Abstract
Purpose
Androgen receptor (AR) and AR signaling pathways are thought to play a role in breast cancer (BC) and are potentially related to treatment responses and outcomes. Ankyrin 3 (ANK3) is associated with AR stability in cancer cells. In the present study, we investigated the clinicopathological utility of ANK3 expression with emphasis on AR and its associated signalling pathway at transcriptomic and proteomic phases.
Patients and methods
The Molecular Taxonomy of Breast Cancer International Consortium (METABRIC) cohort (n = 1980) and The Cancer Genome Atlas (TCGA) dataset (n = 1039) were used to assess the expression and significance of ANK3 mRNA and other AR signalling pathway-associated gene signature. Using immunohistochemistry, ANK3 protein expression was evaluated in large (n = 982) cohort of early-stage BC with long-term follow-up and compared with clinicopathological characteristics and its prognostic value in the whole cohort and the subgroups stratified by AR protein expression.
Results
An AR-related gene signature was developed, comprising 20 genes, which included ANK3. This AR-related gene signature was significantly associated with AR mRNA expression, oestrogen receptor, human epidermal growth factor receptor 2 (HER2) status and the patients’ outcomes. In tumours with high AR protein expression (n = 614), high ANK3 protein expression was significantly associated with progesterone receptor positivity and it was independently associated with the good outcomes (p = 0.025).
Conclusions
This study indicates that ANK3 is related to AR signalling pathway and is associated with BC prognosis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Background
Treatments of breast cancer (BC) are generally determined on the basis of the molecular phenotype of the primary tumour [1, 2]. However, the biological heterogeneity of BC constitutes an important determinant of treatment sensitivity, success and outcomes. Hormone-dependent pathways, including androgen receptor (AR) signalling pathways, are thought to play an important role in BC cell proliferation [3, 4]. Previous studies have indicated that AR and AR signaling pathways are associated with treatment resistance and prognosis of BC [5, 6]. In previous research, we found that approximately 55% of BC had high AR expression, which was observed in 42% of human epidermal growth factor receptor 2 (HER2)-positive tumours and in 20% of triple negative BC (TNBC) [7]. Some studies indicate that high AR expression is a good prognostic factor in BC [7, 8]. However, in HER2-positive and TNBC subtypes, AR signalling pathways are considered to play an important role in tumour progression. He et al. suggested that AR promotes the growth of HER2-positive BC via crosstalk with the intracellular HER2 downstream pathway [9]. The luminal-AR BC subtype, a molecular subtype of TNBC, not only expresses AR but also has enriched hormone-dependent pathways, as demonstrated at the global transcriptomic level [10, 11]. It has also been shown in oestrogen receptor (ER)-positive and HER2-negative BC that aberrant AR-related oncogenic pathway activation is associated with resistance to endocrine therapy [12].
Ankyrin 3 (ANK3), a member of the ankyrin family of membrane-associated proteins, is believed to link integral membrane proteins to cytoskeletal components. Ankyrins are associated with cytoplasmic structures and are also necessary in the regulation of cell migration and adhesion and for the maintenance of cellular membrane domains [13,14,15]. ANK3 has been suggested to play a role in regulating the stability and turnover of AR and is closely associated with AR genomic activities [16]. AR signaling pathway promotes cancer cell proliferation by increasing cyclin-dependent kinase activity [17, 18] and ANK3 regulates the expression of cell cycle components as cyclins A and B [16]. Hence, ANK3 may play an important role in AR signaling pathway in cancer. However, the association between ANK3 expression and AR signaling pathway in BC remains poorly defined. In this study, ANK3 was first evaluated as a component of the AR signaling pathway in BC, utilising well-characterised large cohort transcriptomic databases. The clinicopathological and prognostic significance of ANK3 protein expression levels was assessed using immunohistochemistry (IHC) in a large series of BC patients’ specimens.
Materials and methods
Cluster analysis of AR-signaling-pathway-associated genes
Gene Ontology (GO) Consortium is the large genomic annotation project and widely used as biological databases for annotating genes to the previous evidence regarding their biological role [19, 20]. GO terms are divided into three categories as biological process, molecular function and cellular component [21]. In GO terms of the biological process, gene symbols related to ‘Regulation Of Androgen Receptor Signaling Pathway (GO: 0060765)’ were accessed using the online database Gene Set Enrichment Analysis (http://software.broadinstitute.org/gsea/msigdb/cards/GO_REGULATION_OF_ANDROGEN_RECEPTOR_SIGNALING_PATHWAY) [22, 23]. The mRNA expression data of these genes, including ANK3, together with the clinicopathological characteristics and outcomes of patients with BC, were collected from the Molecular Taxonomy of Breast Cancer International Consortium (METABRIC) dataset [24, 25] (n = 1980) and the Cancer Genome Atlas (TCGA) [26] dataset (n = 1039) provided by cBioPortal [27].
The normalisation method of mRNA expression in the METABRIC cohort was previously described [24]. TCGA mRNA data were log2 transformed prior to cluster analysis. For cluster analysis [28] and heat mapping construction, Cluster 3.0 and Java Treeview was used [29]. Data were filtered to remove all genes that did not have at least one observation with absolute values greater than 2.0 or whose maximum minus minimum values were less than 2.0.
ANK3 protein expression
A total of 982 BC patients who underwent surgery at Nottingham City Hospital in the UK between 1987 and 1998 (referred to as the Nottingham Primary Breast Cancer Series) were included in this study. All patients had undergone breast-conserving surgery or modified radical mastectomy without any neoadjuvant treatment. The availability and assessment of hormone receptors [AR, ER and progesterone receptor (PR)], HER2 and Ki67 were described in previous studies [7, 30,31,32,33,34,35,36,37]. The cohort was stratified on the basis of AR expression [7], with 614 patients (62.5%) with high and 368 patients (37.5%) with low AR expression (Supplementary Table 1).
ANK3 protein expression was assessed by IHC using an anti-ANK3 antibody (HPA055643; Merck, Darmstadt, Germany) diluted 1:300 as previously described [38,39,40]. To evaluate the pattern of ANK3 protein expression, 15 full-face BC tissue sections were assessed prior to staining the whole cohort (n = 982) prepared as tissue microarrays (TMAs). Immunostained TMA sections were digitally scanned using a NanoZoomer (Hamamatsu Photonics, Tokyo, Japan). Cytoplasmic staining of ANK3 in cancer cells was assessed using the H-score method on the basis of intensity scoring (0 = negative, 1 = weak, 2 = moderate, 3 = strong) and proportion scoring (0–100) as previously reported [41, 42].
Statistical analysis
Statistical analyses were conducted using SPSS v24.0 (IBM, Armonk, NY, USA). The relationship between ANK3 mRNA with ANK3 protein expression and AR mRNA expression was examined using Pearson’s correlation coefficient test. To assess the associations between AR mRNA expression and groups stratified by the AR-related gene signature, the Mann–Whitney U test was used. The Chi square test as univariate analysis and the logistic regression test as multivariate analysis were used to assess several clinicopathological factors, including tumour size, lymph node status, histological grade, ER, PR, HER2 and molecular subtypes, stratified by groups based on AR-related gene signature and levels of ANK3 protein expression. To assess the prognostic utility of ANK3 expression, Kaplan–Meier survival curves was used. In univariate and multivariate analyses, to assess the associations between clinicopathological factors, including ANK3 expression, and prognosis, 95% confidence intervals (CIs) were assessed using the Cox proportional hazards regression model. In these survival analyses, the median value (H-score = 120) was used as a cut-off point to divide the samples into high and low expression groups.
Results
ANK3 mRNA expression and AR signaling pathway gene signature
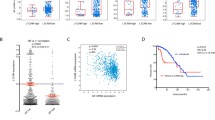
High ANK3 mRNA expression was significantly associated with high AR mRNA expression (METABRIC: r = 0.019, p = 0.39; TCGA: r = 0.28, p < 0.0001) in TCGA cohort. An AR-related gene signature was developed using genomic data filtering, and this comprised 20 genes, including ANK3 and 19 other relevant genes available in the databases: ARRB2, BUD31, DAB2, DDX5, EP300, FOXP1, HDAC1, HDAC6, HEYL, PARK7, PHB, PIAS2, PRMT2, RNF14, RNF6, SFRP1, SIRT1, SMARCA4 and TRIM68 (Supplementary Table 2). Using the dendrogram of cluster analysis, the METABRIC and TCGA cohorts were stratified into two groups on the basis of the AR-signaling-pathway-associated genes (Fig. 1a, b), where tumours in group 1 had significantly lower AR mRNA expression than that in Group 2 (p < 0.0001). Group 1 tumours included 899 (45%) from the METABRIC and 541 (52%) from TCGA cohort.
Prognostic utility of an androgen receptor (AR)-related gene signature, including ANK3 mRNA expression. Heat map of the AR-related gene signature for the a METABRIC and b TCGA cohorts generated by unsupervised cluster analysis, showing a clear division of cases between Group 1 and Group 2 on the basis of the AR-related gene expression. The overall survival of patients with breast cancer with the AR-related Group 2 gene signature was significantly worse than that of those with the Group 1 gene signature in the c METABRIC and d TCGA cohorts
In the METABRIC and TCGA cohorts, multivariate analysis indicated that the AR-related gene signature in group 2 was significantly associated with lower grade (p = 0.0070, and p = 0.0093 respectively), ER positivity (p < 0.0001, and p < 0.0001 respectively), and HER2 positivity (p < 0.0001 and p < 0.0001; Table 1). In the METABRIC cohort, the AR-related gene signature was significantly associated with molecular subtype (p < 0.0001), with 83% of the basal-like tumours in group 1 and 90% of the luminal B tumours in group 2 (Table 1). Although the expression of ANK3 and AR mRNA was not a significant independent prognostic factor in BC (Supplementary Fig. 1), there was an association between AR-related gene signature subgroups and patients’ outcomes, where patients with the AR-related gene signature group 2 showed significantly worse outcome than those with Group 1 tumours [METABRIC: hazard ratio (HR) 1.25, 95% CI 1.09–1.43, p = 0.0013; TCGA: HR 1.61, 95% CI 1.11–2.32, p = 0.011; Fig. 1c, d. On multivariate analysis, AR-related gene signature group 2 was an independent prognostic factor predicting poor outcomes in both cohorts (METABRIC: HR 1.23, 95% CI 1.06–1.42, p = 0.0066; TCGA: HR 1.82, 95% CI 1.08–3.06, p = 0.026; Table 2).
Immunohistochemical expression of ANK3 protein
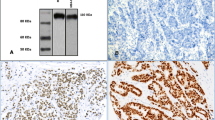
The assessment of ANK3 in full-face tissue sections indicated that the pattern of ANK3 expression in cancer cells was homogeneous, but it differed from that in normal mammary glands (Fig. 2a–c). ANK3 expression was observed in the normal glandular and luminal epithelial cells, where it was stronger than the surrounding myoepithelial cells. ANK3 immunopositivity was observed in the cytoplasm of invasive cancer cells and was typically weaker than in the adjacent normal epithelial cells (Fig. 2d–f).
Morphological characteristics of ANK3 immunohistochemistry in breast cancer tissue. a ANK3 immunoreactivity differs between invasive cancer cells and adjacent normal mammary glandular tissues (black arrow: invasive cancer cells; white arrow: normal mammary gland). Immunoreactivity in normal mammary gland cells is stronger than that in invasive cancer cells (magnification: ×100). b Invasive cancer cells showing uniform ANK3 immunoreactivity primarily in the cytoplasm (magnification: ×200). c ANK3 immunoreactivity is uniformly strong in normal epithelial cells and weaker in myoepithelial cells than in glandular cells (magnification: × 400). Tissue microarray images of breast cancer tissue samples immunohistochemically stained for ANK3, showing d no staining, e weak staining and f strong staining in the cytoplasm of cancer cells (magnification: × 200)
In 198 cases in the METABRIC dataset, which overlapped with the Nottingham Primary Series, ANK3 mRNA and ANK3 protein expression were significantly correlated (r = 0.15, p = 0.039). In the Nottingham series, 579 (59%) tumours had low ANK3 expression (H-score ≤ 120) and 403 (41%) had high ANK3 expression (H-score > 120). High AR expression was present in 614 (63%) tumours and low AR expression was present in 368 (37%). Among those with high AR expression, 250 (41%) also had high ANK3 expression. A similar proportion (153, 42%) had high ANK3 expression in the low AR expression group (n = 368). AR expression was not associated with ANK3 expression on proteomic analysis (p = 0.79). When all 982 cases were combined (i.e. not stratified according to AR expression), ANK3 was not a significant prognostic factor (Supplementary Fig. 2).
In tumours with high AR expression, high ANK3 expression was significantly associated with PR positivity (p = 0.014; Supplementary Table 3). In terms of BC-specific survival, high AR protein expression was a significant good prognostic factor (HR 0.66, 95% CI 0.52–0.84, p = 0.00066; Supplementary Fig. 3). Low ANK3 protein expression was a poor prognostic factor in patients with high AR expression [HR 1.49, 95% CI 1.07–2.09, p = 0.020; Fig. 3a–e, but not in those whose tumours had low AR expression (HR 0.89, 95% CI 0.62–1.28, p = 0.53; Supplementary Fig. 4). In high-AR-expressing BC patients, univariate analysis using the Cox proportional hazards regression analysis identified low ANK3 expression, large tumour size (HR 2.61, p < 0.0001), positive nodal status (HR 2.84, p < 0.0001) and high histological grade (HR 3.27, p < 0.0001) as poor prognostic factors. On multivariate analysis, low ANK3 protein expression was an independent prognostic factor predicting poor outcomes in BC with high AR expression (HR 1.47, p = 0.025; Table 3).
ANK3 protein expression in breast cancer and cumulative survival rates stratified by ANK3 expression. a–d ANK3 and AR expression in breast cancer. Case 1: high ANK3 (a) and high AR (b) expression. Case 2: low ANK3 (c) and high AR (d) expression (magnification: × 200 for all images). e With high AR expression, BC-specific survival was significantly worse in those with low than high ANK3 expression
Discussion
AR expression is a crucial factor in the progression of BC, as it controls the expression of various genes and proteins through a genomic pathway [5, 6]. In this pathway, AR mediates intracellular steroid hormone-related signaling pathways to regulate the transcription of target genes in conjunction with other transcription factors, such as signal transducers and activators of transcription [43, 44]. As a mechanism involved in the development of BC, AR expression might be involved in the crosstalk with epidermal growth factor receptor pathways, such as human epidermal growth factor receptor 1 (EGFR) and HER2 signaling [45]. In this study, there were a significant correlation between ANK3 and AR mRNA and ANK3 was one of the gene component of the AR-related gene signature. When BC was classified into 2 groups based on the expression of AR-related gene signature, the group 2 gene signature, which was associated with high AR mRNA expression and present in 90% of luminal B tumours, was a significant prognostic factor indicating poor outcomes in BC. This finding suggests that aberrant AR-related oncogenic pathway activation is associated with a number of factors that portend a poor BC outcome.
In a previous study using microarray gene expression analysis, the downregulation of ANK3 was included in an 11-gene signature associated with poor prognosis in patients with various cancers including BC [46]. In a meta-analysis of gene expression signatures in BC, the downregulation of ANK3 appeared to enhance cancer cell differentiation, proliferation and metastasis [47]. Previous research using microarray data of prostate cancer suggested that low ANK3 expression is related to positivity for ERG, member of the erythroblast transformation-specific family [48]. ERG is correlated with AR activity [49], transcriptional stability [50] and stem cell maintenance [51] in multiple cancers. Prostate cancer cells with ANK3 knockdown exhibit significant increases in cell invasion through an AR-dependent mechanism as a regulator of AR protein stability [16]. In the present study, the association between ANK3 protein expression and outcomes was highly significant in BC with high AR expression. In addition, high ANK3 protein expression was associated with PR positivity. These findings suggest that ANK3 may play an important role in the maintenance of hormonal activity, and AR stabilisation by ANK3 may, therefore, be related to the improved outcomes in BC patients with high AR expression. A proportion of ER-negative BC are generally considered to retain active AR signaling [6, 52]. Several prospective clinical trials of AR-targeted therapies have been conducted on TNBC with high AR expression. These trials indicated that treatment with an AR inhibitor is feasible, with a clinical benefit rate of approximately 20% in TNBC [53,54,55]. The upregulation of ANK3 may increase AR stability and improve the response to an AR inhibitor in TNBC. Further functional and translational research is necessary to explore the association of ANK3 with AR stability with the efficacy of treating BC with an AR inhibitor.
In conclusion, the AR signaling pathway and ANK3 mRNA expression are associated with AR mRNA expression and BC prognosis. High ANK3 protein expression is an independent prognostic factor in BC with high AR expression. Overall, these findings indicate that ANK3 may play an important role in breast tumour progression and, in conjunction with AR, may be related to BC outcomes.
References
Coates AS, Winer EP, Goldhirsch A, Gelber RD, Gnant M, Piccart-Gebhart M, Thürlimann B, Senn HJ, Panel Members (2015) Tailoring therapies-improving the management of early breast cancer: St Gallen International Expert Consensus on the Primary Therapy of Early Breast Cancer 2015. Ann Oncol 26:1533–1546
Curigliano G, Burstein HJ, Winer PE, Gnant M, Dubsky P, Loibl S, Colleoni M, Regan MM, Piccart-Gebhart M, Senn HJ, Thürlimann B, St. Gallen International expert consensus on the primary therapy of early breast cancer 2017 (2017) De-escalating and escalating treatments for early-stage breast cancer: the St Gallen International Expert Consensus Conference on the primary therapy of early breast cancer 2017. Ann Oncol 28:1700–1712
Kurozumi S, Matsumoto H, Hayashi Y, Tozuka K, Inoue K, Horiguchi J, Takeyoshi I, Oyama T, Kurosumi M (2017) Power of PgR expression as a prognostic factor for ER-positive/HER2-negative breast cancer patients at intermediate risk classified by the Ki67 labelling index. BMC Cancer 17:354
Hayashi S, Yamaguchi Y (2008) Estrogen signalling pathway and hormonal therapy. Breast Cancer 15:256–261
Iacopetta D, Rechoum Y, Fuqua SA (2012) The role of androgen receptor in breast cancer. Drug Discov Today Dis Mech 9:e19–e27
Rampurwala M, Wisinski KB, O’Regan R (2016) Role of the androgen receptor in triple-negative breast cancer. Clin Adv Hematol Oncol 14:186–193
Aleskandarany MA, Abduljabbar R, Ashankyty I, Elmouna A, Jerjees D, Ali S, Buluwela L, Diez-Rodriguez M, Caldas C, Green AR, Ellis IO, Rakha EA (2016) Prognostic significance of androgen receptor expression in invasive breast cancer: transcriptomic and protein expression analysis. Breast Cancer Res Treat 159:215–227
Vera-Badillo FE, Templeton AJ, de Gouveia P, Diaz-Padilla I, Bedard PL, Al-Mubarak M, Seruga B, Tannock IF, Ocana A, Amir E (2014) Androgen receptor expression and outcomes in early breast cancer: a systematic review and meta-analysis. J Natl Cancer Inst 106:319
He L, Du Z, Xiong X, Ma H, Zhu Z, Gao H, Cao J, Li T, Li H, Yang K, Chen G, Richer JK, Gu H (2017) Targeting androgen receptor in treating HER2 positive breast cancer. Sci Rep 7:14584
Lehmann BD, Bauer JA, Chen X, Sanders ME, Chakravarthy AB, Shyr Y, Pietenpol JA (2011) Identification of human triple-negative breast cancer subtypes and preclinical models for selection of targeted therapies. J Clin Invest 121:2750–2767
Masuda H, Baggerly KA, Wang Y, Zhang Y, Gonzalez-Angulo AM, Meric-Bernstam F, Valero V, Lehmann BD, Pietenpol JA, Hortobagyi GN, Symmans WF, Ueno NT (2013) Differential response to neoadjuvant chemotherapy among 7 triple-negative breast cancer molecular subtypes. Clin Cancer Res 19:5533–5540
Fujii R, Hanamura T, Suzuki T, Gohno T, Shibahara Y, Niwa T, Yamaguchi Y, Ohnuki K, Kakugawa Y, Hirakawa H, Ishida T, Sasano H, Ohuchi N, Hayashi S (2014) Increased androgen receptor activity and cell proliferation in aromatase inhibitor-resistant breast carcinoma. J Steroid Biochem Mol Biol 144:513–522
Lambert S, Bennett V (1993) From anemia to cerebellar dysfunction. A review of the ankyrin gene family. Eur J Biochem 211:1–6
Bennett V (1992) Ankyrins. Adaptors between diverse plasma membrane proteins and the cytoplasm. J Biol Chem 267:8703–8706
De Matteis MA, Morrow JS (1998) The role of ankyrin and spectrin in membrane transport and domain formation. Curr Opin Cell Biol 10:542–549
Wang T, Abou-Ouf H, Hegazy SA, Alshalalfa M, Stoletov K, Lewis J, Donnelly B, Bismar TA (2016) Ankyrin G expression is associated with androgen receptor stability, invasiveness, and lethal outcome in prostate cancer patients. J Mol Med (Berl) 94:1411–1422
Ramos-Montoya A, Lamb AD, Russell R, Carroll T, Jurmeister S, Galeano-Dalmau N, Massie CE, Boren J, Bon H, Theodorou V, Vias M, Shaw GL, Sharma NL, Ross-Adams H, Scott HE, Vowler SL, Howat WJ, Warren AY, Wooster RF, Mills IG, Neal DE (2014) HES6 drives a critical AR transcriptional programme to induce castration-resistant prostate cancer through activation of an E2F1-mediated cell cycle network. EMBO Mol Med 6:651–661
Pietri E, Conteduca V, Andreis D, Massa I, Melegari E, Sarti S, Cecconetto L, Schirone A, Bravaccini S, Serra P, Fedeli A, Maltoni R, Amadori D, De Giorgi U, Rocca A (2016) Androgen receptor signaling pathways as a target for breast cancer treatment. Endocr Relat Cancer 23:R485–R498
Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM, Davis AP, Dolinski K, Dwight SS, Eppig JT, Harris MA, Hill DP, Issel-Tarver L, Kasarskis A, Lewis S, Matese JC, Richardson JE, Ringwald M, Rubin GM, Sherlock GB (2000) Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat Genet 25:25–29
The Gene Ontology Consortium (2017) Expansion of the gene ontology knowledgebase and resources. Nucleic Acids Res 45:D331–D338
Rhee SY, Wood V, Dolinski K, Draghici S (2008) Use and misuse of the gene ontology annotations. Nat Rev Genet 9:509–515
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, Paulovich A, Pomeroy SL, Golub TR, Lander ES, Mesirov JP (2005) Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA 102:15545–15550
Mootha VK, Lindgren CM, Eriksson KF, Subramanian A, Sihag S, Lehar J, Puigserver P, Carlsson E, Ridderstråle M, Laurila E, Houstis N, Daly MJ, Patterson N, Mesirov JP, Golub TR, Tamayo P, Spiegelman B, Lander ES, Hirschhorn JN, Altshuler D, Groop LC (2003) PGC-1alpha-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat Genet 34:267–273
Curtis C, Shah SP, Chin SF, Turashvili G, Rueda OM, Dunning MJ, Speed D, Lynch AG, Samarajiwa S, Yuan Y, Gräf S (2012) The genomic and transcriptomic architecture of 2,000 breast tumours reveals novel subgroups. Nature 486:346–352
Pereira B, Chin SF, Rueda OM, Vollan HK, Provenzano E, Bardwell HA, Pugh M, Jones L, Russell R, Sammut SJ, Tsui DW (2016) The somatic mutation profiles of 2,433 breast cancers refines their genomic and transcriptomic landscapes. Nat Commun 10:11479
Cancer Genome Atlas Network (2012) Comprehensive molecular portraits of human breast tumours. Nature 490:61–70
Cerami E, Gao J, Dogrusoz U, Gross BE, Sumer SO, Aksoy BA, Jacobsen A, Byrne CJ, Heuer ML, Larsson E, Antipin Y (2012) The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov 2:401–404
Eisen MB, Spellman PT, Brown PO, Botstein D (1998) Cluster analysis and display of genome-wide expression patterns. Proc Natl Acad Sci USA 95:14863–14868
De Hoon MJL, Imoto S, Nolan J, Miyano S (2004) Open source clustering software. Bioinformatics 20:1453–1454
Rakha EA, Agarwal D, Green AR, Ashankyty I, Ellis IO, Ball G, Alaskandarany MA (2017) Prognostic stratification of oestrogen receptor-positive HER2-negative lymph node-negative class of breast cancer. Histopathology 70:622–631
Rakha EA, Elsheikh SE, Aleskandarany MA, Habashi HO, Green AR, Powe DG, El-Sayed ME, Benhasouna A, Brunet JS, Akslen LA, Evans AJ (2009) Triple-negative breast cancer: distinguishing between basal and nonbasal subtypes. Clin Cancer Res 15:2302–2310
Green AR, Powe DG, Rakha EA, Soria D, Lemetre C, Nolan CC, Barros FF, Macmillan RD, Garibaldi JM, Ball GR, Ellis IO (2013) Identification of key clinical phenotypes of breast cancer using a reduced panel of protein biomarkers. Br J Cancer 109:1886–1894
Rakha EA, Soria D, Green AR, Lemetre C, Powe DG, Nolan CC, Garibaldi JM, Ball G, Ellis IO (2014) Nottingham Prognostic Index Plus (NPI +): a modern clinical decision making tool in breast cancer. Br J Cancer 110:1688–1697
Habashy HO, Powe DG, Rakha EA, Ball G, Paish C, Gee J, Nicholson RI, Ellis IO (2008) Forkhead-box A1 (FOXA1) expression in breast cancer and its prognostic significance. Eur J Cancer 44:1541–1551
Habashy HO, Powe DG, Glaab E, Ball G, Spiteri I, Krasnogor N, Garibaldi JM, Rakha EA, Green AR, Caldas C, Ellis IO (2011) RERG (Ras-like, oestrogen-regulated, growth-inhibitor) expression in breast cancer: a marker of ER-positive luminal-like subtype. Breast Cancer Res Treat 128:315–326
Aleskandarany MA, Rakha EA, Ahmed MA, Powe DG, Ellis IO, Green AR (2011) Clinicopathologic and molecular significance of phosphor-Akt expression in early invasive breast cancer. Breast Cancer Res Treat 127:407–416
Aleskandarany MA, Rakha EA, Ahmed MA, Powe DG, Paish EC, Macmillan RD, Ellis IO, Green AR (2010) PIK3CA expression in invasive breast cancer: a biomarker of poor prognosis. Breast Cancer Res Treat 122:45–53
Joseph C, Macnamara O, Craze M, Russell R, Provenzano E, Nolan CC, Diez-Rodriguez M, Sonbul SN, Aleskandarany MA, Green AR, Rakha EA (2018) Mediator complex (MED) 7: a biomarker associated with good prognosis in invasive breast cancer, especially ER + luminal subtypes. Br J Cancer 118:1142–1151
Kurozumi S, Joseph C, Sonbul S, Aleskandarany MA, Pigera M, Alsaleem M, Alsaeed S, Kariri Y, Nolan CC, Diez-Rodriguez M, Johnston S, Mongan NP, Fujii T, Shirabe K, Martin SG, Ellis IO, Green AR, Rakha EA (2018) Clinicopathological and prognostic significance of Ras association and pleckstrin homology domains 1 (RAPH1) in breast cancer. Breast Cancer Res Treat 25:236. https://doi.org/10.1007/s10549-018-4891-y
Kurozumi S, Joseph C, Sonbul S, Gorringe KL, Pigera M, Aleskandarany MA, Diez-Rodriguez M, Nolan CC, Fujii T, Shirabe K, Kuwano H, Storr S, Martin SG, Ellis IO, Green AR, Rakha EA (2018) Clinical and biological roles of Kelch-like family member 7 in breast cancer: a marker of poor prognosis. Breast Cancer Res Treat 170:525–533
McCarty KS Jr, Miller LS, Cox EB, Konrath J, McCarty KS Sr (1985) Estrogen receptor analyses. Correlation of biochemical and immunohistochemical methods using monoclonal antireceptor antibodies. Arch Pathol Lab Med 109:716–721
Detre S, Saclani Jotti G, Dowsett MA (1995) “Quickscore” method for immunohistochemical semiquantitation: validation for oestrogen receptor in breast carcinomas. J Clin Pathol 48:876–878
Dimitrakakis C, Bondy C (2009) Androgens and the breast. Breast Cancer Res 11:212
Garay JP, Karakas B, Abukhdeir AM, Cosgrove DP, Gustin JP, Higgins MJ, Konishi H, Konishi Y, Lauring J, Mohseni M, Wang GM, Jelovac D, Weeraratna A, Sherman Baust CA, Morin PJ, Toubaji A, Meeker A, De Marzo AM, Lewis G, Subhawong A, Argani P, Park BH (2012) The growth response to androgen receptor signaling in ERα-negative human breast cells is dependent on p21 and mediated by MAPK activation. Breast Cancer Res 14:R27
Chia KM, Liu J, Francis GD, Naderi A (2011) A feedback loop between androgen receptor and ERK signaling in estrogen receptor-negative breast cancer. Neoplasia 13:154–166
Glinsky GV, Berezovska O, Glinskii AB (2005) Microarray analysis identifies a death-from-cancer signature predicting therapy failure in patients with multiple types of cancer. J Clin Invest 115:1503–1521
Gröger CJ, Grubinger M, Waldhör T, Vierlinger K, Mikulits W (2012) Meta-analysis of gene expression signatures defining the epithelial to mesenchymal transition during cancer progression. PLoS ONE 7:e51136
Bismar TA, Alshalalfa M, Petersen LF, Teng LH, Gerke T, Bakkar A, Al-Mami A, Liu S, Dolph M, Mucci LA, Alhajj R (2014) Interrogation of ERG gene rearrangements in prostate cancer identifies a prognostic 10-gene signature with relevant implication to patients’ clinical outcome. BJU Int 113:309–319
Yu J, Yu J, Mani RS, Cao Q, Brenner CJ, Cao X, Wang X, Wu L, Li J, Hu M, Gong Y, Cheng H, Laxman B, Vellaichamy A, Shankar S, Li Y, Dhanasekaran SM, Morey R, Barrette T, Lonigro RJ, Tomlins SA, Varambally S, Qin ZS, Chinnaiyan AM (2010) An integrated network of androgen receptor, polycomb, and TMPRSS2-ERG gene fusions in prostate cancer progression. Cancer Cell 17:443–454
Shah AV, Birdsey GM, Randi AM (2016) Regulation of endothelial homeostasis, vascular development and angiogenesis by the transcription factor ERG. Vascul Pharmacol 86:3–13
Polson ES, Lewis JL, Celik H, Mann VM, Stower MJ, Simms MS, Rodrigues G, Collins AT, Maitland NJ (2013) Monoallelic expression of TMPRSS2/ERG in prostate cancer stem cells. Nat Commun 4:1623
Vranic S, Feldman R, Gatalica Z (2017) Apocrine carcinoma of the breast: A brief update on the molecular features and targetable biomarkers. Bosn J Basic Med Sci 17:9–11
Gucalp A, Tolaney S, Isakoff SJ, Ingle JN, Liu MC, Carey LA, Blackwell K, Rugo H, Nabell L, Forero A, Stearns V, Doane AS, Danso M, Moynahan ME, Momen LF, Gonzalez JM, Akhtar A, Giri DD, Patil S, Feigin KN, Hudis CA, Traina TA, Translational Breast Cancer Research Consortium (TBCRC 011) (2019) Phase II trial of bicalutamide in patients with androgen receptor-positive, estrogen receptor-negative metastatic Breast Cancer. Clin Cancer Res 19:5505–5512
O’Shaughnessy J, Campone M, Brain E, Neven P, Hayes D, Bondarenko I, Griffin TW, Martin J, De Porre P, Kheoh T, Yu MK, Peng W, Johnston S (2016) Abiraterone acetate, exemestane or the combination in postmenopausal patients with estrogen receptor-positive metastatic breast cancer. Ann Oncol 27:106–113
Traina TA, Miller K, Yardley DA, Eakle J, Schwartzberg LS, O’Shaughnessy J, Gradishar W, Schmid P, Winer E, Kelly C, Nanda R, Gucalp A, Awada A, Garcia-Estevez L, Trudeau ME, Steinberg J, Uppal H, Tudor IC, Peterson A, Cortes J (2018) Enzalutamide for the treatment of androgen receptor-expressing triple-negative breast cancer. J Clin Oncol 36:884–890
Acknowledgements
We thank the Nottingham Health Science Biobank and Breast Cancer Now Tissue Bank for the provision of tissue samples.
Funding
This study was funded by the University of Nottingham (Nottingham Life Cycle 6).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
Ibraheem Alshankyty is a consultant/advisory board in Molecular Diagnostics Lab, College of Applied Med. Sci., KAU. All authors of this work declare that they have no conflict of interest.
Ethical approval
This study was approved by the Nottingham Research Ethics Committee 2 (Reference title: Development of a molecular genetic classification of breast cancer). All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki Declaration and its later amendments or comparable ethical standards.
Informed consent
Informed consent was obtained from the participants included in the study.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Kurozumi, S., Joseph, C., Raafat, S. et al. Utility of ankyrin 3 as a prognostic marker in androgen-receptor-positive breast cancer. Breast Cancer Res Treat 176, 63–73 (2019). https://doi.org/10.1007/s10549-019-05216-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-019-05216-w