Abstract
Protein FadR is known as a fatty acid metabolism global regulator that sustains cell envelope integrity by changing the profile of fatty acid. Here, we present its unique participation in the defense against reactive oxygen species (ROS) in the bacterium. FadR contributes to defending extracellular ROS by maintaining the permeability of the cell membrane. It also facilitates the ROS detoxification process by increasing the expression of ROS neutralizers (KatB, KatG, and AhpCF). FadR also represses the leakage of ROS by alleviating the respiratory action conducted by terminal cytochrome cbb3-type heme-copper oxidases (ccoNOQP). These findings suggest that FadR plays a comprehensive role in modulating the bacterial oxidative stress response, instead of merely strengthening the cellular barrier against the environment. This study sheds light on the complex mechanisms of bacterial ROS defense and offers FadR as a novel target for ROS control research.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
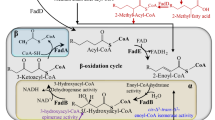
Reactive oxygen species (ROS) are oxygen-containing molecules that can exist independently in the cytoplasm. This group of molecules includes oxygen free radicals, such as peroxide (-O–O-), superoxide (O2−), hydroxyl radical (·OH), singlet oxygen (1O2), and ozone (O3) (Jakubczyk et al. 2020). ROS are by-products of oxygen metabolism and play important roles in cell metabolism, such as signaling and tissue homeostasis (Ray et al. 2012). However, under the duress of certain circumstances (for example, UV or heat exposure), cells usually produce excess ROS, leading to oxidative stress (Schieber and Chandel 2014). Oxidative stress can adversely affect cell modification and cause damage to virtually all biomolecules such as DNA, RNA, lipids, and proteins. For example, ROS would cause lipid peroxidation (LPO), the oxidation of unsaturated fatty acids (UFA). The LPO process produces malignant molecules that contribute to many diseases and pathologies (Nam 2011). Under normal circumstances, the generation and peroxidation of lipids is in dynamic equilibrium. However, when cells encounter excess ROS, the balance of LPO is disrupted and cell components that constitute high contents of UFA are damaged by ROS, such as cell membranes and lipoproteins (Pizzino et al. 2017). Therefore, the consumption of unsaturated fatty acids and the corresponding electron transfer process in the respiratory chain is closely related to the generation and detoxification of reactive oxygen species (ROS). However, conclusive evidence endorsing this hypothesis is still missing and awaits experimental validation.
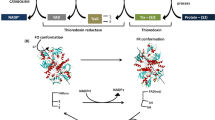
Cells have evolved many strategies to remove excessive ROS, such as the expression of scavenging enzymes like hydroperoxidase, superoxide dismutase (SOD), catalase (KAT), ascorbate peroxidase (APX) (Fath et al. 2001). These enzymes convert free ROS to water and oxygen independently or cooperatively. For example, the alkyl hydroperoxidase reductase system ahpCF that contains ahpC and ahpF detoxifies H2O2 in many species. The residue Cys46 in AhpC is oxidized by H2O2 to form a disulfide bond (Cha et al. 2015). As is generally accepted, when the cellular concentration of H2O2 surpasses 20 nM, the scavenging ability of AhpCF meets its limit, and the OxyR system is activated. OxyR is a regulator that directly senses H2O2 by oxidizing two conserved cysteine residues and forming an intramolecular disulfide bond (Cha et al. 2015). In E. coli or Salmonella enterica serovar, OxyR also acts as an activation regulator to induce downstream catalase expression (Hahn et al. 2002). Not only the scavenging enzymes but also the bacterial terminal oxidases are reported to participate in the control of cellular ROS levels, like quinol oxidase and heme-copper oxidase (Borisov et al. 2021).
To elucidate the possible correlation between fatty acid metabolism and ROS defense in bacteria, we investigated these processes in the Shewanella oneidensis MR-1. This bacterium belongs to the phylum Gammaproteobacteria, a facultative anaerobe widely distributed in marine and freshwater environments. MR-1 is an ideal model organism for this study as it thrives in redox-stratified environments with excess ROS (Ikeda et al. 2021) and it contains multiple respiratory pathways for a variety of organic and inorganic substances to generate excessive ROS and electron leakages. The other advantage of using S. oneidensis MR-1 as the research subject is its thoroughly studied fatty acid metabolism. This advantage provides a valuable foundation for modifying fatty acid synthesis and understanding its implications in the ROS defense mechanism.
A key factor for modifying the bacterial fatty acids metabolism system is the regulator FadR. It binds to specific DNA sequences and controls the expression of the genes involved in the synthesis, degradation, and transport of fatty acids (Cronan et al. 1998; Zhang et al. 2015). For instance, FadR down-regulates several genes in the fatty acid degradation pathway, including fadE, fadBA, fadH, and fadIJ (Feng and Cronan 2009) in E. coli. It also down-regulates fadD that involved in the conversion of fatty acids to acyl-CoAs, and fadL which transports fatty acids across the cell membrane (Cronan 2021). FadR up-regulates fabA and fabB which count for the biosynthesis of UFA (Campbell and Cronan 2001). The deletion of fadR represses the UFA biosynthesis and enhances the fatty acid degradation, thus reducing the total UFA concentrations (Nunn et al. 1983). Correspondingly, the overexpression of FadR could slow down the fatty acid degradation and enhance UFA biosynthesis, thus increasing the accumulation of cellular UFA (Luo 2014).
As shown above, the modification of fadR is a feasible approach for changing the cellular UFA concentration, and this study will be focused on the physiological consequences of the ROS-related metabolism processes brought by the fadR modifications. The results showed that the fadR knockout MR-1 mutant (ΔfadR) is more sensitive to H2O2, and this sensitivity was related to the compromised membrane permeability of MR-1, the enhanced respiration intensity, and the down-regulated ROS-scavenging enzymes (AhpCF, KatB, and KatG). This is the first evidence of FadR’s impact on the overall bacterial ROS defense system.
Materials and methods
Strains, plasmids, and chemicals
The strain Shewanella oneidensis MR-1, the plasmid pHGEI01-lacZ, and the plasmid pHGEPtac were kind gifts given by Dr. Haichun Gao from Zhejiang University. The strains DH5α and WM3064 were commercial cells purchased from manufacturer TsingkeBiotechnology Co., Ltd. The rest of the strains and plasmids used in this study were constructed in-house. The abbreviations of the strains with genetic modifications are explained in detail in Table 1. All the chemicals used in this study were purchased from Sinopharma Co., Ltd.
Disk diffusion assay
The disk diffusion assay was used to test the sensitivity to hydrogen peroxide of S. oneidensis MR-1 strains with different genetic modifications. The cells in the log phase were collected at 4000 rpm for 2 min and adjusted to 109 cells/ml. The cell mixture was diluted 5 times with fresh LB medium before being spread on the LB plates containing IPTG (0.1 mM) at 30 °C for 24 h. Two hundred microliters of cell culture in the mid-exponential phase were spread on LB plates. After 6 h inoculation, a circular paper disk of 6 mm diameter soaked with 10 μl H2O2 was placed on the bacterial lawn at 30 °C for 16 h.
Droplet assay
The droplet assay was used to evaluate the growth-inhibition effects of SDS on the S. oneidensis MR-1 strains with different genetic modifications. S. oneidensis MR-1 cells in the log phase were collected by centrifugation (4000 rpm) and adjusted to 109 cells/ml. Then, a serial dilution by tenfold was performed by fresh LB medium. Diluted cell culture (5 μl) was dropped onto the LB plates supplemented with IPTG (0.1 mM) and SDS of different concentrations and inoculated at 30 °C for 24 h.
β-Galactosidase activity assay
The β-galactosidase activity was determined with the E. coli lacZ integrated reporter gene pHGEI01 (Meng et al. 2018). Briefly, we amplified a sequence approximately 500 bp upstream of the DNA sequence that might contain a promoter for the gene of interest on the 5’-end of the lacZ gene. The vector was constructed in E. coli DH5α and transferred into S. oneidensis MR-1 through the conjugation of E. coli WM3064. S. oneidensis MR-1 cells in log-phase (optical density 600 nm OD600 ~ 0.4) were collected by centrifugation, washed with PBS, and then subjected to o-nitrophenyl-β-D-galactopyranoside (ONPG) -based assays as described previously. The β-galactosidase activity was determined by monitoring the color development at 420 nm using a TECAN microplate reader, and the results are presented as Miller Units.
Minimum inhibitory concentration (MIC) assay
MR-1 was cultured overnight, and 30 µl of the bacterial solution was transferred to a 3 ml LB liquid medium and incubated at the corresponding temperature with a shaking machine to OD600 ~ 0.5 to ensure that the strain was in the logarithmic growth phase. Take a certain amount of bacterial liquid in the test tube, add LB liquid medium, and dilute it 5000 times for use. The H2O2 was diluted in a new 96-well plate according to a two-fold concentration gradient, and then the bacteria solution was added and mixed to set up a control group. The 96-well plates were placed in the corresponding temperature incubator for 16 to 24 h, and then the absorbance value was measured and recorded by a microplate reader.
Microscopy
MR-1 was cultivated to the mid-logarithmic (OD600 ~ 0.4), mixed 1:1 with 0.2 M H2O2, and spotted onto a glass slide containing LB medium. LW300LHT phase contrast microscope was employed to observe cell morphology. Micrographs were captured with a camera.
Cytochrome oxidase activity assay
The Nadi test was used for visual analysis of cytochrome cbb3 oxidase activity (Wan et al. 2017). Three microliters of each culture at the mid-log phase under test were dropped onto LB plates, and the plates were incubated for 24 h. A solution of 0.5% α-naphthol in 95% ethanol and 0.5% N, N-dimethyl-ρ-phenyleneidiamine monohydrochloride (DMPD) was applied to cover the droplets developed. The formation of indophenols blue was timed as an indicator of cytochrome cbb3 activity.
FOX assay
The ferrous oxidation-xylenol orange (FOX) assay was employed to test the residual H2O2 on the outside of S. oneidensis MR-1 cells after H2O2 treatment (Feng et al. 2020). The reagent I contained 100 mM mannitol and 125 μM dimethylthiophenol. The reagent II contained 25 mM ferrous ammonium sulfate and 2.5 M sulfuric acid. The FOX working reagent was made by mixing reagent I and reagent II at a ratio of 100:1. The standard curve was made by mixing 20 μl H2O2 (0.05, 0.1, 0.15, 0.2, 0.25, 0.3, 0.4, 0.5 mM) with 180 μl FOX reagent. Cultivate different strains until OD600 reaches 0.5 and lyse the cells with ultrasonication. Then, clarify 50 μl samples with a 25 μm filter tip at 0, 1, 5, and 10 min. Mix 20 μl filtered sample with 180 μl FOX reagent in a 96-well plate and incubate at 37 °C for 30 min. Afterward, take the absorbance of the mixture at a wavelength of 560 nm. The H2O2 concentration can be calculated according to the standard curve.
Results and discussion
FadR-mediated fatty acids synthesis is directly related to the defense against ROS by MR-1
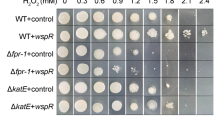
As is shown in Fig. 1A, with the treatment of H2O2, a larger bacteriostatic zone appeared on the cell lawn of mutant MR-1 strain ΔfadR (with the fadR knockout) than the WT (wild-type MR-1 strain). When the ΔfadR was complemented by a plasmid pHGEPtac-fadR that expresses FadR (ΔfadR/Ptac-fadR), its H2O2 resistance was recovered, if not enhanced (Fig. 1A, B, C). This means that the defense mechanism in S. oneidensis MR-1 against extracellular ROS is enhanced by fadR. A similar observation was made during the investigation on fabB (3-oxoacyl-(acyl-carrier-protein) synthase) that catalyzes a key reaction in UFA synthesis, the elongation of the cis-3-decenoyl-ACP (Feng and Cronan 2011). Like fadR knockout, ΔfabB also manifested deteriorated tolerance against H2O2, and this defect was remedied by overexpressing the fabB gene on pHGEPtac plasmid in the strain ΔfabB/Ptac-fabB (Fig. 1A, B, C). These observations manifest that the synthesis of fatty acids controlled by fadR helps MR-1 cells defend ROS from the outside.
Genes fadR and fabB facilitate the bacterial resistance against ROS. A The inhibition effects of H2O2 on the cell lawn of Shewanella oneidensis MR-1 with different genetic modifications. The abbreviations of the strains with genetic modifications are explained in detail in Table 1. B The diameter of the inhibition zone formed by H2O2 treatment to these strains. C The H2O2 minimum inhibitory concentration (MIC) of these strains. D The residual concentration of H2O2 generated by these strains. Biological triplicates were performed with the data presented as means ± SEM
FadR participates in ROS defense by maintaining the permeability of cell membrane
As fadR functions to increase intracellular fatty acid concentration and maintain the cell membrane integrity, the cell permeability towards ROS should also be subjected to fadR regulation. Phase-contrast microscopy images revealed no noticeable abnormality in cell morphology for wild-type MR-1 cells in the presence of 10 mM H2O2 (Fig. 2). However, when the fadR gene was knocked out, the MR-1 cell became longer and an increased rate of cell envelope rupture (from 22.8% to 39.3%) was observed in the presence of 10 mM H2O2 (Fig. 2). When fadR was complemented back to ΔfadR, the ΔfadR/Ptac-fadR cell manifested reduced rupture rate than the ΔfadR (4.8%). Since the ΔfadR cells were longer than the WT, to rule out the possibility that the compromised ROS resistance was due to the changed cell morphology or cell wall components, we knocked out the amiB (N-acetylmuramoyl-L-alanine amidase) gene from MR-1. AmiB belongs to the hydrolase’s protein family, it specifically hydrolyzes the carbon–nitrogen bonds and cleaves the link between N-acetyl muramyl residues and amino acid residues in cell wall glycopeptides, it participates in the synthesis of bacterial cell wall (Yakhnina et al. 2015). The mutant ΔamiB also manifests longer cell length, but the ΔamiB was not more vulnerable to ROS than the WT with no obvious cell rupture observed under the microscope (Fig. 2). These results suggest it is the cell membrane permeability that plays a central role in the ROS defense mechanism, instead of cell morphology or cell wall. This theory is also validated in the defense mechanism of MR-1 against sodium dodecyl sulfate (SDS). Like the H2O2 treatment, treatment of SDS at different concentrations has also manifested stronger harm to ΔfadR than the WT or ΔfadR/Ptac-fadR (Fig. 3). These results showed that fadR contributes to the ROS defense mechanism in M-1 by maintaining the integrity and permeability of cell membrane.
Gene fadR mediates cell permeability by changing the contents of the cell membrane. The upper/lower panels show the cell morphology of the genetically modified MR-1 strains without/with the treatment of hydroperoxide. The strains are named in Fig. 1
FadR represses the expression of terminal oxidases CcoNOQP to reduce ROS generation
Not only the defense efficiency against extracellular ROS was enhanced under fadR regulation, but the cellular ROS generation was also reduced. The FOX assay was used to measure the concentration of hydroperoxide generated by MR-1 (Banerjee et al. 2003). The results showed that the concentration of H2O2 generated by MR-1 increased with the knockout of fadR or fabB, and decreased when fadR or fabB was complemented back (Fig. 1D). This means that the function of fadR in the ROS defense is not only controlling the cell membrane permeability, but it also impacts the cellular ROS metabolism.
S. oneidensis MR-1 has three terminal oxidases for respiration: bd-type quinol oxidase (cydABX), caa3-type heme-copper oxidases (SO4606-9), and cbb3-type heme-copper oxidases (ccoNOQP) (Laz et al. 2014; Kouzuma et al. 2012). The electron transfer process is closely related to the activity of terminal oxidase. Electron leakage occurs during the electron transfer process, and higher terminal oxidase activity could generate more leaked electrons, resulting in more accumulation of ROS (Schönfeld and Wojtczak 2008). Under microaerobic and aerobic conditions, the predominant oxidase on the cell membrane of MR-1 is CcoNOQP (Laz et al. 2014). The Nadi reaction was performed to test the terminal oxidase activity in WT and mutant MR-1 strains (Yu et al. 2021). After five minutes of Nadi reaction, the cell lawn of ΔfadR formed a distinct dark blue circle, while the color of WT or ΔfadR/Ptac-fadR was shallower, indicating a stronger respiration in ΔfadR (Fig. 4A). Since the activity of the promoter of ccoNOQP (PcooNOQP) was higher in ΔfadR than the WT, the enhanced respiration activity was possibly due to the increased expression level of ccoNOQP (Fig. 4A). As a result, the fadR knockout induced an enhanced respiration pathway that generated more ROS, which has been observed in Fig. 1D. Thus, in the WT MR-1, fadR represses respiration activity by reducing the expression of ccoNOQP to control the overall ROS generation.
Gene fadR mediates the ROS defense mechanism by repressing the generation and enhancing the detoxification of ROS. A The activity of the ROS-generating cytochrome cbb3 type oxidase and its promoter PcooNOQP was increased in ΔfadR. B-D The promoters of the detoxification genes kcatB, kcatG, and aphCF were repressed in ΔfadR, and inductive effects of H2O2 toward PahpCF were eliminated. Biological triplicates were performed with the data presented as means ± SEM
FadR enhances ROS detoxification by promoting the expression of ROS-scavenging enzymes
We next investigated the regulation of FadR on the ROS scavenging enzymes KatB, KatG, and AhpCF (Feng et al. 2020; Toporek, et al. 2023). As is shown in Fig. 4B-D, the promoter activity of these three genes was significantly enhanced in the WT in the presence of 10 mM H2O2, endorsing the participation of these genes in ROS metabolism. When fadR was knocked out, the activity of these promoters was lower than the WT in the absence of H2O2, suggesting the potentially positive regulatory effects of fadR on these genes under normal conditions (Fig. 4B-D). Interestingly, the H2O2 stress did not increase the promoter activity of ahpCF in ΔfadR while the other two promoters were upregulated (Fig. 4D). This result implied that the regulatory effects of fadR on the ROS scavenging enzyme AhpCF were higher than that of KatB or KatG under ROS stress.
To sum up, this study revealed that fatty acid metabolism regulator FadR participates in the ROS defense mechanism in S. oneidensis MR-1 by preserving the cell membrane permeability, repressing the terminal cytochrome cbb3-type heme-copper oxidases, and activating the ROS scavenging enzymes katB, katG, and ahpCF.
References
Jakubczyk K et al (2020) Reactive oxygen species - sources, functions, oxidative damage. Pol Merkur Lekarski 48(284):124–127
Ray PD, Huang BW, Tsuji Y (2012) Reactive oxygen species (ROS) homeostasis and redox regulation in cellular signaling. Cell Signal 24(5):981–990
Schieber M, Chandel NS (2014) ROS function in redox signaling and oxidative stress. Curr Biol 24(10):R453–R462
Nam TG (2011) Lipid peroxidation and its toxicological implications. Toxicol Res 27(1):1–6
Pizzino G et al (2017) Oxidative Stress: Harms and Benefits for Human Health. Oxid Med Cell Longev 2017:8416763
Fath A, Bethke PC, Jones RL (2001) Enzymes that scavenge reactive oxygen species are down-regulated prior to gibberellic acid-induced programmed cell death in barley aleurone. Plant Physiol 126(1):156–166
Cha MK et al (2015) Characterization of two alkyl hydroperoxide reductase C homologs alkyl hydroperoxide reductase C_H1 and alkyl hydroperoxide reductase C_H2 in Bacillus subtilis. World J Biol Chem 6(3):249–264
Hahn JS, Oh SY, Roe JH (2002) Role of OxyR as a peroxide-sensing positive regulator in Streptomyces coelicolor A3(2). J Bacteriol 184(19):5214–5222
Borisov VB et al (2021) ROS Defense Systems and Terminal Oxidases in Bacteria. Antioxidants 10(6):839
Ikeda S et al (2021) Shewanella oneidensis MR-1 as a bacterial platform for electro-biotechnology. Essays Biochem 65(2):355–364
Cronan J, John E, Subrahmanyam S (1998) FadR, transcriptional co-ordination of metabolic expediency. Mol Microbiol 29(4):937–943
Zhang H et al (2015) Binding of Shewanella FadR to the fabA fatty acid biosynthetic gene: implications for contraction of the fad regulon. Protein Cell 6(9):667–679
Feng Y, Cronan JE (2009) Escherichia coli Unsaturated Fatty Acid Synthesis: complex transcription of the FabA gene and in vivo identification of the essential reaction catalyzed by FabB. J Biol Chem 284(43):29526–29535
Cronan JE (2021) The Escherichia coli FadR transcription factor: Too much of a good thing? Mol Microbiol 115(6):1080–1085
Campbell JW, Cronan JE (2001) Escherichia coli FadR positively regulates transcription of the fabB fatty acid biosynthetic gene. J Bacteriol 183(20):5982–5990
Nunn WD et al (1983) Role for fadR in unsaturated fatty acid biosynthesis in Escherichia coli. J Bacteriol 154(2):554–560
Luo Q et al (2014) Transcription factors FabR and FadR regulate both aerobic and anaerobic pathways for unsaturated fatty acid biosynthesis in Shewanella oneidensis. Front Microbiol 5:736
Meng Q, Liang H, Gao H (2018) Roles of multiple KASIII homologues of Shewanella oneidensis in initiation of fatty acid synthesis and in cerulenin resistance. Biochim Biophys Acta Mol Cell Biol Lipids 1863(10):1153–1163
Wan F, Shi M, Gao H (2017) Loss of OxyR reduces efficacy of oxygen respiration in Shewanella oneidensis. Sci Rep 7(1):42609
Feng X, Guo K, Gao H (2020) Plasticity of the peroxidase AhpC links multiple substrates to diverse disulfide-reducing pathways in Shewanella oneidensis. J Biol Chem 295(32):11118–11130
Feng Y, Cronan JE (2011) Complex binding of the FabR repressor of bacterial unsaturated fatty acid biosynthesis to its cognate promoters. Mol Microbiol 80(1):195–218
Yakhnina AA, McManus HR, Bernhardt TG (2015) The cell wall amidase AmiB is essential for P seudomonas aeruginosa cell division, drug resistance and viability. Mol Microbiol 97(5):957–973
Banerjee D et al (2003) Measurement of plasma hydroperoxide concentration by FOX-1 assay in conjunction with triphenylphosphine. Clin Chim Acta 337(1–2):147–152
Le Laz S et al (2014) A biochemical approach to study the role of the terminal oxidases in aerobic respiration in Shewanella oneidensis MR-1. PLoS ONE 9(1):e86343
Kouzuma A, Hashimoto K, Watanabe K (2012) Influences of aerobic respiration on current generation by Shewanella oneidensis MR-1 in single-chamber microbial fuel cells. Biosci Biotechnol Biochem 76(2):270–275
Schönfeld P, Wojtczak L (2008) Fatty acids as modulators of the cellular production of reactive oxygen species. Free Radic Biol Med 45(3):231–241
Yu Q, Sun W, Gao H (2021) Thiosulfate oxidation in sulfur-reducing Shewanella oneidensis and its unexpected influences on the cytochrome c content. Environ Microbiol 23(11):7056–7072
Toporek Y et al (2023) Probing cytoplasmic peroxide metabolism in Shewanella oneidensis. FEMS Microbiol Lett 370:fnad075
Funding
This work was supported by the National Natural Science Foundation of China (no. 32300018), and the Basic Public Welfare Research Program of Zhejiang Province (LQ23C010003 and LQ22C010004).
Author information
Authors and Affiliations
Contributions
P. Q. and Q. M. contributed to the funding used in this work. Q.M. and P.Q. conceived the scope of this study and designed the experiments. Y.X., L.D., and X.G. performed the data collection. P.Q. drafted the first manuscript. P.Q., Q.M., and Y.X. revised, edited, and polished the manuscript. All authors approved the final version for publication and agreed to be accountable for all aspects of the work.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Supplementary file 1 (mp4 44515 KB)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Meng, Q., Xu, Y., Dai, L. et al. Regulation of fadR on the ROS defense mechanism in Shewanalla oneidensis. Biotechnol Lett 46, 691–698 (2024). https://doi.org/10.1007/s10529-024-03487-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-024-03487-y