Fig. 1b and Table 1). These include ispinesib (SB-715992) which has been recently shown to target both treatment resistant glioblastoma CSCs and non-CSCs [38]; YM155 which inhibits lung and breast CSCs through attenuation of EGFR and NFκB pathways [39, 40]; and nanchangmycin which exhibits apoptotic and anti-proliferative activities against MCF-7 breast CSCs [41]. These findings independently validate the results of our primary screens.
Next, we sought to investigate whether the hits belong to compound classes that share common molecular targets or structure similarity. We ranked the targets of the hits using the RSA method and identified Bcl-2, mTOR, CDK, HDAC, and EGFR as the top five targets that when inhibited, elicit growth inhibitory effects against both breast CSCs and non-CSCs of MDA-MB-468 (Table 2). Importantly, most of the identified hits (with the exception of EGFR inhibitors) are structurally diverse and do not share chemotype similarity to compounds within the same target class (Fig. 1c). These findings suggest that the observed inhibitory effects are likely to be driven by the inhibition of the molecular targets and not by the chemotype similarity. Indeed, some of the top ranking targets, such as mTOR and CDK, have also been previously implicated in the regulation of cell survival in both CSCs and non-CSCs in breast cancer, indicating that these pathways are required the survival of both CSCs and non-CSCs [3, 5, 6, 42, 43].
HDAC inhibitors synergize chemotherapeutic sensitivity in breast CSCs and non-CSCs
Since recent reports have shown that epigenetic mechanisms can influence breast cancer stemness, and the utility of HDACi as epigenetic drugs for targeting both CSCs and non-CSCs have been demonstrated in hematological and other solid malignancies [44,45,46], we sought to investigate whether HDACi could synergize conventional chemotherapeutic agents in targeting both CSCs and non-CSCs in breast cancer.
We selected six HDACi, including quisinostat, trichostatin A, givinostat, entinostat, belinostat, and vorinostat (SAHA), for further testing. Quisinostat, trichostatin A, givinostat, belinostat, and vorinostat are hydroxamate-based pan-HDACi, whereas entinostat is a benzamide-based class I-specific HDACi [45,46,47]. Of note, vorinostat and belinostat have been approved by FDA for treatment of peripheral T-cell lymphoma, while quisinostat, entinostat, and givinostat are currently under phase 2 clinical trials [48].
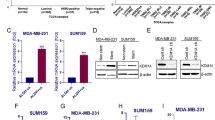
Consistent with previous studies, breast CSCs conferred marked resistance towards cisplatin (approximately threefold), doxorubicin (approximately fourfold), and paclitaxel (approximately 25-fold) in MDA-MB-468, HCC38, MDA-MB-231, and MCF-7 cells (Supplemental Figure 1 and Supplemental Table 1). Interestingly, combination with HDACi synergizes doxorubicin (Figs. 2 and 3; Table 3) and, to a lesser extent, cisplatin sensitivity in both MDA-MB-468 CSCs and non-CSCs (Fig. 4 and Table 4). In contrast, combinations of HDACi and paclitaxel exhibited selective synergism in the non-CSCs but not in CSCs of MDA-MB-468 cells (Supplemental Figure 2 and Supplemental Table 2).
Combinatory effects of HDACi and doxorubicin in breast CSCs and non-CSCs. a The effects of HDACi and doxorubicin alone or in combination on the viability of breast CSCs and non-CSCs were determined 72 h following treatment. Points represent mean ± S.D. of at least three independent experiments. b The Fa-CI plots of HDACi and doxorubicin combination on breast CSCs and non-CSCs was generated using the Chou–Talalay’s CI method [27]. The plots showed the CI versus the fraction of breast CSCs and non-CSCs that were inhibited by the combined treatment of HDACi and doxorubicin at the stated concentration ratio. The combinations were synergistic when CI values were < 1
Synergistic effects of HDACi and doxorubicin on MDA-MB-468 breast CSCs and non-CSCs. MDA-MB-468 breast CSCs and non-CSCs were treated with doxorubicin and/or HDACi for 72 h. Dose–response surface curves and synergy of each combination was assessed using the HSA model (effect-based approach), as implemented in Combenefit software [32]. Level of synergism (blue) or antagonism (red) at each concentration is represented by color scale bar. All experiments were conducted at least three times
Synergistic effects of HDACi and cisplatin on MDA-MB-468 breast CSCs and non-CSCs. MDA-MB-468 breast CSCs and non-CSCs were treated with cisplatin and/or HDACi for 72 h. Dose–response surface curves and synergy of each combination was assessed using the HSA model (effect-based approach) using Combenefit software [32]. Level of synergism (blue) or antagonism (red) at each concentration is represented by color scale bar. All experiments were conducted at least three times
Quisinostat synergizes doxorubicin sensitivity in different subtypes of breast CSCs and non-CSCs
Given that recent clinical studies demonstrated that quisinostat in combination with chemotherapeutic agents exhibits high efficacy and good tolerability in treatment of various advanced solid tumors [49,50,51], we investigated whether CSCs and non-CSCs population is affected by treatment of doxorubicin and/or quisinostat. We showed that treatment of MDA-MB-468 cells with doxorubicin alone induced significant reduction in the number of non-CSCs (p < 0.01, Student’s t test), while the total number of CSCs remained unchanged, suggesting that doxorubicin target mainly the non-stem breast cancer cells (Fig. 5). In contrast, treatment of cells with quisinostat reduced both the CSCs and non-CSCs of MDA-MB-468. Importantly, the combination of doxorubicin and quisinostat further reduced the number of CSCs and non-CSCs compared to single agent alone, suggesting that the combination might exert synergistic effects against both cell populations simultaneously.
Synergistic effects of quisinostat on doxorubicin sensitivity in both CSCs and non-CSCs. MDA-MB-468 cells were treated with 50 nM doxorubicin and/or 50 nM quisinostat for 72 h followed by CD44 and CD24 flowcytometry. a Representative dot plot (20,000 events) of CD44 and CD24 flowcytometry. Numbers accompanying inset boxes indicate the percentage of CD44+/CD24− breast CSCs population (mean ± S.D.) in MDA-MB-468 cells following drug treatment. b Percentage and total number of CSCs and non-CSCs following doxorubicin and/or quisinostat treatment in MDA-MB-468 cells. Bars represent the mean ± S.D. of 3 independent experiments. * Indicate statistical significance compared with control cells (p < 0.01, Student’s t test). #Represents statistically significant improvement with the combination compared with the single agent alone (p < 0.01, Student’s t test)
To test this hypothesis, we investigated whether quisinostat will synergize doxorubicin sensitivity in CSCs and non-CSCs derived from different subtypes of breast cancers using the combination index method [27, 28]. Indeed, combination of quisinostat and doxorubicin exhibited significant synergism in both CSCs and non-CSCs derived from the basal-like HCC38 cells, the mesenchymal-like MDA-MB-231 cells, and the luminal-like MCF-7 cells (Table 5).
Together, our results demonstrated that quisinostat could enhance the doxorubicin-induced cytotoxicity in both breast CSCs and non-CSCs, regardless of the breast cancer subtypes. Given the favorable DRI trends, our data also indicated that such combination regimen could be exploited for the dose reduction potentials of doxorubicin and quisinostat in breast cancer (Supplemental Table 3).
Discussion
In this study, we identified 193 small inhibitors that could target both breast CSCs and non-CSCs. This list includes inhibitors targeting Bcl-2, mTOR, CDK, HDAC, and EGFR signaling. We demonstrated that HDACi synergize cisplatin and doxorubicin sensitivity in both CSCs and non-CSCs derived from distinct subtypes of breast cancer cells.
HDACs are important epigenetic enzymes that catalyze the removal of acetyl groups from lysine residues enzymes in histone, thereby inducing chromatin condensation and transcriptional repression [45, 52]. To date, a total of 18 mammalian HDACs have been identified and classified into five phylogenetic classes: class I (HDAC1, HDAC2, HDAC3, HDAC8), class IIA (HDAC4, HDAC5, HDAC7, HDAC9), class IIB (HDAC6, HDAC10), class III (Sirtuins 1–7), and class IV (HDAC11) [53]. Previous studies have shown that different HDACs are differentially regulated in various cancers and the aberrant recruitment of HDACs by oncogenic DNA-fusion proteins or repressive transcription factors can drive tumorigenesis [46].
Indeed, HDAC1, HDAC2, HDAC3, and HDAC6 have shown to be overexpressed in breast cancer [54,55,56,57], while HDAC1 and HDAC7 are found to be specifically overexpressed in CSCs when compared to non-CSCs in breast and ovarian cancers [56]. Furthermore, knockdown of individual HDACs can inhibit the proliferation and survival of tumor cells, as well as retard the aggressiveness of breast cancer cells [56, 58, 59].
Given the important role of HDAC in regulating the CSC phenotype in cancers, it is not surprising that a large number of structurally diverse HDACi have been developed in recent years to target the epigenetic abnormalities associated with refractory cancers. In general, HDACi can be classified as either pan-HDACi or class-specific HDACi [45, 60]. The pan-HDACi targets HDACs from class I, II, and IV, whereas the class-specific HDACi targets only HDACs from either class I or class II [60]. To date, a large number of HDACi have been developed, many of which are undergoing clinical testing, and some which have been approved for clinical use. For example, romidepsin is approved by the FDA for the treatment of cutaneous T-cell lymphoma (CTCL) and peripheral T-cell lymphoma (PTCL), vorinostat for the treatment of CTCL, belinostat for the treatment of PTCL, and panobinostat for the treatment of multiple myeloma [47, 61].
It has also been reported that a number of broad-spectrum HDACi suppresses the CSCs population in different cancer cell lines through various mechanisms. It has been shown that AR-42 (OSU-HDAC42), a pan-HDACi, induces apoptosis in leukemic stem cells by inhibiting NFκB and HSP90 functions, but not in the normal hematopoietic stem and progenitor cells [62]. Vorinostat has been shown to reduce the self-renewal capacity of pancreatic CSCs by inhibiting of miR-34a-Notch and epithelial–mesenchymal transition (EMT) signaling [63], and reverse cisplatin resistance in head and neck CSCs by downregulating targeting Nanog expression [64]. It has also been reported that abexinostat, another pan-HDACi, reduces the breast CSCs that have low abundance of the long non-coding RNA Xist by inducing cellular differentiation into non-CSCs [65]. More recently, it was shown that the newly developed pan-HDACi, MC1742, and MC2625 are effective in inducing growth arrest, apoptosis, and CSC differentiation in sarcomas [66].
Despite these advances, the mechanism by which HDACi suppresses the CSCs has not been fully elucidated [46]. Mechanistically, the antitumor activity of HDACi arises due to their effects on epigenetic regulation, leading to the reprogramming of gene expression in cancer cells in a manner which promotes growth arrest, differentiation, and apoptosis [67]. However, how these changes affect the CSCs remains to be elucidated. One hypothesis is that HDACi may suppress the ability to self-renew and promote CSC differentiation, hence increasing CSC sensitivity to chemotherapy/radiotherapy [68]. This is supported by the recent evidence showing that non-CSCs may be induced into drug-resistant CSCs in response to chemotherapy through upregulation of HDAC expression [69]. Hence, inhibition of HDACs may compromise the plasticity of CSC and restore sensitivity to chemotherapeutic drugs [69]. Alternatively, HDACi can also exert their biological effects by regulating the acetylation of a variety of non-histone targets in different signaling pathways relevant to CSC homeostasis [45, 61].
Regardless, it is important to note that non-CSCs are able to undergo EMT and de-differentiate into CSCs [19, 21, 70, 71]. Hence, targeting CSCs alone might lead to initial tumor shrinkage, but eventually relapse if one or more of the non-CSCs are able to de-differentiate into a CSC [19,20,21]. Thus, new drug combinations that kill both CSCs and non-CSCs will be more effective in the long run.
In conclusion, our studies suggest that the combination of HDACi (e.g., quisinostat) and doxorubicin can target both breast CSCs and non-CSCs simultaneously. Therefore, HDACi/doxorubicin combination could be an effective adjuvant therapy for the treatment of refractory or drug-resistant cancers.
References
Batlle E, Clevers H (2017) Cancer stem cells revisited. Nat Med 23(10):1124–1134. https://doi.org/10.1038/nm.4409
Lawson DA, Bhakta NR, Kessenbrock K, Prummel KD, Yu Y, Takai K, Zhou A, Eyob H, Balakrishnan S, Wang CY, Yaswen P, Goga A, Werb Z (2015) Single-cell analysis reveals a stem-cell program in human metastatic breast cancer cells. Nature 526(7571):131–135. https://doi.org/10.1038/nature15260
Zhu Y, Zhang X, Liu Y, Zhang S, Liu J, Ma Y, Zhang J (2012) Antitumor effect of the mTOR inhibitor everolimus in combination with trastuzumab on human breast cancer stem cells in vitro and in vivo. Tumor Biol 33(5):1349–1362
Zhou J, Wulfkuhle J, Zhang H, Gu P, Yang Y, Deng J, Margolick JB, Liotta LA, Petricoin E, Zhang Y (2007) Activation of the PTEN/mTOR/STAT3 pathway in breast cancer stem-like cells is required for viability and maintenance. Proc Natl Acad Sci 104(41):16158–16163
Boyer M, Cheng T (2008) The CDK inhibitors: potential targets for therapeutic stem cell manipulations? Gene Ther 15(2):117
Han YK, Lee JH, Park G-Y, Chun SH, Han JY, Kim SD, Lee J, Lee C-W, Yang K, Lee CG (2013) A possible usage of a CDK4 inhibitor for breast cancer stem cell-targeted therapy. Biochem Biophys Res Commun 430(4):1329–1333
Chang W-W, Lin R-J, Yu J, Chang W-Y, Fu C-H, Lai AC-Y, Yu J-C, Alice LY (2013) The expression and significance of insulin-like growth factor-1 receptor and its pathway on breast cancer stem/progenitors. Breast Cancer Res 15(3):R39
Christopoulos PF, Msaouel P, Koutsilieris M (2015) The role of the insulin-like growth factor-1 system in breast cancer. Mol Cancer Res 14(1):1
Hormones TE, Group BCC (2010) Insulin-like growth factor 1 (IGF1), IGF binding protein 3 (IGFBP3), and breast cancer risk: pooled individual data analysis of 17 prospective studies. Lancet Oncol 11(6):530
Al-Hajj M, Wicha MS, Benito-Hernandez A, Morrison SJ, Clarke MF (2003) Prospective identification of tumorigenic breast cancer cells. Proc Natl Acad Sci USA 100(7):3983–3988. https://doi.org/10.1073/pnas.0530291100
Charafe-Jauffret E, Ginestier C, Iovino F, Wicinski J, Cervera N, Finetti P, Hur MH, Diebel ME, Monville F, Dutcher J, Brown M, Viens P, Xerri L, Bertucci F, Stassi G, Dontu G, Birnbaum D, Wicha MS (2009) Breast cancer cell lines contain functional cancer stem cells with metastatic capacity and a distinct molecular signature. Cancer Res 69(4):1302–1313. https://doi.org/10.1158/0008-5472.CAN-08-2741
Fillmore CM, Kuperwasser C (2008) Human breast cancer cell lines contain stem-like cells that self-renew, give rise to phenotypically diverse progeny and survive chemotherapy. Breast Cancer Res BCR 10(2):R25. https://doi.org/10.1186/bcr1982
Ponti D, Costa A, Zaffaroni N, Pratesi G, Petrangolini G, Coradini D, Pilotti S, Pierotti MA, Daidone MG (2005) Isolation and in vitro propagation of tumorigenic breast cancer cells with stem/progenitor cell properties. Cancer Res 65(13):5506–5511. https://doi.org/10.1158/0008-5472.CAN-05-0626
Soo JS, Ng CH, Tan SH, Malik RA, Teh YC, Tan BS, Ho GF, See MH, Taib NA, Yip CH, Chung FF, Hii LW, Teo SH, Leong CO (2015) Metformin synergizes 5-fluorouracil, epirubicin, and cyclophosphamide (FEC) combination therapy through impairing intracellular ATP production and DNA repair in breast cancer stem cells. Apoptosis 20(10):1373–1387. https://doi.org/10.1007/s10495-015-1158-5
Hirsch HA, Iliopoulos D, Tsichlis PN, Struhl K (2009) Metformin selectively targets cancer stem cells, and acts together with chemotherapy to block tumor growth and prolong remission. Cancer Res 69(19):7507–7511. https://doi.org/10.1158/0008-5472.CAN-09-2994
Vazquez-Martin A, Oliveras-Ferraros C, Del Barco S, Martin-Castillo B, Menendez JA (2011) The anti-diabetic drug metformin suppresses self-renewal and proliferation of trastuzumab-resistant tumor-initiating breast cancer stem cells. Breast Cancer Res Treat 126(2):355–364. https://doi.org/10.1007/s10549-010-0924-x
Janzer A, German NJ, Gonzalez-Herrera KN, Asara JM, Haigis MC, Struhl K (2014) Metformin and phenformin deplete tricarboxylic acid cycle and glycolytic intermediates during cell transformation and NTPs in cancer stem cells. Proc Natl Acad Sci USA 111(29):10574–10579. https://doi.org/10.1073/pnas.1409844111
Iliopoulos D, Hirsch HA, Struhl K (2011) Metformin decreases the dose of chemotherapy for prolonging tumor remission in mouse xenografts involving multiple cancer cell types. Cancer Res 71(9):3196–3201. https://doi.org/10.1158/0008-5472.CAN-10-3471
Remsik J, Fedr R, Navratil J, Bino L, Slabakova E, Fabian P, Svoboda M, Soucek K (2018) Plasticity and intratumoural heterogeneity of cell surface antigen expression in breast cancer. Br J Cancer 118(6):813–819. https://doi.org/10.1038/bjc.2017.497
Gupta PB, Onder TT, Jiang G, Tao K, Kuperwasser C, Weinberg RA, Lander ES (2009) Identification of selective inhibitors of cancer stem cells by high-throughput screening. Cell 138(4):645–659. https://doi.org/10.1016/j.cell.2009.06.034
Shibue T, Weinberg RA (2017) EMT, CSCs, and drug resistance: the mechanistic link and clinical implications. Nat Rev Clin Oncol 14(10):611–629. https://doi.org/10.1038/nrclinonc.2017.44
Soo HC, Chung FF, Lim KH, Yap VA, Bradshaw TD, Hii LW, Tan SH, See SJ, Tan YF, Leong CO, Mai CW (2017) Cudraflavone C Induces Tumor-Specific Apoptosis in Colorectal Cancer Cells through Inhibition of the Phosphoinositide 3-Kinase (PI3 K)-AKT Pathway. PLoS ONE 12(1):e0170551. https://doi.org/10.1371/journal.pone.0170551
König R, Chiang C-y, Tu BP, Yan SF, DeJesus PD, Romero A, Bergauer T, Orth A, Krueger U, Zhou Y (2007) A probability-based approach for the analysis of large-scale RNAi screens. Nat Methods 4(10):847
Tiong KH, Tan BS, Choo HL, Chung FF, Hii LW, Tan SH, Khor NT, Wong SF, See SJ, Tan YF, Rosli R, Cheong SK, Leong CO (2016) Fibroblast growth factor receptor 4 (FGFR4) and fibroblast growth factor 19 (FGF19) autocrine enhance breast cancer cells survival. Oncotarget 7(36):57633–57650. https://doi.org/10.18632/oncotarget.9328
Chung FF, Tan PF, Raja VJ, Tan BS, Lim KH, Kam TS, Hii LW, Tan SH, See SJ, Tan YF, Wong LZ, Yam WK, Mai CW, Bradshaw TD, Leong CO (2017) Jerantinine A induces tumor-specific cell death through modulation of splicing factor 3b subunit 1 (SF3B1). Scientific Rep 7:42504. https://doi.org/10.1038/srep42504
Stone EL, Citossi F, Singh R, Kaur B, Gaskell M, Farmer PB, Monks A, Hose C, Stevens MF, Leong CO, Stocks M, Kellam B, Marlow M, Bradshaw TD (2015) Antitumour benzothiazoles. Part 32: DNA adducts and double strand breaks correlate with activity; synthesis of 5F203 hydrogels for local delivery. Bioorg Med Chem 23(21):6891–6899. https://doi.org/10.1016/j.bmc.2015.09.052
Chou T-C (2010) Drug combination studies and their synergy quantification using the Chou-Talalay method. Cancer Res 70(2):440–446
Er JL, Goh PN, Lee CY, Tan YJ, Hii L-W, Mai CW, Chung FF-L, Leong C-O (2018) Identification of inhibitors synergizing gemcitabine sensitivity in the squamous subtype of pancreatic ductal adenocarcinoma (PDAC). Apoptosis 23:1–13
Voon YL, Ahmad M, Wong PF, Husaini R, Ng WT, Leong CO, Lane DP, Khoo AS (2015) Nutlin-3 sensitizes nasopharyngeal carcinoma cells to cisplatin-induced cytotoxicity. Oncol Rep 34(4):1692–1700. https://doi.org/10.3892/or.2015.4177
Low SY, Tan BS, Choo HL, Tiong KH, Khoo AS-B, Leong C-O (2012) Suppression of BCL-2 synergizes cisplatin sensitivity in nasopharyngeal carcinoma cells. Cancer Lett 314(2):166–175
Wong SW, Tiong KH, Kong WY, Yue YC, Chua CH, Lim JY, Lee CY, Quah SI, Fow C, Chung C (2011) Rapamycin synergizes cisplatin sensitivity in basal-like breast cancer cells through up-regulation of p73. Breast Cancer Res Treat 128(2):301–313
Di Veroli GY, Fornari C, Wang D, Mollard S, Bramhall JL, Richards FM, Jodrell DI (2016) Combenefit: an interactive platform for the analysis and visualization of drug combinations. Bioinformatics 32(18):2866–2868
Dai X, Cheng H, Bai Z, Li J (2017) Breast cancer cell line classification and its relevance with breast tumor subtyping. J Cancer 8(16):3131–3141. https://doi.org/10.7150/jca.18457
Neve RM, Chin K, Fridlyand J, Yeh J, Baehner FL, Fevr T, Clark L, Bayani N, Coppe JP, Tong F, Speed T, Spellman PT, DeVries S, Lapuk A, Wang NJ, Kuo WL, Stilwell JL, Pinkel D, Albertson DG, Waldman FM, McCormick F, Dickson RB, Johnson MD, Lippman M, Ethier S, Gazdar A, Gray JW (2006) A collection of breast cancer cell lines for the study of functionally distinct cancer subtypes. Cancer Cell 10(6):515–527. https://doi.org/10.1016/j.ccr.2006.10.008
Liu K, Newbury PA, Glicksberg BS, Zeng WZD, Paithankar S, Andrechek ER, Chen B (2019) Evaluating cell lines as models for metastatic breast cancer through integrative analysis of genomic data. Nat Commun 10(1):2138. https://doi.org/10.1038/s41467-019-10148-6
Li W, Ma H, Zhang J, Zhu L, Wang C, Yang Y (2017) Unraveling the roles of CD44/CD24 and ALDH1 as cancer stem cell markers in tumorigenesis and metastasis. Scientific Rep 7(1):13856. https://doi.org/10.1038/s41598-017-14364-2
Fillmore CM, Kuperwasser C (2008) Human breast cancer cell lines contain stem-like cells that self-renew, give rise to phenotypically diverse progeny and survive chemotherapy. Breast Cancer Res 10(2):R25
Venere M, Horbinski C, Crish JF, Jin X, Vasanji A, Major J, Burrows AC, Chang C, Prokop J, Wu Q, Sims PA, Canoll P, Summers MK, Rosenfeld SS, Rich JN (2015) The mitotic kinesin KIF11 is a driver of invasion, proliferation, and self-renewal in glioblastoma. Sci Transl Med 7(304):304ra143. https://doi.org/10.1126/scitranslmed.aac6762
Cheng CC, Chang J, Huang SC, Lin HC, Ho AS, Lim KH, Chang CC, Huang L, Chang YC, Chang YF, Wu CW (2017) YM155 as an inhibitor of cancer stemness simultaneously inhibits autophosphorylation of epidermal growth factor receptor and G9a-mediated stemness in lung cancer cells. PLoS ONE 12(8):e0182149. https://doi.org/10.1371/journal.pone.0182149
Vequaud E, Seveno C, Loussouarn D, Engelhart L, Campone M, Juin P, Barille-Nion S (2015) YM155 potently triggers cell death in breast cancer cells through an autophagy-NF-kB network. Oncotarget 6(15):13476–13486. https://doi.org/10.18632/oncotarget.3638
Huang M, Liu B, Liu R, Li J, Chen J, Jiang F, Ding H, Deng Z, Liu T (2018) Aglycone Polyether Nanchangmycin and Its Homologues Exhibit Apoptotic and Antiproliferative Activities against Cancer Stem Cells. ACS Pharmacol Transl Sci 1(2):84–95. https://doi.org/10.1021/acsptsci.8b00007
Yunokawa M, Koizumi F, Kitamura Y, Katanasaka Y, Okamoto N, Kodaira M, Yonemori K, Shimizu C, Ando M, Masutomi K (2012) Efficacy of everolimus, a novel m TOR inhibitor, against basal-like triple-negative breast cancer cells. Cancer Sci 103(9):1665–1671
Wander SA, Hennessy BT, Slingerland JM (2011) Next-generation mTOR inhibitors in clinical oncology: how pathway complexity informs therapeutic strategy. J Clin Investig 121(4):1231–1241
Kretsovali A, Hadjimichael C, Charmpilas N (2012) Histone deacetylase inhibitors in cell pluripotency, differentiation, and reprogramming. Stem cells international 2012:10
Bolden JE, Peart MJ, Johnstone RW (2006) Anticancer activities of histone deacetylase inhibitors. Nat Rev Drug Discov 5(9):769–784. https://doi.org/10.1038/nrd2133
West AC, Johnstone RW (2014) New and emerging HDAC inhibitors for cancer treatment. J Clin Investig 124(1):30–39. https://doi.org/10.1172/JCI69738
Ceccacci E, Minucci S (2016) Inhibition of histone deacetylases in cancer therapy: lessons from leukaemia. Br J Cancer 114(6):605
Chiappinelli KB, Zahnow CA, Ahuja N, Baylin SB (2016) Combining epigenetic and immunotherapy to combat cancer. Cancer Res 76(7):1683–1689. https://doi.org/10.1158/0008-5472.CAN-15-2125
Venugopal B, Baird R, Kristeleit RS, Plummer R, Cowan R, Stewart A, Fourneau N, Hellemans P, Elsayed Y, McClue S, Smit JW, Forslund A, Phelps C, Camm J, Evans TR, de Bono JS, Banerji U (2013) A phase I study of quisinostat (JNJ-26481585), an oral hydroxamate histone deacetylase inhibitor with evidence of target modulation and antitumor activity, in patients with advanced solid tumors. Clin Cancer Res 19(15):4262–4272. https://doi.org/10.1158/1078-0432.CCR-13-0312
He B, Dai L, Zhang X, Chen D, Wu J, Feng X, Zhang Y, Xie H, Zhou L, Zheng S (2018) The HDAC Inhibitor Quisinostat (JNJ-26481585) Supresses Hepatocellular Carcinoma alone and Synergistically in Combination with Sorafenib by G0/G1 phase arrest and Apoptosis induction. Int J Biol Sci 14(13):1845–1858. https://doi.org/10.7150/ijbs.27661
Tjulandin S, Fedyanin M, Vladimirov VI, Kostorov V, Lisyanskaya AS, Krikunova L, Cakana A, Azarova V, Karavaeva O, Vostokova N (2017) A multicenter phase II study of the efficacy and safety of quisinostat (an HDAC inhibitor) in combination with paclitaxel and carboplatin chemotherapy (CT) in patients (pts) with recurrent platinum resistant high grade serous epithelial ovarian, primarily peritoneal or fallopian tube carcinoma cancer (OC). American Society of Clinical Oncology,
Thiagalingam S, Cheng KH, Lee HJ, Mineva N, Thiagalingam A, Ponte JF (2003) Histone deacetylases: unique players in shaping the epigenetic histone code. Ann N Y Acad Sci 983(1):84–100
Arrowsmith CH, Bountra C, Fish PV, Lee K, Schapira M (2012) Epigenetic protein families: a new frontier for drug discovery. Nat Rev Drug Discov 11(5):384–400. https://doi.org/10.1038/nrd3674
Müller BM, Jana L, Kasajima A, Lehmann A, Prinzler J, Budczies J, Winzer K-J, Dietel M, Weichert W, Denkert C (2013) Differential expression of histone deacetylases HDAC1, 2 and 3 in human breast cancer-overexpression of HDAC2 and HDAC3 is associated with clinicopathological indicators of disease progression. BMC Cancer 13(1):215
Seo J, Min SK, Park H-R, Kim DH, Kwon MJ, Kim LS, Ju Y-S (2014) Expression of histone deacetylases HDAC1, HDAC2, HDAC3, and HDAC6 in invasive ductal carcinomas of the breast. J Breast Cancer 17(4):323–331
Witt AE, Lee CW, Lee TI, Azzam DJ, Wang B, Caslini C, Petrocca F, Grosso J, Jones M, Cohick EB, Gropper AB, Wahlestedt C, Richardson AL, Shiekhattar R, Young RA, Ince TA (2017) Identification of a cancer stem cell-specific function for the histone deacetylases, HDAC1 and HDAC7, in breast and ovarian cancer. Oncogene 36(12):1707–1720. https://doi.org/10.1038/onc.2016.337
Zhang Z, Yamashita H, Toyama T, Sugiura H, Ando Y, Mita K, Hamaguchi M, Hara Y, Kobayashi S, Iwase H (2005) Quantitation of HDAC1 mRNA expression in invasive carcinoma of the breast. Breast Cancer Res Treat 94(1):11–16
Glaser KB, Li J, Staver MJ, Wei R-Q, Albert DH, Davidsen SK (2003) Role of class I and class II histone deacetylases in carcinoma cells using siRNA. Biochem Biophys Res Commun 310(2):529–536
Rey M, Irondelle M, Waharte F, Lizarraga F, Chavrier P (2011) HDAC6 is required for invadopodia activity and invasion by breast tumor cells. Eur J Cell Biol 90(2–3):128–135
Delcuve GP, Khan DH, Davie JR (2012) Roles of histone deacetylases in epigenetic regulation: emerging paradigms from studies with inhibitors. Clin Epigenet 4(1):5
Falkenberg KJ, Johnstone RW (2014) Histone deacetylases and their inhibitors in cancer, neurological diseases and immune disorders. Nat Rev Drug Discov 13(9):673
Guzman ML, Yang N, Sharma KK, Balys M, Corbett CA, Jordan CT, Becker MW, Steidl U, Abdel-Wahab O, Levine RL, Marcucci G, Roboz GJ, Hassane DC (2014) Selective activity of the histone deacetylase inhibitor AR-42 against leukemia stem cells: a novel potential strategy in acute myelogenous leukemia. Mol Cancer Ther 13(8):1979–1990. https://doi.org/10.1158/1535-7163.MCT-13-0963
Nalls D, Tang SN, Rodova M, Srivastava RK, Shankar S (2011) Targeting epigenetic regulation of miR-34a for treatment of pancreatic cancer by inhibition of pancreatic cancer stem cells. PLoS ONE 6(8):e24099. https://doi.org/10.1371/journal.pone.0024099
Kumar B, Yadav A, Lang JC, Teknos TN, Kumar P (2015) Suberoylanilide hydroxamic acid (SAHA) reverses chemoresistance in head and neck cancer cells by targeting cancer stem cells via the downregulation of nanog. Genes Cancer 6(3–4):169–181. https://doi.org/10.18632/genesandcancer.54
Salvador MA, Wicinski J, Cabaud O, Toiron Y, Finetti P, Josselin E, Lelievre H, Kraus-Berthier L, Depil S, Bertucci F, Collette Y, Birnbaum D, Charafe-Jauffret E, Ginestier C (2013) The histone deacetylase inhibitor abexinostat induces cancer stem cells differentiation in breast cancer with low Xist expression. Clin Cancer Res 19(23):6520–6531. https://doi.org/10.1158/1078-0432.CCR-13-0877
Di Pompo G, Salerno M, Rotili D, Valente S, Zwergel C, Avnet S, Lattanzi G, Baldini N, Mai A (2015) Novel histone deacetylase inhibitors induce growth arrest, apoptosis, and differentiation in sarcoma cancer stem cells. J Med Chem 58(9):4073–4079. https://doi.org/10.1021/acs.jmedchem.5b00126
Seto E, Yoshida M (2014) Erasers of histone acetylation: the histone deacetylase enzymes. Cold Spring Harb Perspect Biol 6(4):a018713. https://doi.org/10.1101/cshperspect.a018713
Liu N, Li S, Wu N, Cho KS (2017) Acetylation and deacetylation in cancer stem-like cells. Oncotarget 8(51):89315–89325. https://doi.org/10.18632/oncotarget.19167
Doherty MR, Smigiel JM, Junk DJ, Jackson MW (2016) Cancer stem cell plasticity drives therapeutic resistance. Cancers 8(1):E8. https://doi.org/10.3390/cancers8010008
Ye X, Tam WL, Shibue T, Kaygusuz Y, Reinhardt F, Ng Eaton E, Weinberg RA (2015) Distinct EMT programs control normal mammary stem cells and tumour-initiating cells. Nature 525(7568):256–260. https://doi.org/10.1038/nature14897
Chaffer CL, Brueckmann I, Scheel C, Kaestli AJ, Wiggins PA, Rodrigues LO, Brooks M, Reinhardt F, Su Y, Polyak K, Arendt LM, Kuperwasser C, Bierie B, Weinberg RA (2011) Normal and neoplastic nonstem cells can spontaneously convert to a stem-like state. Proc Natl Acad Sci USA 108(19):7950–7955. https://doi.org/10.1073/pnas.1102454108
Ravikumar B, Alam Z, Peddinti G, Aittokallio T (2017) C-SPADE: a web-tool for interactive analysis and visualization of drug screening experiments through compound-specific bioactivity dendrograms. Nucleic Acids Res 45(W1):W495–W500. https://doi.org/10.1093/nar/gkx384
Funding
This study was funded by the Malaysia Ministry of Education Exploratory Research Grant Scheme (LCO; ERGS/1/2013/SKK01/IMU/02/1) and Malaysia Ministry of Education Fundamental Grant Scheme (LCO; FRGS/1/2016/SKK08/IMU/01/1).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
10549_2019_5504_MOESM2_ESM.pptx
Supplementary material 2 (PPTX 1110 kb). Supplementary Figure 1: Breast CSCs are intrinsically resistant to conventional chemotherapeutic agents. Both CSCs and non-CSCs derived from MDA-MB-468, MDA-MB-231, HCC38, and MCF-7 breast cancer cell lines were treated with cisplatin, doxorubicin and paclitaxel for 72 h. Points represent mean ± S.D. of at least three independent experiments. Supplemental Figure 2: Combinatory effects of HDACi and paclitaxel on MDA-MB-468 breast CSCs and non-CSCs. MDA-MB-468 breast CSCs and non-CSCs were treated with paclitaxel and/or HDACi for 72 h. Dose–response surface curves and synergy of each combination was assessed using the HSA model (effect-based approach), as implemented in Combenefit software [32]. Level of synergism (blue) or antagonism (red) at each concentration is represented by color scale bar. All experiments were conducted at least three times
Rights and permissions
About this article
Cite this article
Hii, LW., Chung, F.FL., Soo, J.SS. et al. Histone deacetylase (HDAC) inhibitors and doxorubicin combinations target both breast cancer stem cells and non-stem breast cancer cells simultaneously. Breast Cancer Res Treat 179, 615–629 (2020). https://doi.org/10.1007/s10549-019-05504-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-019-05504-5