Abstract
Objectives
To characterize methyltransferases involved in the biosynthesis of benzylisoquinoline alkaloids in Stephania intermedia.
Results
Three N-methyltransferases, SiCNMT1, SiCNMT2, SiCNMT3, and O-methyltransferase SiSOMT were identified in Stephania intermedia. Then, four methyltransferase genes were cloned into the pGEX-6P-1 vector. The recombinant vectors were transformed into Escherichia coli BL21(DE3) for expression and were functionally tested. SiCNMT1, SiCNMT2, and SiCNMT3 could methylate (R)-coclaurine to produce (R)-N-methylcoclaurine. SiCNMT2 further methylated the product of (R)-N-methylcoclaurine to produce (R)-magnocurarine. Similarly, (R)-norcoclaurine was continuously catalyzed to yield (R)-N-methylnorcoclaurine and (R)-N, N-dimethylnorcoclaurine by SiCNMT2. Furthermore, SiSOMT was shown to catalyze the conversion of (S)-scoulerine to (S)-tetrahydropalmatine.
Conclusions
The key methyltransferases, which were in the last step biosynthesis of (R)-magnocurarine, (R)-N, N-dimethylnorcoclaurine and (S)-tetrahydropalmatine were revealed and their activities were verified in vitro. Four novel methyltransferases will be promising candidates for methylation of benzylisoquinoline alkaloids.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Stephania intermedia H. S. Lo (SI), belonging to the genus Stephania (Menispermaceae), is an important medicinal herbaceous vine plant that allows for treatment of pain. SI contains many types of benzylisoquinoline alkaloids (BIAs), such as the analgesic (S)-tetrahydropalmatine ((S)-THP), the anti-microbial berberine and (S)-corydalmine, which reduces morphine tolerance (Facchini and De Luca 2008; Zuo et al. 2011). (S)-THP has been used for more than 40 years as an analgesic in China and exhibits hepatoprotective, anti-arrhythmic, and anti-inflammatory activities (Kang et al. 2016; Sun et al. 2018). (S)-corydalmine, the main metabolite of (S)-THP in vivo, substantially reduces morphine tolerance and relieves bone cancer pain (Dai et al. 2016, 2017; Tang et al. 2016).
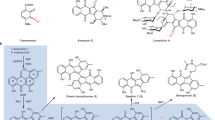
In recent years, researchers have focused on several plant species to provide insight into major BIAs, including morphine found in Papaver somniferum, berberine in Coptis japonica, and sanguinarine in Eschscholzia californica (Hagel and Facchini 2010, 2013; Purwanto et al. 2017; Yamada et al. 2016). Other species that accumulate a wide-range of BIAs, such as SI, have not been reported yet. SI is rich in (S)-THP, suggesting its suitability as a plant material for the purpose of this study, to elucidate the biosynthetic pathway of (S)-THP. However, the genome of SI has not been sequenced and the transcriptome data resources are also scarce. A previous study found that, (R)-N-methylnorcoclaurine, (R)-N, N-dimethylnorcoclaurine, (S)-THP, and their precursors were detected from SI, respectively (data not available here). Furthermore, some of the methyltransferases (MTs) are capable of successive methylation. For example, pavine N-methyltransferase (PavNMT) found in Thalictrum flavum and reticuline N-methyltransferase in P. somniferum (RNMT) could catalyze pavine and reticuline, respectively (Morris and Facchini 2016; Torres et al. 2016). It was, therefore, proposed that N-methylation and O-methylation reactions were mediated by N-methyltransferase and O-methyltransferase, respectively, in the biosynthetic pathway of (R)-N-methylnorcoclaurine, (R)-N, N-dimethylnorcoclaurine, and (S)-THP. Furthermore, due to abundant (S)-THP in SI, we predicted that special biosynthetic pathways, highly efficient or highly expressed enzymes are different in activity from other plants. Based on the above reasons, we proposed the biosynthetic pathway of (S)-THP (Fig. 1; Table 1).
Proposed pathways for BIAs biosynthesis in SI. Enzymes abbreviated in red text have been characterized in this study. NCS norcoclaurine synthase, 6OMT norcoclaurine 6-O-methyltransferase, SiCNMT2 coclaurine N-methyltransferase, NMCH N-methylcoclaurine 3′-hydroxylase, 4′OMT 4′-O-methyltransferase, BBE berberine bridge enzyme, SiSOMT (S)-scoulerine 9-O-methyltransferase
Herein, we have identified four novel MTs (SiCNMT1, SiCNMT2, SiCNMT3 and SiSOMT) from the transcription data of SI. Four MTs were cloned into the pGEX-6P-1 expression vector. Then, the recombinant vectors were transformed into E. coli BL21 (DE3) and induced with IPTG for expression. The fusion proteins were purified using GST-tag Protein Purification Kit and their properties were characterized in detail.
Materials and methods
Materials
Six wild tuberous roots of SI from different plants (each weighed roughly 500 g, and were 10 cm in diameter) were collected in 2017 from the Yunnan province of China. The voucher specimens (S2017070) were deposited in the school of Traditional Chinese Pharmacy, China Pharmaceutical University. Tuberous roots were transplanted into the field. The next year, leaf and root, were harvested separately and frozen using liquid nitrogen before storing at − 80 °C until RNA extraction.
(R)-coclaurine, (S)-tetrahydrocolumbamine, (R)-norcoclaurine, (S)-scoulerine, (S)-tetrahydropalmatrubine, Norlaudanosine and (S)-THP were purchased from Yuan Ye Biotechnology Co., Ltd. S-Adenosyl-l-methionine (SAM) was purchased from Sigma–Aldrich. Acetonitrile, methanol, and formic acid (HPLC grade) for LC analysis were purchased from Merck (Darmstadt, Germany). Ethanol and isopropanol were purchased from Nanjing chemical reagent co. LTD.
RNA extraction
Total RNA was extracted from the samples using Plant RNA Kit (Omega Bio-tek, Norcross, GA, USA). First strand cDNA synthesis was performed on 2 μg of RNA using HiScript III 1st Strand cDNA Synthesis Kit (Vazyme, Nanjing, China). MTs open reading frames were amplified from cDNA using 2 × Phanta Max Master Mix DNA polymerase (Vazyme, Nanjing, China).
Identification of candidate genes
The leaf and root transcriptomes of SI were searched to identify the sequence encoding the N-methyltransferase and O-methyltransferase candidates. The assembled unigenes were blasted against public databases (E value < 1 × 10−5), including the non-redundant protein (NR) database, Nucleotide collection (NT), the SwissProt database, the Kyoto Encyclopedia of Genes and Genomes (KEGG) database, Cluster of Orthologous Groups of proteins (COG) and Gene Ontology (GO) (He et al. 2018). Four unigenes of MTs were isolated from the SI transcriptome data. Three unigenes were highly similar to CNMT and were named as SiCNMT1, SiCNMT2 and SiCNMT3, respectively. One unigene was highly similar to SOMT, which was named as SiSOMT.
Phylogenetic analysis
To characterize the evolutionary relationships between four MTs from SI and MTs from other plant species, 20 amino acid sequences of N-methyltransferases and 22 amino acid sequences of O-methyltransferases were used to construct the phylogenetic tree of N-methyltransferase and the phylogenetic tree of O-methyltransferase, respectively. Molecular phylogenetic trees were constructed by the neighbor joining algorithm method and bootstrap frequencies for each clade were based on 1,000 iterations (Chang et al. 2015; He et al. 2017). Abbreviations and GenBank accession numbers for sequences were used to construct the phylogenetic tree (shown in Supplementary Table 1).
Expression and purification of recombinant protein
SiCNMT1 (GenBank accession number, MK749412), SiCNMT2 (GenBank accession number, MK749413), SiCNMT3 (GenBank accession number, MK749414) and SiSOMT (GenBank accession number, MK749415) genes were obtained from the cDNA of SI by PCR amplification using primers (Supplementary Table 2) and cloned into the pGEX-6P-1 vector, respectively. Recombinant plasmids were transformed into the E. coli BL21(DE3) (TransGen Biotech, China) for protein expression. The 5 μL of recombinant plasmid was added into a tube containing 50 μL of chemically competent cell on the ice and mixed them gently. The mixture was on ice for 30 min and then heated for 45 s at 42 °C. The tube was quickly transferred to the ice for 2 min. Next, 500 μL LB medium was added to the tube, mixed well and placed under 37 °C while spun at 200 rpm for 1 h to revive the bacteria. Plasmid-transformed E. coli BL21(DE3) cells were inoculated in TB medium containing 100 mg/l ampicillin and the expression cultures were grown at 37 °C and shaken at 200 rpm. When the OD at 600 reached 0.6, the cells were induced with 0.1 mM IPTG and the cultured at 16 °C while shaking at 200 rpm for 18 h. The cells were harvested by centrifugation (6000×g, 3 min, at 4 °C) and resuspended in binding buffer, and the suspension was subsequently homogenized for 20 min and applying 200-W sonication (JY99-IIDN Sonicator; scientz, China). Cell debris was subsequently removed by centrifugation at 6000×g for 10 min. After centrifugation, the supernatant was purified using GST-tag Protein Purification Kit (Beyotime, China). The purity of the GST-tagged protein was analyzed by SDS-polyacrylamide gel electrophoresis (SDS-PAGE) with Coomassie Brilliant Blue staining.
Enzyme assays
The enzyme assay for SiSOMT activity was performed using a reaction mixture in 100 μL of 100 mM Tris-HCl (pH 9.0), 5 mM SAM, 25 mM sodium ascorbate, 10% (v/v) glycerol, 1 mM β-mercaptoethanol, 0.5 mM potential alkaloid substrate, and 10 μg of purified recombinant enzyme. SiCNMT1, SiCNMT2, and SiCNMT3 activities were performed using a reaction mixture in 100 μL of 100 mM Tris-HCl (pH 7.0), 5 mM SAM, 25 mM sodium ascorbate, 10% (v/v) glycerol, 1 mM β-mercaptoethanol, 0.5 mM potential alkaloid substrate, and 10 μg of purified recombinant enzyme. Kinetic parameters were determined at 37 °C in a Thermomixer (200 rpm) for 2–20 min by varying alkaloid substrate concentrations from 10 μM to 500 μM. Reactions were extracted by acetonitrile. The products were confirmed by HPLC-Q-TOF-MS/MS and the products amounts were determined from the conversion ratio using area under-peak of product from the HPLC chromatograms. Kinetic constants were determined by fitting the initial velocity-versus-substrate concentration to the Michaelis–Menten equation using GraphPad Prism 5.
HPLC-Q-TOF-MS/MS analysis of enzyme assays
The analysis of the products was carried out using an Agilent 1260 high-performance liquid chromatography system. Substrates and their biocatalytic products were separated on a reversed-phase C18 column (250 mm × 4.6 mm, 5 μm, Zorbax, Agilent). The mobile phase consisted of water that contained 0.1% formic acid and 1 mM ammonium formate (phase A) and acetonitrile (phase B) with the following gradient system: 15% B at 0–5 min, 15–20% B at 5–10 min, 20–22% B at 10–15 min, 22% B at 15–17 min, 22–30% B at 17–20 min, 30–95% B at 20–22 min, 95–15% B at 22–23 min, 15% B at 23–25 min. The flow rate was kept at 1 mL/min with an injection volume of 10 μL. Detection was performed at 280 nm. An Agilent 6530 Q-TOF mass spectrometer (Agilent Technologies, USA) equipped with an electrospray ionization source was used to perform the MS analysis. The acquisition parameters were as follows: drying gas (N2) flow rate, 10.0 L/min; drying gas temperature, 350 °C; capillary voltage, 4 kV; OCT RFV, 750 V; fragmentor voltage, 120 V; and nebulizer, 35 psig. The mass range was recorded from m/z 100 to 1500 with collision energies that range from 10 to 50 eV. Peaks were detected in positive ionization mode for MS and MS/MS detection. To guarantee mass accuracy, the TOF mass spectrometer was calibrated by means of a calibrant solution before sample analysis. The calibrant solution contains the internal reference masses of purine (C5H4N4) at m/z 121.0509 and HP-921 [hexakis-(1H, 1H, 3H-tetrafluoro-pentoxy) phosphazene] (C18H18O6N3P3F24) at m/z 922.009. All operations, data acquisition, and data analysis were carried out by Agilent Mass Hunter Workstation software (version B.07.00).
Results
Isolation and characterization of methyltransferases
The full-length cDNAs of MT genes were obtained from the leaf and root transcriptome databases of SI. The candidate selection strategy was based on a cutoff of 40% amino acid sequence identity, compared with at least one functionally characterized MTs involved in BIAs metabolism (Chang et al. 2015). The open reading frames of SiCNMT1, SiCNMT2, SiCNMT3, and SiSOMT genes were 1065, 1065, 1077, and 1050 bp, respectively. ExPASy database (http://expasy.org/tools/) was used for predicting molecular weight and isoelectric point of the protein (pI). Protein analysis indicated that SiCNMT1, SiCNMT2, SiCNMT3, and SiSOMT had a predicted molecular weight of 41.60 kDa, pI of 6.43; 41.14 kDa, pI of 5.11; 41.08 kDa, pI of 5.98; and 37.67 kDa, pI of 5.55, respectively.
Phylogenetic analysis
To characterize the evolutionary relationships between MTs from SI and known MTs from other plant species, 20 amino acid sequences of N-methyltransferases and 22 amino acid sequences of O-methyltransferases were used to construct the phylogenetic tree of N-methyltransferase and the phylogenetic tree of O-methyltransferase, respectively. Phylogenetic analysis showed that SiCNMT1, SiCNMT2, SiCNMT3, and SiSOMT formed separate clades with characterized MTs (Fig. 2), whereas SiCNMT1 and CjCNMT from C. japonica formed a new clade. SiCNMT3 and TfCNMT from T. flavum formed a new clade. SiCNMT2 formed a new clade and showed 59% sequence identities with PsTNMT4 and TfpavTNMT. SiSOMT showed 81% sequence identities with CjSOMT and TfSOMT.
Phylogenetic tree of MTs (A NMTs; B OMTs). Phylogenetic tree was constructed on the basis of the deduced amino acid sequences for the MTs (black circle marker) and other plant MTs. Bootstrap frequencies for each clade were based on 1000 iterations. The scale bar corresponds to 0.2 amino acid substitutions per site
Purification and in vitro characterization of methyltransferases
The complete coding sequences of MTs were cloned into the pGEX-6P-1 expression vector with a N-terminal GST-tagged translational fusion. Recombinant MTs were purified from the total protein extracts using GST-tag Protein Purification Kit (Beyotime, China). All purified recombinant enzymes were analyzed by 10% SDS-PAGE. (Fig. 3). Enzyme assays were conducted in 100 mM Tris–HCl, pH 7.0 and 9.0, using 0.5 mM alkaloid, 5 mM SAM, and 10 μg of recombinant proteins to screen methylation activity. Mixtures were incubated for 18 h at 37 °C and quenched with the addition of 100 μL of acetonitrile (Morris and Facchini 2016). Several similar substrates were tested to determine the substrate specificity of SiCNMT1, SiCNMT2, SiCNMT3, and SiSOMT. SiCNMT1, SiCNMT2, and SiCNMT3 exhibited activity with (R)-coclaurine (conversion, 30%, 60%, and 45%, respectively) (Supplementary Table 3). Interestingly, SiCNMT2 could further methylate the product of N-methylcoclaurine to form (R)-magnocurarine. SiSOMT showed the differential activities of three substrates from the seven tested substrates. (S)-scoulerine, which displayed 100% conversion, was the preferred substrate. (S)-tetrahydropalmatrubine (90.2%) and (S)-tetrahydrocolumbamine (58.2%) were also converted efficiently. Next, the catalyzation and kinetic parameters were measured to test the reaction activity of MTs. SiCNMT1, SiCNMT2, and SiCNMT3 were determined by (R)-coclaurine, and SiSOMT was determined through (S)-scoulerine. The results showed that Km of SiCNMT1, SiCNMT2, and SiCNMT3 for (R)-coclaurine were 92.2, 74.6.5, and 97.1 μΜ, respectively. SiCNMT2 had better affinity with (R)-coclaurine than SiCNMT1 and SiCNMT3. SiSOMT displayed Km of 53.6 μΜ for (S)-scoulerine (Supplementary Table 4; Supplementary Fig. 1).
GST-tagged purified recombinant MTs on 10% SDS-PAGE gel. Lane1: marker; Lane 2: pGEX-6P-1 (vector control); Lane 3: crude enzyme of SiCNMT1; Lane 4: purified SiCNMT1; Lane 5: crude enzyme of SiCNMT2; Lane 6: purified SiCNMT 2; Lane 7: crude enzyme of SiCNMT3; Lane 8: purified SiCNMT3; Lane 9: crude enzyme of SiSOMT; Lane 10: purified SiSOMT
Reaction product identification
HPLC-Q-TOF–MS/MS was performed to identify the reaction products of recombinant MTs. (R)-coclaurine was catalyzed by SiCNMT1, SiCNMT2, and SiCNMT3, which yielded the same peak with m/z 300.1584 at 12.12 min (Supplementary Fig. 2). The characteristic fragmentation behavior refers to the dissociation of the NH2CH3-generated m/z 269.1141 by retro-Diels–Alder (RDA) cleavage. This behavior showed the existence of one − CH3 group at nitrogen atom (He et al. 2016; Oh et al. 2018; Xiao et al. 2018). The product was further identified as (R)-N-methylcoclaurine through major fragment ions (Supplementary Fig. 2; Supplementary Table 5). Furthermore, SiCNMT2 yielded another peak m/z 314.1758 at 11.11 min, and the characteristic fragment at m/z 269.1114 was formed by losing NH2(CH3)2 group. Thus, the product was identified as (R)-magnocurarine by its major fragment ions. Similarly, (R)-norcoclaurine was catalyzed by SiCNMT2, which yielded two peaks with m/z 286.1449 at 10.82 min and m/z 300.1595 at 9.80 min. Two peaks were identified as (R)-N-methylnorcoclaurine and (R)-N, N-dimethylnorcoclaurine by their characteristic fragment ions, respectively (Supplementary Fig. 3). Tetrahydroprotoberberine alkaloids possessed the indicative ions formed by RDA fragmentation at the C-ring and direct B-ring cleavage, and the isoquinoline fragments generated by RDA fragmentation were stronger than those generated by B-ring cleavage (Shangguan et al. 2018). Positions of new O-methyl groups could be inferred from the increased mass-to-charge ratio (m/z; in multiples of 14 Da) of dissociated isoquinoline and benzyl ion fragments. (S)-scoulerine was catalyzed by SiSOMT and yielded two major peaks at m/z 342.1707 (10.27 min) and m/z 356.1857 (11.18 min) in addition to the substrate peak m/z 328.1532 (8.84 min, Supplementary Fig. 4). These findings suggested the occurrence of single and double O-methylation events (Chang et al. 2015). Two major peaks were identified as (S)-tetrahydrocolumbamine and (S)-THP based on their retention time and characteristic ions with the standard compound (Lin et al. 2018).
(S)-scoulerine has two O-methylation sites, namely, C-9 and C-2 (Supplementary Fig. 4A). The catalysis sequence at the C-9 and C-2 positions was clarified by SiSOMT. Incubation of SiSOMT with (S)-tetrahydrocolumbamine (m/z 342.1702) yielded one major peak at m/z 356.1872. The product was inferred as (S)-THP based on the detection of major fragment ions and authentic standard (Supplementary Fig. 4B; Supplementary Table 5). (S)-tetrahydropalmatrubine (m/z 342.1701), as a substrate with SiSOMT, generated a peak at 11.28 min (m/z 356.1871) (Supplementary Fig. 4C), which was inferred as (S)-THP based on the detection of major fragment ions of m/z 178 (isoquinoline moiety). These data indicated that SiSOMT regio-selectivity catalyzes O-methylation at C-9 and C-2, and O-methylation events occurred at C-9 and then at C-2.
Discussion
In this study, it was reported, for the first time, identification and characterization of four MTs from Stephania intermedia, including three N-methyltransferases (SiCNMT1, SiCNMT2, and SiCNMT3) and one O-methyltransferase (SiSOMT). SiCNMT1, SiCNMT2, and SiCNMT3 could methylate (R)-coclaurine to produce (R)-N-methylcoclaurine. Notably, SiCNMT2 could further methylate N-methylcoclaurine to form (R)-magnocurarine. Similarly, (R)-norcoclaurine could be catalyzed by SiCNMT2, yielded (R)- N-methylnorcoclaurine and (R)-N, N-dimethylnorcoclaurine, respectively. Furthermore, SiSOMT exhibited high catalytic efficiency and continuously catalyzed (S)-scoulerine to form (S)-THP, which might preliminary reveal the reason for large accumulation of (S)-THP in SI.
Phylogenetic analysis indicated that SiCNMT1 and CjCNMT formed a new clade. In addition, SiSOMT showed 81% sequence identities with CjSOMT and TfSOMT. The four MTs derived from SI have high homology with the CNMT and SOMT of C. japonica, and there were many similarities in catalytic activity (He et al. 2018; Takashi et al. 2010a, b). The phylogenetic relationship between four MTs and other plant MTs provides new insights into the evolutionary recruitment of enzymes in plant alkaloid pathways.
The kinetic parameters of SiCNMT1, SiCNMT2, and SiCNMT3 were determined at pH 7.0 and 37 °C, while SiSOMT was determined at pH 9.0 and 37 °C. SiCNMT1, SiCNMT2, and SiCNMT3 exhibited a Km value for (R)-coclaurine lower than that for (S)-norcoclaurine (265 μΜ), as was reported for EcCNMT in E. californica (Bennett et al. 2018). This indicated that SiCNMT1, SiCNMT2, and SiCNMT3 had better affinity for (R)-coclaurine than for (S)-norcoclaurine (Choi et al. 2001). SiSOMT exhibited a Km value for (S)-scoulerine higher than that determined for SOMT from Opium Poppy and Glaucium flavum (Chang et al. 2015; Dang and Facchini 2012). The catalytic efficiencies (kcat/Km, 47,761 M−1 S−1) of SiSOMT were similar in G. flavum.
Three related NMTs have been characterized, including a coclaurine NMT (CNMT) (Kum-Boo et al. 2002), tetrahydroprotoberberine NMT (TNMT) (Liscombe and Facchini 2007), pavine NMT (PavNMT) (Liscombe et al. 2010). CNMT exhibits little stereoselectivity and can methylate both (R)-coclaurine and (S)-coclaurine, which is in agreement with previous studies. Whereas (R)-coclaurine was the optimal substrate for enzyme activity (Choi et al. 2001; Kum-Boo et al. 2002). The catalyzation and kinetic parameters were measured by (R)-coclaurine and SiSOMT was determined by (S)-scoulerine. CNMT is mainly responsible for the N-methylation of the central 1-benzylisoquinoline intermediate coclaurine (Torres et al. 2016), whereas SiCNMT2 primarily accepts (R)-Coclaurine, (R)-Norcoclaurine and possess successive methylation. Furthermore, SiSOMT which possess successive methylation, reflects the actions of SOMT in Glaucium flavum and Papaver somniferum (Dang and Facchini 2012; Limei et al. 2015).
In this study four MTs have been identified and characterized from Stephania intermedia. The novel MTs provide new possibilities to biocatalytic synthesis of (R)-magnocurarine, (R)-N, N-dimethylnorcoclaurine and (S)-tetrahydropalmatine.
References
Andreas W, Franz H, Kutchan TM, Anton G, Peter M (2006) Biochemical evidence that berberine bridge enzyme belongs to a novel family of flavoproteins containing a bi-covalently attached FAD cofactor. J Biol Chem 281:21276–21285
Bennett MR, Thompson ML, Shepherd SA, Dunstan MS, Herbert AJ, Smith DRM, Cronin VA, Menon BRK, Levy C, Micklefield J (2018) Structure and biocatalytic scope of coclaurine N-methyltransferase. Angew Chem Int Ed Engl 57:10600–10604
Chang L, Hagel JM, Facchini PJ (2015) Isolation and characterization of O-methyltransferases involved in the biosynthesis of glaucine in Glaucium flavum. Plant Physiol 169:1127–1140
Choi KB, Morishige T, Sato F (2001) Purification and characterization of coclaurine N-methyltransferase from cultured Coptis japonica cells. Phytochemistry 56:649–655
Dai WL, Xiong F, Yan B, Cao ZY, Liu WT, Liu JH, Yu BY (2016) Blockade of neuronal dopamine D2 receptor attenuates morphine tolerance in mice spinal cord. Sci Rep 6:38746
Dai WL, Yan B, Jiang N, Wu JJ, Liu XF, Liu JH, Yu BY (2017) Simultaneous inhibition of NMDA and mGlu1/5 receptors by levo-corydalmine in rat spinal cord attenuates bone cancer pain. Int J Cancer 141:805–815
Dang TTT, Facchini PJ (2012) Characterization of three O-methyltransferases involved in noscapine biosynthesis in opium poppy. Plant Physiol 159:618–631
Facchini PJ, De Luca V (2008) Opium poppy and Madagascar periwinkle: model non-model systems to investigate alkaloid biosynthesis in plants. Plant J 54:763–784
Facchini PJ, Penzes C, Johnson AG, Bull D (1996) Molecular characterization of berberine bridge enzyme genes from opium poppy. Plant Physiol 112:1669–1677
Hagel JM, Facchini PJ (2010) Dioxygenases catalyze the O-demethylation steps of morphine biosynthesis in opium poppy. Nat Chem Biol 6:273–275
Hagel JM, Facchini PJ (2013) Benzylisoquinoline alkaloid metabolism: a century of discovery and a brave new world. Plant Cell Physiol 54:647–672
He J, Liu Y, Kang Y, Yang P, Wang Y, Guo J, Huang J (2016) Identification of alkaloids in stephania hainanensis by liquid chromatography coupled with quadrupole time-of-flight mass spectrometry. Phytochem Anal 27:206–216
He SM, Song WL, Cong K, Wang X, Dong Y, Cai J, Zhang JJ, Zhang GH, Yang JL, Yang SC, Fan W (2017) Identification of candidate genes involved in isoquinoline alkaloids biosynthesis in Dactylicapnos scandens by transcriptome analysis. Sci Rep 7:9119
He SM, Liang YL, Cong K, Chen G, Zhao X, Zhao QM, Zhang JJ, Wang X, Dong Y, Yang JL, Zhang GH, Qian ZL, Fan W, Yang SC (2018) Identification and characterization of genes involved in benzylisoquinoline alkaloid biosynthesis in coptis species. Front Plant Sci 9:731
Isabel DP, Facchini PJ (2012) Systematic silencing of benzylisoquinoline alkaloid biosynthetic genes reveals the major route to papaverine in opium poppy. Plant J 72:331–344
Kang DW, Moon JY, Choi JG, Kang SY, Ryu Y, Jin BP, Lee JH, Kim HW (2016) Antinociceptive profile of levo-tetrahydropalmatine in acute and chronic pain mice models: role of spinal sigma-1 receptor. Sci Rep 6:37850
Kum-Boo C, Takashi M, Nobukazu S, Kazufumi Y, Fumihiko S (2002) Molecular cloning and characterization of coclaurine N-methyltransferase from cultured cells of Coptis japonica. J Biol Chem 277:830–835
Limei C, Hagel JM, Facchini PJ (2015) Isolation and characterization of O-methyltransferases involved in the biosynthesis of glaucine in Glaucium flavum. Plant Physiol 169:1127–1140
Lin L, Liu XB, Zhou QY, Liu SS, Zuo MT, Zeng JG, Liu ZY (2018) Characterization of in vitro metabolites of three tetrahydroprotoberberine alkaloids in rat liver S9 by high-performance liquid chromatography/quadrupole time-of-flight mass spectrometry. Rapid Commun Mass Spectrom 32:1540–1548
Liscombe DK, Facchini PJ (2007) Molecular cloning and characterization of tetrahydroprotoberberine cis-N-methyltransferase, an enzyme involved in alkaloid biosynthesis in opium poppy. J. Biol. Chem. 282:14741–14751
Liscombe DK, Ziegler J, Schmidt J, Ammer C, Facchini PJ (2010) Targeted metabolite and transcript profiling for elucidating enzyme function: isolation of novel N-methyltransferases from three benzylisoquinoline alkaloid-producing species. Plant J 60:729–743
Menendez-Perdomo IM, Facchini PJ (2018) Benzylisoquinoline alkaloids biosynthesis in sacred lotus. Molecules 23.
Morishige T, Tsujita T, Yamada Y, Sato F (2000) Molecular characterization of the S-adenosyl-l-methionine:3'-hydroxy-N-methylcoclaurine 4'-O-methyltransferase involved in isoquinoline alkaloid biosynthesis in Coptis japonica. J Biol Chem 275:23398–23405
Morris JS, Facchini PJ (2016) Isolation and characterization of reticuline N-methyltransferase involved in biosynthesis of the aporphine alkaloid magnoflorine in opium poppy. J Biol Chem 291:23416–23427
Nailish S, Facchini PJ (2002) Purification and characterization of norcoclaurine synthase. The first committed enzyme in benzylisoquinoline alkaloid biosynthesis in plants. J Biol Chem 277:33878–33883
Oh JH, Ha IJ, Lee MY, Kim EO, Park D, Lee JH, Lee SG, Kim DW, Lee TH, Lee EJ, Kim CK (2018) Identification and metabolite profiling of alkaloids in aerial parts of Papaver rhoeas by liquid chromatography coupled with quadrupole time-of-flight tandem mass spectrometry. J Sep Sci 41:2517–2527
Purwanto R, Hori K, Yamada Y, Sato F (2017) Unraveling additional O-methylation steps in benzylisoquinoline alkaloid biosynthesis in California poppy (Eschscholzia californica). Plant Cell Physiol 58:1528–1540
Robin AY, Giustini C, Graindorge M, Matringe M, Dumas R (2016) Crystal structure of norcoclaurine-6-O-methyltransferase, a key rate-limiting step in the synthesis of benzylisoquinoline alkaloids. Plant J 87:641–653
Shangguan Y, He J, Kang Y, Wang Y, Yang P, Guo J, Huang J (2018) Structural characterisation of alkaloids in leaves and roots of stephania kwangsiensis by LC-QTOF-MS. Phytochem Anal 29:101–111
Sun R, Song Y, Li S, Ma Z, Deng X, Fu Q, Qu R, Ma S (2018) Levo-tetrahydropalmatine attenuates neuron apoptosis induced by cerebral ischemia–reperfusion injury: involvement of c-Abl activation. J Mol Neurosci 1–9.
Takashi M, Emilyn D, Kum-Boo C, Kazufumi Y, Fumihiko S (2010a) Molecular cloning of columbamine O-methyltransferase from cultured Coptis japonica cells. Eur J Biochem 269:5659–5667
Takashi M, Masanori T, Tomoya T, Fumihiko S (2010b) Molecular characterization of O-methyltransferases involved in isoquinoline alkaloid biosynthesis in Coptis japonica. Proc Jpn Acad 86:757–768
Takayuki I, Ken-Ichi T, Nanae F, Takashi M, Fumihiko S (2007) Overexpression of Coptis japonica norcoclaurine 6-O-methyltransferase overcomes the rate-limiting step in Benzylisoquinoline alkaloid biosynthesis in cultured Eschscholzia californica. Plant Cell Physiol 48:252–262
Tang X, Di X, Zhong Z, Xie Q, Chen Y, Wang F, Ling Z, Xu P, Zhao K, Wang Z, Liu L, Liu X (2016) In vitro metabolism of l-corydalmine, a potent analgesic drug, in human, cynomolgus monkey, beagle dog, rat and mouse liver microsomes. J Pharm Biomed Anal 128:98–105
Torres MA, Hoffarth E, Eugenio L, Savtchouk J, Chen X, Morris JS, Facchini PJ, Ng KK (2016) Structural and functional studies of pavine N-methyltransferase from Thalictrum flavum reveal novel insights into substrate recognition and catalytic mechanism. J. Biol. Chem. 291:23403–23415
Xiao J, Song N, Lu T, Pan Y, Song J, Chen G, Sun L, Li N (2018) Rapid characterization of TCM Qianjinteng by UPLC-QTOF-MS and its application in the evaluation of three species of Stephania. J Pharm Biomed Anal 156:284–296
Yamada Y, Yoshimoto T, Yoshida ST, Sato F (2016) Characterization of the promoter region of biosynthetic enzyme genes involved in berberine biosynthesis in Coptis japonica. Front Plant Sci 7:1352
Zuo AX, Li LI, Rao GX (2011) Studies on the alkaloids from the ethno-remedy plants Stephania Intermedia H. S. Lo. J Yunnan Univ Natl 20:17–19
Acknowledgements
This project was supported by the Grant sponsor: Major Scientific and Technological Specialized Project for “New Drugs Development”; Grant Number: 2012ZX09J12110-06B; “Double First-Class” University project (CPU2018GY32).
Supporting information
Supporting Information 1
Supplementary Table 1—The list of abbreviations and GenBank accession numbers for sequences used to construct the phylogenetic tree.
Supplementary Table 2—List of primers used for cloning in this study.
Supplementary Table 3—Substrate range of four Methyltransferases.
Supplementary Table 4—Kinetic data for SiCNMT1, SiCNMT2, SiCNMT3 and SiSOMT.
Supplementary Table 5—HPLC-DAD-Q-TOF MS/MS data for the reaction products of recombinant methyltransferases assay with various substrates.
Supplementary Figure 1—Steady-state enzyme kinetics of affinity-purified recombinant SiCNMT1 (A), SiCNMT2 (B), SiCNMT3(C) for (R)-coclaurine and SiSOMT for (S)-scoulerine (D). Kinetic constants were determined by fitting initial velocity versus substrate concentration to the Michaelis-Menten equation.
Supplementary Figure 2—Total ion chromatograms (TICs) of the reaction products by SiCNMT1, SiCNMT2, SiCNMT3. For each sample, the top (control) corresponds to boiled enzyme (10 min) as the negative control (A), the methylation activity of SiCNMT1 (B), SiCNMT2 (C), and SiCNMT3 (D) on (R)-coclaurine.
Supplementary Figure 3—TICs of the reaction products by SiCNMT2. For each sample, the top (control) corresponds to boiled enzyme (10 min) as the negative control (A), the methylation activity of SiCNMT2 (B) on (R)-norcoclaurine.
Supplementary Figure 4—TICs of the reaction products by SiSOMT. The top (control) corresponds to boiled enzyme (10 min) as the negative control (A), the methylation activity of SiSOMT on (S)-scoulerine, (S)-tetrahydrocolumbamine (B), and (S)-tetrahydropalmatrubine (C).
Supporting Information 2
Nucleotide and predicted protein sequence of four methyltransferases.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflicts of interest
The authors declare that there are no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zhao, W., Shen, C., Zhu, J. et al. Identification and characterization of methyltransferases involved in benzylisoquinoline alkaloids biosynthesis from Stephania intermedia. Biotechnol Lett 42, 461–469 (2020). https://doi.org/10.1007/s10529-019-02785-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-019-02785-0