Abstract
Objective
To develop an endo-β-1,4-xylanase with high specificity for production of prebiotic xylooligosaccharides that optimally works at moderate temperature desirable to reduce the energy cost in the production process.
Results
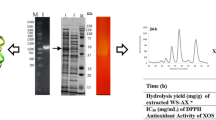
The xylB gene, encoding for a glycosyl hydrolase family 11 xylanase from a thermoresistant fungus, Aspergillus niger BCC14405 was expressed in a methylotrophic yeast P. pastoris KM71 in a secreted form. The recombinant XylB showed a high specific activity of 3852 and 169 U mg−1 protein on beechwood xylan and arabinoxylan, respectively with no detectable side activities against different forms of cellulose (Avicel Ò PH101 microcrystalline cellulose, phosphoric acid swollen cellulose and carboxymethylcellulose). The enzyme worked optimally at 45 °C, pH 6.0. It showed a specific cleavage pattern by releasing xylobiose (X2) as the major product from xylooligosaccharides (X3 to X6) substrates. The highest XOS yield of 708 mg g−1 substrate comprising X2, X3 and X6 was obtained from beechwood xylan hydrolysis.
Conclusion
The enzyme is potent for XOS production and for saccharification of lignocellulosic biomass.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Xylan is one of the major hemicellulosic polysaccharides in plant cell wall. It exists as a branched heteropolysaccharide containing a β-(1,4)-D-xylose backbone with substituting groups comprising different pentose and hexose sugars and sugar acids (Beg et al. 2001). Xylans can be broadly classified as homoxylans, arabinoxylans, glucuronoxylans and arabinoglucuronoxylans according to their main sugar unit (Polizeli et al. 2005). Biodegradation of xylan requires a group of xylanolytic enzymes with different specificities working co-operatively. Among them, endoxylanase (EC 3.2.1.8) is the most important enzyme that cleaves the internal β-1,4 bonds in the xylan main chain into xylooligosaccharides of different lengths (Biely 1985).
Microbial xylanases represent a major group of industrial enzymes applicable to degrade plant derived xylan with potent applications in saccharification of lignocellulosic substrates to xylose and to a promising prebiotic xylooligosaccharides (XOS) (Dodd and Cann 2009). In addition, they can be used in animal feed (Abelilla and Stein 2019), pulp and paper (Walia et al. 2017), textile (Singh et al. 2020) and food industries (Adiguzel et al. 2019). XOS are oligosaccharides of xylose residues that are linked by β-(1,4)-glycosidic bonds. The degree of polymerization (DP) of XOS can vary from 2 to 6 xylose units and their chemical side chains and substitution pattern can vary depending on xylan sources and the production process (Moure et al. 2006; Poletto et al. 2020; Samanta et al. 2015).
A number of xylanases from various bacteria, yeast and fungi have been isolated and characterized (Alokika and Singh 2019; Ding et al. 2018; Liu et al. 2018). Filamentous fungi are widely used as sources of xylanases due to their higher enzyme production levels in comparison with bacteria and yeasts (Ahmed et al. 2009). However, xylanases are produced and secreted together with other hydrolytic enzymes, making it difficult to obtain an endoxylanase with no other enzymes contributing to unwanted side activities. Heterologous expression of fungal endoxylanases with varying biochemical properties and specificities from a number of fungal origins, for example, in genera Aspergillus, Trichoderma, and Penicillium have been reported in various and heterologous hosts (Jun et al. 2009; Asano et al. 2005; Sriprang et al. 2006; Zhuo et al. 2018). Searching for new xylanases with high specificity for XOS production at moderate temperature in a suitable range to inhibit microbial growth while allowing high solubility of substrates is of great interest with an advantage to save the energy cost in production process (Choudhary et al. 2016).
In this study, we further explore the application of the recombinant endoxylanase originated from a thermoresistant fungus A. niger BCC14405 (Asano et al. 2005) and previously expressed in Pichia pastoris (Ruanglek et al. 2007). The recombinant xylanase XylB was further characterized for its biochemical properties with the focus on its specificities on production of XOS. The enzyme showed high endoxylanase activity with high specificity towards production of xylobiose to xylohexaose (X2–X6) preferred for prebiotic application (Aachary and Prapulla 2011; Samanta et al. 2015) while showed no side activities on cellulose. The enzyme is considered potent for further application in production of xylooligosaccharides and saccharification of hemicellulose fraction in lignocellulosic biomass.
Materials and methods
Materials, strains, plasmids and chemicals
pPICZαA (Invitrogen, Carlsbad, CA) was used as the expression vector. Escherichia coli DH5α (Invitrogen, Carlsbad, CA) was used as a host strain for DNA cloning. P. pastoris KM71 (Invitrogen, Carlsbad, CA) was used for recombinant protein expression. Beechwood xylan and carboxymethyl cellulose (CMC) were purchased from Sigma-Aldrich (St. Louis, MO). Wheat arabinoxylan and xylooligosaccharides (X2-X6) were purchased from Megazyme (Wicklow, Ireland). Avicel PH101 was purchased from Fluka (Milwaukee, WI) and used for preparing phosphoric acid swollen cellulose (PASC) (Zhang et al. 2006). All chemicals and reagents were analytical grade and obtained from major chemical suppliers (Sigma, Merck and Fluka).
Construction of recombinant plasmid
The xylB gene was amplified from pET28/xylB plasmid (Sriprang et al. 2006) using primers XylBF/EcoRI (5′-GCGAATTCTCGACCCCGAGCTCGACC-3′) and XylBR/XbaI (5′-GCTCT AGAGCCTGAACAGTAATGGAGGA-3′). Restriction sites for cloning are underlined. The gene was cloned into pPICZαA expression vector and transformed into E. coli DH5α. The transformed clones were selected on LB agar containing 25 µg mL−1 Zeocin. The plasmid harboring the inserted gene was extracted from the selected clone and then confirmed by sequencing using AOX forward and AOX reverse primers. The recombinant plasmid pPICZαA-xylB was linearized with PmeI and transformed into P. pastoris KM71 using electroporation and then selected on YPD agar with 100 µg mL−1 Zeocin. The transformed clone was confirmed by colony PCR using both AOX primers and gene-specific primers.
Protein expression
Expression of the recombinant enzyme from P. pastoris KM71 was performed according to the method described in the supplier’s protocol (MAN0000035). P. pastoris KM71 containing xylB gene was cultured overnight in 5 mL YPD broth at 30 °C and used as a starter. For protein expression, 0.2% (v/v) starter was inoculated into 500 mL BMGY medium in a 2-L flask and then cultured at 30 °C with 200 rpm shaking for 16 h (OD600 = 5–8). The cells were collected by centrifugation at 5000×g for 5 min. The cell pellet was then resuspended in 100 mL BMMY medium and transferred to a 1-L flask. The culture was induced in BMMY medium at 30 °C with 200 rpm shaking. Methanol was added to 0.3% final concentration at 24 h intervals until 72 h. The crude enzyme was harvested by centrifugation at 5000×g at 4 °C for 10 min. The crude enzyme preparation was stored at 4 °C for further experiments. The protein concentration was determined by Bradford assay using the Bio-Rad Protein assay solution (Bio-Rad, Hercules, CA).
Enzyme activity assay
The enzyme activities were determined by measuring the released reducing sugar based on dinitrosalicylic acid (DNS) assay (Miller 1959) using xylose as a standard. The standard reaction contained crude enzyme XylB at an appropriate concentration and 1% (w/v) of the substrate selected from beechwood xylan dissolved in 100 mM sodium phosphate buffer, pH 6.0. The reactions were incubated at 45 °C for 10 min or otherwise specified. After incubation, the reactions were stopped by addition of twofold volumes of DNS and boiled for 10 min. The released reducing sugar was determined by measuring the absorbance at 540 nm. The absorbance data were used to calculate the enzyme activities in which 1 unit of enzyme was defined as the amount of enzyme that produces 1 µ mole of product from substrate per 1 min under the specified condition.
Biochemical characterization
In order to determine the effect of temperature on enzyme activity, the XylB was incubated with 1% (w/v) beechwood xylan dissolved in 100 mM sodium phosphate buffer (pH 6.0) for 10 min. The temperatures of reaction were varied from 30 to 80 °C to determine the optimal temperature. To investigate the optimal pH, XylB was incubated with 1% (w/v) beechwood xylan dissolved in 100 mM buffers with varying pH range, including sodium acetate buffer (pH 4.0–5.5), sodium phosphate buffer (pH 6.0–8.0) and Tris–HCl buffer (pH 8.0–9.0) at 45 °C for 10 min.
To examine thermostability, XylB was pre-incubated in 100 mM sodium phosphate buffer (pH 6.0) at 40–60 °C for 0, 15, 30, 45 and 60 min. The incubated XylB was then added to the reaction containing 1% (w/v) of beechwood xylan dissolved in 100 mM sodium phosphate buffer (pH 6.0) for 10 min. For substrate specificity, various substrates including beechwood xylan, Avicel: PH101 microcrystalline cellulose, carboxymethyl cellulose (CMC), wheat arabinoxylan, phosphoric acid swollen cellulose (PASC) were used to examine the substrate specificity of XylB. Enzyme kinetics of XylB was studied using 0–10 mg mL−1 xylan as the substrate based on Michaelis–Menten equation with 0–120 min of incubation.
Effect of metal ions and chemical surfactants
To investigate the effect of metal ions and chemical surfactants, various ions and surfactants were added to the enzyme reactions containing 1% (w/v) beechwood xylan dissolved in 100 mM sodium phosphate buffer (pH 6.0) with the final concentration of 1 mM for metal ions or 0.1% (v/v) for surfactant and chemicals at 45 °C for 10 min. The metal ions and surfactants used in this study included Na+ (NaCl), K+ (KCl), Zn2+ (ZnCl2), Co2+ (CoCl2 · 6H2O), Mn2+ (MnCl2), Li+ (LiCl), Mg2+ (MgCl2·6H2O), Cu2+ (CuCl2·2H2O), Ni2+ (NiCl2 · 6H2O), Ca2+ (CaCl2·2H2O), Tween 20 (% v/v), Triton X-100 (% v/v), β-mercaptoethanol (% v/v), urea (% w/v) and sodium dodecyl sulphate (SDS) (% w/v). Differences in means among the treatments were identified using independent-samples T-test (p < 0.05) using SPSS 16.0.
Cleavage pattern analysis
The product profile of XylB on hydrolysis of xylooligosaccharides (X2-X6) was studied. The reaction mixture containing 1 mg mL−1 of xylooligosaccharides dissolved in 100 mM sodium phosphate buffer (pH 6.0) and 100 μg of XylB was incubated in Thermomixer C (Eppendorf, Hamburg, Germany) at 45 °C with shaking at 1000 rpm for 24 h. The sugar products released into the supernatant were analyzed using Dionex3000 high performance liquid chromatography (Thermo Fisher Scientific, Waltham, MA) equipped with Aminex HPX-87H column and refractive index detector (Shodex, Kyoto, Japan) using 5 mM sulfuric acid as a mobile phase at 65 °C with a flow rate 0.5 mL min−1.
Production of xylooligosaccharides (XOS) from beechwood xylan
Enzymatic hydrolysis was studied in 1 mL reaction containing 1% (w/v) of beechwood xylan dissolved in 100 mM sodium phosphate (pH 6.0) with an enzyme dosage of 2.5, 5 and 10 mg g−1 substrate. The reaction was incubated at 45 °C with shaking at 200 rpm for 24 h. Profiles of xylooligosaccharides were analyzed using HPLC with the condition described above. The experiments were carried out in triplicates.
Results
Heterologous expression of xylB in P. pastoris KM71
The xylB gene (546 bp) encoding for xylanase (GH11) was previously isolated from A. niger BCC14405. The mature gene was expressed as an intracellular soluble protein with the approximate molecular weight of 21 kDa in E. coli BL21 (Sriprang et al. 2006). However, the recombinant XylB expressed in E. coli required further downstream processes including cell disruption and purification step. Therefore, in this study, the xylB gene was cloned and expressed as a secreted protein in P. pastoris KM71 with a molecular weight of approximately 25 kDa after induction at 24–72 h (Fig. 1). The enzyme was expressed as a single major band on SDS-PAGE with > 95% homogeneity. No activity of xylanase was detected in P. pastoris KM71 strain carrying an empty vector (pPICZαA without XylB gene) used as the negative control. The molecular weight of the protein obtained is larger than the prediction (21 kDa). This could be due to post-translational glycosylation (Thak et al. 2020). The yield of enzyme production obtained in shake flask scale was 116 mg protein L−1 of culture medium. Due to its remarkably high homogeneity, XylB was further characterized with no additional purification step.
Biochemical properties
The enzyme activity of XylB was tested against beechwood xylan under various conditions in order to investigate the effect of pH and temperature. For the optimal pH, enzyme activity was determined under various pH ranging from pH 4.0 to 9.0. XylB exhibited the highest xylanase activity in 100 mM sodium phosphate pH 6.0. Remarkably, more than 80% remaining activity was observed at pH 5.0–6.0 (Fig. 2a). For the optimal temperature of XylB, enzyme activity was determined using beechwood xylan as a substrate by comparing the activity at various temperatures ranging from 30 to 80 °C. According to the results, XylB exhibited the highest activity at 45 °C with more than 60% of its maximal activity at a temperature between 40 and 50 °C (Fig. 2b). For thermostability, more than 80% and 40% remaining activity were observed after the enzyme was incubated without substrate for 15–60 min at 40 °C and 50 °C, respectively. However, the activity substantially decreased after incubation at 60 °C for 15 min and was completely lost after incubation for 45 min (Fig. 2c).
Effects of pH and temperature on XylB activity. a Effect of pH was analyzed in the reaction containing 1% (w/v) of beechwood xylan in 100 mM sodium acetate buffer (pH 4.0–5.5), 100 mM sodium phosphate buffer (pH 6.0–8.0) and 100 mM Tris–HCl buffer (pH 8.0–9.0) at 45 °C. b Effect of temperature was investigated in the reaction containing 1% (w/v) of beechwood xylan in100 mM sodium phosphate buffer pH 6.0 at 30–80 °C. c Thermostability was investigated by incubated the enzyme in 100 mM sodium phosphate buffer pH 6.0 at 40, 50 and 60 °C for 15–60 min
The effect of metal ions (Ca2+, Co2+, Cu2+, K+, Li+ Mg2+, Mn2+, Na+, Ni2+ and Zn2+), surfactants and chemicals (β-mercaptoethanol, SDS, urea, Triton X-100 and Tween 20) on XylB activity were evaluated. According to the results, Cu2+, Mg2+, Ni2+, Na+, Ca2+, Zn2+, K+, Li+ enhanced XylB activity with 8%-28.4% increase, whereas XylB activity drastically decrease with 47% loss of activity in the reaction contain Mn2+ (Table 1). Moreover, Mn2+, β-mercaptoethanol, urea and SDS led to 50% -90% reduction in enzyme activity in the respective conditions. Triton X-100 and Tween 20 did not influence the activity of the enzyme.
Substrate specificity on different polysaccharides
In order to characterize the specificity of XylB, the enzyme activity was tested against various hemicellulosic (beechwood xylan and wheat arabinoxylan) and cellulosic substrates (carboxymethyl cellulose (CMC), phosphoric acid swollen cellulose (PASC) and Avicel). The highest specific activity was detected against beechwood xylan (3852 U mg−1 protein) with a weak activity against wheat arabinoxylan (169 U mg−1 protein) (Table 2). In addition, XylB did not degrade any type of cellulose polysaccharides used in this experiment at a detectable level. The kinetic study of XylB was then determined using beechwood xylan and arabinoxylan as the substrates.
Product profile from xylooligosaccharide and beechwood xylan hydrolysis
In order to investigate the cleavage pattern of recombinant XylB, the hydrolysis products obtained from xylooligosaccharides (X2-X6) hydrolysis were analyzed. Based on the results, X3-X6 were mainly hydrolyzed into xylose (X1), xylobiose (X2) and xylotriose (X3) by XylB (Fig. 3). However, XylB was unable to cleave xylobiose under the experimental conditions. The hydrolysis of xylan from beechwood was then performed. According to the results, xylooligosaccharides released from beechwood xylan hydrolysis comprised xylobiose (X2), xylotriose (X3) and xylohexaose (X6) with the highest XOS yield of 708 mg g−1 substrate (equivalent to 70% conversion yield) using an enzyme dosage of 10 mg g−1 substrate (Fig. 4).
Product profile obtained from hydrolysis of xylooligosaccharides. The reactions contained 1 mg mL−1 of xylooligosaccharides (X2-X6) dissolved in 100 mM sodium phosphate buffer pH 6.0 and 100 µg mg−1 substrate of XylB. The reactions were incubated at 45 °C for 24 h. A Standard sugars (X1-X6); B Xylobiose; C Xylotriose; D Xylotetraose; E Xylopentaose and F Xylohexaose
Discussion
In our previous study, the gene encoding for the endoxylanase XylB was isolated from A. niger BCC14405 and successfully heterologously expressed as an intracellular protein in E. coli BL21 (DE3) (pLysS) expression system (Asano et al. 2005; Sriprang et al. 2006). According to the previous results, the wild type and recombinant xylanases exhibited the highest specificity towards xylan from birchwood with the activity of 5870 and 7663 U mg−1, respectively. This demonstrated the potential application of XylB for several biotechnological industries. However, the XylB was expressed as an intracellular protein that required additional cell disruption and protein purification steps. The production of recombinant enzyme in a secreted form is one of the important properties for economically feasible production in an industrial scale. Several reports demonstrated that P. pastoris has been successfully used as an expression host for production of extracellular recombinant proteins (Bunterngsook et al. 2017; Ding et al. 2018; Li et al. 2018; Zhuo et al. 2018). Expression of this enzyme in P. pastoris and optimization of the fermentation conditions for enzyme production was previously reported with demonstration of its application as an additive in animal feed (Ruanglek et al. 2007). However, the recombinant enzyme has not been characterized in detail for its biochemical properties and its application on other biotechnological purposes has not been studied.
In this work, the xylB gene was expressed in P. pastoris KM71 system by fusing with the alpha factor secretion signal under the control of methanol inducible promoter. The recombinant XylB was highly expressed as a secreted protein with a production yield of 116 mg protein L−1 of culture medium. Based on its biochemical characteristics, XylB showed high specificity against beechwood xylan (3852 U mg−1) and arabinoxylan (168 U mg−1) without any detectable cellulolytic activity on celluloses similar to the purified xylanase from A. niger BCC14405 reported previously (Asano et al. 2005). The structure of Endo-1,4-β-xylanase GH11 called jelly-roll scaffold contains two parts at the active binding site that make the enzyme specific to xylose-based polysaccharides. The first part is the thumb-loop of GH11, which makes the cleft narrow in subsites and is designed for binding xylose-based polysaccharides (Sulzenbacher et al. 1999) but not for glucose-based substrates (Paës et al. 2012). The second part is the highly conserved Val or Leu residue at position 35 of GH11, which binds only to xylan and prevents cellulose-binding because of the lack of hydrogen bonds with the O6 of glucose residue (Sidhu et al. 1999; Sulzenbacher et al. 1999). According to amino acid sequence alignment of XylB compared with the data in SWISS-MODEL (https://swissmodel.expasy.org/), the results suggested that XylB show similarities to Xylanase 11C (Endo-1,4-β-xylanase) from Talaromyces cellulolyticus (PDB ID: 3wp3) (Kataoka et al. 2014) at 67% identity and 78% homology. The structure prediction revealed the presence of both conserved parts in XylB, consistent with its specificity on xylan polysaccharides. This specificity towards xylan is a unique feature of endo-xylanase belonging to GH11 (Paës et al. 2012). For example, an endoxylanase from Trichoderma sp. exhibited xylanase activity towards xylan from beechwood, wheat arabinoxylan and rye arabinoxylan, without hydrolytic activity on CMC and Avicel (Fu et al. 2019). A xylanase purified from Myceliophthora heterothallica F.2.1.4 was active only on beechwood xylan (de Oliveira Simões et al. 2019). However, XylB obtained in this work revealed relatively low optimal temperature (45 °C) compared to endoxylanase purified from A. niger BCC14405 crude enzyme (55 °C) (Asano et al. 2005) and recombinant XylB produced in E. coli expression system (50 °C) (Sriprang et al. 2006). This could be due to differences in post-translational modifications of protein expressed in different microbial systems. The differences in optimal working conditions and specific activity of XylB to the enzyme expressed in P. pastoris (Ruanglek et al. 2007) could come from the use of different sources of xylan substrates (birchwood and beechwood xylan) and components in enzyme assay reactions. The lower optimal temperature of recombinant XylB in our study is desirable for biotechnological application of the enzyme for XOS production. The optimal temperature range of 45–50 °C was reported to be desirable as it inhibits growth of contaminating microbes while allows high solubility of the hemicellulosic substrates (Choudhary et al. 2016). This can also lead to lower processing temperature compared to the use of thermophilic enzymes (Damásio et al. 2011; Qiao et al. 2014; Sriyapai et al. 2011). The optimal temperature of XylB was relatively low compared to most of the reported GH11 endo-xylanases which work optimally at 40–80 °C, while our enzyme showed significantly higher specific activity compared to most previously reported enzymes originated from various microbial sources (Table 3). Our previous work on XylB expression in the E. coli system (Sriprang et al. 2006) did not report the yield of enzyme production and showed only the specific activity 7663 U mg−1 protein against birchwood xylan. The data thus cannot be directly compared to the use of different substrates. The specific activity against arabinoxylan of XylB was 168 U mg−1 which was higher than commercial xylanases such as endo-1,4-β-Xylanase M4 (Product code: E-XYAN4; Megazyme) from A. niger (80 U mg−1) and endo-1,4-β-Xylanase M3 (Product code: E-XYTR3; Megazyme) from Trichoderma longibrachiatum (100 U mg−1).
According to the effect of metal ions and chemical additives on XylB activity, most of the metal ions including Cu2+, Mg2+, Ni2+, Na+, Ca2+, Zn2+, K+, Li+ enhanced XylB activity while β-mercaptoethanol and SDS led to reduction in enzyme activity under the respective conditions. An inhibitory effect of Mn2+ on xylanase activity was reported (Cheng et al. 2012; Mamo et al. 2006). The reduction in enzyme activity by SDS could be due to its property as an anionic surfactant which could destabilize existing interactions in the protein structure leading to enzyme denaturation (da Silva et al. 2017). The effect of β-mercaptoethanol could be due to protein conformation shifting, resulted from changes in the overall charge and structure of the protein, which can lead to decreased enzyme activity (Ahmed et al. 1996).
Regarding its cleavage pattern on xylooligosaccharide (X2–X6), the recombinant XylB showed a capability to act on xylotriose to xylohexaose and released xylobiose (X2) as the major product. Furthermore, xylobiose was not hydrolyzed by the XylB, suggesting that the XylB acted on oligosaccharides containing at least three xylose units. This was supported by the high saccharification efficiency (70% conversion) of xylan from beechwood in which the XOS (X2, X3 and X6) were released. The remaining X6 in the hydrolysis reaction using xylan as the substrate could represent incomplete digestion of the substrate under the experimental conditions. The unlabeled peaks were the background of sodium phosphate buffer (Fig. 3b–f). Its cleavage patterns were similar to some xylanases in glycosyl hydrolase family 11 that hydrolyzed hemicellulose substrates into XOS with different chain lengths. For example, XynST11 from Streptomyces sp. B6 could not hydrolyze xylobiose and the hydrolysis product of xylotriose was xylobiose (Liu et al. 2020). In the same way, the major products from hydrolysis of xylotetraose and xylopentaose were xylotetraose and xylotriose. Some xylanases from other studies were reported to be incapable of xylotriose hydrolysis (Chang et al. 2017; Valenzuela et al. 2014; Chen et al. 2009). In general, the major XOS products from enzymatic hydrolysis are xylobiose, xylotriose, and xylotetraose (Linares-Pasten et al. 2018). The product profile of XylB agrees well with some GH11 xylanases previously reported. The endoxylanase isolated from bagasse metagenome and its variants were recently reported to hydrolyze beechwood xylan into X3–X6 (Boonyapakron et al. 2020). Berrin and coworkers reported the predominant production of xylobiose and xylotriose as end products from wheat soluble arabinoxylan hydrolysis by PgXynA and PfXynC GH11 xylanases from Penicillium griseofulvum (Berrin et al. 2007). In addition, a GH11 xylanase (XynST11) isolated from Streptomyces sp. B6 produced much less xylose and a higher amount of XOS with a DP from 2 to 4 than the GH10 family enzyme (XynST10) (Liu et al. 2020). Overall, these previous works demonstrate the advantages of GH11 xylanases for XOS production and great potential of XylB for conversion of the plant-derived xylan to oligosaccharides.
Conclusion
In summary, recombinant XylB isolated from A. niger BCC14405 fungal strain was highly expressed in a secreted form in the yeast expression system. XylB is a cellulase-free xylanase that exhibits catalytic capability to hydrolyze xylan from beechwood and produce XOS. The properties of XylB demonstrate its potential applications for XOS production and lignocellulosic biomass saccharification.
References
Aachary AA, Prapulla SG (2011) Xylooligosaccharides (XOS) as an emerging prebiotic: microbial synthesis, utilization, structural characterization, bioactive properties, and applications. Compr Rev Food Sci Food Saf 10(1):2–16. https://doi.org/10.1111/j.1541-4337.2010.00135
Abelilla JJ, Stein HH (2019) Degradation of dietary fiber in the stomach, small intestine, and large intestine of growing pigs fed corn- or wheat-based diets without or with microbial xylanase. J Anim Sci 97(1):338–352. https://doi.org/10.1093/jas/sky403
Adiguzel G, Faiz O, Sisecioglu M, Sari B, Baltaci O, Akbulut S, Genc B, Adiguzel A (2019) A novel endo-beta-1,4-xylanase from Pediococcus acidilactici GC25; purification, characterization and application in clarification of fruit juices. Int J Biol Macromol 129:571–578. https://doi.org/10.1016/j.ijbiomac.2019.02.054
Ahmed SA, McPhie P, Miles EW (1996) Mechanism of activation of the tryptophan synthase alpha2beta2 complex. Solvent effects of the co-substrate beta-mercaptoethanol. J Biol Chem 271(46):29100–29106. https://doi.org/10.1074/jbc.271.46.29100
Ahmed S, Riaz S, Jamil A (2009) Molecular cloning of fungal xylanases: an overview. Appl Microbiol Biotechnol 84(1):19–35. https://doi.org/10.1007/s00253-009-2079-4
Alokika, Singh B (2019) Production, characteristics, and biotechnological applications of microbial xylanases. Appl Microbiol Biotechnol 103(21–22):8763–8784. https://doi.org/10.1007/s00253-019-10108-6
Asano K, Sriprang R, Jarupan G, Eurwilaichitr L, Tanapongpipat S, Kirtikara K (2005) Endo-1,4-beta-xylanase B from Aspergillus cf. niger BCC14405 Isolated in Thailand: purification, characterization and gene isolation. J Biochem Mol Biol 38:17–23. https://doi.org/10.5483/BMBRep.2005.38.1.017
Beg QK, Kapoor M, Mahajan L, Hoondal GS (2001) Microbial xylanases and their industrial applications: a review. Appl Microbiol Biotechnol 56(3–4):326–338. https://doi.org/10.1007/s002530100704
Berrin JG, el Ajandouz H, Georis J, Arnaut F, Juge N (2007) Substrate and product hydrolysis specificity in family 11 glycoside hydrolases: an analysis of Penicillium funiculosum and Penicillium griseofulvum xylanases. Appl Microbiol Biotechnol 74(5):1001–1010. https://doi.org/10.1007/s00253-006-0764-0
Biely P (1985) Microbial xylanolytic systems. Trends Biotechnol 3(11):286–290. https://doi.org/10.1016/0167-7799(85)90004-6
Boonyapakron K, Chitnumsub P, Kanokratana P, Champreda V (2020) Enhancement of catalytic performance of a metagenome-derived thermophilic oligosaccharide-specific xylanase by binding module removal and random mutagenesis. J Biosci Bioeng 131(1):13–19. https://doi.org/10.1016/j.jbiosc.2020.09.008
Bunterngsook B, Laothanachareon T, Natrchalayuth S, Lertphanich S, Fujii T, Inoue H, Youngthong C, Chantasingh D, Eurwilaichitr L, Champreda V (2017) Optimization of a minimal synergistic enzyme system for hydrolysis of raw cassava pulp. RSC Adv 7(76):48444–48453. https://doi.org/10.1039/C7RA08472B
Chang S, Guo Y, Wu B, He B (2017) Extracellular expression of alkali tolerant xylanase from Bacillus subtilis Lucky9 in E. coli and application for xylooligosaccharides production from agro-industrial waste. Int J Biol Macromol 96:249–256. https://doi.org/10.1016/j.ijbiomac.2016.11.032
Chen LL, Zhang M, Zhang DH, Chen XL, Sun CY, Zhou BC, Zhang YZ (2009) Purification and enzymatic characterization of two beta-endoxylanases from Trichoderma sp. K9301 and their actions in xylooligosaccharide production. Bioresour Technol 100(21):5230–5336. https://doi.org/10.1016/j.biortech.2009.05.038
Cheng F, Sheng J, Dong R, Men Y, Gan L, Shen L (2012) Novel xylanase from a holstein cattle rumen metagenomic library and its application in xylooligosaccharide and ferulic acid production from wheat straw. J Agric Food Chem 60(51):12516–12524. https://doi.org/10.1021/jf302337w
Choudhary J, Singh S, Nain L (2016) Thermotolerant fermenting yeasts for simultaneous saccharification fermentation of lignocellulosic biomass. Electron J Biotechnol 21:82–92. https://doi.org/10.1016/j.ejbt.2016.02.007
da Silva RR, de Oliveira LCG, Juliano MA, Juliano L, de Oliveira AHC, Rosa JC, Cabral H (2017) Biochemical and milk-clotting properties and mapping of catalytic subsites of an extracellular aspartic peptidase from basidiomycete fungus Phanerochaete chrysosporium. Food Chem 225:45–54. https://doi.org/10.1016/j.foodchem.2017.01.009
Damásio ARDL, Silva TM, Almeida FBDR, Squina FM, Ribeiro DA, Leme AFP, Segato F, Prade RA, Jorge JA, Terenzi HF, Polizeli, M.d.L.T.M. (2011) Heterologous expression of an Aspergillus niveus xylanase GH11 in Aspergillus nidulans and its characterization and application. Process Biochem 46(6):1236–1242. https://doi.org/10.1016/j.procbio.2011.01.027
de Oliveira Simões LC, da Silva RR, de Oliveira Nascimento CE, Boscolo M, Gomes E, da Silva R (2019) Purification and physicochemical characterization of a novel thermostable xylanase secreted by the fungus Myceliophthora heterothallica F.2.1.4. Appl Biochem Biotechnol 188(4):991–1008. https://doi.org/10.1007/s12010-019-02973-8
Ding C, Li M, Hu Y (2018) High-activity production of xylanase by Pichia stipitis: purification, characterization, kinetic evaluation and xylooligosaccharides production. Int J Biol Macromol 117:72–77. https://doi.org/10.1016/j.ijbiomac.2018.05.128
Dodd D, Cann IK (2009) Enzymatic deconstruction of xylan for biofuel production. Glob Change Biol Bioenergy 1(1):2–17. https://doi.org/10.1111/j.1757-1707.2009.01004.x
Fu LH, Jiang N, Li CX, Luo XM, Zhao S, Feng JX (2019) Purification and characterization of an endo-xylanase from Trichoderma sp., with xylobiose as the main product from xylan hydrolysis. World J Microbiol Biotechnol 35(11):171. https://doi.org/10.1007/s11274-019-2747-1
Gonçalves TA, Damásio AR, Segato F, Alvarez TM, Bragatto J, Brenelli LB, Citadini AP, Murakami MT, Ruller R, Paes Leme AF, Prade RA, Squina FM (2012) Functional characterization and synergic action of fungal xylanase and arabinofuranosidase for production of xylooligosaccharides. Bioresour Technol 119:293–299. https://doi.org/10.1016/j.biortech.2012.05.062
Jeya M, Thiagarajan S, Lee JK, Gunasekaran P (2009) Cloning and expression of GH11 xylanase gene from Aspergillus fumigatus MKU1 in Pichia pastoris. J Biosci Bioeng 108(1):24–29. https://doi.org/10.1016/j.jbiosc.2009.02.003
Jun H, Bing Y, Keying Z, Xuemei D, Daiwen C (2009) Expression of a Trichoderma reesei beta-xylanase gene in Escherichia coli and activity of the enzyme on fiber-bound substrates. Protein Expr Purif 67(1):1–6. https://doi.org/10.1016/j.pep.2008.07.015
Kataoka M, Akita F, Maeno Y, Inoue B, Inoue H, Ishikawa K (2014) Crystal Structure of Talaromyces cellulolyticus (Formerly Known as Acremonium cellulolyticus) GH Family 11 Xylanase. Appl Biochem Biotechnol 174(4):1599–1612. https://doi.org/10.1007/s12010-014-1130-9
Li H, Wu H, Jiang F, Wu J, Xue Y, Gan L, Liu J, Long M (2018) Heterologous expression and characterization of an acidic GH11 family xylanase from Hypocrea orientalis. Appl Biochem Biotechnol 184(1):228–238. https://doi.org/10.1007/s12010-017-2532-2
Li C, Kumar A, Luo X, Shi H, Liu Z, Wu G (2020) Highly alkali-stable and cellulase-free xylanases from Fusarium sp. 21 and their application in clarification of orange juice. Int J Biol Macromol 155:572–580. https://doi.org/10.1016/j.ijbiomac.2020.03.249
Liao H, Zheng H, Li S, Wei Z, Mei X, Ma H, Shen Q, Xu Y (2015) Functional diversity and properties of multiple xylanases from Penicillium oxalicum GZ-2. Sci Rep 5:12631. https://doi.org/10.1038/srep12631
Linares-Pasten JA, Aronsson A, Karlsson EN (2018) Structural considerations on the use of endo-xylanases for the production of prebiotic xylooligosaccharides from biomass. Curr Protein Pept Sci 19(1):48–67. https://doi.org/10.2174/1389203717666160923155209
Liu X, Liu Y, Jiang Z, Liu H, Yang S, Yan Q (2018) Biochemical characterization of a novel xylanase from Paenibacillus barengoltzii and its application in xylooligosaccharides production from corncobs. Food Chem 264:310–318. https://doi.org/10.1016/j.foodchem.2018.05.023
Liu L, Xu M, Cao Y, Wang H, Shao J, Xu M, Zhang Y, Wang Y, Zhang W, Meng X, Liu W (2020) Biochemical characterization of xylanases from Streptomyces sp. b6 and their application in the xylooligosaccharide production from viscose fiber production Waste. J Agric Food Chem 68(10):3184–3194. https://doi.org/10.1021/acs.jafc.9b06704
Maitan-Alfenas GP, Oliveira MB, Nagem RAP, de Vries RP, Guimarães VM (2016) Characterization and biotechnological application of recombinant xylanases from Aspergillus nidulans. Int J Biol Macromol 91:60–67. https://doi.org/10.1016/j.ijbiomac.2016.05.065
Mamo G, Hatti-Kaul R, Mattiasson B (2006) A thermostable alkaline active endo-β-1-4-xylanase from Bacillus halodurans S7: purification and characterization. Enzyme Microb Technol 39(7):1492–1498. https://doi.org/10.1016/j.enzmictec.2006.03.040
Miller GL (1959) Use of dinitrosalicyclic acid reagent for determination of reducing sugar. Anal Chem 5:426–428. https://doi.org/10.1021/ac60147a030
Moure A, Gullón P, Domínguez H, Parajó JC (2006) Advances in the manufacture, purification and applications of xylo-oligosaccharides as food additives and nutraceuticals. Process Biochem 41(9):1913–1923. https://doi.org/10.1016/j.procbio.2006.05.011
Paës G, Berrin J-G, Beaugrand J (2012) GH11 xylanases: Structure/function/properties relationships and applications. Biotechnol Adv 30(3):564–592. https://doi.org/10.1016/j.biotechadv.2011.10.003
Poletto P, Pereira GN, Monteiro CRM, Pereira MAF, Bordignon SE, de Oliveira D (2020) Xylooligosaccharides: transforming the lignocellulosic biomasses into valuable 5-carbon sugar prebiotics. Process Biochem 91:352–363. https://doi.org/10.1016/j.procbio.2020.01.005
Polizeli ML, Rizzatti AC, Monti R, Terenzi HF, Jorge JA, Amorim DS (2005) Xylanases from fungi: properties and industrial applications. Appl Microbiol Biotechnol 67(5):577–591. https://doi.org/10.1007/s00253-005-1904-7
Qiao W, Tang S, Mi S, Jia X, Peng X, Han Y (2014) Biochemical characterization of a novel thermostable GH11 xylanase with CBM6 domain from Caldicellulosiruptor kronotskyensis. J Mol Catal B Enzym 107:8–16. https://doi.org/10.1016/j.molcatb.2014.05.009
Ruanglek V, Sriprang R, Ratanaphan N, Tirawongsaroj P, Chantasigh D, Tanapongpipat S, Pootanakit K, Eurwilaichitr L (2007) Cloning, expression, characterization, and high cell-density production of recombinant endo-1,4-β-xylanase from Aspergillus niger in Pichia pastoris. Enzyme Microb Technol 41(1):19–25. https://doi.org/10.1016/j.enzmictec.2006.11.019
Samanta AK, Jayapal N, Jayaram C, Roy S, Kolte AP, Senani S, Sridhar M (2015) Xylooligosaccharides as prebiotics from agricultural by-products: production and applications. Bioactive Carbohydr Diet Fibre 5(1):62–71. https://doi.org/10.1016/j.bcdf.2014.12.003
Sidhu G, Withers SG, Nguyen NT, McIntosh LP, Ziser L, Brayer GD (1999) Sugar ring distortion in the glycosyl-enzyme intermediate of a family G/11 xylanase. Biochemistry 38(17):5346–5354
Singh A, Varghese LM, Battan B, Patra AK, Mandhan RP, Mahajan R (2020) Eco-friendly scouring of ramie fibers using crude xylano-pectinolytic enzymes for textile purpose. Environ Sci Pollut Res Int 27(6):6701–6710. https://doi.org/10.1007/s11356-019-07424-9
Sriprang R, Asano K, Gobsuk J, Tanapongpipat S, Champreda V, Eurwilaichitr L (2006) Improvement of thermostability of fungal xylanase by using site-directed mutagenesis. J Biotechnol 126(4):454–462. https://doi.org/10.1016/j.jbiotec.2006.04.031
Sriyapai T, Somyoonsap P, Matsui K, Kawai F, Chansiri K (2011) Cloning of a thermostable xylanase from Actinomadura sp. S14 and its expression in Escherichia coli and Pichia pastoris. J Biosci Bioeng 111(5):528–536. https://doi.org/10.1016/j.jbiosc.2010.12.024
Sulzenbacher G, Mackenzie LF, Wilson KS, Withers SG, Dupont C, Davies GJ (1999) The crystal structure of a 2-fluorocellotriosyl complex of the Streptomyces lividans endoglucanase CelB2 at 1.2 A resolution. Biochemistry 38(15):4826–4833. https://doi.org/10.1021/bi982648i
Thak EJ, Yoo SJ, Moon HY, Kang HA (2020) Yeast synthetic biology for designed cell factories producing secretory recombinant proteins. FEMS Yeast Res. https://doi.org/10.1093/femsyr/foaa009
Valenzuela SV, Diaz P, Pastor FI (2014) Xyn11E from Paenibacillus barcinonensis BP-23: a LppX-chaperone-dependent xylanase with potential for upgrading paper pulps. Appl Microbiol Biotechnol 98(13):5949–5957. https://doi.org/10.1007/s00253-014-5565-2
Walia A, Guleria S, Mehta P, Chauhan A, Parkash J (2017) Microbial xylanases and their industrial application in pulp and paper biobleaching: a review. 3 Biotech 7(1):11. https://doi.org/10.1007/s13205-016-0584-6
Wang R, Arioka M (2021) Functional analyses of xylanolytic enzymes involved in xylan degradation and utilization in Neurospora crassa. Int J Biol Macromol 169:302–310. https://doi.org/10.1016/j.ijbiomac.2020.12.079
Xu X, Liu M-Q, Huo W-K, Dai X-J (2016) Obtaining a mutant of Bacillus amyloliquefaciens xylanase A with improved catalytic activity by directed evolution. Enzyme Microb Technol 86:59–66. https://doi.org/10.1016/j.enzmictec.2016.02.001
Yang X, Shi P, Huang H, Luo H, Wang Y, Zhang W, Yao B (2014) Two xylose-tolerant GH43 bifunctional β-xylosidase/α-arabinosidases and one GH11 xylanase from Humicola insolens and their synergy in the degradation of xylan. Food Chem 148:381–387. https://doi.org/10.1016/j.foodchem.2013.10.062
Zhang YHP, Cui J, Lynd LR, Kuang LR (2006) A Transition from cellulose swelling to cellulose dissolution by o-phosphoric acid: evidence from enzymatic hydrolysis and supramolecular structure. Biomacromolecules 7(2):644–648. https://doi.org/10.1021/bm050799c
Zhuo R, Yu H, Qin X, Ni H, Jiang Z, Ma F, Zhang X (2018) Heterologous expression and characterization of a xylanase and xylosidase from white rot fungi and their application in synergistic hydrolysis of lignocellulose. Chemosphere 212:24–33. https://doi.org/10.1016/j.chemosphere.2018.08.062
Acknowledgements
This work was financially supported by National Science and Technology Development Agency [Grant No. P18-52705]. The authors would like to thank Dr. Nattapol Arunrattanamook for manuscript proofreading and comments.
Author information
Authors and Affiliations
Contributions
KA: Methodology, Investigation, Writing—original draft. BB: Conceptualization, Methodology, Investigation, Writing—original draft, Writing—review and editing. HL: Methodology, Investigation, Writing—review and editing. WS: Writing—original draft, Writing—review and editing. PK: Writing—original draft, Writing—review and editing. VC: Conceptualization, Investigation, Writing—review and editing, Supervision, Funding acquisition.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest with the contents of this article.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Aiewviriyasakul, K., Bunterngsook, B., Lekakarn, H. et al. Biochemical characterization of xylanase GH11 isolated from Aspergillus niger BCC14405 (XylB) and its application in xylooligosaccharide production. Biotechnol Lett 43, 2299–2310 (2021). https://doi.org/10.1007/s10529-021-03202-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-021-03202-1