Abstract
Four E. coli-Bifidobacterium shuttle vectors were constructed using Bifidobacterium plasmids, pB44 and pB80. The vectors carry two bifidobacterial promoters, a signal peptide-encoding sequence, sec2, of Bifidobacterium breve, and a transcriptional terminator from hup gene of Bifidobacterium longum. Functionality of the constructs were tested using human FGF-2 gene. The expression of FGF-2 was detected by Western blotting in B. breve transformed with three of the vectors. The highest amount of FGF-2 was produced upon transformation with pESH86, which is a pB80-based plasmid carrying FGF-2 under control of a hup promoter (Phup). Similarly, the level of FGF-2 mRNA transcribed from pESH86 was approximately threefold higher, 882 ± 70 AU (arbitrary units), when compared to those transcribed from pB44-based pESH46 (Phup) (289 ± 65 AU) and pESH47 (Pgap) (282 ± 37 AU). These results suggest the vectors have the potential for production of exported fusion proteins in bifidobacteria and the expression levels can be regulated through the employment of different bifidobacterial promoters and/or replicons.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Genus Bifidobacterium are anaerobic Gram-positive rods that are generally non-pathogenic. They constitute a prominent component of human indigenous intestinal and vaginal microflora that exert multiple beneficial effects toward the host (Biavati and Mattarelli 2006). During the last decade, the commercial interest in bifidobacteria has also grown due to their use as “health-promoting” components of various dairy foods and probiotic formulations. The whole-genome sequencing projects (Schell et al. 2002) started to revolutionize our understating of physiology of the bifidobacteria and the mechanisms underlying their interactions with the host and pathogens. However, the ability to genetically manipulate bifidobacteria is essential for the confirmation of functional genomics data through generation of specific mutants, future systems biology and, possibly, metagenomics studies. The proper tools of genetic access are also required in order to design bifidobacteria-based biotherapeuticals such as vaccines, targeted immunomodulators and antimicrobials, as well as the technologically superior probiotic cultures for the industry. Currently, there are few reports describing the production of foreign proteins in bifidobacteria (Moon et al. 2005; Takata et al. 2006; Reyes Escogido et al. 2007). Apparently, the progress in this area is hampered by lack of convenient and freely available genetic tools, namely versatile expression vectors, high-efficiency transformation protocols, and suitable host strains. Recently, we have sequenced and characterized four cryptic plasmids isolated from infant bifidobacteria and employed them for construction of several sets of shuttle cloning vectors based on two distinct replicons (Smeianov et al. 2002; Gibbs et al. 2006). Here, we describe further improvement of these vectors into a series of modular expression vehicles capable of driving constitutive expression and secretion of cloned human basic Fibroblast Growth Factor (FGF-2) gene in bifidobacteria.
Materials and methods
Bacterial strains and growth conditions
Bifidobacterium breve UCC2003 (MacConaill et al. 2003), Bifidobacterium longum VMKB44 (Korschunov et al. 1998), and Bifidobacterium bifidum ATCC 15696 were propagated anaerobically either in MRS (Difco), or in TPY (Biavati and Mattarelli 2006) media supplemented with 0.05% L-cysteine · HCl and 5 μg erythromycin/ml, when needed. E. coli strain XL-1Blue (Stratagene) was grown in LB medium supplemented with 100 μg ampicillin/ml or 500 μg erythromycin/ml, when needed.
DNA isolation and manipulation
Total DNA from bifidobacteria to be used for PCR was isolated using 5 min boiling lysis in TE buffer (Sambrook and Russell 2001). Plasmid DNA from E. coli was isolated using an alkaline lysis procedure as described elsewhere (Sambrook and Russell 2001). Plasmid DNA from bifidobacteria was isolated using the E. coli protocol modified through the incorporation of a lysozyme treatment step, 30 mg lysozyme/ml in Solution I, 30 min at 37°C, before the addition of Solution II. Restriction enzymes, T4 DNA polymerase, T4 DNA ligase, and Taq/Pfu DNA polymerase mix were purchased from Sibenzyme (Moscow, Russian Federation) and used according to the manufacturer’s protocols. The PCR primers used to amplify B. longum VMKB44 hup promoter/Translation Initiation Region (TIR), B. longum VMKB44 gap promoter/TIR, B. longum VMKB44 hup terminator, and B. breve UCC2003 sec2 signal peptide coding sequences were as follows (underlined are recognition sites for restriction enzymes KpnI, NcoI, BamHI, EcoRI, and NdeI, respectively): Phup-F (CGGTACCTACTGGCTGCGTATTCCG), Phup-R (GCCCATGGAGCATCCTTCTTGGGT), Pgap-F (GCGGTACCTGATGATTCGAGACATTCCT), Pgap-R (TTACCATGGTATTCTCCCTTGTAGGGTGG), Thup-F (CGGATCCTGACCTTCTGCTCGTA), Thup-R (CGAATTCGCTGAACTAGTCCGGA), Sec2-F (GGCCATGGAACACATGAAGATGTTC), Sec2-R (GTACATATGCAATGCCACCCAGTCG). For use in cloning, restriction fragments and PCR products were purified through 1% (w/v) agarose gels followed by gel-elution using DNA extraction kit (Fermentas, Vilnius, Lithuania). The overview of construction of plasmids for this study is given in Fig. 1. Plasmid constructs were verified by sequencing of inserts and junction points on an ABI Prizm 3100 automated sequencer at Pynny Joint Stock Company (Moscow, Russian Federation; www.punny.ru).
Construction of expression/secretion vectors used in the study. P, promoter regions; T, transcriptional terminator regions; ery194, erythromycin resistance gene; bla, β-lactamase gene; repA, replication initiator gene of pB80; repB, replication initiator gene of pB44. Please see Sect. “Results and discussion” section for additional description
Transformation of bacteria
E. coli strains were transformed using the CaCl2 protocol (Sambrook and Russell 2001). B. breve UCC2003 was transformed via electroporation using the following protocol: several well-isolated colonies were inoculated into TPY-cysteine · HCl broth supplemented with 0.5 M mannitol and the culture was incubated for 20 h; cells were pelleted and washed twice in a buffer containing 0.5 M mannitol and 1 mM ammonium citrate (pH 6.0), resuspended in the same buffer, and electroporated at 200 Ω, 25 μF, 12 kV/cm, using GenePulser instrument (BioRad). Transformants were selected on TPY-cystein plates with 0.5 M mannitol (optional) and 2 μg erythromycin/ml. B. bifidum ATCC 15696 was transformed according to Argnani et al. (1996).
Quantitative RT-PCR
Mid-growth cultures of B. breve grown in 5 ml of TPY broth were pelleted and resuspended in 100 μl TES buffer (50 mM NaCl, 100 mM Tris/HCl pH 8.0, 70 mM EDTA) supplemented with 30 mg lysozyme/ml. After incubation at 37°C for 1 h, total RNA was extracted from protoplasts using YellowSolve RNA isolation kit (Clonogene, St.-Petersburg, Russian Federation). The integrity of RNA was confirmed by agarose gel electrophoresis, and the amounts of rRNA in the samples were calculated based on the bands intensity using ImageJ software (available from http://rsb.info.nih.gov/ij/). Synthesis of first strand cDNA was accomplished with RevertAid reverse trascriptase (Fermentas) and FGF2-R primer (see below). Quantitative PCR was performed on an ANK-32 thermocycler (Syntol, Moscow, Russian Federation) using 2.5× EvaGreen premixed PCR components, primers FGF2-F (ATGGCAGCAGGATCAATAAC) and FGF2-R (GTACCAGGAGGTGTATTTAC), and the following program: initial denaturation at 95°C for 300 s; 40 cycles of 94°C for 20 s, 52°C for 20 s, and 72°C for 30 s. Specificity of reaction products was confirmed by melting temperature analysis (from 70 to 95°C with 0.5°C/15 s increments). Quantification was accomplished by comparing C(t) values to the calibration curve generated from serial tenfold dilutions of plasmid pESH46. The obtained values were normalized to the amounts of rRNA present in each sample.
Western blotting
Culture supernatants were filtered through 0.22 μm cellulose acetate membranes and precipitated by the addition of trichloroacetic acid to give 15% (w/v). Pellets obtained after centrifugation were washed with acetone, dried and resuspended in 1× SDS gel-loading buffer. SDS-PAGE electrophoresis and western blotting were performed as described elsewhere (Sambrook and Russell 2001). Proteins were transferred onto Hybond P (GE-Amersham) and, after blocking nonspecific binding in PBS containing 0.2% (v/v) of Tween20, were sequentially hybridized with goat anti-human FGF-2 antibodies (sc1390, SantaCruz Biotechnology, SantaCruz, CA) and HRP-conjugated rabbit anti-goat IgG antibodies (sc2768, SantaCruz). Blots were developed using ECL kit (GE-Amersham).
Results and discussion
Construction of FGF-2 expression/secretion cassettes using pB44- and pB80-based vectors
Our choice of human FGF-2 (bFGF) as a model protein was due to its relatively small size and stability at acidic pH, which is characteristic for bifidobacterial culture supernatant. Construction of the FGF-2-producing Bifidobacterium strains may also have therapeutic applications in treatment of inflammatory bowel disease and acute intestinal radiation injury since these condition can be ameliorated by the bFGF treatment in the murine experimental models (Paris et al. 2001; Kojima et al. 2007).
Previously, we have utilized two bifidobacterial plasmids, pB44 and pB80, to construct a series of E. coli-Bifidobacterium shuttle vectors (Smeianov et al. 2002; Gibbs et al. 2006; GenBank accession numbers AY066026 and DQ305402). Two of these vectors, pSUW64/123 and pESH80, derived from pB44 and pB80, respectively, both of which harbor ery194 as a selectable marker, were employed in the construction of the expression vectors.
To obtain the first constitutive expression cassette, promoter/TIR and terminator regions of hup gene encoding the histone-like protein HU (Takeuchi et al. 2002) were PCR-amplified from B. longum VMKB44 total DNA. These two PCR products were cloned into pESH44 resulting in pESH45 (Fig. 1). The former plasmid, pESH44, is pSUW64/123 from which the BamHI site was removed by T4-polymerase treatment of the BamHI-linearized plasmid followed by the blunt-end self-ligation. PCR-amplification of a region encoding for a signal peptide and the first 11 N-terminal amino acids of a mature polypeptide of the bifidobacterial Sec2 secreted protein (MacConaill et al. 2003) was performed using B. breve UCC2003 total DNA as a template. The signal peptide-encoding fragment was inserted into pESH45 simultaneously with a NdeI/BamHI fragment of pλFGFB, which carries the synthetic human FGF-2 gene (Seeger et al. 1995). This construct, named pESH46, contains a chimeric open reading frame coding for Sec2-FGF-2 fusion protein under the control of Phup, TIR, and a transcriptional terminator.
To construct a vector with different constitutive promoter, the fragment correspondent to the promoter/TIR of B. longum VMKB44 gene gap (Klijn et al. 2006) was PCR-amplified from the total DNA and was used to replace a Phup/TIR fragment in pESH46 resulting in the pESH47 vector.
Our prior experiments showed that pESH80, albeit larger than pSUW64/123, demonstrates higher segregational stability, and replicates at higher copy numbers in B. breve UCC2003 (Shkoporov et al., submitted). To obtain the pESH80-based expression constructs, the Sec2-FGF-2 cassettes along with ery194 gene were transferred from pESH46 and pESH47 into pESH80 as ClaI/EcoRI fragments, resulting in plasmids pESH86 and pESH87, respectively.
Transformation of bifidobacteria and expression of FGF-2
All four vectors constructed, i.e., pESH46, pESH47, pESH86, and pESH87 transformed B. breve UCC2003 to erythromycin resistance (Fig. 2a, b), and pESH46 (contains Phup/TIR-Sec2-FGF2 cassette) also transformed B. bifidum ATCC 15696. However, B. breve UCC2003 (pESH87) showed impaired growth, possibly due to toxicity associated with the higher expression level, and was excluded from the further study. One may further hypothesize that this, in turn, is due to the higher copy number of the pESH80 backbone as compared to that of pSUW64/123.
Transformation of Bifidobacterium breve UCC2003 with Sec2-FGF-2 fusion protein-encoding plasmids and expression of target protein therein (a) Colony PCR analysis of transformants using primers FGF2-F and FGF2-R; lanes 1–3, transformation with pESH46; lanes 4–6, transformation with pESH47; lanes 7–9 transformation with pESH86; lane 10, negative control. (b) Isolation of plasmid DNA from transformed B. breve: lane 1, supercoiled DNA ladder (Invitrogen); lanes 2–4, pESH86; lane 5 pESH47; lane 6, pESH46. (c) Immunoblotting analysis of FGF-2 expression in the culture supernatants of transformed strains; Lanes 1–2, pESH46; lanes 3–4, pESH47; lanes 5–6, pESH80 (negative control); lanes 7–8, pESH86
The results of immunodetection of FGF-2 in bifidobacteria are shown in Fig. 2c. The FGF-2 immunoreactivity was only found in supernatants of B. breve UCC2003, but not in the cytoplasmic fraction. The target protein was not detected in either supernatant or cytoplasm of B. bifidum ATCC 15696. The expression level of FGF-2 appeared to be the highest in pESH86-transformed B. breve UCC2003, followed by the pESH47- and pESH46-transformed cells. In the supernatants of pESH86 transformants, a large portion of the immunoreactivity was represented by a band of ca. 19 kDa that corresponds to calculated molecular weight of a mature fusion protein. However, an additional, lower molecular weight band of ca. 14 kDa was also detected in all cases, and this form was a predominant one detected in the pESH46 transformants. The reason for appearance of the 14 kDa aberrant form is not clear, but it may be resulted from either premature translation termination or posttranslational degradation of the fusion protein. When the pSUW64/123 was used as the backbone, Pgap/TIR (pESH47) led to the higher production of FGF-2 than hup regulatory sequences (pESH46).
The results of quantitative RT-PCR for FGF-2 transcript in B. breve transformants are shown on Fig. 3. In line with immunoblotting results, the cells transformed with pESH86 generated the highest numbers of FGF-2 transcripts. The expression levels in the pESH46 and pESH47 transformants were more than 3-fold lower than in the case of pESH86, and were very close to each other. Therefore, it appears that the higher amount of the immunoreactive protein observed in the pESH47 transformants may be due to the higher activity of gap TIR as compared to hup TIR present in pESH46.
Quantitative RT-PCR analysis of Bifidobacterium breve transformed with three Sec2-FGF-2 expression plasmids (a) Calibration curve generated from serial tenfold dilutions of plasmid pESH46 (b) Determination of relative abundance of cDNA generated from FGF-2 transcripts in B. breve transformed with three different plasmids. Amounts of the transcripts normalized to rRNA levels were 289 ± 65 arbitrary units (AU) (pESH46 transformants), 282 ± 37 AU (pESH47 transformants), and 882 ± 70 AU (pESH86 transformants). The experiments were performed in triplicate; bars on the graphs indicate standard deviation
In summary, the results show that the pSUW64/123- and pESH80-derived expression vectors are able to drive the production of human FGF-2 fused to the Sec2 signal peptide in B. breve UCC2003. The reason for the absence of detectable FGF-2 expression in the B. bifidum ATCC 15696 requires further investigation and emphasizes the importance of the host strain choice for the production of foreign proteins in bifidobacteria.
References
Argnani A, Leer RJ, van Luijk N, Pouwels PH (1996) A convenient and reproducible method to genetically transform bacteria of the genus Bifidobacterium. Microbiology 142:109–114
Biavati B, Mattarelli P (2006) The family Bifidobacteriaceae. In: Dworkin M, Falkow S, Rosenberg E, Schleifer KH, Stackebrandt E (eds) The prokaryotes, vol 3. Springer, New York, pp 322–382
Gibbs MJ, Smeianov VV, Steele JL, Upcroft JA, Efimov BA (2006) Two families of Rep-like genes that probably originated by inter-species recombination, are represented in viral, plasmid, bacterial and parasitic protozoan genomes. Mol Biol Evol 23:1097–1100
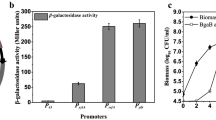
Klijn A, Moine D, Delley M, Mercenier A, Arigoni F, Pridmore RD (2006) Construction of a reporter vector for the analysis of Bifidobacterium longum promoters. Appl Environ Microbiol 72:7401–7405
Kojima T, Watanabe T, Hata K, Nagawa H (2007) Basic fibroblast growth factor enema improves experimental colitis in rats. Hepatogastroenterology 54:1373–1377
Korschunov VM, Efimov BA, Smeianov VV, Bainov NA, Pikina AP (1998) Treatment of experimental acute radiation disease in mice with probiotics, quinolones and general gnotobiological isolation. Armed Forces Radiobiology Research Institute, Defense Nuclear Agency, Bethesda, USA. Available at http://stinet.dtic.mil/oai/oai?verb=getRecord&metadataPrefix=html&identifier=ADA359101
MacConaill LE, Fitzgerald GF, Van Sinderen D (2003) Investigation of protein export in Bifidobacterium breve UCC2003. Appl Environ Microbiol 69:6994–7001
Moon GS, Pyun YR, Park MS, Ji GE, Kim WJ (2005) Secretion of recombinant pediocin PA-1 by Bifidobacterium longum, using the signal sequence for bifidobacterial alpha-amylase. Appl Environ Microbiol 71:5630–5632
Paris F, Fuks Z, Kang A, Capodieci P, Juan G, Ehleiter D, Haimovitz-Friedman A, Cordon-Cardo C, Kolesnick R (2001) Endothelial apoptosis as the primary lesion initiating intestinal radiation damage in mice. Science 293:293–297
Reyes Escogido ML, De León Rodríguez A, Barba de la Rosa AP (2007) A novel binary expression vector for production of human IL-10 in Escherichia coli and Bifidobacterium longum. Biotechnol Lett 29:1249–1253
Sambrook J, Russell DW (2001) Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory Press, Cold Spring Harbor
Schell MA, Karmirantzou M, Snel B, Vilanova D, Berger B, Pessi G, Zwahlen MC, Desiere F, Bork P, Delley M, Pridmore RD, Arigoni F (2002) The genome sequence of Bifidobacterium longum reflects its adaptation to the human gastrointestinal tract. Proc Natl Acad Sci USA 99:14422–14427
Seeger A, Schneppe B, McCarthy JEG, Deckwer WD, Rinas U (1995) Comparison of temperature- and isopropyl-beta-D-thiogalactopyranoside-induced synthesis of basic fibroblast growth factor in high-cell-density cultures of recombinant Escherichia coli. Enzyme Microb Technol 17:947–953
Smeianov VV, Efimov BA, Korschunov VM, Steele JL (2002) Construction of the E. coli-Bifidobacterium shuttle vectors based on two distinctive Bifidobacterium replicons. American Society for Microbiology 102nd General Meeting, Salt Lake City, Utah:H-7
Takata T, Shirakawa T, Kawasaki Y, Kinoshita S, Gotoh A, Kano Y, Kawabata M (2006) Genetically engineered Bifidobacterium animalis expressing the Salmonella flagellin gene for the mucosal immunization in a mouse model. J Gene Med 8:1341–1346
Takeuchi A, Matsumura H, Kano Y (2002) Cloning and expression in Escherichia coli of a gene, hup, encoding the histone-like protein HU of Bifidobacterium longum. Biosci Biotechnol Biochem 66:598–603
Acknowledgements
Authors would like to thank Dr. Ursula Rinas for providing plasmid pλFGFB and Dr. Douwe van Sinderen for strain B. breve UCC2003. We are also thankful to Ryan Algino for his help in preparing the manuscript. This work was supported in part by Federal agency of Health Care and Social Development, Russian Federation (grant #521-PD).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Shkoporov, A.N., Efimov, B.A., Khokhlova, E.V. et al. Production of human basic fibroblast growth factor (FGF-2) in Bifidobacterium breve using a series of novel expression/secretion vectors. Biotechnol Lett 30, 1983–1988 (2008). https://doi.org/10.1007/s10529-008-9772-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-008-9772-8