Abstract
Apoptosis is a fundamental process for the elimination of damaged or unwanted cells, and is a key aspect of development. It is triggered by pro-apoptotic genes responding to the intrinsic pathway that senses cell stress or the extrinsic pathway that responds to signals from other cells. The disruption of these genes can therefore lead to developmental defects and disease. Pro-apoptotic genes have been studied in detail in the fruit fly Drosophila melanogaster, a widely-used developmental model. However, little is known about the corresponding genes in its relative D. suzukii, a pest of soft fruit crops that originates from Asia but is now an invasive species in many other regions. The characterization of D. suzukii pro-apoptotic genes could lead to the development of transgenic sexing strains for pest management. Here, we describe the isolation and characterization of the pro-apoptotic genes reaper (Dsrpr), head involution defective (Dshid) and grim (Dsgrim) from a laboratory strain of D. suzukii. We determined their expression profiles during development, revealing that all three genes are expressed throughout development but Dsrpr is expressed most strongly, especially at the pupal stage. Functional analysis was carried out by expressing single genes or pairs (linked by a 2A peptide) in S2 cell death assays, indicating that Dsgrim and Dshid are more potent pro-apoptotic genes than Dsrpr, and the lethality can be significantly enhanced by co-expression of two genes. Therefore, the binary or multiple expression of different pro-apoptotic genes can be considered to build an efficient transgenic sexing system in D. suzukii.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Apoptosis (a form of programmed cell death) is an evolutionarily conserved process that eliminates unwanted cells during development as well as cells damaged by stress [1]. Apoptotic cells are characterized by plasma membrane blebbing, cytoplasmic condensation and shrinkage, progressive chromatin degradation, and nuclear fragmentation, before the apoptotic bodies are engulfed by cells of the immune system. The process can be triggered by intracellular stress-response signals via the mitochondria (intrinsic pathway) or ligands released by other cells (extrinsic pathway). These converge on a small group of pro-apoptotic genes, which in Drosophila melanogaster include the closely linked loci reaper (rpr), head involution defective (hid), grim and sickle (skl) on chromosome 3 [2, 3]. The corresponding proteins contain an N-terminal RHG motif and an internal GH3 domain, which activate different but synergistic downstream pathways [4]. The RHG motif binds to inhibitor of apoptosis proteins (IAPs) such as Diap1, which normally sequesters caspases in an inactive complex. The binding of RHG proteins to Diap1 displaces the caspases, including three (Dronc, Dcp-1 and DrICE) that trigger apoptosis [5]. The conserved RHG motif is therefore also described as an IAP-binding motif (IBM) [6, 7].
Although pro-apoptotic genes have been studied in detail in D. melanogaster, this is not the case for the related species D. suzukii, a pest of soft-skinned fruits. D. suzukii is native to Southeast Asia but has spread throughout much of North America and Europe [8,9,10,11]. Adult females pierce the soft skin of fruits with their ovipositor, laying eggs which hatch into larvae that feed on the fruit pulp, causing large-scale damage [12, 13]. D. suzukii has a short generation time and a population can thus grow rapidly [14]. Insecticides have been developed and can control D. suzukii efficiently [15] but species-specific and sustainable ones have not been reported. Thus, several approaches have been followed to environment friendly control methods for D. suzukii [14, 16]. The sterile insect technique (SIT) is one of those alternative and environmentally friendly strategies that can be used as a targeted biocontrol measure. For traditional SIT program, large numbers of flies sterilized with gamma radiation are released into the field to mate with their wildtype counterpart, and lead to no viable offspring therefore reducing the population size [17, 18]. The SIT has already been used to successfully control the Mediterranean fruit fly, Ceratitis capitata [19], new world screwworm Cochliomyia hominivorax, and other tephritid fruit flies, tsetse flies and various lepidopteran pests [20,21,22]. Efficient SIT strategies require the production of genetic sexing strains to facilitate the mass separation of sterilized males and females so that only males are released in the field [23, 24]. In C. capitata, this was achieved through classical genetics and sex-linked markers generating a heat-inducible female-specific lethal system [19]. This system could not be transferred to other species yet, because the phenotypic markers identified are genetically unknown so far. Therefore, other approaches were followed to produce transgenic embryonic sexing systems (TESS) in which active pro-apoptotic genes containing a sex-specific intron are expressed during early embryonic development to kill females [25, 26]. In addition, redundant or multi-lethal systems were recommended to improve strain stability under mass-rearing conditions and reduce the risk of resistance in the field if fertile males were to be released [27, 28]. It was also reported that endogenous genes are more efficient than exogenous ones when generating transgenic sexing strains [29, 30]. Therefore, we wanted to isolate the endogenous genes from D. suzukii for the development of efficient TESS.
Here, we isolated the pro-apoptotic genes Dsrpr, Dshid, and Dsgrim from D. suzukii that are described as apoptosis inducing in D. melanogaster and identified their conserved functional motifs by comparing the orthologs of different insect species. The fourth member, sickle (skl) [31, 32], that enhances but doesn’t induce apoptosis, was not isolated. The expression profiles of the genes were verified by Reverse-Transcriptase (RT) and quantitative Real-Time (qRT) PCR, confirming similar patterns to those from D. melanogaster. We further tested the activity of each gene in S2 cell death assays, as well as the co-expression of pairs of apoptotic genes using a 2A peptide containing vector [33]. These experiments allowed us to select appropriate candidate genes for the development of TESS strategies for the control of D. suzukii in the future.
Methods
Insect rearing and sample collection
Wild-type D. suzukii flies (USA strain) were maintained at 25 °C and 60% humidity with a 12-h photoperiod. Embryos were collected over a duration of 60 min, and were allowed to develop on grape juice agar plates (1% agar, 30% grape juice) as previously described [34]. The larvae and pupae were collected from stock vials at the desired age. Adult males and females were isolated immediately after they emerged, and were sampled 1 or 5 d later.
Gene sequence isolation and analysis
A high-quality D. suzukii reference genome sequence [35] is available at SWDbase (https://spottedwingflybase.org/). The coding sequences of the D. melanogaster genes Dmrpr (FBgn0011706), Dmhid (FBgn0003997) and Dmgrim (FBgn0015946) were obtained from FlyBase (https://flybase.org/) and used as tBLASTx search queries against SWDbase. Based on the hits recovered for each search, primers were designed to amplify the full-length coding sequences of the three orthologs from D. suzukii. Total RNA was isolated from adult flies (5 days old) using the ZR Tissue & Insect RNA MicroPrep kit (Zymo Research, USA) and treated with Turbo DNase (Thermo Fisher Scientific, USA). The iScript cDNA Synthesis Kit (Bio-Rad, USA) was used to synthesise cDNA from 0.5 μg of DNA-free total RNA. cDNA was diluted to 1:10 according to the protocol for further use.
The Dsrpr and Dsgrim cDNAs were amplified in 25 μl reactions comprising 0.1 μl Platinum Taq DNA polymerase (Invitrogen, USA), 2.5 µl 10 × PCR buffer, 1 µl 50 mM MgCl2, 2.5 µl 10 mM dNTPs, 0.75 µl 10 µM of each primer, and 1 μl diluted cDNA. The cycling conditions were 95 °C for 2 min, followed by 35 cycles of 95 °C for 30 s, 52 °C for 30 s, and 72 °C for 60 s, and a final extension step at 72 °C for 5 min. The 198-bp Dsrpr amplicon was transferred to the vector pCR4 by TOPO™ TA Cloning™ Kit cloning (Invitrogen GmbH) using primers P9/P10 (all primers are listed in Online Resource 1), resulting in construct V12. The 360-bp Dsgrim product was transferred similarly using primers P135/P136, resulting in construct V34.
The Dshid cDNA was amplified in a 20 µl reaction comprising 10 µl Phusion Flash High-Fidelity PCR Master mix (Thermo Fisher Scientific), 1 µl 10 µM of each primer and 1 µl diluted cDNA. The cycling conditions were 98 °C for 10 s, followed by 30 cycles of 98 °C for 30 s, 55 °C for 5 s and 72 °C for 30 s, and a final extension step at 72 °C for 2 min. The low abundance of Dshid mRNA was overcome by adopting a two-step procedure in which exons 1 and 2 were amplified using primers P81/P41 and exons 3 and 4 by using primers P165/P42 before combining the products and reamplifying with the external primers P41/P42. The 1281 bp Dshid product was transferred to the vector pCR4 by TOPO™ TA Cloning™ Kit cloning (Invitrogen GmbH), using primers P41/P42, resulting in construct V392.
To generate DshidAla4, four potential MAPK phosphorylation sites were identified in the Dshid sequence (Online Resource 2), and the acceptor residues were replaced with alanine to prevent inactivation by phosphorylation [30, 36]. This constitutively active Dshid gene was synthesized by Eurofins (Germany), resulting in construct V45. The integrity of all vectors was confirmed by restriction digestion and sequencing using primers M13/M14, and sequences were analyzed using the Geneious Prime software [37].
Isolation and analysis of the Dshid 3′UTR
A 2260 bp region of the 3′UTR from Dshid was amplified from an embryonic cDNA (2–4 h) pool from D. suzukii by using primers P82/P84 (all primers are listed in Online Resource 1) and in-silico predictions on the Dshid gene. The amplified region was cloned into the pCR4 vector using the TOPO™ TA Cloning™ Kit cloning (Invitrogen), sequenced and resulting sequences analyzed for previously predicted bantam miRNA binding sites [38] (Online Resource 3) using the Geneious Prime software.
Protein sequence alignments from other species
Orthologues of RHG proteins from D. grimshawi (Dg), D. hydei (Dh), D. willistoni (Dw), D. ficusphila (Df), D. biarmipes (Db), D. erecta (Der), D. melanogaster (Dm), D. serrata (Dser), Lucilia cuprina (Lc) and Musca domestica (Md) were downloaded from NCBI as following: DgRPR (XP_001985528.1), DhRPR (XP_023170619.1), DwRPR (XP_023033239.1), DfRPR (XP_017058909.1), DbRPR (XP_016955290.1), DerRPR (XP_001972845.1), DmRPR (NP_524138.1), DserRPR (XP_020801924.1), LcRPR (XP_023291721.1), MdRPR (XP_005184304.1), DgHID (XP_001985533.1), DhHID (XP_023170594.1), DwHID (XP_002067955.1), DfHID (XP_017059087.1), DbHID (XP_016956024.1), DerHID (XP_001972854.1), DmHID (AAA79985.1), DserHID (XP_020801905.1), LcHID (XP_023305760.1), MdHID (XP_005180529.1). DgGRIM (XP_001996719.1), DhGRIM (XP_023170636.1), DwGRIM (XP_002067945.1), DfGRIM (XP_017058891.1), DbGRIM (XP_016955275.1), DerGRIM (XP_001972846.1), DmGRIM (NP_524137.2), Dser (XP_020801932.1), LcGRIM (XP_023291726.1), MdGRIM (XP_019891253.1).
Construction of pIE expression plasmids
The Dsrpr sequence was reamplified from construct V12 (see above) in a 25-μl reaction comprising 0.1 μl Platinum Taq DNA polymerase, 2.5 µl 10 × PCR buffer, 1 µl 50 mM MgCl2, 2.5 µl 10 mM dNTPs, 0.75 µl 10 µM of each primer P150/P149 (all primers are listed in Online Resource 1) and 1 μl 100 ng template DNA. The cycling conditions were 95 °C for 2 min, followed by 35 cycles of 95 °C for 30 s, 52 °C for 30 s and 72 °C for 60 s, and a final extension step at 72 °C for 5 min. The product was digested with NotI and SacII (New England Biolabs, USA) and was transferred to vector pIE4 prepared with the same enzymes, resulting in expression vector V42. The Dsgrim sequence was reamplified in the same manner, except the template was vector V34 (see above). The product was transferred to vector pIE4 as above, resulting in expression vector V44. The Dshid sequence was reamplified from vector V381 using primers P1654/P1655 containing restriction sites for SacII and NotI, respectively. The 25-μl reaction comprised 0.2 μl Platinum Taq DNA polymerase, 2.5 µl 10 × PCR buffer, 0.75 µl 50 mM MgCl2, 1.0 µl 10 mM dNTPs, 1 µM of each primer and 0.5 μl V381. The cycling conditions were 95 °C for 2 min, followed by 35 cycles of 95 °C for 30 s, 52 °C for 30 s and 72 °C for 2 min, then a final extension step at 72 °C for 5 min. The PCR product was transferred to vector pIE4 as above, resulting in expression vector V392. To prepare vector V93_pIE4_DshidAla4, V45_pEX-K4-DshidAla4 was digested with SacII and NotI and the insert was transferred to pIE4, prepared using the same enzymes. The previously reported overexpression plasmids pIE-Dmrpr, pIE-Dmhid and pIE-DmhidAla5 harboring D. melanogaster rpr and hid genes [30] were also included in the assays.
Construction of 2A peptide expression plasmids
The 2A peptide plasmids (Fig. 4a) were constructed by amplifying the Dsrpr sequence from construct V12 using primers P213/P150 (all primers are listed in Online Resource 1), the Dsgrim sequence from construct V44 using primers P313/P162 and the DshidAla4 sequence from construct V45 using primers P314/P315. Pairs of genes were transferred to construct V142 using the restriction enzymes ApaI and NotI to join them via the DrosCV2A peptide, or to construct V145 using the same restriction enzymes to join them via the TaV2A peptide [33].
Construction of RMCE plasmids
Dsrpr, DshidAla4, and Dsgrim were amplified by PCR in a 25-μl reaction comprising 0.1 μl Platinum Taq DNA polymerase, 2.5 µl 10 × PCR buffer, 1 µl 50 mM MgCl2, 2.5 µl 10 mM dNTPs, 0.75 µl 10 µM of each primer and 1 μl 100 ng template DNA. The cycling conditions were 95 °C for 2 min, followed by 35 cycles of 95 °C for 30 s, 55 °C for 30 s and 72 °C for 60 s, and a final extension step at 72 °C for 5 min; on plasmids V163_pIE4_Dsrpr_DrosCV-2A_Dsrpr_SV40, V165_pIE4_Dsrpr_DrosCV-2A_Dsrpr_SV40, and V164_pIE4_Dsrpr_DrosCV-2A_Dsgrim_SV40 by using primers P1068/P1071, P1070/P1071, and P1069/P1071, respectively (all primers are listed in Online Resource 1). PCR fragments were then inserted into AH448_ pSL_loxN-3xP3-PUbDsRed-lox2272 [39] by SmaI and SalI restriction sites to generate V388_pSL_loxN-3xP3-Dsrpr_SV40-PUbDsRed-lox2272, V350_pSL_loxN-3xP3-DshidAla4_SV40-PUbDsRed-lox2272, and V337_pSL_loxN-3xP3-Dsgrim_SV40-PUbDsRed-lox2272, respectively.
Quantitative real-time PCR
SsoAdvanced Universal SYBR Green Supermix (Bio-Rad) was used for qPCR with 100 ng cDNA in a CFX96 Touch Real-Time time PCR Detection System (Bio-Rad). The cycling conditions were 95 °C for 30 s, followed by 40 cycles of 95 °C for 10 s, 60 °C for 20 s, and 65 °C for 5 s, and 95 °C for 0.5 s. Reactions were carried out on three biological replicates each comprising three technical replicates. Samples for all developmental stages were collected for each biological replicate from the same culture. The 2–∆∆Ct method was used for all samples, and the data were normalized to the control gene TBP [40].
Cell culture
We used D. melanogaster Schneider 2 (S2) cells [41] grown in Schneider’s medium at 25 °C with 10% heat-inactivated fetal bovine serum (Hi-FBS) and 1% penicillin/streptomycin in closed capped flasks without CO2. Cells were passaged every 2–3 days unless ≥ 90% viability was achieved. Transient transfection was carried out using Xfectin reagent (Takara, Japan) according to the manufacturer’s instructions. To monitor the cell damage, we seeded 24-well plates, each well lined with a 13-mm TC coverslip (Sarstedt, Germany), with 5 × 105 cells (live cell count) in a volume of 500 µl medium and allowed the cells to settle for 3 h. The cells were then transfected with 1 µg plasmid DNA using 0.3 µl Xfectin and 27.4 µl Xfectin buffer in 270 µl serum-free Schneider’s medium for 4 h. Transfection was stopped by removing the reagents and replenishing the cells with 500 µl Schneider’s medium containing Hi-FBS and penicillin/streptomycin as above. The cells were incubated for a further 16 h at 25 °C before fixing in 4% paraformaldehyde for 15 min and washing twice with 1 × PBS. Morphological images were taken with an inverted microscope (DM IL LED, Leica Microsystems, Wetzlar, Germany). For cell counting, 0.5 μg pIE4-EGFP plasmid was co-transfected with one of the pIE expression plasmids (0.5 μg) or with pIE4-DsRed control vector (0.5 μg) to visualize the successfully transfected cells. Cells expressing EGFP were imaged using M205FA MZ FLIII microscopes (Leica Microsystems, Germany) with EGFP filter sets (λexcitation = 500/20; λemission = 535/30) using consistent settings. TC coverslips containing adhesive fluorescent S2 cells were placed on a slide over a drop of Hi-FBS. We captured 25 images per TC coverslip, and cells were counted using Image J (Fiji) by first converting to 8-bit (threshold 30) inverted images, and then applying the watershed and automated cell count functions. For the comparison of Dmhid to Dshid, Dmrpr to Dsrpr, and DmhidAla5 to DshidAla4, we captured ten images per TC coverslip and fluorescent S2 cells were counted as described before.
Statistical analysis
Statistical analysis was carried out using SigmaPlot v14 for the differences in viability of S2 cells after post transfection of different pIE4 vectors. Data were square root transformed to pass the normality test, and analyzed by one-way ANOVA, and means were separated using the Holm–Sidak method. In total, two transfection experiments were performed, one for constructs expressing the single pro-apoptotic genes Dsrpr, Dshid, DshidAla4, Dmrpr, Dmhid, and DmhidAla5 and a second, with Dsgrim, and all 2A peptide constructs expressing two pro-apoptotic genes. Each experiment was normalized to its control and individual percentages calculated. All data was then statistically compared to generate Fig. 4b. Detailed statistics is provided as Online Resource 4.
Results
D. suzukii pro-apoptotic genes can be isolated by homology-based screening
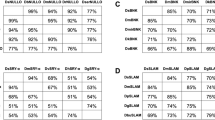
Three D. suzukii pro-apoptotic genes were identified by searching the SWDbase using orthologs from D. melanogaster. The coding sequence of Dsrpr (DS10_00012288) was found to be 198 bp in length, encoding a protein of 66 amino acids. DsRPR (Fig. 1a) was most closely related to its ortholog in D. melanogaster (96.9% similarity, 92.3% identity). The CDS of Dshid (DS10_00012680) was found to be 1281 bp in length, encoding a protein of 427 amino acids. The DsHID protein (Fig. 1b) was more closely related to its ortholog in D. biarmipes (96.5% similarity, 94.6% identity) than its ortholog in D. melanogaster (91.8% similarity and 86.7% identity). The coding sequence of Dsgrim (DS10_00013088) was found to be 360 bp in length, encoding a protein of 120 amino acids. DsGRIM (Fig. 1c) was also most closely related to its ortholog in D. melanogaster (93.4% similarity, 86.8% identity). Phylogenetic analysis using a neighbor-joining algorithm revealed that all three pro-apoptotic genes clustered with their orthologs from several other Drosophila species, notably D. biarmipes (Fig. 2). Canonical RHG/IBM and GH3 domains were identified in all three proteins, suggesting their pro-apoptotic functions are likely to be retained.
Alignment of the D. suzukii RHG proteins with orthologs in other dipterans: a REAPER (RPR), b HEAD INVOLUTION DEFECTIVE (HID) and c GRIM. In each case, the protein from D. suzukii (Ds) is aligned with orthologs from D. melanogaster (Dm), D. grimshawi (Dg), D. hydei (Dh), D. willistoni (Dw), D. ficusphila (Df), D. biarmipes (Db), D. erecta (Der), D. serrata (Dser), Lucilia cuprina (Lc) and Musca domestica (Md). Identical amino acids are shaded in black and conservative changes in gray. The IBM and GH3 domains are shown
Phylogenetic analysis of dipteran RPR, HID and GRIM proteins. Unrooted neighbor-joining trees were constructed with a RPR, b HID and c GRIM amino acid sequences. Bootstrap values (1000 replicates) are shown on the nodes of the trees. Species abbreviations are the same as in Fig. 1
D. suzukii pro-apoptotic genes are expressed throughout development
Apoptosis is initiated during embryonic stage 11 in insects and thereafter becomes widespread and dominant during embryonic and post-embryonic development, as previously reported in D. melanogaster [42]. We analyzed the mRNA profiles of the three D. suzukii genes by RT-PCR and found that Dsgrim and Dshid expression commenced within the first hour of embryogenesis, whereas Dsrpr expression commenced after 4 h. Dsrpr expression levels were highest during late embryogenesis and the pupal stage (Fig. 3a). Similar profiles were revealed by qPCR when the data were normalized to the 0–6 h embryonic stage expression of Dshid. We found that Dshid and Dsrpr expression commences at the embryonic stage, increases throughout development and peaks in late pupae. Comparative analysis indicated that Dsrpr is expressed at a higher level than the other two genes, reaching almost 20-fold higher than Dshid in the pupae, whereas Dsgrim is expressed at the lowest level (Fig. 3b).
Gene expression profiles of Dshid, Dsrpr and Dgrim throughout the development of Drosophila suzukii. a Reverse transcriptase-PCR analysis of Dshid, Dsrpr and Dsgrim. RNAs from embryos collected at different time points after egg laying (in hours), larvae, pre-pupae 24 h, pupae, adult females and males (1 and 5 days old). DsTBP, which is expressed at all stages, is the loading control. M is the molecular weight ladder. The PCR product sizes are 109 bp for Dshid, 146 bp for Dsgrim, 147 bp for Dsrpr, and 182 bp for DsTBP. b Relative expression levels of pro-apoptotic genes at different time points after egg laying (in hours), larvae, pre-pupae 24 h, pupae, adult females, and males (1 and 5 days old) as determined by qRT-PCR. Expression levels were normalized to DsTBP (reference gene), which is expressed at all stages. In addition to that, expression was further normalized to Dshid 0–6 h. The mean and standard error from three replicate experiments are shown
Activity of D. suzukii pro-apoptotic genes in S2 cells
To characterize each D. suzukii pro-apoptotic gene, we generated the single expression constructs V42, V44, V392 and V93, representing the wild-type Dsrpr, Dsgrim and Dshid genes and the constitutive Dshid mutant DshidAla4, respectively (Fig. 4a). The binary constructs in which Dsrpr was paired with another copy of itself (V163, V172), with Dsgrim (V164, V173) or with DshidAla4 (V165, V174), were also developed to test the pro-apoptotic activity from combinations (Fig. 4a). The two distinct constructs representing each pairing correspond to the use of two different picornaviral 2A peptides [33], namely DrosCV-2A (V163, V164, V165) and TaV-2A (V172, V173, V174). After 16 h, the transfected cells from single expression constructs had lost confluence, there was a change in cell shape, and in some cases there was evidence of membrane rupture, and such phenomena was not observed in the cells that transfected with pIE4-DsRed control vector (V125) (Online Resource 2), suggesting expression of each pro-apoptotic gene was able to reduce the fitness of the cells, but not trigger widespread apoptosis. Quantitative analysis by cell counting suggested that single constructs using Dsrpr, Dshid, DshidAla4, and Dsgrim (P < 0.001, One-way ANOVA) significantly reduced the cell number compared to the control (Fig. 4b). In addition, all binary constructs killed more cells compared to single expression constructs (P < 0.001, One-way ANOVA), confirming that the combinations of pro-apoptotic genes were most efficient in reducing the cell viability (Fig. 4b). We also compared cell death activity of Dshid, DshidAla4, and Dsrpr to the previously reported Dmhid, DmhidAla5, and Dmrpr genes [30] (Fig. 4b). No significant difference was observed in the activity of Dmrpr versus Dsrpr (P = 0.868, One-way ANOVA), Dmhid versus Dshid (P = 1.000, One-way ANOVA), and DshidAla4 versus DmhidAla5 (P = 0.014; One-way ANOVA, see Online Resource 4). The comparison was repeated in another set of cells (Online Resource 6) in triplicates to ensure the results from Fig. 4b.
Functional activity of Drosophila suzukii pro-apoptotic genes. a Schematic map of the pIE4 test plasmids. b S2 cells were co-transfected with pIE4-EGFP plasmid and one of the pIE expression plasmids or pIE4-DsRed control plasmid to visualize the successfully transfected cells using M205FA MZ FLIII microscope (Leica Microsystems). Number of EGFP positive cells as survived cells, were counted using Image J (Fiji). The experiment was carried out in three replicates. Mean and standard errors are shown in the figure, bars with different uppercase letters are significantly different at P < 0.050 (one-way ANOVA, Holm–Sidak method for pairwise multiple comparison)
Activity of D. suzukii pro-apoptotic genes in larvae-pupae.
To characterize in vivo activity of pro-apoptotic genes in D. suzukii, we microinjected pro-apoptotic genes Dsrpr, Dshid and Dsgrim incorporated into RMCE donor constructs as V338, V350 and V337, respectively, and AH448 as a control [39] into the embryos of the previously established transgenic D. suzukii landing site line carrying the construct AH443_PUbEGFP-TRE-CctraI-AlhidAla2-loxN-3xP3-AmCyan-lox2272 (Online Resource 7) [39]. 48 h after injections, hatched larvae were counted and screened for transient expression (DsRed or AmCyan). We observed low survival rate for embryos injected with the RHG containing donor constructs (Online Resource 7) compared to the control. There were also no adult survivors for all larvae transiently expressing the RHG containing constructs while 20% of the control larvae survived to adulthood. Because apoptotic genes were driven by the 3xP3 promoter, it can be speculated that 3xP3 lead the expression of apoptotic gene already in larval tissues [43, 44], causing lethality in transient larvae.
Discussion
Apoptosis is a highly regulated mechanism for the targeted and programmed destruction of cells that are damaged or unwanted. In contrast to the events that occur during necrosis, apoptosis is a controlled and beneficial process, with essential roles in development and homeostasis. It can be initiated intrinsically in response to cell stress or extrinsically by extracellular ligands. Still, the pathways converge on a small number of pro-apoptotic genes that directly mediate the cellular-level biochemical and cellular changes involved in programmed cell death, such as cytoplasmic shrinkage, chromatin condensation and nuclear fragmentation [1]. The most important pro-apoptotic genes include the RHG family, named after the founder members rpr, hid and grim. These genes were initially identified in D. melanogaster, and the corresponding proteins are characterized by an N-terminal IBM that interacts with Diap1 to release pro-apoptotic caspases and an internal GH3 domain that induces the mitochondrial death pathway in a caspase-independent manner [7]. Removing either IBM or GH3 motifs from the protein partially or entirely blocked the pro-apoptotic activity of the responsive genes from the Caribbean fruit fly Anastrepha suspensa [30], the Scuttle Fly Megaselia scalaris [45], the primary malaria vector Anopheles gambiae [46], and the silkworm Bombyx mori [47]. Thus, IBM and GH3 motifs are critical features responsible for functional pro-apoptotic genes [48, 49]. The IBM and GH3 motifs in HID and GRIM are identical among D. suzukii, D. melanogaster and several other Drosophila species (Fig. 1a, b). In RPR homologs, the IBM motif is nearly identical among these species, but the GH3 domain is less well conserved (Fig. 1c). It was suggested that the amino acid changes in the GH3 domain, which is potentially a HID and GRIM binding motif, reduces the effectiveness of RPR [48, 50]. Indeed, we observed less pro-apoptotic activity from Dsrpr compared to these from Dshid and Dsgrim in the cell death assays (Fig. 4b), suggesting that Dshid and Dsgrim are better candidate genes for the development of TESS in D. suzukii. In addition to GH3 and IBM domains, we also searched for bantam miRNA binding sites in the Dshid 3′UTR that have been reported in the Dmhid 3′UTR homologous sequence [38]. All five regions are conserved and could also play a role in Dshid regulation (Online Resource 3).
Both D. suzukii and D. biarmipes have similar spots on the wings and are closely related to each other [35, 51], and it was suggested that D. suzukii diverged from D. biarmipes approximately 6 to 9 million years ago [52]. The phylogenetic analysis here also confirmed that the DsHID, DsGRIM and DsRPR are closely related to their orthologs from D. biarmipes (Fig. 2). In D. melanogaster, hid and rpr genes are expressed at moderate (hid) or high (rpr) levels in pupae, moderate levels in embryos and low levels in larvae and adults. In contrast, grim is expressed at low levels through development [53]. Similar patterns were observed for the expression of Dsrpr, Dshid and Dsgrim (Fig. 3), suggesting that also the functional roles and regulation pathways of these genes could be conserved between the two species. In fact, the activity of certain pro-apoptotic genes is well conserved across different species. For example, ectopic expression of pro-apoptotic genes from the European green blow fly L. sericata [50] and M. scalaris [45] caused tightly-regulated cell death in D. melanogaster, and D. melanogaster rpr could efficiently induce apoptosis in A. suspensa [54] and mammalian cells [55]. However, other studies showed that endogenous genes work more efficiently and can be tightly controlled compared to homologous genes from closely related species [29]. Using D. melanogaster S2 cells as a reporter system, we identified that the Dshid and Dsgrim as well as Dmhid and DmhidAla5 are equally effective to reduce the cell viability while rpr homologs and the DshidAla4 are less effective to confer cell death (Fig. 4b). A weaker activity of rpr was reported before [30, 50], but the result from DshidAla4 is unexpected because the mutations in the MAPK phosphorylation sites should avoid downregulation of HID by the Ras signaling pathways [36]. Previous studies showed that the A. suspensa and L. cuprina TESS strains using the phospho-mutated version of hid were more effective at causing cell death [25, 54]. The Ras1/MAPK pathway may affect the cell death-inducing ability of DshidAla4 differently in S2 cells than in analysis conducted on in vivo cell networks. Indeed, injecting a transgenic line with cell death promoting constructs led to a reduction of progeny in a small-scale transient in vivo assay (Online Resource 7).
The efficacy of an SIT program relies on mass-rearing and releasing a large number of insects [17, 22]. For example, more than 15 million sterile C. hominivorax are released per week for efficient containment of this species [21]. It was demonstrated that the spontaneous mutations occur in the pro-apoptotic gene of a TESS strain with a 1 in a million frequency [56]. Uncontrolled breakdown of the TESS during mass rearing due to the loss-of-function mutation could lead to the release of females. Consequently, employing multiple lethal genes or the development of redundant lethal systems would be important for the efficiency and stability of TESS strains [27, 28]. Previous reports also showed that combinations of pro-apoptotic genes caused a higher level of lethality than a single gene [30]. An efficient co-expression system was recently described in D. suzukii using picornaviral self-cleaving 2A peptides [33], which can express two or more genes in a stoichiometric ratio by ribosomal skipping [57]. Thus, two copies of the same gene should express and be translated independently and confer higher lethality levels compared single copy expression systems. The difference in lethality between single versus multiple copies of a pro-apoptotic genes has functionally been shown in D. melanogaster, C. capitata, L. cuprina and A. suspensa flies. There, lethality tests with heterozygous and homozygous transgenic individuals showed that lethality was most effective in homozygous individuals, carrying the double amount of copies [58, 59]. Similarly, while single DsRPR was conferring only low lethality numbers in our setup, the double amount of DsRPR co-expressed by a 2A peptide might have reached the required dosage for cell death. Previous reports showed that the 2A peptide is always cleaved with the upstream protein, not the downstream protein [33, 57, 60]. However, this doesn’t hinder the correct translation of the upstream or downstream proteins [33, 57, 60,61,62,63]. Here, co-expression of rpr and rpr, hid or grim with the help of 2A peptides confirmed the essential interaction of pro-apoptotic cell death genes to confer lethality that was reported with different expression strategies in D. melanogaster before [3, 48]. Our studies verified Dsrpr as an always expressed apoptotic gene in embryonic, larval and pupal stages (Fig. 3b) showing its role as a global regulator of apoptosis. The apoptotic effects were more pronounced when cells were transfected with the bicistronic constructs, with a clear impact on cell growth as well as morphology resulting in 97–98% cell death (Fig. 4b; Online Resource 2). Thus, different pro-apoptotic genes in combination with conditional systems like the tetracycline-controllable Tet-Off system can be developed into TESS strains in D. suzukii. With several copies of apoptotic genes on one construct, the desired effect could be enhanced, and the system could cope with the loss of individual genes due to missense mutations, ensuring that the emergence of resistance would be delayed. However, a strong cell death effector is not preferred for TESS if the lethal effect is leaky [64,65,66]. Consequently, the performance of pro-apoptotic genes needs to be carefully evaluated in the TESS strains for an efficient, safe, and sustainable control program of D. suzukii.
References
Goyal L, McCall K, Agapite J, Hartwieg E, Steller H (2000) Induction of apoptosis by Drosophila reaper, hid and grim through inhibition of IAP function. EMBO J 19:589–597
Chen P, Nordstrom W, Gish B, Abrams JM (1996) Grim, a novel cell death gene in Drosophila. Genes Dev 10:1773–1782
White K, Grether ME, Abrams JM, Young L, Farrell K, Steller H (1994) Genetic control of programmed cell death in Drosophila. Science 264:677–683
Claveria C, Caminero E, Martinez AC, Campuzano S, Torres M (2002) GH3, a novel proapoptotic domain in Drosophila Grim, promotes a mitochondrial death pathway. EMBO J 21:3327–3336
Song Z, McCall K, Steller H (1997) DCP-1, a Drosophila cell death protease essential for development. Science 275:536–540
Freel CD, Richardson DA, Thomenius MJ, Gan EC, Horn SR, Olson MR, Kornbluth S (2008) Mitochondrial localization of Reaper to promote inhibitors of apoptosis protein degradation conferred by GH3 domain-lipid interactions. J Biol Chem 283:367–379
Olson MR, Holley CL, Gan EC, Colón-Ramos DA, Kaplan B, Kornbluth S, Colon-Ramos DA, Kaplan B, Kornbluth S (2003) A GH3-like domain in Reaper is required for mitochondrial localization and induction of IAP degradation. J Biol Chem 278:44758–44768
Calabria G, Maca J, Bachli G, Serra L, Pascual M (2012) First records of the potential pest species Drosophila suzukii (Diptera: Drosophilidae) in Europe. J Appl Entomol 136:139–147
Deprá M, Poppe JL, Schmitz HJ, De Toni DC, Valente VLS (2014) The first records of the invasive pest Drosophila suzukii in the South American continent. J Pest Sci 87:379–383
Hauser M (2011) A historic account of the invasion of Drosophila suzukii (Matsumura) (Diptera: Drosophilidae) in the continental United States, with remarks on their identification. Pest Manag Sci 67:1352–1357
Schwirz J, Fischbach M, Vilcinskas A, Fischer R, Schetelig MF (2016) Monitoring data and future control possibilities for Drosophila suzukii in Germany. Proceedings of the 9th international symposium on fruit flies of economic importance, pp 310–322
Bolda MP, Goodhue R, Zalom F (2010) Spotted wing drosophila: potential economic impact of a newly established pest. Giannini Found Agric Econ 13:5–8
Farnsworth D, Hamby KA, Bolda M, Goodhue RE, Williams JC, Zalom FG (2017) Economic analysis of revenue losses and control costs associated with the spotted wing drosophila, Drosophila suzukii (Matsumura), in the California raspberry industry. Pest Manag Sci 73:1083–1090
Asplen M, Anfora G, Biondi A, Choi D-S, Chu D, Daane K, Gibert P, Gutierrez AP, Hoelmer K, Hutchison W, Isaacs R, Jiang Z-L, Kárpáti Z, Kimura M, Pascual M, Philips C, Plantamp C, Ponti L, Vétek G, Desneux N (2015) Invasion biology of spotted wing drosophila (Drosophila suzukii): a global perspective and future priorities. J Pest Sci 88:469–494
Beers EH, Van Steenwyk RA, Shearer PW, Coates WW, Grant JA (2011) Developing Drosophila suzukii management programs for sweet cherry in the western United States. Pest Manag Sci 67:1386–1395
Cini A, Ioriatti C, Anfora G (2012) A review of the invasion of Drosophila suzukii in Europe and a draft research agenda for integrated pest management. Bull Insectol 65:149–160
Knipling EF (1955) Possibilities of insect control or eradication through the use of sexually sterile males. J Econ Entomol 48:459–462
Lanouette G, Brodeur J, Fournier F, Martel V, Vreysen M, Cáceres C, Firlej A, Caceres C, Firlej A (2017) The sterile insect technique for the management of the spotted wing drosophila, Drosophila suzukii: Establishing the optimum irradiation dose. PLoS ONE 12:1–14
Robinson A (2002a) Genetic sexing strains in medfly, Ceratitis capitata, sterile insect technique programmes. Genetica 116:5–13
Robinson AS (2002b) Mutations and their use in insect control. Mutat Res 511:113–132
Scott MJ, Concha C, Phillips PL, Skoda SR, Welch JB (2017) Review of research advances in the screwworm eradication program over the past 25 years. Entomol Exp Appl 164(3):226–236
Klassen W, Curtis F (2005) History of the sterile insect technique. In: Dyck VA, Hendrichs J, Robinson AS (eds) Sterile insect technique principles and practice in area-wide integrated pest management. Springer, Dordrecht, pp 3–36
Foster G (1991) Chromosomal inversions and genetic control revisited: the use of inversions in sexing systems for higher Diptera. Theor Appl Genet 81:619–623
Franz G (2005) Genetic sexing strains in Mediterranean fruit fly, an example for other species amenable to large-scale rearing for the sterile insect technique. In: Sterile insect technique: principles and practice in area-wide integrated pest management. Springer, Dordrecht, pp 427-451
Yan Y, Scott MJ (2015) A transgenic embryonic sexing system for the Australian sheep blow fly Lucilia cuprina. Sci Rep 5:1–12
Schetelig MF, Handler AM (2012a) A transgenic embryonic sexing system for Anastrepha suspensa (Diptera: Tephritidae). Insect Biochem Mol Biol 42:790–795
Eckermann KN, Dippel S, KaramiNejadRanjbar M, Ahmed HM, Curril IM, Wimmer EA (2014) Perspective on the combined use of an independent transgenic sexing and a multifactorial reproductive sterility system to avoid resistance development against transgenic Sterile Insect Technique approaches. BMC Genet 15(Suppl 2):S17
Handler AM (2016) Enhancing the stability and ecological safety of mass-reared transgenic strains for field release by redundant conditional lethality systems. Insect Sci 23:225–234
Li F, Wantuch HA, Linger RJ, Belikoff EJ, Scott MJ (2014) Transgenic sexing system for genetic control of the Australian sheep blow fly Lucilia cuprina. Insect Biochem Mol Biol 51:80–88
Schetelig MF, Nirmala X, Handler AM (2011) Pro-apoptotic cell death genes, hid and reaper, from the tephritid pest species, Anastrepha suspensa. Apoptosis 16:759–768
Srinivasula SM, Datta P, Kobayashi M, Wu JW, Fujioka M, Hegde R, Zhang Z, Mukattash R, Fernandes-Alnemri T, Shi Y, Jaynes JB, Alnemri ES (2002) sickle, a novel Drosophila death gene in the reaper/hid/grim region, encodes an IAP-inhibitory protein. Curr Biol 12:125–130
Christich A, Kauppila S, Chen P, Sogame N, Ho SI, Abrams JM (2002) The damage-responsive Drosophila gene sickle encodes a novel IAP binding protein similar to but distinct from reaper, grim, and hid. Curr Biol 12:137–140
Schwirz J, Yan Y, Franta Z, Schetelig MF (2020) Bicistronic expression and differential localization of proteins in insect cells and Drosophila suzukii using picornaviral 2A peptides. Insect Biochem Mol Biol 119:103324
Schetelig MF, Handler AM (2013) Germline transformation of the spotted wing drosophilid, Drosophila suzukii, with a piggyBac transposon vector. Genetica 141:189–193
Chiu JC, Jiang X, Zhao L, Hamm CA, Cridland JM, Saelao P, Hamby KA, Lee EK, Kwok RS, Zhang G, Zalom FG, Walton VM, Begun DJ (2013) Genome of Drosophila suzukii, the spotted wing drosophila. G3 (Bethesda) 3:2257–2271
Bergmann A, Agapite J, McCall K, Steller H (1998) The Drosophila gene hid is a direct molecular target of Ras-dependent survival signaling. Cell 95:331–341
Kearse M, Moir R, Wilson A, Stones-Havas S, Cheung M, Sturrock S, Buxton S, Cooper A, Markowitz S, Duran C, Thierer T, Ashton B, Meintjes P, Drummond A (2012) Geneious Basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28:1647–1649
Brennecke J, Hipfner DR, Stark A, Russell RB, Cohen SM (2003) bantam encodes a developmentally regulated microRNA that controls cell proliferation and regulates the proapoptotic gene hid in Drosophila. Cell 113:25–36
Schetelig MF, Yan Y, Zhao Y, Handler AM (2018) Genomic targeting by recombinase-mediated cassette exchange in the spotted wing drosophila, Drosophila suzukii. Insect Mol Biol 28(2):187–159
Zhai Y, Lin Q, Zhou X, Zhang X, Liu T, Yu Y (2014) Identification and validation of reference genes for quantitative real-time PCR in Drosophila suzukii (Diptera: Drosophilidae). PLoS ONE 9:e106800
Schneider I (1972) Cell lines derived from late embryonic stages of Drosophila melanogaster. J Embryol Exp Morphol 27:353–365
Xu D, Woodfield SE, Lee TV, Fan Y, Antonio C, Bergmann A (2009) Genetic control of programmed cell death (apoptosis) in Drosophila. Fly 3:78–90
Thomas JL, Da Rocha M, Besse A, Mauchamp B, Chavancy G (2002) 3xP3-EGFP marker facilitates screening for transgenic silkworm Bombyx mori L. from the embryonic stage onwards. Insect Biochem Mol Biol 32:247–253
Berghammer AJ, Klingler M, Wimmer EA (1999) A universal marker for transgenic insects. Nature 402:370–371
Yoo S, Lam H, Lee C, Lee G, Park JH (2017) Cloning and functional characterizations of an apoptogenic Hid gene in the Scuttle Fly, Megaselia scalaris (Diptera; Phoridae). Gene 604:9–21
Zhou L, Jiang G, Chan G, Santos CP, Severson DW, Xiao L (2005) Michelob_x is the missing inhibitor of apoptosis protein antagonist in mosquito genomes. EMBO Rep 6:769–774
Bryant B, Zhang Y, Zhang C, Santos CP, Clem RJ, Zhou L (2009) A lepidopteran orthologue of reaper reveals functional conservation and evolution of IAP antagonists. Insect Mol Biol 18:341–351
Sandu C, Ryoo HD, Steller H (2010) Drosophila IAP antagonists form multimeric complexes to promote cell death. J Cell Biol 190:1039–1052
Wang H, Clem RJ (2011) The role of IAP antagonist proteins in the core apoptosis pathway of the mosquito disease vector Aedes aegypti. Apoptosis 16:235–248
Edman RM, Linger RJ, Belikoff EJ, Li F, Sze SH, Tarone AM, Scott MJ (2015) Functional characterization of calliphorid cell death genes and cellularization gene promoters for controlling gene expression and cell viability in early embryos. Insect Mol Biol 24:58–70
Atallah J, Teixeira L, Salazar R, Zaragoza G, Kopp A (2014) The making of a pest: the evolution of a fruit-penetrating ovipositor in Drosophila suzukii and related species. Proc Biol Sci 281:20132840
Ometto L, Cestaro A, Ramasamy S, Grassi A, Revadi S, Siozios S, Moretto M, Fontana P, Varotto C, Pisani D, Dekker T, Wrobel N, Viola R, Pertot I, Cavalieri D, Blaxter M, Anfora G, Rota-Stabelli O (2013) Linking genomics and ecology to investigate the complex evolution of an invasive Drosophila pest. Genome Biol Evol 5:745–757
Graveley BR, Brooks AN, Carlson JW, Duff MO, Landolin JM, Yang L, Artieri CG, van Baren MJ, Boley N, Booth BW, Brown JB, Cherbas L, Davis CA, Dobin A, Li R, Lin W, Malone JH, Mattiuzzo NR, Miller D, Sturgill D, Tuch BB, Zaleski C, Zhang D, Blanchette M, Dudoit S, Eads B, Green RE, Hammonds A, Jiang L, Kapranov P, Langton L, Perrimon N, Sandler JE, Wan KH, Willingham A, Zhang Y, Zou Y, Andrews J, Bickel PJ, Brenner SE, Brent MR, Cherbas P, Gingeras TR, Hoskins RA, Kaufman TC, Oliver B, Celniker SE (2011) The developmental transcriptome of Drosophila melanogaster. Nature 471:473–479
Schetelig MF, Handler AM (2012b) Strategy for enhanced transgenic strain development for embryonic conditional lethality in Anastrepha suspensa. Proc Natl Acad Sci USA 109:9348–9353
Tait SW, Werner AB, de Vries E, Borst J (2004) Mechanism of action of Drosophila Reaper in mammalian cells: Reaper globally inhibits protein synthesis and induces apoptosis independent of mitochondrial permeability. Cell Death Differ 11:800–811
Zhao Y, Schetelig MF, Handler AM (2020) Genetic breakdown of a Tet-off conditional lethality system for insect population control. Nat Commun 11:3095
Donnelly MLL, Luke G, Mehrotra A, Li X, Hughes LE, Gani D, Ryan MD (2001) Analysis of the aphthovirus 2A/2B polyprotein “cleavage” mechanism indicates not a proteolytic reaction, but a novel translational effect: A putative ribosomal “skip.” J Gen Virol 82:1013–1025
Horn C, Wimmer EA (2003) A transgene-based, embryo-specific lethality system for insect pest management. Nat Biotechnol 21:64–70
Schetelig MF, Caceres C, Zacharopoulou A, Franz G, Wimmer EA (2009) Conditional embryonic lethality to improve the sterile insect technique in Ceratitis capitata (Diptera: Tephritidae). BMC Biol 7:4
Donnelly MLL, Hughes LE, Luke G, Mendoza H, Ten Dam E, Gani D, Ryan MD (2001) The “cleavage” activities of foot-and-mouth disease virus 2A site-directed mutants and naturally occurring “2A-like” sequences. J Gen Virol 82:1027–1041
Wang Y, Wang F, Xu S, Wang R, Chen W, Hou K, Tian C, Wang F, Zhao P, Xia Q (2019) Optimization of a 2A self-cleaving peptide-based multigene expression system for efficient expression of upstream and downstream genes in silkworm. Mol Genet Genomics 294:849–859
Wang Y, Wang F, Wang R, Zhao P, Xia Q (2015) 2A self-cleaving peptide-based multi-gene expression system in the silkworm Bombyx mori. Sci Rep 5:16273
Liu Z, Chen O, Wall JBJ, Zheng M, Zhou Y, Wang L, Vaseghi HR, Qian L, Liu J (2017) Systematic comparison of 2A peptides for cloning multi-genes in a polycistronic vector. Sci Rep 7:2193
Schetelig MF, Targovska A, Meza JS, Bourtzis K, Handler AM (2016) Tetracycline-suppressible female lethality and sterility in the Mexican fruit fly, Anastrepha ludens. Insect Mol Biol 25:500–508
Yan Y, Williamson ME, Davis RJ, Andere AA, Picard CJ, Scott MJ (2020) Improved transgenic sexing strains for genetic control of the Australian sheep blow fly Lucilia cuprina using embryo-specific gene promoters. Mol Genet Genomics 295:287–298
Yan Y, Linger RJ, Scott MJ (2017) Building early-larval sexing systems for genetic control of the Australian sheep blow fly Lucilia cuprina using two constitutive promoters. Sci Rep 7:2538
Acknowledgements
We thank Bashir Hosseini for technical assistance and Carl Stein for the help with RT-PCR. S2 cells were thankfully received by Dr. Denise Salzig (THM). This work was supported by the Emmy Noether program of the German Research Foundation (SCHE 1833/1-1; to MFS), the Fraunhofer Attract program (‘Applications for population control of D. suzukii’; to MFS), and the LOEWE Center for Insect Biotechnology and Bioresources of the HMWK.
Author information
Authors and Affiliations
Contributions
SAJ, YY and JS performed the research. YY and MFS conceived the study. SAJ, YY and MFS analyzed data and wrote the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
We confirm that no author has any conflict of interest to disclose, all authors have approved the version submitted for publication, the work in this article is original and has not been published previously, and the article is not under consideration by any other journal.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The GenBank accession numbers are as follows: Dshid mRNA: MN982930; Dsgrim mRNA: MN982931; and Dsrpr mRNA: MN982932.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Jaffri, S.A., Yan, Y., Schwirz, J. et al. Functional characterization of the Drosophila suzukii pro-apoptotic genes reaper, head involution defective and grim. Apoptosis 25, 864–874 (2020). https://doi.org/10.1007/s10495-020-01640-2
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10495-020-01640-2