Abstract
The cloud forest of the Andean Region contains a high biodiversity. Unfortunately, human land use has caused most of the forest to become fragmented, negatively impacting many species due to the reduction of and constant change within the local habitat. In Colombia, these fragmentation triggers can include agriculture, livestock, and corridors for tourism. Conservation strategies focusing on keystone species could have more impact and better results to recover ecosystem dynamics. The wax palm Ceroxylon quindiuense (C. quindiuense) is an endemic and keystone species in cloud forests with a distribution across the three cordilleras of Colombia. Despite its ecological, economic and social importance, most forests of C. quindiuense are endangered; the most severely affected residing in small isolated populations in Central Cordillera. Nevertheless, these populations seem to retain a high genetic diversity. Because of this, the goal of conservation strategies should focus on retaining genetic diversity instead of increasing it. Because it can take as long as 80 years for C. quindiuense to reach maturity, our approach entails the introduction of juveniles (around 30 years) with genetic profiles similar to wild populations in order to augment population size, connect isolated populations, and avoid outbreeding. We evaluated the genetic makeup of three neighboring ex situ collections of living palms and compared them with the genetic profile of three wild populations of Central Cordillera. Multivariate analysis was used to assess patterns of genetic similarity and assign individuals to infer genetic clusters between collections and wild populations. Expected heterozygosity (He) of ex situ collections was lower (0.56) than wild populations (0.63), and the percentage of private alleles was higher in the wild populations (25%) than ex situ collections (10%). Collections Milan and Botanic Garden show genetic similarity with the Cocora and La Linea populations while the Toche and Roso collections were the most genetically distinct among the ones studied. Our results are that conservation programs should consider each population as a different evolutionary unit and protect them as such.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The damaging effects of human activities on wild populations such as reduction of habitat, poaching and deforestation are well-known and strategies to mitigate these effects are in progress for many of the species (Santos et al. 2016). One such strategy along with habitat restoration, is the recovery and maintenance of the genetic diversity of individual populations (Kashimshetty et al. 2015; Mijangos et al. 2015; Santos et al. 2016). In order to preserve the adaptive potential of species to changing environments such as in tropical regions a powerful conservation strategy is to focus on maintaining the genetic diversity of such species. (Ruber et al. 2001; Mijangos et al. 2015; Santos et al. 2016). However, there could be some pitfalls to this approach if implemented without knowledge of the ecological and biological characteristics of each individual species. These include (1) Introducing different genetic profiles that could lead to contamination with foreign alleles and (2) Introducing closely related individuals which could lead to increased endogamy (Kashimshetty et al. 2015). One of the reasons why most conservation strategies have failed is when implemented broadly without taking into account the underlying differences in life history traits (LHT) between distinct species. Therefore, programs which include species LHT, ecological status and preservation of genetic diversity are needed to improve conservation practices.
The tropical mountain cloud forest is the ecosystem with the highest biodiversity globally (Brummit and Lughada 2003). The cloud forest encompasses 11.7% of mountain tropical forests and represents 2.5% of the area of forests globally (Bubb et al. 2004). In the 1980s, the highly endangered central American and Caribbean cloud forests were disappearing more rapidly than any other type of forest (LaBastille and Pool 1978; Bruijnzeel et al. 2010; Armenteras et al. 2017). Colombia has been known as one of the most megadiverse countries in the world, and economic activities have centered in the Andean region where cloud forests subsist. Since pre-Colombian times, this area has undergone large transformations of the natural ecosystem primarily due to exponential human population growth and extensive land use occurring throughout the eighteenth and nineteenth centuries (Etter and Van Wyngaarden 2000). The demands of the growing economic market have transformed over 90% of the habitats into pastures and crops, and in later decades, this came along with lowered governability and the expansion of illicit crops (Etter and Van Wyngaarden 2000; Doumenge et al. 1995; Armenteras et al. 2007, 2017). Despite being poorly studied, the effects of deforestation in the tropical mountain cloud forests have been shown to be highly detrimental (Svenning 1998; Goerner et al. 2007; Svenning et al. 2009). Even though, the landscape of the Colombian Andes still contains small resilient fragments of forest that could hold the specific genetic pool from which modern repopulation strategies could derive.
Ceroxylon quindiuense (C. quindiuense), the Colombian national tree, is an endemic and umbrella species in cloud forests of the Tropical Andes of Colombia and Peru and grows in dense but isolated populations along the three cordilleras of Colombia (Sanín and Galeano 2011; Sanín et al. 2013). Unfortunately, threats to this species are numerous and the conservation status is endangered (EN) according to the IUCN criteria (IUCN 1987; Galeano and Bernal 2005).
The leaves are used for many religious celebrations, the stems for constructions and extracting wax and the juveniles are used as food for live-stock among many other uses. Pastures and crop plantation have taken over the landscape, replacing natural forest and leading to reduced wild populations and exposing juveniles to direct light which causes the leaves to burn. Additionally, a more recent threat to the successful growth of these habitats is the massive increase in local tourism (Bernal et al. 2015). The remaining individuals have challenging conditions for growth, including this species’ long-life cycle. First, it takes around 3 years for seedlings to emerge and produce the first leaves; this is a small plant with a height no larger than 20 cm which are easily confused with grass. This can mean a higher susceptibility to herbivory and damaging direct light exposure. Second, individuals expand around 30 years to become juveniles with a height between 12 and 19 m (Bernal and Sanín 2013). Although this is large enough for the plant to survive and grow, it takes another 50 years to achieve the reproductive adult stage (Bernal and Sanín 2013; Sanín et al. 2013). Third, although C. quindiuense still retains a high genetic diversity in many populations found within the three Cordilleras, these forests remain isolated and highly structured (i.e. discrete populations) which could represent a drastic loss of genetic diversity in the future (Sanín et al. 2013; Bernal et al. 2015).
Our working model includes the C. quindiuense populations in the Cocora-Quindio region of Colombia (Quindío wax palm). Cocora holds one of the most fragile populations of C. quindiuense. After colonization and up to the mid-twentieth century approximately, the distribution of the species became scattered throughout Central Cordillera. Nevertheless, the Cocora population today possesses a high genetic diversity despite the high structure, reproductive isolation, small remaining populations and reduction of gene flow among populations, but still the persistence of the species is threatened (Sanín 2013; Sanín et al. 2017). Despite the ecological and economic importance of C. quindiuense, only a few conservation studies have approach strategies that take into account the current ecological status of the species. Wild populations contain a high percentage of older adults and smaller populations of juveniles and seedlings that could continue with the generational replacement (Bernal and Sanín 2013).
Several populations of C. quindiuense found within natural forest regions, including Cocora, have been declared protected under government protection, preventing more habitat degradation. Despite this, the extremely small and highly isolated populations are still at high risk for becoming endangered or extinct. First, isolated wild populations do not have enough juveniles that could replace older plants, so small populations are already at risk of endogamy. Second, due to the species’ long-life cycle, connections between populations and recovery of habitat would be slow. Lastly, although the introduction of individuals could accelerate the recovery process, it may take a long time before any positive or negative results could even be evaluated.
An increasing concern remains for the fate of C. quindiuense, especially the Quindío wax palm, but several conservation strategies have been suggested, such as regeneration sites. Juveniles and adults are resilient to intermediate disturbance and can grow well under moderate shade, therefore a vast and intensive repopulation program could improve the current status of C. quindiuense (Anthelme et al. 2011; Bernal and Sanín 2013; Sanín et al. 2013; Bernal et al. 2015). Specifically, conservation strategies point to augmentation of the wild populations supplemented with juveniles from near nurseries (Bernal et al. 2015). These private ex situ collections are a source of seed and seedling, and most of them are formed by individuals from Cocora, Toche and Linea, wild populations in the Central Cordillera.
The genetic characterization of the populations and the ex situ juveniles to be introduced to these wild populations should be a priority before augmentation programs initiate (Hoban and Schlarbaum 2014; Mijangos et al. 2015). For this reason, the current study focuses on two main goals: (1) Restore the original genetic profile of Cocora by the introduction of individuals that resemble the original genetic diversity and genetic profile of the natural population. This would aim to accelerate the recovery process of wild populations by the introduction and establishment of juveniles closer to a reproductive state and reduce the chances of introducing genetic material that could lead to outbreeding depression. (2) Evaluate ex situ private collections in terms of allele capture and the effects of kinship in collection composition. Due to the vast distribution of C. quindiuense and the fact that collections do not have a conservation purpose, individuals from living ex situ collections are likely to come from the same parental line. Hence, results from this study could not only lead to better practices for establishment of ex situ collections (i.e. kinship reduction) and also act as a bridge between the community and environment by the use of local and private nurseries. Together, results of this analysis will be used to provide conservation management recommendations that both protect the local gene pools of wild populations and enhance diversity within ex situ collections.
Materials and methods
Sampling and PCR conditions
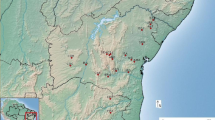
Sampling consisted of 142 individuals, divided into two main groups. The first group included three wild populations from Central Cordillera of Colombia: (1) Cocora Valley, Quindío at 2300 to 2900 masl, an area of 600 ha of which nearly 360 ha have been converted to pastures (N = 30); (2) Toche, Tolima at 2160 to 3000 masl, an area of 4500 ha consisting mostly of pastures (N = 26); (3) La Línea, Tolima at 2500 to 2700 masl, that underwent total deforestation (N = 29) (Fig. 1). The second group included three local ex situ plant collections: (1) Quindío Botanical Garden (N = 30) and (2) two commercial ex situ plant collections Milan (N = 11), and Roso (N = 16).
Map of the wild populations. Lower corner shows a map of Colombia and the three Cordilleras of the Andes. Dark circle shows the location on the central Cordillera of the three populations evaluated in this study. Red circles show the location of the wild population Cocora, Toche and La Linea in a bigger scale
We collected 10 g of young leaf tissue of juvenile individuals and the material was saved and dried in silica gel. The DNA was extracted by using Qiagen DNeasy Plant Mini Kit (Qiagen, Ltd.) following the manufacturer’s instructions. DNA quality was checked in 1.5% agarose gels and quantified by using a NanoDrop 1000 Spectrophotometer V3.
Ten microsatellite loci previously developed for C. alpinum and C. sasaimae (Gaitán 2003) were amplified in 142 individuals, which were CA7, CA16, CA17, CS24, CA25, CA20, CA30, CS30, CS5, and CS3. Polymerase Chain reaction (PCR) were 25 µl in volume, consisting of 5 µl of DNA (5 ng/µl), 12.5 µl of GoTaq® Green Master Mix (Promega, Ltd.), 1 µl of each primer and 5.5 µl of free nuclease water (Promega, Ltd.) The PCR program was 94 °C (4 min), [94 °C (15 s), AT °C (15 s), 72 °C (5 min)] for 35 cycles, 72 °C (5 min), 10 °C (∞), as described in Sanín (2013). The PCR products were visualized in polyacrylamide gels (4%) with positive and negative controls. To identify the size of the fragments, we used a Thermo Fisher 10-bp DNA ladder, and score reads were carried out by two independent operators to generate the allele profile per sample.
Analysis for genetic diversity for wild and ex situ populations
The restoration strategy through augmentation of the wild populations focuses primary in the Cocora region. Nevertheless, individuals from the collections expected to be introduced could come from any population across the Central Cordillera in Colombia. We evaluated the genetic profile of three wild populations of the Central Cordillera, including Cocora, and three ex situ collections from the area, two of them belonging to a private party (Milan and Roso) and one to a public botanical garden. Genetic diversity statistics were compared for every population. We calculated deviations from Hardy–Weinberg equilibrium (HWE), heterozygote deficiency, and linkage disequilibrium were computed in GENEPOP on the Web (Rousset 1995, 2008). Markov chain parameters were set to 5000 iterations, according to Gou and Thompson (1992). Standard molecular diversity indices were estimated in Arlequin v.3.5 (Excoffier and Lischer 2010) including number of alleles per population and per locus (Ap, Al respectively), effective number of alleles (Ae), Allelic richness (Ar), number of rare alleles (alleles with frequency < 0.01), private alleles (alleles found in only in one population), observed heterozygosity (Ho) and expected heterozygosity (He), genetic diversity (H). Allele frequencies were calculated in GenAlEx v.6.4.1 (Peakall and Smouse 2006) and diveRsity package (Keenan et al. 2013).
Genetic structure analysis of wild populations and ex situ plant collections
The Discriminant Analysis of Principal Components (DAPC) was used to identify the number of clusters or gene pools and identify the genetic profile of the ex situ collections that could resemble the genetic profile of wild populations, focusing primarily on Cocora (Pritchard et al. 2000; Jombart et al. 2010). DAPC analysis was performed as an individual assignment and infer implied genetic clusters based on k-means clustering. DAPC analysis was done in R (R development Core Team 2013), with the Adegenet package (Jombart et al. 2010). DAPC analysis does not rely on a particular population genetics model and is tolerant of deviations from HWE, linkage equilibrium, and null alleles (Jombart et al. 2010). The find.clusters function was used to run k-means clustering to select the best-supported number of clusters using the Bayesian Information Criterion (BIC) for values of K. Additionally, automated clustering was performed using a fix cluster number of 5 based on BIC and a cross-validation to select the correct number of PCs to retain. Assignment probabilities were based on the retained discriminant functions as indicators of how clear-cut genetic clusters were.
We separated the data into two groups: wild populations and ex situ collections. To validate the genetic variation components among, within, and between groups, an Analysis of Molecular Variance (AMOVA) was performed using FST-like statistics in Arlequin program (Excoffier and Heckel 2006; Excoffier and Lischer 2010). To elucidate relationships between groups, two different approaches were used to estimate genetic differentiation, FST (Wright 1969; Weir and Cockerham 1984) and DST (Jost 2008, 2009). Pairwise FST was tested with log likelihood ratio (test G) to assess population differentiation by Markov chains algorithms in the GENEPOP software and pairwise DST was performed in DEMEtics package in R (Gerlach et al. 2010). Both statistics were assumed under a significance of 95% bootstrap confidence intervals (1000 iterations).
Assignment test based on ten microsatellite loci
To infer the origin of the ex situ plant collections, we performed an assignment test using Bayesian method Rannala and Mountain (1997) criterion with the ‘leave one out’ procedure on the basis of microsatellite variation with the program GeneClass2 (Piry et al. 2004). This test calculates the probability that an individual’s multilocus genotype occurs on each reference population. Here we took each wild population as a reference and compared them with each ex situ collection. After we found the population with highest probability, we evaluated rejection and acceptance of membership of the genotype to the population by the Monte Carlo resampling Frequency method of Paetkau et al. (2004). The probability of given individual multilocus genotype was compared to the distribution of probabilities of multilocus genotypes (10,000 replicates) generated based on the allele frequencies of the population, and if the value was below α < 0.01, the individual was rejected from the population. Such a combination of an assignment test and exclusion-method is used for other studies (Manel et al. 2005; Frantz et al. 2006; Hare et al. 2006). Additionally, we used likelihood values to test for presence of first-generation migrants on each sample. We used L_home criteria which is the likelihood of individual genotype from the population where it was sample (Paetkau et al. 2004).
Evaluation of relatedness analysis and eligible material for augmentation programs
Kinship analysis was performed using Nei genetic distances (Nei and Feldman 1972). Values were obtained with the dist.genpop function in the Adegenet package. Analysis of hierarchical clustering was performed with bootstrap confidence intervals of 1000 interactions and plotted in R with the Dendextend package (Galili 2015). To assess relatedness among individuals as a proxy of intrapopulation genetic diversity, we used the Demerelate package (Kraemer and Gerlach 2013) in R to evaluate the levels of half and full siblings based on the model of shared genotypes (Mxy) (Blouin et al. 1996), and 1000 iterations. Subsequently, individuals with a genetic profile similar to wild populations were taken as eligible material for a restoration program.
Results
All ten loci were amplified in all populations, and all sampled loci were polymorphic. 136 alleles were found in the ten loci, ranging from 2 to 21 alleles per locus. Significant deviations from the Hardy–Weinberg equilibrium were found in all populations, but no evidence of linkage disequilibrium, p < 0.001. Cocora had the highest values for expected heterozygosity and allelic richness among all population, and levels of expected heterozygosity among collections were similar to each other (Table 1). Botanical Garden and Milan were very similar in their diversity measures. Roso, La Linea, Toche and Cocora harbored larger and similar amounts of expected heterozygosity. However, both groups differ in their observed heterozygosity levels. Total number of alleles, rare alleles and private alleles were measured. Between wild populations, Cocora had the highest number of total, private, and rare alleles, while among ex situ collections, Roso had the highest number in all previous categories (Fig. 2). In general, wild populations showed a higher number of alleles (total, rare and private) compared to collections. Although a Chi square test suggests that all categories are homogeneous across populations, it does not imply a similar genetic makeup between wild and ex situ collections.
Pairwise comparisons with both FST and DST among wild populations and ex situ collections showed significant levels of differentiation. FST was 0.30 and DST was 0.76, both with a p < 0.001 (Table 2). Genetic differentiation between the two groups (wild populations and ex situ collections) was also reflected in the AMOVA results, indicating that 34.85% of the variation was among populations within groups (FIT = 0.35, p < 0.001), 3.07% among individuals within populations (FIS = 0.04, p < 0.001) and 64.56% within individuals (FSC = 0.34, p < 0.001). The variation among groups was not significant (− 2.49% p > 0.05). Multivariate DAPC analysis indicated that populations of C. quindiuense from wild and ex situ collections were divided into three distinct clusters identified by k-means clustering (Fig. 3). Botanical Garden, Milan, La Linea and Cocora are very close, suggesting potential admixture. Conversely, Toche and Roso are each genetically distinct, and none of the ex situ collections appears to capture the diversity from Toche with limited evidence of admixtures (Fig. 3). A second DAPC encompassing just wild populations showed a high genetic structure between populations (Supplementary Fig. 1). Additionally, GeneClass2 results based on Bayesian methods assigned each individual to the population from which they were sampled with a 100% score (Supplementary Table 1). Frequency-based method (Paetkau et al. 2004) using L_home provide no evidence of first-generation migrants among any of the groups we sampled (p value > 0.01). Only three individuals from Cocora, one from Toche, two from Roso, and three from Botanic Garden were below threshold of 0.01 (Supplementary Table 2).
Assignment analysis of C. quindiuense individuals. a Discriminant analysis of principal components (DAPC) with 4 K-mean clusters. Confidence interval (95%) represented by the ellipses. Each line represents and individual and the shapes represent its assigned genetic cluster. Eigenvalues show the genetic information of each component. Axis X and Y correspond to the first and second principal component respectively. b Membership probabilities of individual wax palm for all individuals. Each line represents and individual and the colors represent genetic clusters. Color segments for each individual are proportional to the probability of assignment to a genetically unique cluster
The Nei genetic distance showed a close relation among the Botanical Garden, Milan, and the group from Cocora and Toche also with low genetic distance. On the other hand, Roso was the most distant from the rest of the wild population and the other two collections (Fig. 4). Results of kinship analysis indicated that collections had significant levels of full-siblings (p < 0.001); Milan and Botanical Garden levels are over 50%, different from Roso with equal percentages of full and half-siblings. Wild populations had lower levels of full-siblings than collections, and Cocora had the lowest percentage of full siblings (5%, p < 0.001) (Fig. 4). A negative correlation (R2 = − 0.82) was obtained between genetic diversity and percentage of full-siblings in each population and collection evaluated (p < 0.005).
a Hierarchical clustering between wild populations and ex situ collections of C. quindiuense in the Central Cordillera of Colombia evaluated by Nei distances. All nodes are supported by bootstrap > 75. Kinship analysis. b Bars represent the percentage of full (blue) and half siblings (red) for each population. Kinship values were calculated using 1000 randomizations by population (p < 0.001). Wild populations present a lower percentage of full siblings when compared to collections
Discussion
The microsatellites used in this analysis were polymorphic and informative and allowed us to characterize both ex situ and wild populations. As expected, these suggest significant genetic structure between the three wild populations (Cocora, La Línea, Toche) in the Central Cordillera of Colombia. Regarding genetic provenance and representativeness, the ex situ plant collections Botanical Garden and Milan showed genetic similarity only with Cocora, and the collection Roso did not show genetic concordance with any of the three wild populations. The topics presented in this study will focus on (1) conservation state of the wild populations Cocora, La Linea, Toche and (2) the use of the evaluated ex situ plant collections for augmentation of these wild populations and (3) the effectiveness of ex situ plant collections for conservation purposes.
Conservation status of wax palm (C. quindiuense) in the Andean region
Wax palms in general, and C. quindiuense in particular, face several conservation problems. Total population extension and connectivity is lost to deforestation by cattle raising and crops, areas in which ecological population structure is lost (Sanín et al. 2013). Secondly, even when deforestation took place decades before, the low tolerance of seedlings to radiation in open areas, limited seed dispersal, and the destruction of seeds and seedlings by trampling and livestock grazing prevent generation replacement (Anthelme et al. 2011). The establishment of these forest systems imposes long time lapses with no seed recruitment and leads to an inevitable population scenario wherein all large wax palm forests are solely composed of very old adults (Bernal and Sanín 2013). The resilience of these old adults and the longevity of the species provide a conservation opportunity. As a standing generation derived from better forests, these old adults hold high genetic diversity and continue to reproduce massively. However, their seeds and seedlings have to be protected within areas where human activities prevail or should be taken to protected areas where seeds may develop into the more resilient juveniles (rosette fase, see Sanín et al. 2013).
Evidence of a high level of genetic diversity was found across the three populations and agrees with previous findings showing that populations in this region have the highest level of diversity across the entire distribution (Sanín et al. 2017). Evidence of genetic structuring and inbreeding was found within wild populations, although, few studies have evaluated the genetic diversity and population structure of C. quindiuense, it has been shown that Cocora and Toche have some level of structure compare to other wild populations (Sanín et al. 2017; González-Rivillas et al. 2018). This structure pattern could potentially reduce genetic diversity of the already isolate populations creating a worse scenario for C. quindiuense persistence.
Small, close and isolate populations are more susceptible to inbreeding and loss of genetic diversity (Wright 1969; Frankham et al. 2014). Anthropogenic impacts have reduced the connectivity between populations, thus, genetic composition is based on chance and forged by inbreeding and drift. There is generally a tradeoff between genetic diversity and local population fitness, and both are correlated with population size (Leimu et al. 2006; Reed and Frankham 2003; Szczecińska et al. 2016). Additionally, for this species, dispersion is carry on by small/medium mammals (i.e. bats) or by air (Bernal et al. 2015), nevertheless, in disturbed habitats the dispersal ability is reduced, and this correlates negatively with FST values (Leimu et al. 2006; Frankham et al. 2014).
In the Central Cordillera, C. quindiuense populations are small and isolated which means they may become more susceptible to inbreeding (Fis values Table 1). Although inbreeding is not very strong, and the genetic diversity still high wild population remain at risk. These results are screenshots of old generations of palms which could differ from new generations (juveniles and seedlings) which are more affected by habitat reduction and isolation. Future studies that explore genetic diversity differences between the three palm generations (adults, juveniles, seedings) are needed. Genetic rescue could be an optimal management strategy for C. quindiuense conservation. Augmenting population size with material that resembles the original genetic makeup of the populations and connecting isolated populations would provide a low risk for outbreeding depression within the crosses and an increase of gene flow, respectively (Frankham et al. 2011; Frankham 2015; Ralls et al. 2017). Although genetic rescue might be a sound strategy for species with a marked reduction in local genetic diversity, all populations that were investigated here and previously studied by Sanín et al. (2017) show high levels of genetic diversity despite the apparent isolation. Also, fruit production and germination remain high, even in extremely small populations suggesting sexual reproduction from local material is still an option. Moreover, historical population divergence was found between wild populations (Sanín et al. 2017) which could entail local adaptation, but this has not yet been studied. Therefore, we believe the key issue is population size as a consequence of habitat availability and that genetic diversity is still there to support population growth and permanence.
Use of ex situ plant collections for augmentation of the studied wild populations
Aiming at the conservation of natural habitats and of natural biological interactions, (re)introduction has become important for the restoration of endangered plant species that require augmentation of existing populations (Robert et al. 2007; Godefroid et al. 2011; Maschinski et al. 2012; Menges et al. 2016). In an ideal scenario, individuals for reintroduction are available in ex situ plant collections that are enrolled in conservation projects. These are especially important in cases where it is difficult or impractical to employ ex situ strategies through standard methods (i.e. seed or tissue banks; Namoff et al. 2010; Pence 2011; Griffith et al. 2015). Introducing juveniles from these collections (Fig. 5) would save a quarter to half a decade for conservation efforts which takes into account that wax palms are long-lived and only reach maturity at about 80-year-old according to developmental age estimates from Sanín et al. (2013).
Life stages of C. quindiuense individuals taking from Sanín et al. (2013): Seedling, Juvenile 1, Juvenile 2, Juvenile 3 and Adult. The bottom left number point the estimated age by the end of that developmental phase. Photograph on the right shows a juvenile from Botanical Garden collection
Given the marked genetic structuring between wild populations and ex situ collections, material for reintroduction programs should be locally sourced. A high level of genetic diversity is advantageous for conservation, but the introduction of non-local genotypes to enrich genetic diversity could lead to increased non-adapted characteristics as a result of exogamic depression or expression of intermediate genotypes not favored locally (Mckay et al. 2005; Menges 2008). Even if two populations are similar phenotypically and experience similar selection pressures, crosses between these populations can produce unfit hybrids from the breakup of the complex of genes (Fenster and Galloway 2000; Mckay et al. 2005), which is a disadvantage especially for species with long generation times, represented by small and isolated populations (Hufford and Mazer 2003; Rice and Emery 2003).
Both the genetic distance and multivariate analyses show that Botanical Garden and Milan collections can be unambiguously traced to Cocora rather than Toche or La Linea wild populations. However, collections are established from only certain genotypes, and genetic relatedness can be even closer than observed (Etisham-Ul-Haq et al. 2001; Li YY et al. 2005). Furthermore, the provenance of Roso collection could not be traced to any of the studied populations. Using genetic assignment methods based in Bayesian, Frequency and multivariate analysis, we have demonstrated clear discrimination between ex situ plant collections and wild populations despite relatively small sample sizes. GeneClass2 using Bayesian approach (Rannala and Mountain 1997) and a Frequency approach (Paetkau et al. 2004) suggested that wild populations were highly unlikely to be source population of any of the collections. Although, no evidence of first-generation migrants on the collections was found, assignation of individuals to the different groups by DAPC showed similar genetic profiles between Cocora Milan and Botanical Garden. Based on these results we suggest that individuals from the collections could be introduce in the wild population Cocora but not on any of the other two wild populations Toche or La Linea. Finally, Roso genetic profile does not match any of the wild populations which suggest unknown wild population source from any of the Cordilleras. Interestingly, this match with few exceptions found by GeneClass2 on Roso, Cocora, Toche and La Linea where few individuals appear to be first generation migrants from unknown sources.
Effectiveness of collections for conservation purposes
Taking into account that the materials from Botanical Garden and Milan collections were unambiguously traced to Cocora wild population, the comparison between ex situ collections and their wild source could also be used to assess the effectiveness of collections in capturing and representing wild genetic diversity (Griffith et al. 2011). Changes in wild populations are also important for the future direction of collections, as the effect of inbreeding on and the reduction of genetic diversity in wild populations would indirectly affect collections. This would mean that genetic information saved in living collections now may not reflect the genetic composition of future wax palm generations (Li-Q et al. 2005; Liu et al. 2008; Yokogawa et al. 2013).
Ex situ plant collections are normally established in the studied area with commercial purposes where a sampling plan is not considered. Some collections, as is the case for Botanical Garden, are not established for business purposes, but their relevance for conservation research is not an issue when the material is retrieved from the source population. Values of genetic diversity were significantly different (p < 0.05). Roso with a small collection size showed high diversity (potentially representative of a wild population), while Milan and Botanical Garden, with a small and big collection size respectively, showed lower genetic diversity and redundant alleles. These results suggest that a high number of full siblings may be an indication of poor management strategies (Li et al. 2005; Yokogawa et al. 2013), whereas a large size or genetically diverse source do not translate into high captured diversity. In all cases collections were developed without a sampling scheme focusing on allelic capture of the wild genetic pool, nonetheless, Roso was the only collection where the material came from distant geographic parents whereas Milan and Botanic Garden material were from the same individual in many cases.
Nevertheless, in general, allelic diversity and heterozygosity are positively correlated with population size, (e.g., Leimu et al. 2006) and one valuable estimate approximation might be to increase the size of collections in terms of half-siblings to enhance allelic capture. Collectors should not exclusively rely on the quantity of individuals but on the sample, these represent (Walters 2004; Namoff et al. 2010). Strategies that intend to improve allelic capture in ex situ plant collections can change depending on the species life history traits (i.e. pollination and dispersal of seeds), demographic history, relevant threats or on their distribution (Namoff et al. 2010; Godefroid et al. 2011; Griffith et al. 2011; Menges et al. 2016; Griffith et al. 2015).
Based on the previous analysis, we could assume that the material from ex situ collections, specifically Botanical Garden and Milan, came mostly from Cocora. Nevertheless, these collections did not recover a high genetic diversity from the wild population. The high percentage of private alleles in the wild population indicates that several alleles were missing from ex situ collections. This is explained by a common practice whereby material for collections (seedlings) is retrieved from a few parentals (i.e. many seeds or seedlings collect from a single female palm). Seed and seedling selection should be scattered to cover the vicinities of many female palms in order to increase the number of half siblings. Otherwise, artificial selection can affect and reduce genetic diversity in collections which would generate genetic differences between them and wild populations (Abbo et al. 2003; Yokogawa et al. 2013; Miao et al. 2015).
In general, Botanic Gardens have progressively focused on ex situ conservation (Raven and Havens 2014), and several works have established guidelines for creating effective and genetically diverse ex situ collections (Namoff et al. 2010; Godefroid et al. 2011; Griffith et al. 2011, 2015; Guerrant et al. 2014). This would help to implement a protocol for high allelic capture in Botanical Garden collections based on the information that we discuss before. Even though sampling protocols may differ from one species to another, we can build from studies in other palms. In studies focusing on Leucothrinax morrisii (Namoff et al. 2010; Griffith et al. 2011), results suggested that collections with 59 plants were able to capture more than 90% of the genetic diversity of a single population and that a collection of 15 plants from three accessions was sufficient to recover a mean of 83% of alleles in the population. Here we found a good example where collections built with different material sources are more efficient to capture wild diversity than collection built with few sources and a higher number of individuals. For our case, 16 individuals in the collection were sufficient to capture a high genetic diversity as long as the parental sources were different.
More important than collection number is a collection procedure that follows general recommendations established by and cited textually from Griffith et al. (2015): (1) use the species biology to inform the collecting strategy; (2) manage each population separately; (3) collect and maintain multiple accessions; and (4) collect over multiple years. Also, guidelines for collection management in Krishnan et al. (2013) for coffee and Cibrian-Jaramillo et al. (2010, 2013) for cycads highlight the way to represent the full range of wild genetic diversity. Wax palm populations from the Central Cordillera are markedly structured and behave as different evolutionary units and should therefore be represented in independent ex situ collections.
Conclusions
Similar to previous studies, we have found that populations of C. quindiuense from the Central Cordillera cover restricted areas but host high genetic diversity (see Sanín et al. 2013; Bernal et al. 2015). This diversity is highly structured, and conservation programs should consider each population as a different evolutionary unit (Sanín et al. 2017) to be genetically represented and protected as such. Genetic rescue could be an ideal plan for small and isolated populations since there is limited evidence of inbreeding, however C. quindiuense still remains at risk if it continues solely within small and isolated populations. Consequently, we propose ecological rescue, focusing on the augmentation of current populations using ex situ living collections ensuring genetic fidelity to source populations. From the three studied ex situ plant collections, only Botanical Garden and Milan represent part of the wild gene pool of Cocora and can be potentially used for restorations programs. Nevertheless, neither of the collections cover the total diversity of the wild population especially concerning private alleles. Furthermore, collection strategies that specify better practices regarding seed and seedling retrieval are needed. Juveniles should be retrieved from distant parental to minimize the probability of obtaining full-siblings. At the time of introduction to the wild, genetic mixture between populations should be avoided, and juveniles should be shuffled within the palm stand to avoid proximity between full siblings.
Although these strategies focus within a local context, we believe the development of new strategies that link conservation and genetic and population analysis are required to optimize conservation strategies. Especially if the strategies have a social impact that could involve local communities that already hold commercial collections by including them in restoration programs where they can contribute with seed, seedlings or juveniles. This would also promote awareness in which people are economically favored and contribute to the conservation of the entire cloud forest ecosystem. As a final and more regional perspective, these practices are feasible within the current socioeconomic context of the region. Colombia has entered a post-conflict period after 76 years of armed conflict, and many possibilities open up for conservation actions in previously inaccessible areas (Castro-Nuñez et al. 2017). Under this new context, the Quindio wax palm becomes a flagship species (Bernal and Sanín 2013) to address new conservation challenges under scientific criteria.
References
Abbo S, Berger J, Turner NC (2003) Viewpoint: evolution of cultivated chickpea: four bottlenecks limit diversity and constrain adaptation. Funct Plant Biol 30:1081–1087
Anthelme F, Lincango J, Gully C, Duarte N, Montúfar R (2011) How anthropogenic disturbances affect the resilience of a keystone palm tree in the threatened Andean cloud forest? Biol Conserv 144:1059–1067
Armenteras D, Cadena VC, Moreno RP (2007) Evaluación del estado de los bosques de niebla y de la meta 2010 en Colombia. Instituto de Investigación de Recursos Biológicos Alexander von Humboldt, Bogotá
Armenteras D, Espelta J, Rodríguez N, Retana J (2017) Deforestation dynamics and drivers in different forest types in Latin America: Three decades of studies (1980–2010). Global Environ Change 46:139–147
Bernal R, Sanín MJ (2013) Los palmares de Ceroxylon quindiuense (Arecaceae) en el Valle de Cocora, Quindío: perspectivas de un ícono escénico de Colombia. Colomb For 16:67–79
Bernal R, Galeano G, Sanín MJ (2015) Plan de conservación, manejo y uso sostenible de la palma de cera del Quindío (Ceroxylon quindiuense), Árbol Nacional de Colombia. Ministerio de Ambiente y Desarrollo Sostenible; Universidad Nacional de Colombia, Bogotá, D.C., Colombia
Blouin MS, Parsons M, Lacaille V, Lotz S (1996) Use of microsatellite loci to classify individuals by relatedness. Mol Ecol 5:393–401
Bruijnzeel L, Kappelle M, Mulligan M, Scatena F (2010) Tropical mountain cloud forest: state of knowledge and sustainability perspectives in a changing world. In: Bruijnzeel L, Kappelle M, Mulligan M, Scatena F (eds) Tropical mountain cloud forests: science for Conservation and Management. Cambridge University Press, Cambridge, p 768
Brummit N, Lughada EI (2003) Biodiversity: where’s hot and where’s not. Conserv Biol 17:1442–1444
Bubb P, May I, Miles L, Sayer J (2004) Cloud forest agenda. UNEP-WCMC, Cambridge
Castro-Nuñez A, Mertz O, Casanova AB, Sosa CC, Lee S (2017) Land related grievances shape tropical forest-cover in areas affected by armed-conflict. Appl Geogr 85:39–50
Cibrian-Jaramillo A, Daly AC, Brenner E, Desalle R, Marler TE (2010) When north and south don’t mix: genetic connectivity of a recently endangered oceanic cycad, Cycas micronesica in Guam using EST-microsatellites. Mol Ecol 19(12):2364–2379
Cibrian-Jaramillo A, Hird A, Oleas N, Ma H, Meerow AW, Francisco-Ortega J, Griffith MP (2013) What is the conservation value of a plant in a botanic garden? Using indicators to improve management of ex situ collections. Bot Rev 79(4):559–577
Doumenge C, Gilmour D, Pérez MR, Blockhus J (1995) Tropical montane cloud forests: conservation status and management issues. In: Tropical montane cloud forests. Springer, New York, NY.
Etisham-Ul-Haq M, Allnutt TR, Smith-Ramirez C, Gardner MF, Armesto JJ, Newton AC (2001) Patterns of genetic variation in and ex situ populations of the threatened Chilean Vine Berberidopsis corallina, detected using RAPD markers. Ann Bot 87:813–821
Etter A, Van Wyngaarden W (2000) Patterns of landscape transformation in Colombia, with emphasis in the Andean region. Ambio 29(7):432–439
Excoffier L, Heckel G (2006) Computer programs for population genetics data analysis: a survival guide. Nat Rev Genet 7:745
Excoffier L, Lischer HE (2010) Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Mol Ecol Resour 10:564–567
Fenster CB, Galloway LF (2000) Population differentiation in an annual legume: genetic architecture. Evolution 54:1157–1172
Frankham R (2015) Genetic rescue of small inbred populations: meta-analysis reveals large and consistent benefits of gene flow. Mol Ecol 24:2610–2618
Frankham R, Ballou JD, Eldridge MD, Lacy RC, Ralls K, Dudash MR, Fenster ChB (2011) Predicting the probability of outbreeding depression. Conserv Biol 25:465–475
Frankham R, Bradshaw CJ, Brook BW (2014) Genetics in conservation management: revised recommendations for the 50/500 rules, red list criteria and population viability analysis. Biol Cons 170:56–63
Frantz AC, Pourtois T, Heuertz M, Schley L, Flamand MC, Krier A, Bertouille S, Chaumont F, Burke T (2006) Genetic structure and assignment tests demonstrate illegal translocation of red deer (Cervus elaphus) into a continuous population. Mol Ecol 15:3191–3203
Gaitán E (2003) Obtención y uso de secuencias microsatélites GA/CA en estudios de diversidad genética en las especies de palmas colombianas Ceroxylon sasaimae, Ceroxylon alpinum y Attalea amygdalina. Dissertation, Universidad Nacional de Colombia, Palmira, Colombia
Galeano G, Bernal R (2005) Palmas. In: Calderón E, Galeano G, García N (eds) Libro Rojo de las Plantas de Colombia, 2nd edn. Palmas, Frailejones y Zamias. Serie de Libros Rojos de Especies Amenazadas de Colombia, Bogotá, Colombia, pp 59–224
Galili T (2015) dendextend: an R package for visualizing, adjusting and comparing trees of hierarchical clustering. Bioinformatics 31:3718–3720
Gerlach G, Jueterbock A, Kraemer P, Deppermann J, Harmand P (2010) Calculations of population differentiation based on GST and D: forget GST but not all of statistics! Mol Ecol 19:3845–3852
Godefroid S, Piazza C, Rossi G, Buord S, Stevens AD, Aguraiuja R, Cowell C, Weekley C, Vogg G, Iriondo J, Jhonson I, Dixon B, Gordon D, Magnannon S, Valentin B, Bjureke K, Kopman R, Vicens M, Virevaire M, Vanderborght T (2011) How successful are plant species reintroductions? Biol Conserv 144:672–682
Goerner A, Gloaguen R, Makeschin F (2007) Monitoring of the Ecuadorian mountain rainforest with remote sensing. J Appl Remote Sens 1:3172–3178. https://doi.org/10.1117/1.2784111
González-Rivillas N, Bohorquez A, Gutierrez JP, Merchan VHG (2018) Diversity and population genetic structure of the wax palm Ceroxylon quindiuense in the Colombian Coffee Region. BioRxiv, 443960
Griffith MP, Lewis C, Francisco-Ortega J (2011) Palm conservation at a botanic garden: a case study of the keys thatch palm. Palms 55(2):93–101
Griffith MP, Calonje M, Meerow AW, Tut F, Kramer AT, Hird A, Magellan TM, Husby ChE (2015) Can a botanic garden cycad collection capture the genetic diversity in a wild population? Int J Plant Sci 176(1):1–10
Guerrant E, Havens K, Vitt P (2014) Sampling for effective ex situ plant conservation. Int J Plant Sci 175(1):11–20
Hare MP, Allen SK Jr, Bloomer P, Camara MD, Carnegie RB, Murfree J, Luckenbach M, Meritt D, Morrison C, Paynter K, Reece KS, Rose CG (2006) A genetic test for recruitment enhancement in Chesapeake Bay oysters, Crassostrea virginica, after population supplementation with a disease tolerant strain. Conserv Genet 7:717–734
Hoban S, Schlarbaum S (2014) Optimal sampling of seeds from plant populations for ex situ conservation of genetic biodiversity, considering realistic population structure. Biol Conserv 177:90–99
Hufford KM, Mazer SJ (2003) Plant ecotypes: genetic differentiation in the age of ecological restoration. Trends Ecol Evol 18(3):147–155
IUCN (1987). living organisms. IUCN Position Statement. The IUCN Red List of Threatened Species. Version 2015-4. <www.iucnredlist.org>. Accessed 27 Feb 2016
Jombart T, Devillard S, Balloux F (2010) Discriminant analysis of principal components: a new method for the analysis of genetically structured populations. BMC Genet 11:94
Jost L (2008) GST and its relatives do not measure differentiation. Mol Ecol 17:4015–4026
Jost L (2009) D vs. GST: response to Heller and Siegismund (2009) and Ryman and Leimar (2009). Mol Ecol 18:2088–2091
Kashimshetty Y, Pelikan S, Rogstad SH (2015) Variable gene dispersal conditions and spatial deforestation patterns can interact to affect tropical tree conservation outcomes. PLoS ONE 10:e0127745
Keenan K, McGinnity P, Cross TF, Crozier WW, Prodöhl PA (2013) diveRsity: an R package for the estimation and exploration of population genetics parameters and their associated errors. Methods Ecol Evol 4:782–788. https://doi.org/10.1111/2041-210X.12067
Kraemer P, Gerlach G. (2013) Package ‘Demerelate’. R software. https://cran.r-project.org/web/packages/Demerelate/Demerelate.pdf. Accessed 24 Dec 2015
Krishnan S, Ranker TA, Davis AP, Rakotomalala JJ (2013) The study of genetic diversity patterns of Coffea commersoniana, an endangered coffee species from Madagascar: a model for conservation of other littoral forest species. Tree Genet Genomes 9(1):179–187
LaBastille A, Pool D (1978) On the need for a system of cloud-forest parks in Middle America and the Caribbean. Environ Conserv 5(3):183–190
Leimu R, Mutikainen P, Koricheva J, Fischer M (2006) How general are positive relationships between plant population size, fitness and genetic variation? J Ecol 94:942–952
Li Q, He T, Xu Z (2005a) Genetic evaluation of the efficacy of in situ and ex situ conservation of Parashorea chinensis (Dipterocarpaceae) in southwestern China. Biochem Genet 43:387–406
Li YY, Chen YX, Zhang X, Wu TY, Ping LUH, Cai YW (2005b) Genetic differences between wild and artificial populations of Metasequoia glyptostroboides: implications for species recovery. Conserv Biol 19:224–231
Liu MH, Chen XY, Zhang X, Shen DW (2008) A population genetic evaluation of ecological restoration with the case study on Cyclobalanopsis myrsinaefolia (Fagaceae). Plant Ecol 197:31–41
Manel S, Gaggiotti OE, Waples RS (2005) Assignment methods: matching biological questions with appropriate techniques. Trends Ecol Evol 20:136–142
Maschinski J, Haskins KE, Raven PH (2012) Plant reintroduction in a changing climate. Island Press, Washington, pp 1–432
McKay JK, Christian CE, Harrison S, Rice KJ (2005) How local is local?—a review of practical and conceptual issues in the genetics of restoration. Restor Ecol 13:432–440
Menges ES (2008) TURNER REVIEW No. 16. Restoration demography and genetics of plants: when is a translocation successful? Aust J Bot 56:187–196
Menges ES, Stacy AS, Carl WW (2016) Adaptive introductions: how multiple experiments and comparisons to wild populations provide insights into requirements for long-term introduction success of an endangered shrub. Plant Divers 38(5):238–246
Miao YC, Su RJ, Zhang ZJ, Lang XD, Liu WD, Li SF (2015) Microsatellite markers indicate genetic differences between cultivated and natural populations of endangered Taxus yunnanensis. Bot J Linn Soc 177:450–461
Mijangos JL, Pacioni C, Spencer P, Craig MD (2015) Contribution of genetics to ecological restoration. Mol Ecol 24(1):22–37
Namoff S, Husby C, Francisco-Ortega J, Noblick L, Lewis C, Griffith P (2010) How well does a botanical garden collection of a rare palm capture the genetic variation in a wild population? Biol Conserv 143:1110–1117
Nei M, Feldman MW (1972) Identity of genes by descent within and between populations under mutation and migration pressures. Theor Popul Biol 3:460–465
Paetkau D, Slade R, Burden M, Estoup A (2004) Genetic assignment methods for the direct, real-time estimation of migration rate: a simulation-based exploration of accuracy and power. Mol Ecol 13:55–65
Peakall ROD, Smouse PE (2006) GENALEX 6: genetic analysis in Excel. Population genetic software for teaching and research. Mol Ecol Notes 6:288–295
Pence VC (2011) Evaluating cost for the in vitro propagation and preservation of endangered plants. In vitro Cellular & Developmental Biology 47(1):176–187
Piry S, Alapetite A, Cornuet JM, Paetkau D, Baudouin L, Estoup A (2004) GENECLASS2: a software for genetic assignment and first-generation migrant detection. J Hered 95:536–539
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155(2):945–959
R Development Core Team (2013) R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing, Vienna. http://www.R-project.org. Accessed 27 Nov 2015
Ralls K, Ballou JD, Dudash MR, Eldridge MD, Fenster ChB, Lacy RC, Sunnucks P, Frankham R (2017) Call for a paradigm shift in the genetic management fo fragment populations. Conserv Lett 11:e12412
Rannala B, Mountain JL (1997) Detecting immigration by using multilocus genotypes. Proc Natl Acad Sci USA 94:9197–9221
Raven P, Havens K (2014) Ex situ plant conservation and cryopreservation: breakthroughs in tropical plant conservation. Int J Plant Sci 175(1):1–2
Reed DH, Frankham R (2003) Correlation between fitness and genetic diversity. Conserv Biol 17(1):230–237
Rice KJ, Emery NC (2003) Managing microevolution: restoration in the face of global change. Front Ecol Environ 9:469–478
Robert A, Couvet D, Sarrazin F (2007) Integration of demographic and genetics in population restorations. Ecoscience 14(4):463–471
Rousset F (1995) GENEPOP (version 1.2): population genetics software for exact tests and ecumenicism. J Hered 86:248–249
Rousset F (2008) genepop’007: a complete re-implementation of the genepop software for Windows and Linux. Mol Ecol Resour 8:103–106
Ruber L, Meyer A, Sturmbauer C, Verheyen E (2001) Population structure in two sympatric species of the Lake Tanganyika cichlid tribe Eretmodini: evidence for introgression. Mol Ecol 10:1207–1225
Sanín MJ (2013) Estudios ecológicos y evolutivos en Ceroxylon (palmae: ceroxyloideae). Dissertation. Universidad Nacional de Colombia
Sanín MJ, Galeano G (2011) A revision of the Andean wax palms, Ceroxylon (Arecaceae). Phytotaxa 34:1–64
Sanín MJ, Anthelme F, Pintaud JC, Galeano G, Bernal R (2013) Juvenile resilience and adult longevity explain residual populations of the Andean Wax Palm Ceroxylon quindiuense after Deforestation. PLoS ONE 8:74139
Sanín MJ, Zapata JP, Pintaud JC, Galeano G, Bohórquez A, Tohme J, Hansen MM (2017) Up and down the blind alley: population divergence with scant gene flow in an endangered tropical lineage of Andean palms (Ceroxylon quindiuense clade: Ceroxyloideae). J Hered. https://doi.org/10.1093/jhered/esx006
Santos AS, Cazetta E, Dodonov P, Faria D, Gaiotto FA (2016) Landscape-scale deforestation decreases gene flow distance of a keystone tropical palm, Euterpe edulis Mart (Arecaceae). Ecol Evol 6:6586–6598
Svenning JC (1998) The effect of land-use on the local distribution of palm species in an Andean rain forest fragment in northwestern Ecuador. Biodivers Conserv 7:1529–1537
Svenning JC, Harlev D, Sorensen MM, Balslev H (2009) Topographic and spatial controls of palm species distributions in a mountain rain forest, southern Ecuador. Biodivers Conserv 18:219–228
Szczecińska M, Sramko G, Wołosz K, Sawicki J (2016) Genetic diversity and population structure of the rare and endangered plant species Pulsatilla patens (L.) Mill in East Central Europe. PLoS One 11(3):e0151730
Walters C (2004) Principles for preserving germplasm in gene banks. In: Guerrant EO, Havens K, Maunder M (eds) Ex situ plant conservation: supporting species survival in the wild. Island Press, Washington, pp 113–138
Weir BS, Cockerham CC (1984) Estimating F-statistics for the analysis of population structure. Evolution 38(6):1358–1370
Wright S (1969) Evolution and the Genetics of populations, 2nd edn. The theory of gene frequencies. The University of Chicago Press, Chicago
Yokogawa M, Kaneko S, Takahashi Y, Isagi Y (2013) Genetic consequences of rapid population decline and restoration of the critically endangered herb Polemonium kiushianum. Biol Conserv 157:401–408
Acknowledgements
This research was funded by Corporación Autónoma Regional del Quindío-CRQ, Colombia. We would like to thank Adriana Bohórquez, Patricia Zapata and Janeth Gutierrez for the big support during the lab work at the International Center for Tropical Agriculture (CIAT) and Johan Carvajal for his big help during the field work. Additionally, we would like to thank the reviwers for their comments that considerable improved this manuscript. Finally, we would like to thank Danny McSweeney, Chitra Naidu, Alicia Knudson and Allan Carrillo-Baltodano for their valuable feedback and improving the English of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by David L. Hawksworth.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article belongs to the Topical Collection: Ex-situ conservation.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Chacón-Vargas, K., García-Merchán, V.H. & Sanín, M.J. From keystone species to conservation: conservation genetics of wax palm Ceroxylon quindiuense in the largest wild populations of Colombia and selected neighboring ex situ plant collections. Biodivers Conserv 29, 283–302 (2020). https://doi.org/10.1007/s10531-019-01882-w
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10531-019-01882-w