Abstract
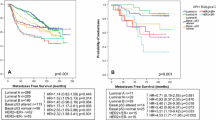
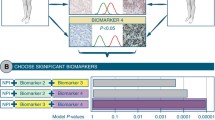
Accurate distant metastasis (DM) prediction is critical for risk stratification and effective treatment decisions in breast cancer (BC). Many prognostic markers/models based on tissue marker studies are continually emerging using conventional statistical approaches analysing complex/dimensional data association with DM/poor prognosis. However, few of them have fulfilled satisfactory evidences for clinical application. This study aimed at building DM risk assessment algorithm for BC patients. A well-characterised series of early invasive primary operable BC (n = 1902), with immunohistochemical expression of a panel of biomarkers (n = 31) formed the material of this study. Decision tree algorithm was computed using WEKA software, utilising quantitative biomarkers’ expression and the absence/presence of distant metastases. Fifteen biomarkers were significantly associated with DM, with six temporal subgroups characterised based on time to development of DM ranging from <1 to >15 years of follow-up. Of these 15 biomarkers, 10 had a significant expression pattern where Ki67LI, HER2, p53, N-cadherin, P-cadherin, PIK3CA and TOMM34 showed significantly higher expressions with earlier development of DM. In contrast, higher expressions of ER, PR and BCL2 were associated with delayed occurrence of DM. DM prediction algorithm was built utilising cases informative for the 15 significant markers. Four risk groups of patients were characterised. Three markers p53, HER2 and BCL2 predicted the probability of DM, based on software-generated cut-offs, with a precision rate of 81.1 % for positive predictive value and 77.3 %, for the negative predictive value. This algorithm reiterates the reported prognostic values of these three markers and underscores their central biological role in BC progression. Further independent validation of this pruned panel of biomarkers is therefore warranted.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Distant recurrence is the major cause of cancer-related deaths in breast cancer (BC) patients [1]. For cancer cells to successfully colonise a secondary site, they have to fulfil specific prerequisites to overcome the vast stresses throughout the metastatic cascade [2]. Collectively, the success of the metastatic process results from integration and contribution of complex molecular pathways controlling cellular proliferation, survival, metabolism, invasion and migration [3].
Accuracy in BC prognosis/prediction, particularly distant recurrence risk assessment, is critical for accurate patients’ stratification and effective treatment decision-making. Many prognostic markers and models based on tissue marker studies are continually emerging; however, very few of them have fulfilled satisfactory evidence for clinical application. Poor study design and misleading statistical analyses have been proposed as to explain discrepancies in research studies generating relevant clinically useful prognostic markers [4].
Mining large datasets regarding the expression patterns of a large number of biomarkers and clinical variables requires stringent statistical approaches to derive robust conclusions. Decision tree is an approach followed to automatically learn, through machine learning, to recognise complex patterns and illustrate relations between observed variables to make intelligent decisions. Therefore, decision tree algorithms could help improving our basic understanding of cancer development and progression [5] which can be used to assist the classification of BC cases by sorting them based on feature or attribute values (e.g. tissue marker expression). Each node in a decision tree represents a feature in a case to be classified, and each branch represents a value that the node can assume. Cases are classified starting at the root node and sorted based on their feature values [6].
Perhaps the most common algorithm in literature for building decision trees is the C4.5 developed by Ross Quinlan. C4.5 is a statistical classifier system which uses the concept of information gain to make a tree of classificatory decisions with respect to a previously chosen target classification (e.g. distant recurrence). Each attribute of the data can be used to make a decision that splits the data into smaller subsets [7]. These outputs are then expressed as models, in the form of decision trees or sets of if-then rules, which can be used to classify new cases, with an emphasis on making the models understandable as well as accurate. In general, it is often possible to prune a decision tree to obtain a simpler and more accurate tree [6, 8].
The aims of this study are to explore biomarkers of greatest impact on distant metastasis (DM) development in BC patients and their combinatorial behavioural expression patterns, and to build a decision tree algorithm for predictive tissue markers of DM which could be used, following validation, in newly diagnosed BC cases.
Materials and methods
Patients and tumours
This study was based on a well-characterised cohort of early stage (I–III) primary operable invasive BC (n = 1902) from patients enrolled into the Nottingham Tenovus Primary Breast Carcinoma Series between 1987 and 1998, and managed in accordance to a uniform protocol and has been comprehensively studied with a broad range of markers [9, 10]. During the follow-up time within this series, distant recurrence had developed in 578/1902 cases (30 %). The median time to DM was 128 months (range 4–247 months).
This study included 31 biomarkers of clinical and biological relevance to BC tumourigenesis and progression [9, 11]. These were hormone receptors [oestrogen receptor (ER), progesterone receptor (PR)], epidermal growth factor receptor family members [HER1 (EGFR), HER2, HER3, HER4], cytokeratins [basal CKs; CK5/6 and CK14, and luminal CKs; CK7/8, 18 and 19], tumour suppressor and cell cycle regulator proteins [p53 and P27] anti-apoptotic BCL2, a proliferation marker (Ki-67/MIB1 clone), cadherin family [E-cadherin, N-cadherin and P-cadherin], markers of key molecular pathways [TGFβ1, PIK3CA, pAkt-S473], transcription factors [phospho-STAT3 and TWIST2], markers reported to be associated with invasiveness and tumour aggressiveness [CTEN, CD44 and CD24] [12, 13], in addition to five markers/proteins encoded by five transcripts/genes significantly expressed between metastatic and non-metastatic BC (TOMM34, ZFN22, KRT23, ST8SIA6, and chromogranin-A). These latter markers resulted from ANN of analysis of cDNA expression data of 128 primary invasive frozen BC samples from the Nottingham Tenovus Primary Breast Carcinoma series previously studied using gene expression profiling (GEP) [14]. This approach stratified the transcripts on their ability to classify samples based on the occurrence of DM (n = 35) compared with those without DM (n = 93), as previously described [15, 16]. These five proteins are encoded by genes/transcripts’ data analysed by artificial neuronal network. These genes were amongst the top 40 differentially expressed genes between metastatic and non-metastatic cases. This research was approved by Nottingham Research Ethics Committee 2 under the title of “Development of a molecular genetic classification of breast cancer”.
Immunohistochemistry (IHC)
Four-μm sections were cut from paraffin-processed block of previously prepared TMAs and mounted on Superfrost slides (Surgipath). MIB1 expression was determined using full face FFPE breast tissue sections as previously described [17]. Tissue sections were deparaffinised in xylene (Genta Medica, York, UK), rehydrated in descending series of ethanol, 10 s each. Heat-induced retrieval of antigen epitopes was carried out using microwave treatment of slides in 10 mM sodium citrate buffer (pH 6.0) for 20 min. Slides were then incubated primary antibody in optimal working dilution (Table 1). Secondary detection system was NovoLink™ Polymer Detection System (Leica, RE7150-K). Reaction was visualised using freshly prepared filtered solution of 3-3′ Diaminobenzidine tetrahydrochloride (DAB, Dako, K3468). Counterstaining was performed with Mayer’s haematoxylin (DAKO, AR106) for 6 min. Sections were dehydrated in alcohol, cleared in xylene and coverslipped using DPX mounting medium (BDH, Poole, UK).
Assessment of IHC staining
Slides were scanned as high resolution digital images (0.45 µm/pixel) using a NanoZoomer slide scanner (Hamamatsu Photonics, Welwyn Garden City, UK) and accessed using a web-based interface (Distiller, SlidePath Ltd, Dublin, Ireland). TMA cores were scored at ×20 magnification using a minimum of 24 high-resolution screen (1920 × 1080).
Scoring of IHC staining of markers was performed using the modified H-score method [18], except MIB1LI and BCL2 which were scored as the percentage of expression. All sections were scored without prior knowledge of the patients’ pathologic or outcome data.
Statistical analysis
To establish a set of rules to determine to which group; presence or absence of DM, a patient is more likely to be assigned using its variables’ values; WEKA software was used to compute the decision tree algorithm C4.5. Wilcoxon test, a non-parametric version of t test, was used to specify those markers appearing to behave differently in the two groups of patients. Results were validated using univariate Cox regression analysis. A p value <0.05 (two-tailed) was considered significant. Box plots were organised to visualise the differential distribution of each marker between those cases with metastatic disease from those without.
Results
Box plots for the distribution of all the 31 markers’ expression within the studied series with relevance to the presence or absence of distant metastases are summarised in Fig. 1a. According to these plots, the variable distribution of markers within both patients’ subsets can be inferred. For instance, the H-score of PR expression for 95 % of cases with DM ranges from “0–150”, median 5, compared with “0–200”, median 90 in cases with no DM.
Box plots for the studied markers in cases with no distant recurrence versus those with distant recurrence. The box for a certain marker represents 95 % of cases. The horizontal bold line inside the box is the median value of the marker. a Distribution of all markers (n = 31) within the studied series. b Box plots of biomarker panel showing significant expression between metastatic versus non-metastatic patients’ subsets
To test those markers which were the drivers of the two groups, Wilcoxon test and univariate Cox regression analysis were performed. This resulted in a panel consisting of 15 biomarkers being significantly associated with distant recurrence (p < 0.001). These markers were Ki67/MIB1LI, ER, PR, HER2, EGFR, p53, BCL2, N-cadherin, P-cadherin, PIK3CA, pSTAT3 nuclear expression, TOMM34, ZFN22, CD44 and Ck5/6. Table 2 displays these markers and the functional group under which they could be classified. Figure 1b displays box plots showing differential expression levels of this panel within metastatic versus non-metastatic groups of the studied series. As these plots display, variable distribution of different markers between the two groups could be appreciated. For instance, the H-score of BCL2 expression for 95 % of cases with distant recurrence lies between “0–75”, median 50, compared with “0–100”, median 70 in cases with no distant recurrence.
Biomarker expression pattern and time to distant metastasis
Based on time to development of DM in cases where distant recurrence occurred (n = 578 patients), six temporal subgroups were characterised, which ranged from less than 1 year of follow-up to more than 15 years. These were (1) <1 year (n = 50), (2) between 1 and 2 years (n = 102), (3) >2 up to 5 years (n = 201), (4) >5 up to 10 years (n = 155), (5) >10 up to 15 years (n = 56) and (6) >than 15 years (n = 14). Box blots were constructed to depict the expression pattern of the 15 biomarker panel significantly associated with DM within these temporal subgroups, Fig. 2. According to box plots of these metastatic subgroups, 10 markers had a significant expression pattern with respect to time of developing DM. However, the remaining five markers did not show this temporal relation with occurrence of DM.
Box plot showing expression pattern of metastatic biomarker panel within the six temporal groups within the metastatic group based on time to DM. a <1 year (n = 50), b from 1–2 years (n = 102), c >2 up to 5 years (n = 201), d >5 up to 10 years (n = 155), e >10 up to 15 years (n = 56) and F >15 years (n = 14). Y axis represents the H-score or percent expression of markers on X axis
Markers with significant expression trend within groups of time to distant recurrence
Within this group of markers, Ki67/MIB1LI, HER2, p53, N-cadherin, P-cadherin, PIK3CA and TOMM34 showed characteristic pattern of differential expression between the six subgroups, where higher expression values were associated with significantly earlier development of distant recurrence and vice versa. For PIK3CA, the expression values were very high for those cases that developed earlier DM (mean H-sore = 210 in those developed DM in less than 1 year compared with 130 in those developed DM >10 years up to 15 years). In contrast, higher expression values of ER, PR and BCL2, were observed to be associated with delayed occurrence of DM.
Markers with no observed expression trend within the time to distant recurrence period
The remaining markers, however, (i.e. EGFR, phospho-STAT3, ZFN22, CD44 and CK5/6), did not show an evident behavioural/differential trend in their expression with relevance to the time to development of DM in the six metastatic subgroups.
Decision tree-calculated metastasis prediction algorithm
To build a distant recurrence risk assessment algorithm for BC patients, decision trees were computed. For the purpose of robustness, only cases with complete values for all biomarkers were used to compute the decision tree. The number of informative cases available for the 15 significant markers was 176 cases, which constituted a test set for building a decision tree algorithm for metastatic recurrence prediction. DM had developed in 64 (36.3 %) cases, with the remaining 112 (63.7 %) cases remaining DM free throughout the period of follow-up. The input data for WEKA software were the expression data of these 15 markers, entered as continuous data “H-score or % expression”, and the metastatic status (Yes/No), with minimum number of cases in each branch to be equal to or more than 4. The resulting tree is as illustrated in Fig. 3.
Decision tree algorithm for predicting distant recurrence. Circles represent the markers in the algorithm (p53, HER2 and BCL2). Rectangles represent feature value tested (DM); Zero = No DM, One = Yes DM and numbers represent subsets of patients correctly classified and misclassified, respectively. Branches emerging from each marker are levels (H-score for p53 and HER2, and percentage of BCL2) of expression below or above which a specific case is to be classified into either zero or one
According to the tree, four groups of patients were characterised, based on the expression of three markers p53, HER2 and BCL2, which were able to predict the probability of DM in the test set. Cut-off points for these three markers were automatically set by the software. Table 3 displays these groups and the number of correctly classified and misclassified cases in each of group.
This ability of this algorithm to classify patients on the basis of probability of DM (i.e. DM Yes and DM No) was significantly associated with tumour size, tumour grade, number of positive axillary lymph nodes and BC molecular subtype as assessed by IHC [19]. Although associations with axillary nodal stage and lymphovascular invasion (LVI) did not reach statistical significance, more proportions of cases with ≥4 positive axillary nodes experienced DM more than those with node negative disease (30 % versus 13.5 %, respectively). Supplementary Table 1 displays the results of these associations. Multivariate Cox regression analysis showed that this algorithm is significantly associated with breast cancer-specific survival (BCSS) and metastasis-free interval [p = 0.001, Hazard ratio (HR) = 3.139, 95 % interval (CI) 1.640–6.011, and HR 2.856, 95 % CI 1.538–5.305, respectively] independently of grade, size, stage, molecular subtype and number of positive axillary lymph nodes.
The precision rate of this algorithm in predicting DM was evaluated using the positive predictive value (PPV) and the negative predictive value (NPV) [20]. For this algorithm, the PPV = 26/32 = 81.1 %, and the NPV = 150/194 = 77.3 %.
Discussion
Within the studied series, DM had developed in 30 % of cases during the period of follow-up and the outcome of this group was markedly reduced compared to patients who did not develop DM (5-year survival rate was 58 % compared to 98 %, respectively). This significant decline in patients’ survival is, to a large extent, attributable to the biological differences in tumours with metastatic potential from those without. Therefore, the molecular factors driving growth and differentiation pathways in tumours with metastases were scrutinised to explore their relative contribution in their non-metastatic tumour counterparts.
For this purpose, the expression pattern of 31 biomarkers with close relevance to BC biology and progression was studied with relevance to the occurrence of DM. Based on their distributions as continuous variables, many of the studied markers displayed variable expression within both patients’ subsets with and without DM, respectively. However, a panel formed of fifteen markers was significantly associated with the occurrence of DM.
Functional categories within this biomarker panel revealed that they belonged to molecular pathways responsible for carcinogenesis and cancer progression including hormonal receptors (ER, PR), epidermal growth factor receptor family members (EGFR, HER2), tumour growth fraction as assessed by MIBL1, tumour suppressor p53, anti-apoptotic BCL2, cell adhesion molecules (N-cadherin, P-cadherin), signalling pathways and transcription factors (PIK3CA, pSTAT3 and ZNF22), basal Ck14, TOMM34 and CD44.
In meta-analysis of publicly available BC GEP studies, Wirapati and colleagues showed that the key biological drivers in nine prognostic signatures were proliferation-related genes, in addition to ER signalling and HER2 amplification [21]. The current IHC marker panel shared ER, PR, HER2 and proliferative fraction as major drivers of progression with GEP studies [21]. Additionally, p53 and BCL2 were among the major contributors of DM in the IHC panel. Inactive TP53, as assessed by positive p53 protein expression, disturbs the functional braking and emergency cell cycle arrest in genetically damaged cells leading ultimately to cell cycle progression [22]. On the other hand, BCL2 is a cellular pro-survival molecule that protects transformed cells from apoptotic cell death. Therefore, from functional point of view, p53 and BCL2 which were more expressed in metastatic cases, lead to enhanced cell proliferation, through cell cycle progression and cell immortalisation [23].
The expression of basal/myoepithelial markers has been previously reported to contribute to the identification of a subset of BC characterised by poor outcome, the basal-like subtype [24]. In the current IHC panel, two markers could be assigned into this category of markers: EGFR and Ck14. These results support recommendations of using these two makers, beside ER, PR and HER2 negativity, as additional surrogates in characterising BCs with basal phenotype [25]. Moreover, the roles played by PIK3CA in BC progression through its downstream effectors, especially through driving an epithelial-to-mesenchymal transition program (EMT) with upregulation of N-cadherin and P-cadherin, have been reported in the same series [26, 27], as well as in others [28].
Metastatic recurrence developed in these patients with primary BC over a time schedule ranged from 4 months to more than 15 years. Ten markers of the IHC biomarker panel displayed differential expression patterns within the temporal groups of time to DM. These include ER, PR, HER2, MIB1LI, p53, BCL2, N-cadherin, P-cadherin, PIK3CA and TOMM34. However, the remaining five markers did not show this sort of trend. Markers of poor prognostic impact [MIB1LI, HER2, p53, N-cadherin, P-cadherin, PIK3CA and TOMM34] were more expressed in cases where DM had manifested earlier, while less expression was observed with prolonged metastasis-free intervals. The reverse was true for biomarkers of good prognostic impact [ER, PR and BCL2]. These findings could be interpreted in view of the concept of tumour dormancy, in which disseminated cancer cells leaving the primary tumour stay dormant for variable periods of times in another anatomical niche that could extend into many years. During this dormancy stage, cells may remain quiescent or form clinically undetectable micrometastases. Entering of these dormant cells or micrometastatic nodules into an overt progressive growth phase leads to the commencement of clinically detectable metastasis. The length of dormancy periods has been determined by balancing cell proliferation and apoptosis [29, 30]. More insight into the differentially expressed markers between early and late metastatic groups reveals, once again, that major drivers of BC progression, especially proliferation and apoptosis regulator, not only play major roles in emergence of recurrences but also in the time of their arousal.
Using the 15 IHC biomarker panel and decision tree, a probabilistic algorithm was computed to be applied for risk assessment of DM in BC patients. According to the resulting algorithm, four risk groups of patients were characterised. Expression levels of p53, HER2 and BCL2 at automatically-generated specific cut-off points were able to predict the probability of distant recurrence in the studied set with satisfactory precision rate 81.1 % PPV and 77.3 % NPV. Therefore, a tumour suppressor, an oncogene and an anti-apoptotic marker could be reliably used in DM risk prediction. In the root node of the decision tree, p53 expression was the first determinant, with distant recurrence expected in cases of high p53 (H-score >60) with HER2 or p53 high, low HER2 and low BCL2. However, low p53 alone or high p53, low HER2 and high BCL2 were associated with lower risk of DM. The ability of this probabilistic model in stratifying patients into DM risk groups was significantly associated with tumour size, grade, number of positive axillary lymph nodes and BC molecular subtype. Moreover, the model was significantly associated with both BCSS and metastasis-free interval, independent of other factors. This algorithm reiterates the documented prognostic values of these three markers and underscoring the central biological role played by each of these three markers in BC progression [31–33]. According to these results, this small or pruned panel of biomarkers could be used with acceptable success in distant recurrence prediction. However, its performance needs to be validated in an independent BC patient series especially on a prospective basis.
Conclusions
Metastatic recurrence in the studied series appears to result from contribution of a molecular biomarker panel controlling the major nodes in carcinogenic and progression pathways including hormonal receptors, growth factors, tumour suppressor, apoptotic regulator, cell adhesion apparatus and transcription factors. A predictive algorithm formed of p53, BCL2 and HER2 IHC expression was able to successfully predict the probability of distant recurrence, which requires independent validation. These findings affirm that metastasis is an inherent early cancer trait that could be predicted from the primary tumour biomarker expression profile.
References
Rabbani F, Koppie TM, Charytonowicz E, Drobnjak M, Bochner BH, Cordon-Cardo C (2007) Prognostic significance of p27(Kip1) expression in bladder cancer. BJU Int 100(2):259–263
Nguyen DX, Massague J (2007) Genetic determinants of cancer metastasis. Nat Rev Genet 8(5):341–352
Pantel K, Brakenhoff RH (2004) Dissecting the metastatic cascade. Nat Rev Cancer 4(6):448–456
McShane LM, Altman DG, Sauerbrei W, Taube SE, Gion M, Clark GM (2005) Reporting recommendations for tumor marker prognostic studies (REMARK). Nat Clin Pract Oncol 2(8):416–422
Cruz JA, Wishart DS (2007) Applications of machine learning in cancer prediction and prognosis. Cancer Inform 11(2):55–77
Kotsiantis SB (2007) Supervised machine learning: a review of classification techniques. Informatica 31:249–268
Salzberg S (1994) C4.5: programs for machine learning by J. Ross Quinlan. Morgan Kaufmann Publishers Inc, 1993. Mach Learn 16(3):235–240
Quinlan JR (1993) C4.5: programs for machine learning. Morgan Kaufmann Publishers, Inc., Los Altos
Abd El-Rehim DM, Ball G, Pinder SE, Rakha E, Paish C, Robertson JF, Macmillan D, Blamey RW, Ellis IO (2005) High-throughput protein expression analysis using tissue microarray technology of a large well-characterised series identifies biologically distinct classes of breast cancer confirming recent cDNA expression analyses. Int J Cancer (Journal International du Cancer) 116(3):340–350
Rakha EA, Elsheikh SE, Aleskandarany MA, Habashi HO, Green AR, Powe DG, El-Sayed ME, Benhasouna A, Brunet J-S, Akslen LA et al (2009) Triple-negative breast cancer: distinguishing between basal and nonbasal subtypes. Clin Cancer Res 15(7):2302–2310
Abd El-Rehim DM, Pinder SE, Paish CE, Bell JA, Rampaul RS, Blamey RW, Robertson JF, Nicholson RI, Ellis IO (2004) Expression and co-expression of the members of the epidermal growth factor receptor (EGFR) family in invasive breast carcinoma. Br J Cancer 91(8):1532–1542
Albasri A, Seth R, Jackson D, Benhasouna A, Crook S, Nateri AS, Chapman R, Ilyas M (2009) C-terminal Tensin-like (CTEN) is an oncogene which alters cell motility possibly through repression of E-cadherin in colorectal cancer. J Pathol 218(1):57–65
Ling LJ, Wang S, Liu XA, Shen EC, Ding Q, Lu C, Xu J, Cao QH, Zhu HQ, Wang F (2008) A novel mouse model of human breast cancer stem-like cells with high CD44 + CD24-/lower phenotype metastasis to human bone. Chin Med J (Engl) 121(20):1980–1986
Naderi A, Teschendorff AE, Barbosa-Morais NL, Pinder SE, Green AR, Powe DG, Robertson JFR, Aparicio S, Ellis IO, Brenton JD et al (2006) A gene-expression signature to predict survival in breast cancer across independent data sets. Oncogene 26(10):1507–1516
Zhang H, Rakha EA, Ball GR, Spiteri I, Aleskandarany M, Paish EC, Powe DG, Macmillan RD, Caldas C, Ellis IO et al (2010) The proteins FABP7 and OATP2 are associated with the basal phenotype and patient outcome in human breast cancer. Breast Cancer Res Treat 121(1):41–51
Naderi A, Teschendorff AE, Barbosa-Morais NL, Pinder SE, Green AR, Powe DG, Robertson JF, Aparicio S, Ellis IO, Brenton JD et al (2007) A gene-expression signature to predict survival in breast cancer across independent data sets. Oncogene 26(10):1507–1516
Aleskandarany MA, Rakha EA, Macmillan RD, Powe DG, Ellis IO, Green AR (2011) MIB1/Ki-67 labelling index can classify grade 2 breast cancer into two clinically distinct subgroups. Breast Cancer Res Treat 127(3):591–599
McCarty KS Jr, Miller LS, Cox EB, Konrath J, McCarty KS Sr (1985) Estrogen receptor analyses. Correlation of biochemical and immunohistochemical methods using monoclonal antireceptor antibodies. Arch Pathol Lab Med 109(8):716–721
Rakha EA, Aleskandarany M, El-Sayed ME, Blamey RW, Elston CW, Ellis IO, Lee AH (2009) The prognostic significance of inflammation and medullary histological type in invasive carcinoma of the breast. Eur J Cancer 45(10):1780–1787
Altman DG, Bland JM (1994) Statistics Notes: diagnostic tests 2: predictive values. BMJ 309(6947):102
Wirapati P, Sotiriou C, Kunkel S, Farmer P, Pradervand S, Haibe-Kains B, Desmedt C, Ignatiadis M, Sengstag T, Schutz F et al (2008) Meta-analysis of gene expression profiles in breast cancer: toward a unified understanding of breast cancer subtyping and prognosis signatures. Breast Cancer Res 10(4):R65
Liu Y, Kulesz-Martin MF (2006) Sliding into home: facilitated p53 search for targets by the basic DNA binding domain. Cell Death Differ 13(6):881–884
Goodsell DS (2002) The molecular perspective: Bcl-2 and apoptosis. Oncologist 7(3):259–260
Rakha EA, Putti TC, El-Rehim DMA, Paish C, Green AR, Powe DG, Lee AH, Robertson JF, Ellis IO (2006) Morphological and immunophenotypic analysis of breast carcinomas with basal and myoepithelial differentiation. J Pathol 208(4):495–506
Cheang MC, Voduc D, Bajdik C, Leung S, McKinney S, Chia SK, Perou CM, Nielsen TO (2008) Basal-like breast cancer defined by five biomarkers has superior prognostic value than triple-negative phenotype. Clin Cancer Res 14(5):1368–1376
Aleskandarany MA, Negm OH, Green AR, Ahmed MA, Nolan CC, Tighe PJ, Ellis IO, Rakha EA (2014) Epithelial mesenchymal transition in early invasive breast cancer: an immunohistochemical and reverse phase protein array study. Breast Cancer Res Treat 145(2):339–348
Aleskandarany MA, Rakha EA, Ahmed MA, Powe DG, Paish EC, Macmillan RD, Ellis IO, Green AR (2010) PIK3CA expression in invasive breast cancer: a biomarker of poor prognosis. Breast Cancer Res Treat 122(1):45–53
Zardavas D, Phillips WA, Loi S (2014) PIK3CA mutations in breast cancer: reconciling findings from preclinical and clinical data. Breast Cancer Res 16(1):201
Holmgren L, O’Reilly MS, Folkman J (1995) Dormancy of micrometastases: balanced proliferation and apoptosis in the presence of angiogenesis suppression. Nat Med 1(2):149–153
Townson JL, Chambers AF (2006) Dormancy of solitary metastatic cells. Cell Cycle 5(16):1744–1750
Abdel-Fatah TM, Powe DG, Agboola J, Adamowicz-Brice M, Blamey RW, Lopez-Garcia MA, Green AR, Reis-Filho JS, Ellis IO (2010) The biological, clinical and prognostic implications of p53 transcriptional pathways in breast cancers. J Pathol 220(4):419–434
Cianfrocca M, Goldstein LJ (2004) Prognostic and predictive factors in early-stage breast cancer. Oncologist 9(6):606–616
Callagy GM, Webber MJ, Pharoah PD, Caldas C (2008) Meta-analysis confirms BCL2 is an independent prognostic marker in breast cancer. BMC Cancer 8:153
Conflict of interest
The authors have no conflicts of interest to declare.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Aleskandarany, M.A., Soria, D., Green, A.R. et al. Markers of progression in early-stage invasive breast cancer: a predictive immunohistochemical panel algorithm for distant recurrence risk stratification. Breast Cancer Res Treat 151, 325–333 (2015). https://doi.org/10.1007/s10549-015-3406-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-015-3406-3