Abstract
Objectives
The objective of the study was to develop a strategy for the identification of new vitamin B12-producing species and to characterize their production capability using a fast and sensitive LC–MS/MS method developed in this study.
Results
Searching for homologues of the bluB/cobT2 fusion gene known to be responsible for the production of the active vitamin B12 form in P. freudenreichii was shown to be a successful strategy for the identification of new vitamin B12-producing strains. The analysis of the identified strains via LC–MS/MS showed the ability of Terrabacter sp. DSM102553, Yimella lutea DSM19828 and Calidifontibacter indicus DSM22967 to produce the active form of vitamin B12. Further analysis of vitamin B12 production capability of Terrabacter sp. DSM102553 in M9 minimal medium and peptone-based media revealed that the highest yield of 2.65 µg of vitamin B12 per g dry cell weight was obtained in M9 medium.
Conclusions
The proposed strategy enabled identification of Terrabacter sp. DSM102553, whose relatively high yields obtained in the minimal medium open new perspectives for the possible application of the strain for biotechnological vitamin B12 production.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Vegetarian and vegan diets have gained popularity as alternative ways of nutrition in large parts of the world in recent years. However, vegans are at a particular risk of vitamin B12 deficiency since this vital nutrient naturally originates only from foods of animal origin or fortified foods fermented with bacteria (Chamlagain et al. 2017).
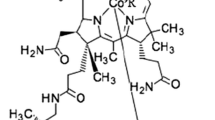
Vitamin B12 (cobalamin) plays a crucial role for the human body since it is used as a cofactor for two important enzymes, methylmalonyl-CoA mutase and methionine synthase (Martens et al. 2002). Methionine synthase is known to be involved in DNA synthesis (Weyden et al. 1973), while methylmalonyl-CoA mutase plays an important role in branched amino acid and odd-chain fatty acid metabolism (Takahashi-Iñiguez et al. 2012). Despite such essential requirement, the human metabolism is not capable of vitamin B12 production, which results in a need for products enriched with vitamin B12. Due to the very complex chemical synthesis of vitamin B12, microbial biosynthesis is applied for its commercial production (Martens et al. 2002). It is known for a long time that several bacteria belonging to different genera are able to synthesize vitamin B12 (Perlman 1959), while other reports have also demonstrated high amounts of vitamin B12 produced by cyanobacteria of the genus Spirulina (Berg et al. 1988). Nevertheless, the commercially available Spirulina supplements mainly contain a corrinoid later identified as pseudovitamin B12 (Watanabe et al. 1999, 2001), a vitamin B12 analogue which cannot be utilized as cofactor by human enzymes. The structures of the vitamin B12 forms active in humans and the pseudovitamin B12 are very similar (Fig. 1).
The lower ligand of the cobalamin molecule is represented by 5,6-dimethylbenzimidazole (DMBI) in the active forms of vitamin B12 or by adenine in pseudovitamin B12, respectively (Watanabe 2007). Due to the biological inactivity of pseudovitamin B12 in humans, determination of the type of the synthesized cobalamin compound during the microbiological production is very important (Chamlagain et al. 2017).
Besides Pseudomonas denitrificans, Propionibacterium freudenreichii (previously designated as P. shermanii) is used for the industrial production of vitamin B12 (Martens et al. 2002) due to the ability to nearly exclusively synthesize the active vitamin B12 (Deptula et al. 2017). The high levels of the produced active vitamin B12 in P. freudenreichii were shown to be the result of the BluB/CobT2 fusion enzyme activity in this strain (Deptula et al. 2015). The study demonstrated that BluB enzyme is responsible for the formation of the lower ligand DMBI, while CobT2 facilitates the activation and the subsequent incorporation of DMBI into the cobalamin molecule.
Based on the data reported by Deptula et al. 2015, we aimed to identify new, previously not described cobalamin-producing species by evaluation of bacterial strains containing a gene encoding the BluB/CobT2 fusion enzyme. In addition, a sensitive and reliable LC–MS/MS method for the identification and quantification of vitamin B12 was used to screen the identified candidates for cobalamin (here termed as the active form of vitamin B12).
Materials and methods
Development of the method for vitamin B 12 analysis
The sample analysis was performed with a triple quadrupole LCMS-8045 (Shimadzu, Germany) and the Lab Solutions Analysis Software (Shimadzu, Germany) was used for data acquisition and analysis. A Luna® Omega 3 µm PS C18 100 Å Column (Phenomenex, Germany) was operated with 0.1% formic acid in water (solvent A) and 0.1% formic acid in acetonitrile (solvent B). All used solvents were LC/MS grade (Carl Roth, Germany). The following LC time program was used: 0–3 min 18%–32% B, 3–3.1 min 32%–95% B, 3.1–4.1 min 95% B, 4.1–4.3 min 95%–18% B, 4.3–7 min 18% B. The flow rate was 0.4 ml min−1 and the column temperature was maintained at 40 °C. The MS analysis was carried out in positive ion mode using electrospray ionization (ESI) under following parameters: nebulizing gas flow 3 l min−1, drying gas flow 10 l min−1, interface temperature 300 °C, desolvation line temperature 250 °C and heat block temperature 400 °C. The mass spectrometer was run in multiple reaction monitoring (MRM) mode, method parameters (collision energies, dwell times and exact m/z values) were optimized for the double-charged cyanocobalamin ion [M + 2H]2+ with 678.40 m/z and for the double-charged pseudocobalamin ion [M + 2H]2+ with 672.75 m/z with the software. The injection volume was 1 µl and the quantification of vitamin B12 in the samples was performed using a calibration curve obtained from a set of cyanocobalamin standards (Merck, Germany). Since pseudovitamin B12 is not commercially available as analytical standard, the cell extract of L. reuteri containing pseudovitamin B12 was used as reference material.
Method validation
The instrumental detection limit (LOD) was determined as the lowest concentration of cyanocobalamin standard corresponding to the first peak that can be integrated and distinguished from zero using the signal-to-noise (S/N) approach (S/N ratio ≥ 3). The limit of quantification (LOQ) was defined as the lowest concentration of cyanocobalamin that can be determined with an acceptable repeatability (relative standard deviation (RSD) between the samples under 10%) using the signal-to-noise (S/N) approach (S/N ratio ≥ 10). The linearity of the method was estimated over the concentration range of 20–2000 nM of CNCbl, a set of 10 concentrations of the standard solution was used, each solution was injected in triplicate. The determination coefficient R2 was calculated to estimate the linearity. To evaluate the selectivity of the method, the chromatograms of vitamin B12-free acetate buffer and E. coli DSM18039 cell extracts were compared with the chromatograms the acetate buffer and E. coli DSM18039 cell extracts spiked with cyanocobalamin standard.
Commercially available vitamin B12 supplements were used for assessing the accuracy of the method. Five different samples containing various amounts of cyanocobalamin were spiked with a known amount of cyanocobalamin standard and the accuracy was estimated through the recovery rates which were calculated as described by Campos-Gimnez et al. 2008:
where Cs is the concentration of vitamin B12 found in the spiked sample, Cn is the concentration of vitamin B12 in the native sample, and Ca is the concentration of cyanocobalamin added in the spiked sample.
The within-day repeatability was evaluated in the range of 100–1000 nM by analysing five different samples, each solution was injected in triplicate, while the intermediate reproducibility (between-day) was determined by analysing a calibration row of five different samples in the range of 100–1000 nM during three consecutive days.
Sequence analysis
The BluB/CobT2 protein sequence [GenBank: CBL56167.1] from P. freudenreichii was used for the identification of bacterial homologues by means of BLAST (National Center for Biotechnology Information). The BLASTp tool was applied for the search in the protein sequence database excluding the genus Propionibacterium (taxid:1743), the maximum number of aligned sequences was set to 1000.
Strains and cultivation conditions
Lactobacillus reuteri DSM20016 used for pseudovitamin B12 purification, E. coli DSM18039 and all strains identified in this work as candidates for vitamin B12 production were obtained from DSMZ (German Collection of Microorganisms and Cell Cultures, Braunschweig, Germany). The cultivation of microorganisms was performed in the media recommended by DSMZ for every specific strain: Terrabacter sp. DSM102553 and Terrabacter sp. DSM102554 in medium 513 (PP), Calidifontibacter indicus DSM22967 and Raineyella antarctica DSM100494 in medium 92, Yimella lutea DSM19828 in medium 65, Blastococcus sp. DSM44272 in medium 714, E. coli DSM18039 in medium 381 and Lactobacillus reuteri DSM20016 in medium 11, respectively. A specific M9 minimal medium (Rüdiger and Schwab 2019) and medium 513 with the doubled amount of all components (2xPP) were also used in the growth experiments with Terrabacter sp. DSM102553. 50 ml cultures were grown in 300 ml shaking flasks at 30 °C under aeration and shaking conditions.
Anaerobic cultivation was applied for the production of pseudovitamin B12 with L. reuteri DSM20016. Briefly, the pre-cultures were inoculated from the glycerol culture stocks (− 80 °C) in 50 mL medium and incubated in septum flasks anaerobically for 24 h at 30 °C.
Cultivation of bacterial strains in BioLector microbioreactor
The pre-cultures of the candidate strains were grown in respective media for 48 h and used for the inoculation of 1 mL medium at the starting OD600 of 0.1. The cultivation was carried out in a BioLector® MB system (m2p-labs, Germany) in MTP-48 FlowerPlates® without pH optodes at 30 °C, 1000 rpm and 95% humidity. The growth of the cultures was monitored online by scattered light signal measurement with an excitation wavelength of 620 nm.
Vitamin B 12 and pseudovitamin B 12 extraction, purification and analysis
For the cobalamin extraction 25 ml of the cultures of the identified candidate strains were harvested by centrifugation at 315×g for 30 min after 5 days of cultivation, while the complete biomass of L. reuteri DSM20016 obtained after 24 h of cultivation was applied for the purification of pseudocobalamin. The cell pellets were resuspended in 10 ml of acetate buffer (4.1 g sodium acetate l−1 adjusted to pH 4.5 with acetic acid) containing 100 µl of 1% KCN. After incubation in a water bath at 98 °C for 30 min the samples were cooled on ice for 30 min and centrifuged again. Vitamin B12 and pseudocobalamin were purified from the obtained supernatants using BAKERBOND spe™ C18 columns JB7020-03 (J. T. Baker) according to the manufacturer’s instructions. The extracts were then syringe sterile filtered (0.2 µm), dried at 60 °C under vacuum, resuspended in 100 µl of deionized H2O and then used directly for the vitamin B12 analysis with the above described LC–MS/MS method.
Results and discussion
LC–MS/MS method development and validation
Since the widely used microbiological assay is not able to differentiate between active forms of vitamin B12 and its analogues (Chamlagain et al. 2015), the first objective of this study was to develop a suitable LC–MS/MS method for vitamin B12 analysis. The parent and fragment ion masses of m/z 678.40, m/z 146.95 and m/z 359.10, which were identified in the spectrum of the active cobalamin have already been demonstrated in previous reports on LC–MS/MS-based methods for vitamin B12 determination (Lu et al. 2008; Luo et al. 2006; Schwertner et al. 2012; Szterk et al. 2012; Lee et al. 2015; Zironi et al. 2014). The demonstrated LOQ was comparable with that of the method described by Yi et al. 2012 which was also based on the tandem triple quadrupole mass spectrometry in MRM mode. In comparison to previously described LC–MS/MS methods by Schwertner et al. 2012, Lee et al. 2015, Szterk et al. 2012, the method reported in this study provides a faster analytical procedure and can, therefore, enable a higher sample throughput. Moreover, only few micrograms of cell material were sufficient for the successful B12 analysis with the developed method, while earlier reported LCMS/MS-based methods applied 15–30 g of sample material for vitamin B12 purification for subsequent analysis (Szterk et al. 2012; Luo et al. 2006).
The method was linear within the tested range over 10 calibration levels with R2 = 0.9997. The selectivity of the method was evaluated by comparing the chromatograms of the buffer and vitamin B12-free E. coli MG1655 cell matrix blanks with the same samples spiked with cyanocobalamin standard. The comparison of the blank samples with the spiked samples showed a high selectivity of the method, since no peaks with the fragmentation pattern and the retention time corresponding to cyanocobalamin were detected in the controls whereas all spiked samples demonstrated cobalamin peaks. Although an unidentified peak was observed in the chromatogram of the cell extract of E. coli DSM18039 (Supplementary Fig. 1), the fragment signals with m/z 456.75 and m/z 359.10 characteristic for cyanocobalamin were not detected. This allows to exclude that the observed peak corresponds to CNCbl and is rather evidence for a substance with the same precursor ion mass as cyanocobalamin but another chemical structure.
The accuracy of the method was determined through recovery by spiking of five different commercially available vitamin B12 supplements with known amounts of CNCbl standard. The average recovery was estimated in technical triplicates by determination of the peak areas of the samples before and after spiking. The method demonstrated high accuracy, since the average recovery from spiked samples was between 94.4% and 103.4% (Supplementary Table 1), which was comparable with the previously reported LC–MS/MS-based methods for vitamin B12 determination (Lu et al. 2008; Luo et al. 2006).
The within-day repeatability was evaluated in the range of 100–1000 nM by analyzing five different samples, measured as technical triplicates. The relative standard deviation in the repeatability experiment was less than 4%. The intermediate precision (between-day or day-by-day) was determined by analyzing calibration series of five different samples in the range of 100–1000 nM during three consecutive days. The RSD in the intermediate reproducibility was below 4%.
Identification of strains containing a bluB/cobT2 fusion gene
Activity of the fusion enzyme BluB/CobT2, which produces and directly activates DMBI, is the main mechanism responsible for the high levels of active vitamin B12 in P. freudenreichii cells, which has been shown in an earlier study (Deptula et al. 2015). Furthermore, the metabolite channeling feature of fusion enzymes can contribute to high flux through a biosynthetic pathway by protection of reactive intermediates and provision of high local substrate concentration (Kummer et al. 2021). Therefore, we aimed at the investigation of microorganisms with a bluB/cobT2 fusion gene for their vitamin B12 production capabilities.
The results of a respective BLAST search were analyzed and the sequences showing an identity of more than 50% to the BluB/CobT sequence from P. freudenreichii were chosen. From the resulting list of 338 sequences identified at the time of the investigation, we randomly selected six non-pathogenic organisms (risk group 1) for further analysis (Supplementary Table 2). The identified candidate strains C. indicus DSM22967, R. antarctica DSM100494, Y. lutea DSM19828, Blastococcus sp. DSM4427 and also Terrabacter sp. DSM102553 and Terrabacter sp. DSM102554 for the selected Terrabacter sp. were obtained from DSMZ.
Determination of vitamin B 12 and pseudovitamin B 12 synthesis by the selected strains
To investigate the ability of the identified strains to produce cobalamins, the cell extracts were analyzed with the developed LC–MS/MS method and compared with cyanocobalamin and pseudocobalamin reference compounds. Since various microorganisms can produce different B12 analogues, where DMBI is replaced by other benzimidazoles, purines or phenolic compounds (Crofts et al. 2013), we aimed to identify only those producing the active vitamin B12. As the active and inactive forms differ in their respective ligands (Fig. 1), their fragmentation patterns are characteristic and can be used do differentiate between the two forms. The spectrum of the cyanocobalamin reference compound demonstrated a parent ion with m/z 678.40 and fragment signals with m/z 146.95 [DMBI + H]+ and m/z 359.10 [DMBI + sugar + PO3 + H]+, while a parent ion with m/z 672.75 and fragment signals with m/z 136.05 [adenine + H]+ and m/z 348.05 [adenine + sugar + PO3 + H]+ were detected in the spectrum of the pseudovitamin B12 (Fig. 2). These results correlate with the data reported in previous studies (Chamlagain et al. 2017; Bernhardt et al. 2019).
Identification of the active and inactive cobalamin forms in the cell extracts of the selected producing strains. Shown are TICs (total ion currents) and traces corresponding to the fragment ions originating from the identified parent mass obtained from the cyanocobalamin reference compound, cell extracts of the selected strains and of the pseudocobalamin produced by L. reuteri
Among the selected strains, cobalamins were detected in the extracts of Y. lutea, C. indicus and Terrabacter sp. DSM102553 (Fig. 2), while LC–MS/MS analysis of Terrabacter sp. DSM102554, R. antarctica DSM100494 and Blastococcus sp. DSM44272 revealed no production of the active cobalamin (Supplementary Fig. 2). Nevertheless, further investigations can be performed to analyze, whether the mentioned microorganisms can produce vitamin B12 under certain other conditions or in other cultivation media. For example, supplementation with FMNH2 necessary for the formation of DMBI (Deptula et al. 2015) might facilitate synthesis of the vitamin B12. The lack of cobalamin production in Terrabacter sp. DSM102554, R. antarctica DSM100494 and Blastococcus sp. DSM44272 might be due to the presence of an incomplete biosynthetic way or to transcriptional repression of vitamin B12 synthesis gene expression. In future research, these strains can be examined for the presence of the other genes of the vitamin B12 synthesis pathway and the expression levels of the corresponding genes can be analyzed.
In the case of the three producing strains, the active form was prevalent in their cell extracts, since the characteristic retention time and fragmentation pattern observed for the cyanocobalamin standard were also detected in the cell extracts of these examined candidates. Moreover, the retention time and fragmentation pattern of the second minor peak identified in the chromatogram of Y. lutea demonstrated production of low amounts of pseudovitamin B12. Additionally, an unidentified peak was detected in the chromatogram of C. indicus, while no other compounds except the active vitamin B12 were detected for Terrabacter sp. DSM102553.
Growth comparison of vitamin B 12 -producing strains in complex media
To get a first impression of the growth behavior of the vitamin B12-producing strains, their growth curves in respective complex media recommended by DSMZ were determined. As Fig. 3 shows, remarkable differences in the growth behaviour of the investigated strains could be observed. The cultures of C. indicus demonstrated a lag-phase of approx. 20 h, which was followed by the exponential growth phase after which a linear biomass accumulation continued. The growth of Y. lutea under the tested conditions was characterized by a prolonged lag-phase if compared to C. indicus and Terrabacter sp. DSM102553 and the maximum scattered light signal of approx. 25 a. u. after 80 h. Among three tested strains, Terrabacter sp. DSM102553 demonstrated the highest maximum scattered light signal of 42 a. u. already after 25 h of cultivation.
Comparison of Y. lutea, C. indicus and Terrabacter sp. DSM102553 growth curves. Respective complex media recommended by DSMZ were used (medium 65 for Y. lutea, medium 92 for C. indicus, medium 513 for Terrabacter sp. DSM102553). The data points represent the mean values and standard deviations of three biological replicates
Since the same media were applied for the precultures and main cultivation experiment, which was started at the same OD600 for all strains, these differences cannot be explained by the adaptation of the cells to growth in new cultivation medium but are rather due to characteristic features of each specific strain. Therefore, the above demonstrated ability to accumulate higher biomass during a shorter time period indicates that Terrabacter sp. DSM102553 is the most appropriate strain for vitamin B12 production among the identified candidates under the tested conditions.
Moreover, since no pseudovitamin B12 or unidentified peaks were detected in the chromatograms of Terrabacter sp. DSM102553 extracts (Fig. 2), the strain was selected for further investigation.
Production of active vitamin B 12 with Terrabacter sp. DSM102553 in different media
In order to improve cell density and thereby volumetric vitamin B12 productivity of Terrabacter sp. DSM102553, we tested two-fold concentrated PP medium. However, as peptones are one of the most expensive components of microbial media, we aimed to find a suitable cheap minimal medium. For this purpose, precultures grown in the standard PP medium were inoculated in PP, 2xPP and M9 medium and the growth of Terrabacter sp. DSM102553 was compared. As shown in Fig. 4a, the maximal cell density in 2xPP and M9 medium was about 1.5 times higher in comparison to PP. Although a prolonged lag phase was observed in M9 medium, the maximum cell density in M9 medium was comparable with that reached in 2xPP. Our further investigations have also shown that biotin and thiamin have no influence on the growth of the strain (Supplementary Fig. 3), which is why they were excluded from the medium composition in the vitamin B12 quantification experiments.
Growth and vitamin B12 production capability of Terrabacter sp. DSM102553 in different media A Growth curves of Terrabacter sp. DSM102553 in PP, 2xPP and M9 medium. B Vitamin B12 content in cells cultured in PP, 2xPP and M9 medium. The data points represent the mean values and standard deviations of three biological replicates
The analysis of vitamin B12 production was performed in PP, 2xPP and M9 medium after 100 h of cultivation as higher amounts of produced cobalamin at later growth stages have already been reported for P. freudenreichii (Deptula et al. 2017). Among the three media, the highest vitamin B12 production level was observed in M9 medium (Fig. 4b). The amounts of 2.65 µg of vitamin B12 per g dry cell weight (DCW) obtained with this medium were two-fold and five-fold higher than those produced in PP or 2xPP medium, respectively. In comparison to PP medium, faster growth and higher cell density, but lower vitamin B12 concentrations were observed in 2xPP medium, which shows that under these conditions the production of vitamin B12 was not linked to biomass formation in Terrabacter sp. DSM102553.
By using the presence of bluB/cobT2 fusion genes as a potential indicator for efficient production of active vitamin B12, we were able to identify strains which produce vitamin B12 under aerobic conditions. Since oxygen is necessary for DMBI formation and active vitamin B12 synthesis in P. freudenreichii (Deptula et al. 2015), a two-step cultivation including aerobic and anaerobic stages is applied for vitamin B12 production with these microaerophilic strains (Chamlagain et al. 2015, 2017; Deptula et al. 2017). The omission of the anaerobic cultivation step can be an attractive advantage since it makes the synthesis procedure more easy-to-handle and time-effective. So far, the vitamin B12 contents reported in this manuscript cannot compete with the values of approx. 6–15 µg per g wet cell mass reported for P. freudenreichii (Deptula et al. 2017) in a comparable cultivation in shaking flasks, but the achieved levels obtained with minimal medium are promising. Future optimization steps can involve further medium optimization, for example via supplementation of DMBI (Chamlagain et al. 2016) or cobalt (Mohammed et al. 2014) as well as identification of mutants with higher productivity.
Conclusions
Our strategy for the identification of new vitamin B12 producing microorganisms resulted in the description of bacteria with the ability to produce active vitamin B12 under aerobic conditions. Cobalamin analysis was performed with a sensitive optimized LC–MS/MS method with a wide concentration range developed in this study which not only allows vitamin B12 quantification, but also enables clear discrimination between its active and inactive form and simplifies selective screening processes for potential producers. The identified ability of Terrabacter sp. DSM102553 to synthesize active cobalamin in the minimal medium demonstrated in our work opens new opportunities for further optimization of the newly identified strain and its possible application for biotechnological vitamin B12 synthesis in a fairly simple aerobic manner.
Data availability
The data generated during the is available in the manuscript and in the Supplementary Information.
References
Bernhardt C, Zhu X, Schütz D, Fischer M, Bisping B (2019) Cobalamin is produced by Acetobacter pasteurianus DSM 3509. Appl Microbiol Biotechnol 103:3875–3885. https://doi.org/10.1007/s00253-019-09704-3
Campos-Gimnez E, Fontannaz P, Trisconi M-J, Kilinc T, Gimenez C, Andrieux P (2008) Determination of Vitamin B12 in food products by liquid chromatography/UV detection with immunoaffinity extraction: single-laboratory validation. J AOAC Int 91:786–793. https://doi.org/10.1093/jaoac/91.4.786
Chamlagain B, Edelmann M, Kariuloto S, Ollilainen V, Piironen V (2015) Ultra-high performance liquid chromatographic and mass spectrometric analysis of active vitamin B12 in cells of Propionibacterium and fermented cereal matrices. Food Chem 166:630–638. https://doi.org/10.1016/j.foodchem.2014.06.068
Chamlagain B, Deptula P, Edelmann M, Kariluoto S, Grattepanche F, Lacroix C, Varmanen P, Piironen V (2016) Effect of the lower ligand precursors on vitamin B12 production by food-grade Propionibacteria. LWT—Food Sci Technol 72:117–124. https://doi.org/10.1016/j.lwt.2016.04.023
Chamlagain B, Sugito T, Deptula P, Edelmann M, Kariuloto S, Varmanen P, Piironen V (2017) In situ production of active vitamin B12 in cereal matrices using Propionibacterium freudenreichii. Food Sci Nutr 6:67–76. https://doi.org/10.1002/fsn3.528
Crofts T, Seth E, Hazra A, Taga M (2013) Cobamide structure depends on both lower ligand availability and CobT substrate specificity. Chem Biol 20:1265–1274. https://doi.org/10.1016/j.chembiol.2013.08.006
Deptula P, Kylli P, Chamlagain B, Holm L, Kostiainen R, Piironen V, Savijoki K, Varmanen P (2015) BluB/CobT2 fusion enzyme activity reveals mechanisms responsible for production of active form of vitamin B12 by Propionibacterium freudenreichii. Microb Cell Fact. https://doi.org/10.1186/s12934-015-0363-9
Deptula P, Chamlagain B, Edelmann M, Sangsuwan P, Nyman T, Savijoki K, Piironen V, Varmanen P (2017) Food-Like Growth Conditions Support Production of Active Vitamin B12 by Propionibacterium freudenreichii 2067 without DMBI, the Lower Ligand Base, or Cobalt Supplementation. Front Microbiol 8:368. https://doi.org/10.3389/fmicb.2017.00368
Kummer MJ, Lee YS, Yuan M, Alkotaini B, Zhao J, Blumenthal E, Minteer SD (2021) Substrate channeling by a rationally designed fusion protein in a biocatalytic cascade. JACS Au 1:1187–1197. https://doi.org/10.1021/jacsau.1c00180
Lee J, Shin J, Park J, Kim H, Ahn J, Kwak B, Kim J (2015) Analytical determination of Vitamin B12 content in infant and toddler milk formulas by liquid chromatography tandem mass spectrometry (LC-MS/MS). Korean J Food Sci Anim Resour 35:765–771. https://doi.org/10.5851/kosfa.2015.35.6.765
Lu B, Ren Y, Huang B, Liao W, Cai Z, Tie X (2008) Simultaneous determination of four water-soluble vitamins in fortified infant foods by ultra-performance liquid chromatography coupled with triple quadrupole mass spectrometry. J Chromatogr Sci 46:225–232. https://doi.org/10.1093/chromsci/46.3.225
Luo X, Chen B, Ding L, Tang F, Yao S (2006) HPLC-ESI-MS analysis of Vitamin B12 in food products and in multivitamins-multimineral tablets. Anal Chim Acta 562:185–189. https://doi.org/10.1016/j.aca.2006.01.073
Martens J, Barg H, Warren M, Jahn D (2002) Microbial production of vitamin B12. Appl Microbiol Biotechnol 58:275–285. https://doi.org/10.1007/s00253-001-0902-7
Mohammed Y, Lee B, Kang Z, Du G (2014) Development of a two-step cultivation strategy for the production of vitamin B12 by Bacillus megaterium. Microb Cell Fact 13:102. https://doi.org/10.1186/s12934-014-0102-7
Perlman D (1959) Microbial Synthesis of Cobamides 1:87–122. doi: https://doi.org/10.1016/s0065-2164(08)70476-3
Rüdiger J, Schwab W (2019) Improving an Escherichia coli-based biocatalyst for terpenol glycosylation by variation of the expression system. J Ind Microbiol Biotechnol 46:1129–1138. https://doi.org/10.1007/s10295-019-02184-4
Schwertner H, Valtier S, Bebarta V (2012) Liquid chromatographic mass spectrometric (LC/MS/MS) determination of plasma hydroxocobalamin and cyanocobalamin concentrations after hydroxocobalamin antidote treatment for cyanide poisoning. J Chromatogr B Analyt Technol Biomed Life Sci 905:10–16. https://doi.org/10.1016/j.jchromb.2012.07.012
Szterk A, Roszko M, Małek K, Czerwonka M, Waszkiewicz-Robak B (2012) Application of the SPE reversed phase HPLC/MS technique to determine vitamin B12 bio-active forms in beef. Meat Sci 91:408–413. https://doi.org/10.1016/j.meatsci.2012.02.023
Takahashi-Iñiguez T, García-Hernandez E, Arreguín-Espinosa R, Flores ME (2012) Role of vitamin B12 on methylmalonyl-CoA mutase activity. J Zhejiang Univ Sci B 13:423–437. https://doi.org/10.1631/jzus.B1100329
van der Berg H, Daqnelie P, van Staweren W (1988) Vitamin B12 and seaweed. Lancet 1:242–243. https://doi.org/10.1016/s0140-6736(88)91093-8
Watanabe F (2007) Vitamin B12 sources and bioavailability. Exp Biol Med (maywood) 232:1266–1274. https://doi.org/10.3181/0703-MR-67
Watanabe F, Katsura H, Takenaka S, Fujita T, Abe K, Tamura Y, Nakatsuka T, Nakano Y (1999) Pseudovitamin B(12) is the predominant cobamide of an algal health food, spirulina tablets. J Agric Food Chem 47:4736–4741. https://doi.org/10.1021/jf990541b
Watanabe F, Miyamoto E, Nakano Y (2001) Inactive corrinoid-compound significantly decreases in Spirulina platensis grown in a cobalt-deficient medium. J Agric Food Chem 49:5685–5688. https://doi.org/10.1021/jf010733i
Weyden MVD, Cooper M, Firkin B (1973) Defective DNA synthesis in human megaloblastic bone marrow: effects of hydroxy-B12 5′-deoxyadenosyl-B12 and methyl-B12. Blood 41:299–308. https://doi.org/10.1182/blood.V41.2.299.299
Yi S, Seth EC, Men Y-J, Stabler SP, Allen RH, Alvarez-Cohen L, Taga ME (2012) Versatility in corrinoid salvaging and remodeling pathways supports corrinoid-dependent metabolism in Dehalococcoides mccartyi. Appl Environ Microbiol 78:7745–7752. https://doi.org/10.1128/AEM.02150-12
Zironi E, Gazzotti T, Barbarossa A, Farabegoli F, Serraino A, Pagliuca G (2014) Determination of Vitamin B12 in dairy products by ultra performance liquid chromatography-tandem mass spectrometry. Ital J Food Saf 3:4513. https://doi.org/10.4081/ijfs.2014.4513
Supporting information
Supplementary Fig. 1—Selectivity of the method to cyanocobalamin in the presence of the acetate buffer and cell matrix. Figure shows the chromatograms corresponding to (A) buffer blank, (B) buffer spiked with CNCbl standard, (C) cell matrix blank obtained from the non producing strain E. coli DSM18039 and (D) cell matrix spiked with CNCbl standard.
Supplementary Fig. 2—LC-MS/MS chromatograms of the cyanocobalamin reference compound and the cell extracts of R. antarctica DSM100494, Blastococcus sp. DSM44272 and Terrabacter sp. DSM102554. Shown are representative LC-MS/MS chromatograms of the cyanocobalamin reference compound and of the cell extracts of the selected strains.
Supplementary Fig. 3—Growth curves of Terrabacter sp. DSM102553 in M9 medium with lowered concentrations of biotin and thiamin or without biotin and thiamin supplementation. The data points represent the mean values and standard deviations of three biological replicates.
Funding
Open Access funding enabled and organized by Projekt DEAL. This study was supported by the LOEWE project AROMAplus of the State of Hessen (Germany).
Author information
Authors and Affiliations
Contributions
DD carried out literature review, performed the experiments, analyzed the data and wrote the first draft of the manuscript, DH and SM supervised the work and revised the manuscript, MB performed BLAST analysis of genome data, supervised the work and revised the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no relevant financial or non-financial interests to disclose.
Ethical approval
This article does not contain any studies with human participants or animals.
Consent to participations
Not applicable.
Consent for publications
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Dudko, D., Milker, S., Holtmann, D. et al. Identification of vitamin B12 producing bacteria based on the presence of bluB/cobT2 homologues. Biotechnol Lett 45, 563–572 (2023). https://doi.org/10.1007/s10529-023-03362-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-023-03362-2