Abstract
Itaconic acid is an excellent polymeric precursor with a wide range of industrial applications. The efficient production of itaconate from various renewable substrates was demonstrated by engineered Escherichia coli. However, limitation in the itaconic acid precursor supply was revealed by finding out the key intermediate of the tricarboxylic acid in the itaconic acid pathway. Efforts of enhancing the cis-aconitate flux and preserving the isocitrate pool to increase itaconic acid productivity are required. In this study, we introduce a synthetic protein scaffold system between CadA and AcnA to physically combine the two enzymes. Through the introduction of a synthetic protein scaffold, 2.1 g L−1 of itaconic acid was produced at pH 7 and 37 °C. By fermentation, 20.1 g L−1 for 48 h of itaconic acid was produced with a yield of 0.34 g g−1 glycerol. These results suggest that carbon flux was successfully increased itaconic acid productivity.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Itaconic acid is a white crystalline unsaturated C5 dicarbonic acid, which is widely used in the industrial synthesis of resins like polyesters and plastics, and the total market size of itaconic acid is about 30,000 ton per year (Steiger et al. 2013). In the itaconic acid production, due to high cost and relatively low yields, chemical synthesis of itaconic acid has been less competitive compared to the microbial fermentation processes. Among various itaconic acid producing strains, Aspergillus terreus is one of the dominant itaconic acid production hosts considering the high itaconic acid concentration (129 g L−1) (Kuenz et al. 2012; Okabe et al. 2009). However, some characteristics of A. terreus such as the filamentous phenotype, slow growth rate, spore-forming life cycle and difficulty in genetic modification hinder the further engineering of A. terreus strain and process development.
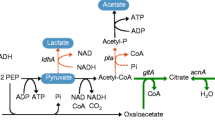
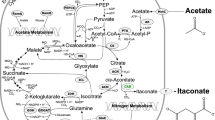
The itaconic acid biosynthesis mechanism consists of the enzymatic decarboxylation of cis-aconitate to itaconic acid (Fig. 1). cis-Aconitate is an intermediate of the tricarboxylic acid cycle, and the itaconic acid synthesis is catalyzed by cis-aconitic acid decarboxylase (CadA). Since 2008, the sequence of the CadA of A. terreus was characterized, and the enhanced itaconic acid production studies were reported by overexpression of the heterologous CadA. For example, the overexpression CadA in Saccharomyces cerevisiae lead 0.17 g L−1 itaconic acid production from glucose (Blazeck et al. 2014). CadA has been introduced into various strains and the constructed recombinant strains were cultured with various carbon substrates for the itaconic acid production. Recombinant Yarrowia lipolytica produced 4.6 g L−1 itaconic acid from glucose (Blazeck et al. 2015), recombinant Candida lignohabitans produced 4 g L−1 from glucose (Bellasio et al. 2015), recombinant Escherichia coli produced 7 g L−1 from glycerol (Jeon et al. 2016), recombinant Corynebacterium glutamicum produced 7.8 g L−1 from glucose (Otten et al. 2015) and recombinant Synechocystis sp. PCC6803 produced 0.015 g L−1 from CO2 (Chin et al. 2015). In spite of the intensive studies, the overall itaconic acid titer obtained from the recombinant hosts still remain low level compare with A. terreus. Therefore, development of the novel itaconic acid production strategy was required for further improvement of itaconic acid production. E. coli have many advantages as a production host, including well-established molecular tools for genetic modification and relatively easy operating conditions for fermentation. Recently E. coli is considered as promising non-natural hosts that produces itaconic acid (Becker and Wittmann 2016; Harder et al. 2016).
Enzyme co-localization is a novel strategy used to engineer the intracellular metabolism via protein–protein interaction between the domains and ligands. The co-localizing metabolic pathway enzymes can lead the decrease of the intermediates transit time, the prevent of intermediates loss to diffusion or competing pathways and the prevention of unstable intermediates (Miles et al. 1999). Therefore, the cellular metabolic flux can be redirected toward desired metabolic networks and metabolites. The enzyme co-localization strategy can improve metabolite production with low expression levels of pathway enzymes, and it has been used for the development of various metabolite production systems (Kuenz et al. 2012). For example, the concentration of glucaric acid in the recombinant E. coli was increased by 5 folds via employing enzyme co-localization strategy (Moon et al. 2010).
In this work, we employed the enzyme co-localization strategy to for the enhanced itaconic acid production from various carbon source in recombinant E. coli. CadA and aconitase (AcnA) were co-localized, and the constructed recombinant srtain was cultured for itaconic acid production. Itaconic acid production conditions were optimized, and the significant increase of itaconic acid concentration was archived in fed-batch fermentation study.
Materials and methods
Strains and culture conditions
The bacterial strains used in this study are listed in Table 1. The single colonies were picked from Luria–Bertani (LB) plates and inoculated overnight in 10 mL LB medium (10 g L−1 bacto-tryptone, 5 g L−1 bacto-yeast extract and 5 g L−1 NaCl) at 37 °C in a rotary shaker (250 rpm), then inoculated (1%, v v−1) into 100 mL LB supplemented with antibiotics (100 g mL−1 ampicillin) until the optical density of the suspension reaches 0.6 at 600 nm wavelength (OD600). For expressing genes, 0.5% arabinose was added to the culture medium. pH, temperature and, glucose (10 g L−1) was tested to optimize the culture conditions.
For itaconic acid production, M9 medium (12.8 g L−1 Na2HPO4·7H2O, 3 g L−1 KH2PO4, 0.5 g L−1 NaCl, 1 g L−1 NH4Cl, 1 mmol MgSO4, 1 mg L−1 vitamin B1 and 0.1 mmol CaCl2) was also used with various carbon sources (10 g L−1 of xylose or 10 g L−1 of glycerol).
Fed-batch fermentation was performed in the 5 L stirred-tank bioreactor. Initially, 2.5 L of the modified M9 medium was used with the addition of 10 g L−1 of glycerol, 5 g L−1 of yeast extract. The fresh medium was inoculated with 100 mL of the overnight pre-culture grown in M9, 0.5% arabinose was added at the time of inoculation. Dissolved oxygen (DO) was maintained at 10% with respect to air saturation by raising the stirrer speed (from 300 to 800 rpm). The aeration was set to 1 vvm (volume of gas per volume of liquid per minute). The pH was maintained at 7 by the automatic addition of 1 M NaOH. At the 12 h, 24 h, 36 h 48 h, and 60 h, 15.6 mL of glycerol feed with 625 g L−1 were fed to the bioreactor. All chemicals and reagents were purchased from Sigma–Aldrich (South Korea) unless otherwise specified.
Plasmid construction
The oligonucleotides used in this study were listed in Table 2. The codon-optimized cadA gene (Gene bank ID: KM464677) (Vuoristo et al. 2015) was chemical synthesized and acnA from E. coli genome were amplified using the Expand high-fidelity polymerase chain reaction (PCR) system (Roche Molecular Biochemicals, Mannheim, Germany) from XB chromosomal DNA. SH3 domain and SH3 ligand genes were PCR amplified from the laboratory stock plasmid. Then, the SH3 domain gene was fused to the C-terminus of acnA by overlap PCR to make acnA-SH3D, The SH3 ligand was attached to the C-terminus gene to make cadA-SH3L. The acnA-SH3D genes were cloned into pBAD30C using SacI and XmaI restriction sites. The cadA-SH3L gene was cloned downstream of acnA-SH3D genes using XmaI and XbaI restriction sites to construct the pBAC expression plasmids. Restriction enzymes, T4 DNA polymerase were obtained from New England Biolabs (Ipswich, MA). Oligonucleotides were purchased from Genotech (Daejeon, South Korea).
Analytical method
The concentrations of substrates and organic acids were determined by HPLC using the Aminex HPX-87H column (300 \(\times\) 7.8 mm, Bio-Rad). Samples were centrifuged at 12,000 rpm for 5 min. Then, the 1 mL supernatant was filtered through a 0.2 μm millipore filter and analyzed on HPLC system using RID detector. 0.08 N H2SO4 was used as a mobile phase. The temperature of the column was set at 35 °C and the flow rate of the mobile phase was 0.6 mL min−1. The standard curves were determined using the same procedure for seven standard solutions: 0.1, 0.5, 1, 2, 3, 5, and 10 g L−1 of glucose, xylose, glycerol and organic acids (Sigma, Missouri, USA).
Result and discussion
Construction of the itaconic acid production strain
To overcome the low productivity of itaconic acid and achieve the economic feasibility of itaconic acid production process, the novel strategy allowing the redirection of pathway intermediate to itaconic acid network should be developed (Chang et al. 2017). In the TCA cycle, citrate is converted to isocitrate via cis-aconitate through two sequential reactions which are catalyzed by a single enzyme, AcnA. For the efficient itaconic acid production, the intermediate cis-aconitate need to redirected to itaconic acid by CadA (Fig. 1). If CadA was co-localized with AcnA by ligand-domain interaction, cis-Aconitrate produced by AcnA will have more chance to contact with CadA and can be converted to itaconic acid with higher chance.
To realize this idea, the codon-optimized cadA gene with SH3 ligand and acnA with SH3 domain were cloned to construct the plasmid pBAC, then introduced into E. coli.
By the overexpression of scaffold consist with AcnA-SH3D and CadA-SH3L under the control of araC promoter, the recombinant E. coli strain produced about 2.1 g L−1 of itaconic acid in the culture in 48 h (Fig. 2). The control strain without scaffold produced 0.19 g L−1 of itaconic acid. These data showed that itaconic acid production was increased by 11 folds by introduction of protein scaffold between AcnA and CadA, and TCA cycle carbon flux was redirected to itaconic acid pathway.
Itaconic acid production by the strains with and without the AcnA and CadA scaffold. Itaconic acid concentration obtained with control strain E. coli (pBNC) (unfilled circle). Concentrations of itaconic acid (filled circle), glucose (filled square), and cell (filled triangle) obtained with E. coli (pBAC)
Optimizing condition of itaconic acid production
In order to optimize the conditions for itaconic acid production, the various pH (5, 6, 7, 8 and 9) and temperature (25, 30 and 37 °C) were tested. As shown in Fig. 3a, the strain cultured in 30 °C generally provided higher itaconic acid concentration. The strain cultured in pH 5 and 25 °C did not showed the increase of itaconic acid concentration (0.1 g L−1) compare with wild type strain. When the E. coli (pBAC) was cultured in pH 7.0 at 30 °C, the highest itaconic acid concentration of 2.1 g L−1 was achieved in 48 h with 10 g L−1 glucose as a carbon source (Fig. 3a).
To evaluate the effect of culture media, E. coli (pBAC) strain was cultured in M9 minimal media supplemented with glucose (10 g L−1). During culture, the itaconic acid concentration gradually increased and reached 3.6 g L−1 at 48 h (Fig. 3b). This result supports that M9 minimal media is more suitable for itaconic acid than LB media.
To improve itaconic acid production, various carbon sources were tested since different carbon source can affect intracellular metabolism by producing different amounts of metabolites including NADH, ATP, etc. (Fig. 4). Glycerol is converted to glyceraldehyde-3-P (G3P) via glycerol-3-P, and then joins glycolysis pathway and further converted to itaconic acid. Xylose is converted to fructose-6-P and G3P via pentose phosphate pathway, and enters glycolysis pathway. When the strain was cultured in M9 medium supplemented with 10 g L−1 of glycerol, 4.8 g L−1 of final itaconic acid concentration was achieved. Xylose and glucose supplemented medium provided 3.4 and 3.5 g L−1 of itaconic acid concentration, respectively (Fig. 4). Time profiles of cell densities were also compared. When glycerol was used as substrate, cell density (OD600) was increased to 12.4 at the end of culture while 15.9 of OD600 was achieved with glucose. In combination with itaconic acid concentration data, it can be deduced that intracellular carbon flux was directed toward itaconic acid pathway more efficiently when glycerol was used carbon substrate. Therefore, it was suggested that glycerol is good substrate candidate for itaconic acid biosynthesis in E. coli.
The effect of glycerol in organic acid production has been investigated in various studies for long time (Gao et al. 2016; Le Meur et al. 2014; Zambanini et al. 2016). It has been reported that glycerol can be considered as suitable substrate for the production of certain product considering its highly reduced state (Dharmadi et al. 2006). Glycerol is also an inexpensive and abundant carbon source, which makes glycerol more attractive carbon source.
Fed-batch fermentation for itaconic acid production with glycerol
To achieve higher itaconic acid concentration, fed-batch fermentation study was carried out in 5 L reactor using glycerol as carbon source (Fig. 5). Since it has been reported that controlling pH is important for itaconic acid production, fed-batch fermentation was carried out at pH 7. During 60 h of fermentation, itaconic acid concentration continuously increased as well as cell density. The highest itaconic acid concentration of 20.1 g L−1 was obtained at 48 h, and it decreased after that point. Through fed-batch fermentation study, 5 times higher cell concentration was obtained compare with flask culture and 4 times higher itaconic acid concentration was obtained (Fig. 5). These results suggest that itaconic acid concentration can be elevated by feeding of carbon substrate such as glycerol.
Conclusion
In this study, the synthetic scaffold was employed as a novel strategy to construct more efficient itaconic acid producing recombinant strain. Through the synthetic protein scaffold strategy, a stable complex of several enzymes can be formed and co-localized in specific space. That leads to decreasing the diffusion of the product and increase metabolite production rate and yield. By scaffold, CadA is physically connected to AcnA. Therefore, cis-aconitate produced by AcnA can have more chance to react with CadA than other competing enzymes and can be converted to itaconic acid more efficiently. By the introduction of synthetic protein scaffold, itaconic acid concentration was increased 90.9% in the E. coli XB (pBAC) strain compared with that obtained with E. coli XB (pBNC), which does not have synthetic protein scaffold. By fed-batch fermentation, itaconic acid concentration was further increased to 20.1 g L−1. The strategy developed in this study can be applied for the bio-refinery host improvements, and might enable the generation of an E. coli strain as an industrial itaconic acid producer.
Availability of data and materials
The datasets generated during and/or analysed during the current study are available from the corresponding author on reasonable request.
References
Becker J, Wittmann C (2016) Systems metabolic engineering of Escherichia coli for the heterologous production of high value molecules—a veteran at new shores. Curr Opin Biotechnol 42:178–188. https://doi.org/10.1016/j.copbio.2016.05.004
Bellasio M, Mattanovich D, Sauer M, Marx H (2015) Organic acids from lignocellulose: Candida lignohabitans as a new microbial cell factory. J Ind Microbiol Biotechnol 42:681–691. https://doi.org/10.1007/s10295-015-1590-0
Blazeck J, Miller J, Pan A, Gengler J, Holden C, Jamoussi M, Alper HS (2014) Metabolic engineering of Saccharomyces cerevisiae for itaconic acid production. Appl Microbiol Biotechnol 98:8155–8164. https://doi.org/10.1007/s00253-014-5895-0
Blazeck J, Hill A, Jamoussi M, Pan A, Miller J, Alper HS (2015) Metabolic engineering of Yarrowia lipolytica for itaconic acid production. Metab Eng 32:66–73. https://doi.org/10.1016/j.ymben.2015.09.005
Chang P, Chen GS, Chu H-Y, Lu KW, Shen CR (2017) Engineering efficient production of itaconic acid from diverse substrates in Escherichia coli. J Biotechnol 249:73–81. https://doi.org/10.1016/j.jbiotec.2017.03.026
Chin T, Sano M, Takahashi T, Ohara H, Aso Y (2015) Photosynthetic production of itaconic acid in Synechocystis sp. PCC6803. J Biotechnol 195:43–45. https://doi.org/10.1016/j.jbiotec.2014.12.016
Dharmadi Y, Murarka A, Gonzalez R (2006) Anaerobic fermentation of glycerol by Escherichia coli: a new platform for metabolic engineering. Biotechnol Bioeng 94:821–829. https://doi.org/10.1002/bit.21025
Gao C, Yang X, Wang H, Rivero CP, Li C, Cui Z, Qi Q, Lin CSK (2016) Robust succinic acid production from crude glycerol using engineered Yarrowia lipolytica. Biotechnol Biofuels 9:179. https://doi.org/10.1186/s13068-016-0597-8
Harder B-J, Bettenbrock K, Klamt S (2016) Model-based metabolic engineering enables high yield itaconic acid production by Escherichia coli. Metab Eng 38:29–37. https://doi.org/10.1016/j.ymben.2016.05.008
Jeon H-G, Cheong D-E, Han Y, Song JJ, Choi JH (2016) Itaconic acid production from glycerol using Escherichia coli harboring a random synonymous codon-substituted 5′-coding region variant of the cadA gene. Biotechnol Bioeng 113:1504–1510. https://doi.org/10.1002/bit.25914
Kuenz A, Gallenmüller Y, Willke T, Vorlop K-D (2012) Microbial production of itaconic acid: developing a stable platform for high product concentrations. Appl Microbiol Biotechnol 96:1209–1216. https://doi.org/10.1007/s00253-012-4221-y
Le Meur S, Zinn M, Egli T, Thöny-Meyer L, Ren Q (2014) Improved productivity of poly (4-hydroxybutyrate) (P4HB) in recombinant Escherichia coli using glycerol as the growth substrate with fed-batch culture. Microb Cell Fact 13:131–131. https://doi.org/10.1186/s12934-014-0131-2
Miles EW, Rhee S, Davies DR (1999) The molecular basis of substrate channeling. J Biol Chem 274:12193–12196. https://doi.org/10.1074/jbc.274.18.12193
Moon TS, Dueber JE, Shiue E, Prather KLJ (2010) Use of modular, synthetic scaffolds for improved production of glucaric acid in engineered E. coli. Metab Eng 12:298–305. https://doi.org/10.1016/j.ymben.2010.01.003
Okabe M, Lies D, Kanamasa S, Park EY (2009) Biotechnological production of itaconic acid and its biosynthesis in Aspergillus terreus. Appl Microbiol Biotechnol 84:597–606. https://doi.org/10.1007/s00253-009-2132-3
Otten A, Brocker M, Bott M (2015) Metabolic engineering of Corynebacterium glutamicum for the production of itaconate. Metab Eng 30:156–165. https://doi.org/10.1016/j.ymben.2015.06.003
Steiger M, Blumhoff M, Mattanovich D, Sauer M (2013) Biochemistry of microbial itaconic acid production. Front Microbiol. https://doi.org/10.3389/fmicb.2013.00023
Vuoristo KS, Mars AE, Sangra JV, Springer J, Eggink G, Sanders JPM, Weusthuis RA (2015) Metabolic engineering of itaconate production in Escherichia coli. Appl Microbiol Biotechnol 99:221–228. https://doi.org/10.1007/s00253-014-6092-x
Zambanini T, Sarikaya E, Kleineberg W, Buescher JM, Meurer G, Wierckx N, Blank LM (2016) Efficient malic acid production from glycerol with Ustilago trichophora TZ1. Biotechnol Biofuels 9:67. https://doi.org/10.1186/s13068-016-0483-4
Funding
This work was supported by the 2024 Research Fund of the University of Ulsan, Republic of Korea.
Author information
Authors and Affiliations
Contributions
Kim-Ngan T. Tran and Hong Soon Ho designed the study. Kim-Ngan T. Tran and Jaehoon Jeong performed the study. Kim-Ngan T. Tran, Jaehoon Jeong and Hong Soon Ho performed the study commented on draft versions of the article. All authors have approved the final article.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no relevant financial or non-financial interests to disclose.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Consent to participate
This article does not contain any studies with human participants or animals performed by any of the authors.
Consent to publish
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Tran, KN.T., Jeong, J. & Hong, S.H. Engineering of itaconic acid pathway via co-localization of CadA and AcnA in recombinant Escherichia coli. Biotechnol Lett 46, 593–600 (2024). https://doi.org/10.1007/s10529-024-03496-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-024-03496-x