Abstract
Purpose
Obesity and weight gain are associated with comorbidities including a higher risk of tumor recurrence and cancer-related deaths among breast cancer (BC) survivors; however, the underlying mechanisms linking obesity and cancer are poorly understood. Given the lack of clinically validated BC biomarkers, obesity and weight-loss studies utilize serum biomarkers as the intermediary outcomes of tumor recurrence. Studies have indicated microRNAs (miRNA)s are reliable biomarkers for cancer. We hypothesized that miRNA expression correlates with obesity and weight loss amongst BC survivors. This would yield insight into the biological pathways by which this association occurs, enabling more precise development of therapeutics.
Patients and methods
We correlated baseline body mass index (BMI) with serum miRNA expression in 121 BC survivors enrolled in the Hormones and Physical Exercise (HOPE) trial. We then analyzed expression of the 35 most abundant miRNAs from HOPE in a six-month randomized controlled weight-loss trial (Lifestyle, Exercise, and Nutrition; LEAN) in 100 BC survivors. Ingenuity pathway analysis (IPA) software was used to identify biological pathway targets of the BMI-associated and intervention-responsive miRNAs using predictive biomarkers.
Results
Pearson correlations in HOPE identified eight miRNAs associated with BMI, including miR-191-5p (r = − 0.22, p = 0.016) and miR-122-5p (r = 0.25, p = 0.0048). In the LEAN validation study, levels of miR-191-5p significantly increased during the six-month intervention (p = 0.082). Ingenuity Pathway Analysis identified “Estrogen-mediated S-phase entry” (HOPE p = 0.003; LEAN p < 0.001) and “Molecular mechanisms of cancer” (HOPE p = 0.02; LEAN p < 0.001) as the top canonical pathways that significantly correlated with BMI-associated and intervention-responsive miRNAs and contain obesity and cancer-relevant genes including the E2F family of transcription factors and CCND1, which have been implicated in sporadic BC.
Conclusion
While the association between obesity and BC recurrence and mortality has been demonstrated in the literature, mechanisms underlying the link between weight gain and cancer are unclear. Using two independent clinical trials, we identified novel miRNAs associative to BMI and weight loss that contribute to the development of cancer. Predictive modeling of miRNA targets identified multiple canonical pathways associated with cancer, highlighting potential mechanisms explaining the link between BMI and increased cancer risk.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In the United States, over 200,000 women will be diagnosed with invasive BC annually and is the second leading cause of cancer-related death in women [1,2,3]. Only 10% of BC cases are due to a genetic predisposition, indicating that a number of environmental and anthropometric factors influence BC risk [4,5,6,7,8]. Lifestyle factors associated with higher BC risk include poor diet, alcohol intake, and smoking [9,10,11]. Studies consistently indicate post-menopausal women who are obese have an increased risk for breast cancer-specific mortality as compared to women with a normal BMI both prior to and post-diagnosis [12,13,14]. This increased risk is particularly concerning since obesity rates are increasing faster in post-menopausal women with cancer than amongst women without a history of cancer (3.01% vs. 2.31% increase annually within the past 10 years) [15]. Adult weight gain is also associated with a 64% increased risk of BC death among survivors [16]. Elucidating the molecular factors associated with the rise in obesity will play an important role in developing biomarkers associated with obesity-related BC recurrence.
The proposed mechanisms through which obesity and weight change affect cancer risk and survival involve an abundance of metabolically active adipose tissue that results in elevated levels of blood glucose, insulin, free estradiol, and inflammatory cytokines [16,17,18,19]. The heightened levels of these cytokines support a hyper-proliferative state of the surrounding epithelium, and therefore, a number of regulatory checkpoints are maintained to inhibit this process. miRNAs, small non-coding RNAs, are one such entity that regulates various cellular processes [20,21,22,23] and are highly dysregulated in chronic diseases, such as cancer [24,25,26,27,28,29]. For instance, miR-21, miR-155, and miR-10b are all deregulated in BC tissue [30]. Additionally, forced expression of these miRNAs in vitro promotes cell invasion, proliferation, and pro-survival phenotypes, by targeting a number of tumor suppressor genes including PDCD4, PTEN, SOCS1, and HOXD10. miRNAs are also expressed in the serum of BC patients, and function as reliable biomarkers for the disease [31]. Few studies have assessed how miRNAs regulate the cellular pathways controlling weight gain and metabolic homeostasis. For instance, serum levels of miR-122 and miR-519d are strongly correlated with obesity [32,33,34], and pathway analysis indicates these miRNAs target regulators of survival and proliferation pathways important in BC tumorigenesis. Understanding, which miRNAs correlate to BMI and weight-loss intervention will provide a unique insight into the mechanisms underlying the geneticlinks between these factors and BC recurrence.
In the present study, we tested the correlation between BMI and cancer-related miRNAs with obesity and a weight-loss intervention within two completed randomized control trials among BC survivors. First, we used baseline serum samples from BC survivors enrolled in the HOPE study [35] to identify miRNAs associated with BMI in this population; second, we utilized serum samples from the LEAN [36] trial to assess the effect of weight-loss intervention and the change in miRNA expression over six months comparing intervention and usual care study arms. We also tested miRNA expression in relation to body composition and serum biomarkers. We hypothesized that certain miRNAs and the pathways they regulate would correlate with BMI and change in response to weight-loss intervention, offering some insight into the mechanisms linking obesity to BC recurrence, and perhaps even identifying novel targets to improve survival outcomes for these patients.
Methods
Participants and study background
The methodologies of the HOPE and LEAN trials have been previously described [35, 36]. In brief, 121 inactive, post-menopausal BC survivors with joint pain were recruited to the HOPE trial between 2010 and 2012 from hospitals in Connecticut through the Rapid Case Ascertainment Shared Resource of the Yale Cancer Center [35]. Participants had a history of hormone receptor-positive stage I to III BC and received aromatase inhibitor (AI) adjuvant therapy for at least six months. Assessments of HOPE data in this manuscript evaluated blood draws and anthropometric measurements at the baseline (pre-intervention) time-point.
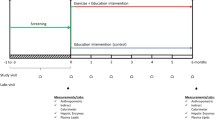
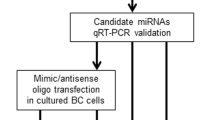
The LEAN trial [36, 37] was a three-arm weight-loss randomized study involving 100 BC survivors comparing (1) in-person counseling, (2) telephone counseling, versus (3) usual care. Participants included in the study were diagnosed with stage 0–III BC survivors, had a BMI > 25.0 kg/m2, and completed chemo- and/or radiation-therapy (Fig. 1). Since no significant weight loss occurred between the in-person and telephone counseling arm, these arms were combined into an intervention group for our analysis, while arm three remained the usual care group. LEAN weight-loss intervention strategies were adapted from US Dietary Guidelines as well as the American Institute for Cancer Research (AICR) nutritional and physical activity guidelines [38,39,40,41,42]. The usual care group was described in Irwin et al. [37]. Assessments of LEAN data in this manuscript evaluated blood draws and anthropometric measurements at both baseline and 6 months post-intervention.
For both HOPE and LEAN trials, institutional review board protocol was approved by the Yale School of Medicine Human Investigation Committee, and the Connecticut Department of Public Health Human Investigation Committee. Certain data used in this study were obtained from the Connecticut Tumor Registry located in the Connecticut Department of Public Health. The author(s) assumes full responsibility for analyses and interpretation of these data. The clinical characteristics of HOPE and LEAN trials are outlined in Table 1.
Primary and secondary measures
For participants enrolled in HOPE and LEAN, height, weight, and BMI (kg/m2) were determined as previously described [35, 36]. In LEAN, weight change was additionally calculated using the baseline and six-month weight measures. In HOPE, dual-energy x-ray absorptiometry scans were performed to assess body fat, lean body mass, and bone mineral density. For biomarker analysis, fasting blood draws were obtained, and serum was barcoded, aliquoted, and stored at − 80 °C until assayed. Circulating insulin, leptin, glucose, adiponectin concentrations, as well as IL-6, TNF-α, and C-reactive protein (CRP) levels were also measured as previously described [35,36,37, 43]. Sample specimens were assayed at study endpoint and measured in duplicate, with laboratory technicians blinded to treatment assignment. Coefficient of variation for samples was under 10%.
RNA collection and isolation
In brief, 150 µl of serum from patients under fasting conditions enrolled in HOPE with a BMI of 18.5–44 kg/m2 was arrayed onto 96-well plates and assayed for miRNA expression using the Firefly platform [44]. A total of 68 BC-specific miRNAs were assayed in multiplex on 121 samples arrayed onto two assay plates, with each plate containing control wells for post-hybridization analysis. Data were processed as described below. After assessment of HOPE, miRNA expression was then assessed from patients enrolled in LEAN. Specifically, 100 baseline samples and 85 6-month post-intervention samples were analyzed for 35 robustly detectable BC-specific miRNAs. Samples were processed similarly for Firefly detection.
RNA processing and miRNA expression analysis
In HOPE, baseline serum samples were analyzed for miRNA expression. In LEAN, miRNA expression was assessed at both baseline and six months following intervention [35, 36]. Firefly methodology was employed as previously described [44,45,46]. Samples were scanned on an EMD Millipore Guava 6HT flow cytometer, and flow cytometer output was analyzed with Firefly Analysis Workbench software (Abcam plc., Cambridge, MA). Target-specific background subtraction was performed to remove the signal from each target contributed by the negative control wells. For normalization, the geNorm [47] algorithm was used to calculate probe stability and determine expression of all targets above a defined threshold across all samples. Probes that did not meet these standards were removed. For quantification, raw signal values from each assay were assessed for expression above the calculated noise threshold of the assay, which was a raw mean fluorescent intensity (RMFI) of eighty, log2 scale. Expression values underwent background subtraction and “geNorm+average analysis”, to normalize the expression of miRNA candidates within the entire sample dataset across the average signal from the 20 most invariant-expressed miRNAs. Hierarchal clustering analysis using a root-mean square clustering metric with complete linkage analysis was used to assess quality of the miRNA expression dataset.
The criterion for candidate miRNA inclusion was as follows, (1) to be associative with known breast tumorigenic processes [25, 48,49,50]; and/or (2) known to regulate pathways associated with metabolism [51,52,53], adipogenesis, and/or obesity [54,55,56]; and/or (3) previously identified to be detectable in circulating biofluids such as plasma or serum [57,58,59]. We identified 68 miRNAs eligible for analysis; these miRNAs were assessed in crude serum isolates from the HOPE trial utilizing the probe-hybridization multiplex profiling assay mentioned above [44]. This procedure reduces the risk of error associated with RNA extraction of clinical samples from small sample volumes.
When testing miRNA expression from samples in the LEAN trial, we used the top 50% most robust and consistently expressed miRNA probes from the original 68. This was done because a number of miRNAs were not consistently detectable across a number of samples in the HOPE dataset (see Figure S1 and Figure S2), as well as in patients with a BMI 30 kg/m2. Given all samples in LEAN were from overweight patients, the inclusion of those miRNAs in future analyses was not performed. See Figure S3 for the list of the 35 miRNAs used in the LEAN cohort analysis.
Target miRNA analyses
miRNA targets were compared to the clinical endpoints and secondary measures from both HOPE and LEAN, as described above. Pearson correlation coefficients between each miRNA and clinical endpoint were computed using SAS software [60]. p < 0.1 indicated statistical significance for BMI-associated miRNAs in HOPE. In LEAN, 6-month changes in miRNA expression were compared to the randomized control cohort. The changes in miRNA abundance were determined by a least square means analysis, with a ± 95% confidence interval as a cut-off. For exploratory purposes, p < 0.1 indicated a statistically significant cutoff for intervention-associated miRNAs in LEAN.
Biological pathway analyses
The miRNAs identified in HOPE associated with BMI (p < 0.05; N = 8) and in LEAN associated with changes post-intervention (p < 0.05; N = 6) were uploaded into Ingenuity Pathway Analysis (IPA) Suite (Qiagen Inc., Redwood City, CA) for biological pathway analyses. IPA utilizes miRNA seed sequence binding with cognate mRNA targets to identify miRNA:mRNA target interactions across multiple available bioinformatic data sources. Since, miR-93_5p and miR-17_5p, two BMI-associated miRNAs, have identical seed sequences, miR-93_5p was considered a duplicate miRNA for predictive modeling. Therefore, the final miRNA number used for BPA predictions was an N of 7 for HOPE and an N of 6 for LEAN. We utilized a conservative approach within the IPA microRNA Target Filter for mRNA target prediction, whereby only experimentally verified interactions, as determined by miRTarBase [61], and/or highly conserved target sites containing 8-mer seed binding, as determined by Target Scan [62] algorithms, were selected. This approach produced a HOPE-specific gene target list of 994 mRNAs, and a LEAN-specific gene target list of 1292 mRNAs. Each gene list was used by IPA to determine enriched canonical pathways targeted by BMI-associated and intervention-responsive miRNAs. Representation of each canonical pathway was determined using a fisher exact test of miRNA-targeted genes within our dataset, as compared to the total number of genes in each IPA pathway.
Results
HOPE analysis
To identify BMI-associated miRNAs, baseline serum from those enrolled in HOPE was assayed for miRNA expression. We calculated Pearson correlation coefficients to identify miRNAs correlated with baseline BMI in HOPE (Table 2). We identified eight miRNAs significantly associated with BMI with the strongest positive associations as miR-22_3p (r = 0.26, p = 0.004) and miR-122_5p (r = 0.25, p = 0.005), both of which are implicated in BC metabolism [63,64,65]. miRNAs with the strongest negative BMI association were miR-191_5p (r = − 0.22, p = 0.016) and miR-17_5p (r = − 0.22, p = 0.017), which are implicated in influencing BC progression [66, 67].
To determine the genetic pathways these miRNAs regulate, we assessed the top ten target mRNA genes for each BMI-associated miRNA via TargetScan (Table S1). A number of these target genes are members of either pro-inflammatory and Notch signaling pathways, or glycolytic and lipid metabolic pathways. Utilizing formal canonical pathway analysis, we identified that mRNAs targeted by these BMI-associated miRNAs also had a biological relevance for cancer. Of note, two of the eight miRNAs identified, miR-17_5p and miR-93_5p, have the same seed sequence and therefore the same mRNA targets. Probes for both miRNAs negatively associated with BMI and had the same magnitude of association, as was expected. Thus, seven miRNA targets of interest were used for pathway analysis. The most significant pathways altered were the “Estrogen mediated S-phase entry”(p = 0.003) and the “P53 signaling” pathways (p = 0.006). Specific genes in these pathways are listed in Table 3, and include a number of genes that regulate the process of cell proliferation, survival, and motility.
LEAN analysis
We identified miRNAs responsive to weight-loss intervention by assessing the change in miRNA expression at both baseline and six months in the diet and exercise intervention group, as compared to the usual care group within LEAN using a least square means analysis, with a ± 95% confidence interval. Six miRNAs were significantly different (p < 0.10) between the intervention and usual care arms (Table 4). Most notably, miR-106_5p decreased in expression in the intervention arm [effect change − 1.0 (− 9.0 to 6.9)] and increased in the usual care arm [effect change 12.5 (2.3–22.7)], as compared to baseline, p = 0.041. This is in line with the finding that miR-106b_5p is a prognostic indicator of BC recurrence and also targets cell-cycle regulators in BC cells [68, 69]. Another miRNA, miR-191_5p, had the largest absolute difference in change between usual care and intervention arms, and the greatest increase in expression in the intervention arm [effect change 32.1 (6.6–57.6)], as compared to baseline, p = 0.082. miR-191_5p was also significantly negatively associated with BMI in the HOPE analysis indicating this miRNA functions as a predictor of weight loss in overweight BC survivors.
To determine the genetic pathways these miRNAs regulate, we assessed the top 10 target genes for each intervention-responsive miRNA (Table S2). These target genes control either Notch signaling or apoptosis pathways, indicating these miRNAs regulate a distinct set of pathways from that of HOPE-associated miRNAs. Canonical pathway analysis was performed to determine the molecular underpinnings of weight-loss responsiveness. The most significant pathways identified in LEAN included the “Molecular mechanisms of cancer” (p = 6.8 × 10−7) and “Estrogen mediated S-phase entry” (p = 1.4 × 10−6). Specific genes in these pathways are listed in Table 5 and include a number of genes that control cancer-related processes.
While assessing the molecular pathways regulated by BMI-associated and intervention-responsive miRNAs in HOPE and LEAN, we noticed “Estrogen mediated S-phase entry” (HOPE, p = 0.003; LEAN, p = 1.4 × 10−6) and the “Molecular mechanisms of cancer” (HOPE, p = 0.0016; LEAN, p = 6.8 × 10−7) were the most significantly modified pathways (Table S3). When assessing gene targets relevant to these pathways, we found the presence of the E2F family of transcription factors, which controls cell cycle and in tumor suppressor proteins. Other targets identified included genes related to cell-cycle progression at G1, such as CDKN1A and CCND1, both of which have been linked to sporadic BC [70,71,72].
Secondary analyses (HOPE)
A secondary analysis was also performed to determine if any of the BMI-associated miRNAs in HOPE correlated to intermediary outcomes such as inflammatory serum biomarkers, and body composition (Table 6). Among the five biomarkers tested, miR-22_3p positively correlated with circulating CRP (p = 0.005), IL6 (p < 0.001), glucose (p < 0.001), insulin (p = 0.003), and leptin (p < 0.001) levels. However, BMI-associated miRNAs such as miR-17_5p negatively correlated with IL6 (p = 0.001) and leptin (p = 0.001) levels, while miR-191_5p negatively correlated with IL6 levels (p = 0.05) indicating these miRNAs predict BMI independent of glycolytic-related pathways. Similar results were observed with miR-93_5p, as expected given this miRNA has a similar seed sequence to miR-17_5p.
Analysis of body composition measures (e.g., bone mineral content, total fat, etc.) yielded a distinct subset of miRNAs not linked with BMI (Table 6). For instance, miR-27a_3p was significantly negatively associated with four of the five body measures assessed, and target genes such as NOVA1 which controls RNA splicing and is found in the serum of patients with paraneoplastic opsoclonus-ataxia and BC [73, 74]. While miR-122_5p did not correlate with any body composition measures, there was an association with insulin (p < 0.001) and IL6 (p < 0.001), consistent with the reported literature [75].
Discussion
High BMI and weight gain are associated with poor outcomes amongst BC survivors, yet the underlying mechanisms explaining this association are unclear. Studies indicate free fatty acids, glucose, and eicosanoids promote cell-cycle proliferation, inhibit the activity of pro-apoptotic pathways, and/or induce changes in cellular lipid architecture that enhances cellular migration [76,77,78,79]. Activities of these processes are regulated by the transcription factors E2F2 and PPARγ. PPARγ is a transcription factor that hetero-dimerizes with RXR to induce the expression of genes essential for cellular metabolism. PPARγ expression is altered in a variety of solid carcinomas [80, 81], and in mouse models of sporadic BC, PPARγ controls cell-cycle progression by regulating checkpoint genes such as CDKN1A and CCND1 [82, 83]. E2F2 belongs to a family of nine transcription factors that control cellular proliferation and apoptosis, and are also suppressed during tumorigenesis [84, 85]. E2F2 is central to the cellular timing of G1/S phase transitions by controlling the expression of cyclins, FGF2, and SOX2 [86].
E2F2, PPARγ, and other BC-associated genes are regulated via phosphorylation of E2F-interacting proteins such as Rb and CCND2, and are tightly controlled at the epigenetic level through histone methyltransferase activity, direct methylation of the gene promoter, and/or post-transcriptional regulation via interactions with non-coding RNA [87,88,89,90,91]. For instance, miR-10b and miR-122 are bona-fide regulators of PPARγ and regulate the pathogenesis of non-alcoholic fatty liver disease (NAFLD) by controlling hepatic stenosis [92, 93], while the miR-17/92 family regulates the expression of E2F, and subsequently the activity of the E2F/MYC signaling axis [91].
miRNAs control many functions of BC etiology as well as the metabolic pathways that support BC tumor growth and survival [94]. However, few studies have directly assessed the role of noncoding RNAs as an indicator of obesity and weight-loss intervention. A limiting factor has been the ability to detect circulating miRNAs in biofluids such as serum. In this study, we used an miRNA detection system that obviates the use of PCR to detect the copy number of a particular miRNA species from crude serum extract [44]. Using this system, we identified miR-122_5p as a BMI-associated miRNA, which was expected given miR-122_5p is essential in regulating lipid metabolism, and is associated with circulating insulin and leptin levels [57]. In fact, the original in vivo antisense studies targeted miR-122 in the adult liver of obese mice using a diet-induced model, effectively reducing plasma cholesterol levels and improving liver steatosis [32].
We also identified miRNAs such as miR-191_5p and miR-17_5p to be significantly inversely correlated with BMI, both of which are involved in tumorigenic processes. For instance, the miR-17_5p/miR-93_5p family was also inversely correlated with inflammatory markers such as CRP, IL6, and leptin, and is known to inhibit tumor growth by suppressing MYC-induced cellular proliferation [66, 95]. This miRNA cluster post-transcriptionally inhibits MYC gene expression, while E2F and MYC induce the transcription of miR-17_5p/miR-92_5p, forming a negative feedback loop resulting in a sharp “on/off” state for pro-tumorigeneic-related protein expression. A related miR-17_5p/miR-93_5p family member, miR-106b_5p, was unexpectedly found to be the most significant miRNA that decreased in response to diet and exercise intervention. However, a number of groups have shown that miRNAs of the same family can mediate various cellular responses through non-seed base-pairing interactions with target mRNAs [96]. In support of this, miR-106b_5p is upregulated in both tissue and plasma from BC patients, and promotes cell-cycle progression by targeting CDKN1A [68, 69]. Additional experiments are required to elucidate the molecular mechanisms behind the negative association of miR-17-5p with BMI, and the negative association of miR-106 with weight-loss intervention.
We identified that low circulating miR-191_5p levels are associated with high BMI and high circulating miR-191_5p levels are predictive of successful weight-loss post diet- and exercise-intervention. The mechanism for this is unclear given miR-191_5p associates with relatively few secondary measures in HOPE (IL6 = r − 0.182; p = 0.046). However, pathway analysis, which included miR-191_5p target genes, indicated potential mechanisms may involve 17β-estradiol signaling and S-phase entry by targeting genes such as CCND1, E2F2, RB1, and IRS1. While it is not known if miR-191_5p directly inhibits ESR1, it is known that miR-191_5p is dysregulated in ERα-positive BC, a 17β-estradiol-dependent tumor [67, 97]. Furthermore, 17β-estradiol promotes the expression of miR-191_5p and protects ERα-positive tumors from hormone depletion-induced apoptosis. This is relevant given all patients in HOPE and half of the patients in LEAN were on AI therapy, which blocks the synthesis of 17β-estradiol. Therefore, miR-191_5p may function as an early indicator of 17β-estradiol-specific signaling due to reduced adipose burden during weight-loss intervention, rather than as a measure of late-stage fluctuations in inflammatory and/or metabolic activity.
Circulating miRNAs are extremely stable biomarkers that can be prognostic indicators of disease onset as well as predictive biomarkers for drug response. A recent study measured 13 a priori selected miRNAs for change in response to a 16-week diet and exercise weight-loss intervention trial (N = 89 men and women), and identified miR-221_3p and miR-223_3p as increasing in both low- and high-weight-loss responders in response to intervention, as well as miR-140 as increasing in the low responder group [98]. None of the miRNAs identified in these studies overlapped with miRNAs captured from the LEAN and HOPE datasets. The likely explanation for this is that all patients enrolled in our trials were either overweight BC survivors or on AI therapy.
The overall strengths of our study include the utilization of two trials: one validating that our method can detect miRNAs previously associated with BMI in a population of BC survivors, and two measuring the change in miRNA expression within a separate BC population over a six-month weight-loss intervention. Our study is limited in that only a single intervention study was performed. Therefore, follow-up weight-loss intervention trials will be required to identify circulating miRNAs and target pathways associated with weight loss so as to provide opportunities to develop clinical biomarkers for a physiological response to weight-loss intervention. We also identified miRNAs related to hormone regulation; further studies are warranted to assess how miRNAs can be indicators of effective endocrine therapies in BC survivors.
References
Desantis C, Siegel R, Bandi P et al (2011) Breast cancer statistics, 2011. Cancer 61:409–418
Eheman CR, Shaw KM, Ryerson AB et al (2009) The changing incidence of in situ and invasive ductal and lobular breast carcinomas: United States, 1999–2004. Cancer Epidemiol Biomarkers Prev 18:1763–1769
Youlden DR, Cramb SM, Dunn NAM et al (2012) The descriptive epidemiology of female breast cancer: an international comparison of screening, incidence, survival and mortality. Cancer Epidemiol 36:237–248
Martin AM, Weber BL (2000) Genetic and hormonal risk factors in breast cancer. J Natl Cancer Inst 92:1126–1135
Cordell HJ, Clayton DG (2005) Genetic association studies. Lancet 366:1121–1131
Amadou A, Hainaut P, Romieu I (2013) Role of obesity in the risk of breast cancer: Lessons from anthropometry J. Oncol. https://doi.org/10.1155/2013/906495
Friedenreich CM (2001) Review of anthropometric factors and breast cancer risk. Eur J Cancer Prev 10:15–32
Turnbull C, Rahman N (2008) Genetic predisposition to breast cancer: past, present, and future. Annu Rev Genomics Hum Genet 9:321–345
Castello A, Martin M, Ruiz A et al (2015) Lower breast cancer risk among women following the world cancer research fund and american institute for cancer research lifestyle recommendations: EpiGEICAM case-control study. PLoS ONE 10:e0126096
Shapira N (2017) The potential contribution of dietary factors to breast cancer prevention. Eur J Cancer Prev 26(5):385
Nomura SJO, Inoue-Choi M, Lazovich D et al (2016) WCRF/AICR recommendation adherence and breast cancer incidence among postmenopausal women with and without non-modifiable risk factors. Int J Cancer 138:2602–2615
Sternfeld B, Weltzien E, Quesenberry CP et al (2009) Physical activity and risk of recurrence and mortality in breast cancer survivors: findings from the LACE study. Cancer Epidemiol Biomarkers Prev 18:87–95
Kroenke CH, Chen WY, Rosner B et al (2005) Weight, weight gain, and survival after breast cancer diagnosis. J Clin Oncol 23:1370–1378
Neuhouser ML, Aragaki AK, Prentice RL et al (2015) Overweight, obesity, and postmenopausal invasive breast cancer risk. JAMA Oncol 1:611
Greenlee H, Shi Z, Sardo Molmenti CL et al (2016) Trends in obesity prevalence in adults with a history of cancer: results from the US national health interview survey, 1997 to 2014. J Clin Oncol 34:3133–3140
Caan BJ, Kwan ML, Shu XO et al (2012) Weight change and survival after breast cancer in the after breast cancer pooling project. Cancer Epidemiol Biomarkers Prev 21:1260–1271
Chen X, Lu W, Zheng W et al (2010) Obesity and weight change in relation to breast cancer survival. Breast Cancer Res Treat 122:823–833
De Pergola G, Silvestris F (2013) Obesity as a major risk factor for cancer. J Obes. https://doi.org/10.1155/2013/291546
Stephenson GD, Rose DP (2003) Breast cancer and obesity: an update. Nutr Cancer 45:1–16
Baer C, Claus R, Plass C (2013) Genome-wide epigenetic regulation of miRNAs in cancer. Cancer Res 73:473–477
Croce CM, Calin GA (2005) miRNAs, cancer, and stem cell division. Cell 122:6–7
Visone R, Croce CM (2009) MiRNAs and Cancer. Am J Pathol 174:1131–1138
Fabbri M, Calin GA (2010) Epigenetics and miRNAs in Human Cancer. Adv Gene 70:87–99
Esquela-Kerscher A, Slack FJ (2006) Oncomirs—microRNAs with a role in cancer. Nat Rev Cancer 6:259–269
Adams BD, Kasinski AL, Slack FJ (2014) Aberrant regulation and function of microRNAs in cancer. Curr Biol 24(16):R762–R776
Adams B, Parsons C, Walker L et al (2017) Targeting noncoding RNAs in disease. J Clin Investig 127(3):761–771
Iorio MV, Croce CM (2009) MicroRNAs in cancer: small molecules with a huge impact. J Clin Oncol 27:5848–5856
Slack FJ, Weidhaas JB (2006) MicroRNAs as a potential magic bullet in cancer. Futur Oncol 2:73–82
Trang P, Weidhaas JB, Slack FJ (2009) MicroRNAs as potential cancer therapeutics. Oncogene 27:S52–S57
van Schooneveld E, Wildiers H, Vergote I et al (2015) Dysregulation of microRNAs in breast cancer and their potential role as prognostic and predictive biomarkers in patient management. Breast Cancer Res 17:21
Adams BD, Parsons C, Slack FJ (2016) The tumor-suppressive and potential therapeutic functions of miR-34a in epithelial carcinomas. Expert Opin Ther Targets 20:737–753
Esau C, Davis S, Murray SF et al (2006) miR-122 regulation of lipid metabolism revealed by in vivo antisense targeting. Cell Metab 3:87–98
Heneghan HM, Miller N, Kerin MJ (2010) MiRNAs as biomarkers and therapeutic targets in cancer. Curr Opin Pharmacol 10:543–550
McGregor RA, Choi MS (2011) microRNAs in the regulation of adipogenesis and obesity. Curr Mol Med 11:304–316
Irwin ML, Cartmel B, Gross CP et al (2015) Randomized exercise trial of aromatase inhibitor-induced arthralgia in breast cancer survivors. J Clin Oncol 33:1104–1111
Harrigan M, Cartmel B, Loftfield E et al (2016) Randomized trial comparing telephone versus in-person weight loss counseling on body composition and circulating biomarkers in women treated for breast cancer: the lifestyle, exercise, and nutrition (LEAN) study. J Clin Oncol 34:669–676
Irwin ML, Alvarez-Reeves M, Cadmus L et al (2009) Exercise improves body fat, lean mass, and bone mass in breast cancer survivors. Obesity 17:1534–1541
Knowler WC, Barrett-Connor E, Fowler SE et al (2002) Reduction in the incidence of type 2 diabetes with lifestyle intervention or metformin. N Engl J Med 346:393–403
Rock CL, Doyle C, Meyerhardt W et al (2012) Nutrition and Physical Activity Guidelines for Cancer Survivors. CA Cancer J Clin 62:243–274
US Department of Health and Human Services and US Department of Agriculture (2015) Dietary guidelines for Americans, 2015–2020. vol 232, 8th edn p 71 http://health.gov/dietaryguidelines/2015/guidelines/
Rogers D, Leonberg BL, Broadhurst CB et al (2002) 2000 Commission on dietetic registration dietetics practice audit. J Am Diet Assoc 102:270–292
Griswold K, Rogers D, Sauer KL et al (2016) Entry-level dietetics practice today: results from the 2015 commission on dietetic registration entry-level dietetics practice audit. J Acad Nutr Diet 116:1632–1684
Jones SB, Thomas GA, Hesselsweet SD et al (2013) Effect of exercise on markers of inflammation in breast cancer survivors: the yale exercise and survivorship study. Cancer Prev Res 6:109–118
Pregibon DC, Toner M, Doyle PS (2007) Multifunctional encoded particles for high-throughput biomolecule analysis. Science 315(5817):1393–1396
Adams BD, Wali VB, Cheng CJ (2015) miR-34a Silences c-SRC to attenuate tumor growth in triple-negative breast cancer. Cancer Res 76(4):927–939
Chapin SC, Appleyard DC, Pregibon DC et al (2011) Rapid microRNA profiling on encoded gel microparticles. Angew Chemie—Int Ed 50:2289–2293
Vandesompele J, De Preter K, Pattyn F et al (2002) Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol 3:34
Metheetrairut C, Adams BD (2016) Nallur Scel-mir-237 and its homologue, hsa-miR-125b, modulate the cellular response to ionizing radiation. Oncogene 36(4):512
Adams BD, Cowee DM, White BA (2009) The Role of miR-206 in the epidermal growth factor (EGF) induced repression of estrogen receptor-α (ERα) Signaling and a luminal Phenotype in MCF-7 Breast Cancer Cells. Mol Endocrinol 23:1215–1230
Adams B, Guttilla I, White B (2008) Involvement of MicroRNAs in breast cancer. Semin Reprod Med 26(6):522–536
Ling H, Li X, Yao CH et al (2013) The physiological and pathophysiological roles of adipocyte miRNAs. Biochem Cell Biol 91:195–202
Trajkovski M, Lodish H (2013) MicroRNA networks regulate development of brown adipocytes. Trends Endocrinol Metab 24:442–450
Chen L, Song J, Cui J et al (2013) Micrornas regulate adipocyte differentiation. Cell Biol Int 37:533–546
Cui J, Zhou B, Ross SA et al (2017) Nutrition, microRNAs, and human health. Adv Nutr An Int Rev J 8:105–112
Sliwinska A, Kasinska MA, Drzewoski J (2017) MicroRNAs & metabolic disorders—where are we heading? Arch Med Sci 13:885–896
Nardelli C, Iaffaldano L, Pilone V (2017) Changes in the MicroRNA profile observed in the subcutaneous adipose tissue of obese patients after laparoscopic adjustable gastric banding. J Obes. https://doi.org/10.1155/2017/6754734
Ortega FJ, Mercader JM, Catalán V et al (2013) Targeting the circulating microRNA signature of obesity. Clin Chem 59:781–792
Blenkiron C, Goldstein LD, Thorne NP et al (2007) MicroRNA expression profiling of human breast cancer identifies new markers of tumor subtype. Genome Biol 8:R214
Witwer KW (2012) XenomiRs and miRNA homeostasis in health and disease: evidence that diet and dietary miRNAs directly and indirectly influence circulating miRNA profiles. RNA Biol 9:1147–1154
SAS Institute Inc (2014) SAS software 9.4. SAS Inst Inc Mark Co pp 1–25
Da H, Lin FM, Wu WY (2011) MiRTarBase: a database curates experimentally validated microRNA-target interactions. Nucleic Acids Res 39:D163–D169
Bartel DP (2009) MicroRNAs: target recognition and regulatory functions. Cell 136:215–233
Fong MY, Zhou W, Liu L et al (2015) Breast-cancer-secreted miR-122 reprograms glucose metabolism in premetastatic niche to promote metastasis. Nat Cell Biol 17:183–194
Pola C (2013) Cancer: miR-22 attacks on several fronts. Nat Med 19:980
Koufaris C, Valbuena GN, Pomyen Y et al (2016) Systematic integration of molecular profiles identifies miR-22 as a regulator of lipid and folate metabolism in breast cancer cells. Oncogene 35:2766–2776
Fan M, Sethuraman A, Brown M et al (2014) Systematic analysis of metastasis-associated genes identifies miR-17-5p as a metastatic suppressor of basal-like breast cancer. Breast Cancer Res Treat 146:487–502
Di Leva G, Piovan C, Gasparini P (2013) Estrogen mediated-activation of miR-191/425 cluster modulates tumorigenicity of breast cancer cells depending on estrogen receptor Status. PLoS genet 9(3):e1003311
Zheng R, Pan L, Gao J et al (2015) Prognostic value of miR-106b expression in breast cancer patients. J Surg Res 195:158–165
Ivanovska I, Ball AS, Diaz RL et al (2008) MicroRNAs in the miR-106b family regulate p21/CDKN1A and promote cell cycle progression. Mol Cell Biol 28:2167–2174
Wei CY, Tan QX, Zhu X et al (2015) Expression of CDKN1A/p21 and TGFBR2 in breast cancer and their prognostic significance. Int J Clin Exp Pathol 8:14619–14629
Bièche I, Olivi M, Noguès C et al (2002) Prognostic value of CCND1 gene status in sporadic breast tumours, as determined by real-time quantitative PCR assays. Br J Cancer 86:580–586
Kenemans P, Verstraeten RA, Verheijen RHM (2004) Oncogenic pathways in hereditary and sporadic breast cancer. Maturitas 49(1):34–43
Stich O, Rasiah C, Rauer S (2009) Paraneoplastic antibody during follow-up of a patient with anti-Ri-associated paraneoplastic neurological syndrome. Acta Neurol Scand 119:338–340
Zhang YA, Liu HN, Zhu JM et al (2016) RNA binding protein Nova1 promotes tumor growth in vivo and its potential mechanism as an oncogene may due to its interaction with GABAA Receptor-gamma2. J Biomed Sci 23:71
Small EM, Olson EN (2011) Pervasive roles of microRNAs in cardiovascular biology. Nature 469:336–342
Spiegelman BM, Flier JS (1996) Adipogenesis and obesity: rounding out the big picture. Cell 87:377–389
Jeffery E, Wing A, Holtrup B et al (2016) The adipose tissue microenvironment regulates depot-specific adipogenesis in obesity. Cell Metab 24:142–150
de sá PM, Richard AJ, Hang H et al (2017) Transcriptional regulation of adipogenesis. Compr Physiol 7:635–674
Cao Y (2007) Angiogenesis modulates adipogenesis and obesity. J Clin Invest 117:2362–2368
Kersten S, Desvergne B, Wahli W (2000) Roles of PPARs in health and disease. Nature 405:421–424
Tachibana K, Yamasaki D, Ishimoto K (2008) The role of PPARs in cancer. PPAR Res. https://doi.org/10.1155/2008/102737
Dabydeen SA, Furth PA (2014) Genetically engineered ERα-positive breast cancer mouse models. Endocr Relat Cancer 21(3):R195–R208
Yao PL, Morales JL, Zhu B et al (2014) Activation of peroxisome proliferator-activated receptor-β/δ (PPAR-β/δ) inhibits human breast cancer cell line tumorigenicity. Mol Cancer Ther 13:1008–1017
Ho GH, Calvano JE, Bisogna M et al (2001) Expression of E2F-1 and E2F-4 is reduced in primary and metastatic breast carcinomas*. Breast Cancer Res Treat 69:115–122
Bracken AP, Ciro M, Cocito A et al (2004) E2F target genes: unraveling the biology. Trends Biochem Sci 29:409–417
Julian LM, Blais A (2015) Transcriptional control of stem cell fate by E2Fs and pocket proteins. Front Genet 6:161
Wang L, Xu S, Lee JE et al (2012) Histone H3K9 methyltransferase G9a represses PPARγ expression and adipogenesis. EMBO J 32:45–59
Portius D, Sobolewski C, Foti M (2017) MicroRNAs-dependent regulation of PPARs in metabolic diseases and cancers. PPAR Res. https://doi.org/10.1155/2017/7058424
Johnson KC, Koestler DC, Cheng C et al (2014) Age-related DNA methylation in normal breast tissue and its relationship with invasive breast tumor methylation. Epigenetics 9:268–275
Narita M, Nũnez S, Heard E et al (2003) Rb-mediated heterochromatin formation and silencing of E2F target genes during cellular senescence. Cell 113:703–716
Emmrich S, Pützer BM (2010) Checks and balances: E2F-microRNA crosstalk in cancer control. Cell Cycle 9:2555–2567
Gatfield D, Le Martelot G, Vejnar CE et al (2009) Integration of microRNA miR-122 in hepatic circadian gene expression. Genes Dev 23:1313–1326
Zheng L, Lv G, Sheng J et al (2010) Effect of miRNA-10b in regulating cellular steatosis level by targeting PPAR-Î ± expression, a novel mechanism for the pathogenesis of NAFLD. J Gastroenterol Hepatol 25:156–163
Hatziapostolou M, Polytarchou C, Iliopoulos D (2013) MiRNAs link metabolic reprogramming to oncogenesis. Trends Endocrinol Metab 24:361–373
Aguda BD, Kim Y, Piper-Hunter MG et al (2008) MicroRNA regulation of a cancer network: consequences of the feedback loops involving miR-17-92, E2F, and Myc. Proc Natl Acad Sci 105:19678–19683
Helwak A, Kudla G, Dudnakova T et al (2013) Mapping the human miRNA interactome by CLASH reveals frequent noncanonical binding. Cell 153:654–665
Nagpal N, Kulshreshtha R (2014) miR-191: An emerging player in disease biology. Front Genet 5:99
Milagro FI, Miranda J, Portillo MP (2013) High-throughput sequencing of microRNAs in peripheral blood mononuclear Cells: identification of potential weight loss biomarkers. PLoS One 8(1):e54319
Acknowledgements
This work was supported by grants to B.D. Adams from NIH P50 CA196530, a Firefly Pilot Grant award, and from start-up funds through The RNA Institute, and The State University of New York. B.D. Adams is also President and CEO of The Brain Institute of America (brian.adams@braininstitueamerica.com) and holds patent interests with AUM LifeTech. B. Cartmel is a consultant/advisor for Pfizer. F. Li is supported by Yale CTSA grant UL1TR000142, and Yale Cancer Center Support Grant(CCSG/P30). M. Harrigan and T. Sanft are supported by grant NIH 1R01CA207753-01A1. T. Sanft is also a consultant/advisor to bioTheranostics. C.J. Cheng is currently an employee of Alexion Pharmaceuticals. L. Pusztai is supported by a Breast Cancer Research Foundation Award. M.L. Irwin is supported by grants from NCI R01CA132931, the American Institute for Cancer Research, and by the Breast Cancer Research Foundation, as well as through a Yale Cancer Center Support Grant P30CA016359, and a Clinical and Translational Science Award NCATS UL1TR000142. We thank Jessica Tytell, Irene G. Reed, and Elizabeth Posey for the critical reading of this manuscript. We thank Mike Tackett at Firefly for answering questions regarding miRNA detection within serum samples. Other authors have declared that no conflict of interest exists. This study analyzed data obtained from clinical trials NCT02056067 and NCT02109068.
Author information
Authors and Affiliations
Contributions
Conception and design: Brian D. Adams, Melinda L. Irwin. Financial support: Brian D. Adams, Melinda L. Irwin. Provision of study materials or patients: Melinda L. Irwin. Collection and assembly of data: Brian D. Adams, Monica J. Hubal, Melinda L. Irwin. Data analysis and interpretation: all authors. Manuscript writing: all authors
Corresponding author
Ethics declarations
Conflicts of interest
B.D. Adams is President and CEO of The Brain Institute of America and holds patent interests with AUM LifeTech. Other authors have declared no conflicts of interest exist.
Clinical trial
This study analyzed data obtained from clinical trials Hormones and Physical Exercise (HOPE) Study; NCT02056067, https://clinicaltrials.gov/ct2/show/NCT02056067; and Lifestyle, Exercise and Nutrition (LEAN) Study; NCT02109068, https://clinicaltrials.gov/ct2/show/NCT02109068.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Adams, B.D., Arem, H., Hubal, M.J. et al. Exercise and weight loss interventions and miRNA expression in women with breast cancer. Breast Cancer Res Treat 170, 55–67 (2018). https://doi.org/10.1007/s10549-018-4738-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-018-4738-6