Abstract
The genus Cyclamen (family Myrsinaceae) contains about 20 species, most of which occur in the Mediterranean region. Turkey has critically important Cyclamen genetic resources. Molecular characterization of plant materials collected from different regions of Turkey in which Cyclamen species grow naturally, namely Adana, Antalya, Aydın, Muğla, İzmir, Denizli, Kahramanmaraş, Osmaniye, Eskişehir, Trabzon, and Rize provinces, was performed using RAPD and SRAP markers. DNA was successfully amplified by 30 RAPD primers and 14 SRAP primer pairs. Among the 470 bands generated by the RAPD primers, 467 were polymorphic. The number of bands detected by a single primer set ranged from 11 to 22 (average of 15.6). The percentage polymorphism was 99.3 % based on the RAPD data. In the SRAP analysis, a total of 216 bands were generated, showing 100 % polymorphism. The number of bands detected by a single primer set ranged from 9 to 22 (average of 15.4). All data were scored and UPGMA dendrograms were constructed with similar results in both marker systems, i.e., different species from nine provinces of Turkey were separated from each other in the dendrograms with the same species being clustered together.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cyclamen (Cyclamen spp.; Myrsinaceae) is among the most popular flowering pot plants, and many cultivars derived from C. persicum are grown commercially (Kitamura et al. 2012). The 20 cyclamen taxa originate from the Mediterranean region (Aka Kacar et al. 2013) and under natural conditions typically grow under trees and bushes. Ten Cyclamen species grow naturally in Turkey, five of which are endemic to Turkey, which is a genetic epicenter of many plant species. The genus Cyclamen is a particularly interesting and traceable genus to examine phylogeographic patterns of differentiation. Although traditionally placed in the Primulaceae, molecular evidence suggested that the genus Cyclamen may actually be nested in the Myrsinaceae (Kallersjo et al. 2000). There are 20 species in the genus, 17 of which are circum-Mediterranean (Grey-Wilson 1997). In the Mediterranean Basin, several species have a widespread distribution with allopatric populations on different islands and continental land masses (Gielly et al. 2001).
There are few studies that have investigated the genetic relationships of Cyclamen genotypes by using molecular markers. Naderi et al. (2009) used randomly amplified polymorphic DNA (RAPD) markers to determine inter- and intra-specific genetic diversity among cyclamen accessions collected from different parts of Iran. The genetic diversity of natural Turkish C. alpinum populations was investigated by using RAPD markers (Taskin et al. 2012). In another study, in order to make classifications of the genus Cyclamen, three regions of DNA were selected for sequencing from two genomic compartments: cpDNA trnL intron, nrDNA ITS1, and ITS2 regions. The resulting molecular data were also combined with published morphological data (Compton et al. 2004). In addition to molecular studies, there are several reports on morphological data on Cyclamen (Anderberg 1994; Debussche and Thompson 2002; Aalaey et al. 2007; Curuk et al. 2015).
RAPD markers, which result from the polymerase chain reaction (PCR) amplification of genomic DNA fragments using short oligonucleotides (usually 10-mers) of arbitrary sequence as primers, provide a fast and easy approach for many purposes in plant genetic analysis (Aka Kacar et al. 2005). Sequence-related amplified polymorphism (SRAP), a PCR-based marker system (Li and Quiros 2001), is a simple and efficient marker system that can be adapted for a variety of purposes in different crops, including map construction, gene tagging, genomic and cDNA fingerprinting, and map-based cloning (Amar et al. 2011). It has several advantages over other systems: it is simple, has a reasonable throughput rate, discloses numerous co-dominant markers, targets open-reading frames (ORFs), and allows easy isolation of bands for sequencing (Uzun et al. 2009).
The aim of this study was to determine the genetic diversity of cyclamen genotypes collected from nine provinces of Turkey using RAPD and SRAP markers.
Materials and Methods
This study was carried out at the Department of Horticulture, Faculty of Agriculture, University of Çukurova, Sarıçam, Turkey.
Plant Materials
Plant material was collected from Adana, Antalya, Aydın, Muğla, İzmir, Denizli, Kahramanmaraş, Osmaniye, Eskişehir, Trabzon, and Rize provinces of Turkey (Table 1). A total of 95 cyclamen genotypes were collected from different Cyclamen species (C. pseudibericum, C. cilicium, C. persicum, C. graecum, C. mirabile, C. hederifolium, C. alpinum, C. coum, C. intaminatum, and C. parviflorum).
DNA Isolation
Leaves from all samples were immediately frozen in liquid nitrogen and stored at −80 °C. High molecular weight genomic DNA was extracted from the leaf of each sample following the CTAB protocol for minipreps (Edwards et al. 1991). DNA concentration was measured using a NanoDrop (ND 100) spectrophotometer (NanoDrop Technologies, Inc., Wilmington, DE, USA) and gel electrophoresis. DNA was diluted in water to a final concentration of 50 ng/μL and stored at −20 °C.
RAPD Analysis
Fifty RAPD 10-mer primers (Operon Technologies, Almeda, CA, USA) were initially tested. Primers that produced polymorphic bands were used to amplify all the 95 genotypes studied. Thirty primers which were found to be polymorphic (Table 2) were used to generate RAPD markers. Amplification reactions were performed in 9-μL volumes containing 2X PCR Mastermix (Fermentas K0171, Waltham, MA, USA), 1 U Taq DNA polymerase (Fermentas EP0402), 25 mM MgCl2, 30 ng of the primer, and 15 ng of cyclamen DNA. Mixtures were assembled at 0 °C, transferred to a thermal cycler, and then precooled to 4 °C. The amplification was carried out in a model Master Gradient thermal cycler (Eppendorf, Hauppauge, NY, USA) using an optimized in-house program consisting of an initial denaturation step of 2 min at 94 °C, and then 45 cycles of 2 min at 94 °C, 1 min at 37 °C, 2 min at 72 °C, followed by a 10-min elongation step at 72 °C. PCR products were stored at 4 °C before analysis. Amplification products were separated by electrophoresis on 1.5 % agarose gels and 0.5 g/mL ethidium bromide in 1X TAE buffer (40 mM Tris–acetate, 1 mM EDTA, pH 8.0) for 3 h at 70 V. The fragment patterns were photographed under UV light for further analysis. A 1-kb DNA ladder (Fermentas) was used to determine the fragment size.
SRAP Analysis
All SRAP primer combinations (Table 3) were initially screened with the 95 genotypes. The 14 primer combinations producing scorable polymorphic bands were used to amplify all 95 genotypes (Table 1). Amplification reactions were done in volumes of 22 μL containing 2× PCR Mastermix (Fermentas K0171), 1 U Taq DNA polymerase (Fermentas EP0402), MgCl2, 25 mM of each primer, and 125 ng of cyclamen DNA. The mixtures were assembled at 0 °C and then transferred to a thermal cycler, precooled to 4 °C. The amplification was carried out in a model Master Gradient thermal cycler (Eppendorf) using a program consisting of an initial denaturation step of 5 min at 94 °C, and then 5 cycles of 1 min at 94 °C, 1 min at 35 °C, 2 min at 72 °C, and then 35 cycles of 1 min at 94 °C, 1 min at 50 °C, and 2 min at 72 °C, followed by a 10-min elongation step at 72 °C. PCR products were stored at 4 °C before analysis. Amplification products were separated by electrophoresis on 2.5 % agarose gels and 0.5 g/mL ethidium bromide in 1X TAE buffer for 3.5 h at 110 V. The fragment patterns were photographed under UV light for further analysis. A 100-bp DNA ladder was used as the molecular standard in order to confirm the appropriate SRAP markers.
Data Analysis
Reproducible SRAP and RAPD profiles were scored manually in the binary mode with 1 indicating the presence and 0 indicating the absence of a band, and then the data were used to generate a pair-wise similarity matrix using Jaccard’s coefficient (Jaccard, 1908). The unweighted pair group method using UPGMA was employed to create the clustering dendrograms using the NTSYS-PC program (version 2.02i) (Rohlf 1998). The principle coordinates (PCoA) analysis was performed based on the same similarity matrix using the PAST software (Hammer et al. 2001). Polymorphism information content (PIC) values were calculated according to Smith et al. (1997), using the algorithm for RAPD primers and SRAP primer combinations as follows: PIC = 1−Σfi2, where fi2 is the frequency of the ith allele.
Results
The genetic diversity among the 95 cyclamen genotypes was evaluated by RAPD (Fig. 1a) and SRAP (Fig. 1b) markers. Amplification was successful with 30 RAPD primers and 14 SRAP primer pairs assayed. Ninety-five cyclamen genotypes were screened for RAPD markers using 51 primers in a PCR-based DNA amplification procedure. Thirty primers produced clear and good amplification. All RAPD primers that produced polymorphic bands were used to generate RAPD markers with all genotypes (Table 2). Among the 470 bands generated by the 30 selected RAPD primers, 467 were polymorphic. The number of bands detected by a single primer set ranged from 11 to 22, with an average of 15.6. The rate of polymorphism was calculated as 99.3 % among the 95 cyclamen genotypes based on RAPD data. PIC values ranged between 0.72 (UBC16) and 0.97 (UBC48) for RAPD data.
In SRAP analysis, 49 SRAP primer combinations were screened. In total, 14 SRAP primer combinations were determined and used to differentiate the 95 cyclamen genotypes. From the SRAP analysis, a total of 216 bands were generated and the rate of polymorphism was calculated as 100 %. The number of bands detected by a single primer set ranged from 9 to 22, with an average of 15.4. PIC values ranged between 0.70 (ME3F X EM2R) and 0.95 (ME6F X EM1R) for SRAP data (Table 4). PIC values were higher than 0.6 for both marker systems. The average level of stable polymorphisms was very good, demonstrating that RAPDs and SRAPs markers were useful to discriminate all Cyclamen genotypes.
Discussion
The number of bands detected by a single primer set and the rates of polymorphism for both RAPD and SRAP analysis were higher than those of many previous reports. Taskin et al. (2012) investigated the genetic diversity among six C. alpinum populations in Turkey with 15 RAPD primers. They reported 62.16 % polymorphism with a total of 190 bands, averaging 9.5 bands/locus. In another study, a total of 122 RAPD bands and an average of 11.5 bands/locus were obtained for Iranian C. persicum and C. com (Naderi et al. 2009).
There are only few reports on the molecular characterization of Cyclamen genotypes in the world, but there are many reports explaining the genetic relationship of ornamental and other horticultural plants by RAPD and SRAP markers.
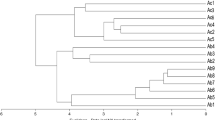
The similarity coefficient ranged from 0.27 to 0.80 in RAPD analysis and from 0.28 to 0.91 in SRAP analysis. Cluster analysis (UPGMA) employing RAPD and SRAP data resulted in dendrograms and PCoA scatters shown in Figs. 2,3 for RAPD and Figs. 4,5 for SRAP, respectively.
Evaluation of the dendrograms indicates that similar results were obtained for both marker systems. Different cyclamen genotypes collected from different regions of Turkey were separated from each other in dendrograms while cyclamen genotypes belonging to same species were clustered together. C. persicum genotypes were clustered together. Similarly, C. coum genotypes were clustered in the same branch, but these two groups of genotypes were separated from each other in both marker systems. In another study (Naderi et al. 2009), a total of 26 Iranian C. persicum and C. coum genotypes were used to investigate the genetic diversity by RAPD markers. The similarity coefficient among the genotypes used in that study was between 0.99 and 0.08. The genotypes were clustered into three groups and the three clusters were 68 % similar. In group A, five wild accessions were separated from other accessions with only 15 % similarity. The lowest similarity in this group was obtained between wild C. coum from Lahijan and other wild accessions from different regions of Chaloos. In group B, there were 18 commercial genotypes with divergent flower color. These monotype commercial plants did not show identical profiles, but were separated from three commercial C. persicum genotypes (group C) from Chardangeh. The similarity matrix indicated that the lowest genetic similarity (0.08) was between a wild accession WL26 and a white commercial accession, Cw14 and the highest similarity (0.99) was between Cp12 (C. persicum cv. ‘Laser Orchid’) and Cp15 (cv. ‘Laser Pink’) as well as Cr16 (cv. ‘Sierra Deep Rose’) and Cr19.
Cyclamen pseudibericum genotypes collected from southern Turkey were also clustered together, but the differences were determined based on location. C. pseudibericum genotypes obtained from Adana districts and Kahramanmaraş genotypes were identified in a separate sub-group. Similarly, C. hederifolium genotypes from geographically different locations were clustered together. C. alpinum genotypes collected from southwestern Turkey were clustered together in both dendrograms. Taskin et al. (2012) studied six natural C. alpinum populations collected from southern and southwestern areas of Turkey by RAPD markers to understand genetic diversity. They determined that C. alpinum genotypes have a narrow genetic diversity. Our findings are in agreement with results of that study. In our study, both RAPD and SRAP markers indicated that C. alpinum genotypes could not be separated from each other, hence their clustering together. One of the C. intaminatum genotypes collected from the central Anatolia region (center) of Turkey was separated from other C. intaminatum in SRAP dendrogram. C. intaminatum genotypes were divided into two different sub-groups in RAPD dendrogram. C. intaminatum genotypes were clustered together with C. cilicium genotypes. C. intaminatum genotypes are morphologically similar to C. cilicium. C. intaminatum has a confusing past. It was first discovered by E.K. Balls and collected as number EKB669a in June 1934. However, he simply described it as C. cilicium var. and although it was briefly erroneously referred to as C. alpinum, it was not until 1978 that it received a formal name as C. cilicium var. intaminatum Meikle. Undoubtedly, the plant is closely allied to C. cilicium and shared the chromosome count 2n = 30, but in 1988, Grey-Wilson recognized that it was distinct and elevated it to specific status as C. intaminatum (Grey-Wilson 1997).
C. graecum genotypes collected from southwest Turkey (Antalya) were placed in the same group in both RAPD- and SRAP-derived dendrograms and showed narrow genetic diversity. C. persicum and C. graecum genotypes were clustered in the same sub-branches for both marker systems. Based on the nrDNA ITS, cpDNA trnL intron sequence data performed by Compton et al. (2004), C. persicum and C. graecum have been detected to be close to each other.
Curuk et al. (2015) investigated a total of 27 phenotypic characters (13 flower, 11 leaf, 2 plant, and 1 tuber) and 13 quantitative traits (7 flower, 5 leaf, and 1 tuber) in C. persicum, C. cilicium, C. coum, and C. pseudibericum species collected from different parts of Turkey. Based on the principal component analysis, the grouping of characters was determined using species-specific clusters, although one or two clusters could not differentiate species, indicating that morphological and cluster analyses alone are not enough for characterizing this complex Cyclamen germplasm and that molecular techniques may reveal more intricate and useful relationships.
Molecular markers have also mainly been used for testing genetic purity in cyclamen seeds (Zhang et al. 1997) and somaclonal variation within C. persicum callus (Laura et al. 2003; Aka Kacar et al. 2013) as well as in molecular characterization studies. However, there are few studies on the molecular characterization of Cyclamen by RAPD. In addition, this study represents the first attempt at the use of SRAP markers for cyclamen. SRAP and RAPD markers were equally powerful tools to separate different cyclamen species and/or genotypes, and in order to determine genetic diversity among the same species, different markers can be used. Especially, SSR markers can be employed for the genetic characterization of Cyclamen species, but there are no currently available SSR markers for cyclamens. The development of SSRs is an important and necessary issue for cyclamen molecular studies.
The conservation of plant genetic resources is an important issue for Turkish Cyclamen. At first, these genetic resources should be accurately characterized allowing them to be preserved in vitro or in situ (Jalali et al. 2012).
References
Aalaey M, Naderi R, Khalighi A (2007) Morphological categorization of some cyclamens from Iran through studying and comparing the quantitative and qualitative traits. Iran J Agr Sci 37:385–393
Aka Kacar Y, Darıcı, HC, Söğüt Z, Solmaz İ, Bozdoğan E, Haspolat G, Teixeira da Silva JA (2013) Molecular characterization and in vitro preservation of Cyclamen Species Grown Naturally in Turkey Project No: 1100102 Final Report (In Turkish), pp 1–157
Aka Kacar Y, Demirel A, Tuzcu O, Yesiloglu T (2005) Preliminary results on fingerprinting lemon genotypes tolerant to mal secco (Phoma tracheiphila Kanc. et Ghik) disease by RAPD markers. Biologia (sect. Mol. Biol.) 60:295–300
Amar MH, Biswas MK, Zhang Z, Guo WW (2011) Exploitation of SSR, SRAP and CAPS-SNP markers for genetic diversity of Citrus germplasm collection. Sci Hortic 128(3):220–227
Anderberg AA (1994) Phylogeny and subgeneric classification of Cyclamen L. (Primulaceae. Kew Bul 49(3):455–467
Compton JA, Clennett JCB, Culham A (2004) Nomenclature in the dock. Overclassification leads to instability: a case study in the horticulturally important genus Cyclamen (Myrsinaceae). Bot J Linn Soc 146:339–349
Curuk P, Sogut Z, Bozdogan E, Izgu T, Sevindik B, Tagipur E, Teixeira da Silva JA, Serce S, Aka Kacar Y, Mendi Y (2015) Morphological characterization of Cyclamen sp. grown naturally in Turkey: part I. S Afr J Bot 100:7–15
Debussche M, Thompson JD (2002) Morphological differentiation among closely related species with disjunct distributions: a case study of Mediterranean Cyclamen L. subgen. Psilanthum Schwarz (Primulaceae). Bot J Linn Soc 139:133–144
Edwards K, Johnstone C, Thompson C (1991) A simple and rapid method for the preparation of plant genomic DNA for PCR analysis. Nucleic Acids Res 19:1349
Gielly L, Debussche M, Thompson JD (2001) Geographic isolation and evolution of Mediterranean endemic Cyclamen: insights from chloroplast trnL (UAA) intron sequence variation. Plant Syst Evol 230:75–88
Grey-Wilson C (1997) Cyclamen. A guide for gardeners, horticulturists and botanists. B.T. Batsford Ltd., London, pp 192
Hammer Ø, Harper DAT, Ryan PD (2001) PAST: paleontological statistics software package for education and data analysis. Palaeontologia Electronica 4(1):9
Jaccard P (1908) Nouvelles recherches sur la distribution florale. Bull Soc Vaud Sci Nat 44:223–270
Jalali N, Naderi R, Ali SG, Teixeira da Silva JA (2012) Cyclamen tissue culture. Sci Hortic 137:11–19
Kallersjo M, Bergqvist G, Anderberg AA (2000) Generic realignment in primuloid families of the Ericales s.l.: a phylogenetic analysis based on DNA sequences from three chloroplast genes and morphology. Am J Bot 87:1325–1341
Kitamura S, Akita Y, Ishizaka H, Narumi I, Tanaka A (2012) Molecular characterization of an anthocyanin-related glutathione S-transferase gene in cyclamen. J Plant Physiol 169:636–642
Laura M, De Benedetti L, Bruna S, Burchi G, Berio T, Giovannini A, Ruffoni B (2003) Cyclamen persicum Mill. Somatic embryogenesis and RAPD analysis of embryogenic callus. Acta Hort 625:137–144
Li G, Quiros CF (2001) Sequence-related amplified polymorphism (SRAP) a new marker system based on a simple PCR reaction: its application to mapping and gene tagging in Brassica. Theor Appl Genet 103:455–461
Naderi R, Alaey M, Khalighi A, Hassani ME, Salami SA (2009) Inter-and intra-specific genetic diversity among cyclamen accessions investigated by RAPD markers. Sci Hortic 122:658–661
Rohlf FJ (1998) NTSYS-PC Numerical taxonomy and multivariate analysis system. Version 2.00 Exeter software Setauket, New York
Smith JSC, Chin ECL, Shu H, Smith OS, Wall SJ, Senior ML, Mitchel SE, Kresorich S, Tiegle J (1997) An evaluation of the utility of SSR loci as molecular markers in maize (Zea mays L.): comparisons with data from RFLPs and pedigree. Theor Appl Genet 95:163–173
Taskin BG, Vardareli N, Doğaç E, Mammadov R, Taskin V (2012) Genetic diversity of natural Cyclamen alpinum populations. Turk J Biol 36(4):413–422
Uzun A, Yesiloglu T, Aka-Kacar Y, Tuzcu O (2009) Genetic diversity and relationships within Citrus and related genera based on sequence related amplified polymorphism markers (SRAPs). Sci Hortic 121:306–312
Zhang JH, McDonald MB, Sweeney PM (1997) Testing for genetic purity in petunia and cyclamen seed using random amplified polymorphic DNA markers. Hort Science 32:246–247
Acknowledgments
This research was funded by the Scientific and Technical Research Council of Turkey (TÜBİTAK, project number 110O102).
Author information
Authors and Affiliations
Corresponding author
Additional information
An erratum to this article is available at http://dx.doi.org/10.1007/s10528-016-9774-5.
Rights and permissions
About this article
Cite this article
Simsek, O., Curuk, P., Aslan, F. et al. Molecular Characterization of Cyclamen Species Collected from Different Parts of Turkey by RAPD and SRAP Markers. Biochem Genet 55, 87–102 (2017). https://doi.org/10.1007/s10528-016-9770-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10528-016-9770-9