Abstracts
Strain Y1T, a Gram-negative, non-spore-forming, rod-shaped bacterium, was isolated from activated sludge. This strain is able to degrade several commonly used chloroacetamide herbicides, such as butachlor, acetochlor and alachlor. Phylogenetic analysis based on 16S rRNA gene sequences revealed that strain Y1T is a member of the genus Sphingomonas and shows high sequence similarities with S. starnbergensis 382T (95.7 %), S. sanxanigenens NX02T (95.7 %) and S. haloaromaticamans A175T (95.3 %), and shows low (<95 %) sequence similarities to all other Sphingomonas species. Chemotaxonomic analysis revealed that strain Y1T possesses Q-10 as the predominant ubiquinone, C14:0 2-OH as the major 2-hydroxy fatty acid and sym-homospermidine as the major polyamine. The main cellular fatty acids of strain Y1T were found to be C18:1 ω7c (38.2 %), C16:1 ω6c/C16:1 ω7c (28.5 %), C16: 0 (10.7 %) and C14:0 2-OH (14.3 %). The main polar lipids were determined to be diphosphatidylglycerol, phosphatidylglycerol, phosphatidylethanolamine, sphingoglycolipids (SGL1-SGL3), phosphatidyl dimethylethanolamine and aminophospholipid. The DNA G+C content was found to be 66 ± 0.4 mol%. Based on phylogenetic analysis, phenotypic characteristics and chemotaxonomic data, strain Y1T is considered to represent a novel species of the genus Sphingomonas, for which the name Sphingomonas chloroacetimidivorans sp. nov. is proposed. The type strain is Y1T (=CCTCC AB 2011178T = KACC 16607T).
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Yabuuchi et al. (1990) first described the genus Sphingomonas, which belongs to the α-proteobacteria (Anzai et al. 2000; Lee et al. 2005). Based on refined phylogenetic, chemotaxonomic and physiological analysis, sphingomonads have recently been divided into four genera: Novosphingobium, Sphingobium, Sphingomonas and Sphingopyxis (Takeuchi et al. 2001). Six additional genera have been classified within the family Sphingomonadaceae, namely Sandarakinorhabdus (Gich and Overmann 2006), Sphingosinicella (Maruyama et al. 2006), Stakelama (Chen et al. 2010), Parasphingopyxis (Uchida et al. 2012), Sphingomicrobium (Kämpfer et al. 2012) and Sphingorhabdus (Jogler et al. 2013). Sphingomonas are Gram-negative, rod-shaped, strictly aerobic, orange-, yellow- or whitish-brown-pigmented bacteria, having sphingoglycolipids in the outer membrane and lacking lipopolysaccharides (White et al. 1996). Chemotaxonomic studies have demonstrated that Sphingomonas strains contain C18:1 ω7c, C16:0 and/or C17:0 as the major fatty acids, C14:0 2-OH as the major hydroxylated fatty acid, Q-10 as the major respiratory quinone and sym-homospermidine as the major polyamine (Takeuchi et al. 2001). The polar lipid pattern comprises diphosphatidylglycerol (DPG), phosphatidylglycerol (PG), sphingoglycolipid (SGL), phosphatidylethanolamine (PE), phosphatidyldimethylethanolamine (PDME) and phosphatidylcholine (PC) (Busse et al. 1999; Wittich et al. 2007). Members of Sphingomonas represent environmental isolates that play important roles in the biodegradation of organic pollutants. Because of their metabolic diversity, pollutant-degrading Sphingomonas strains can serve as potential microbial agents in the remediation of contaminated soils (White et al. 1996). At the time this manuscript was drafted, this genus comprised 69 recognized species (http://www.bacterio.cict.fr/s/sphingomonas.html). Chloroacetamide herbicides are among the most important class of pre-emergence herbicides used in agriculture. Several studies have demonstrated that these herbicides are highly toxic to some aquatic organisms, and the residues in soil consistently injure subsequent rotation crops, particularly in sandy soils with low organic matter (Zhang et al. 2011). Microbes play significant roles in degrading and detoxifying chloroacetamide herbicide residues in the environment (Stamper and Tuovinen 1998).
Materials and methods
In this study, comparative 16S rRNA gene sequences analysis of a newly isolated strain indicated that strain Y1T belongs to the genus Sphingomonas and shows <96 % similarities with other Sphingomonas species. Based on the results of a polyphasic taxonomic study, this strain is considered to represent a novel species of the genus Sphingomonas.
Bacterial strains, isolation and cultivation
While screening for chloroacetamide herbicide-degrading isolates, 5 g of sludge sample collected from activated sludge of a chloroacetamide herbicide-manufacturing wastewater treatment facility in Kunshan, Jiangsu Province, China (N31°22′ E120°56′) was added into 100 mL sterile MSM medium containing 100 mg/L butachlor and incubated in a rotary shaker at 180 rpm at 30 °C for 4 days. 5 mL of the enriched culture was transferred to another 100 mL fresh MSM medium. This procedure was repeated three times. Then the mixture was diluted in a tenfold series and a 100 μL sample of each dilution was spread onto R2A plates with 100 mg/L butachlor. Finally, a strain with the ability to degrade several chloroacetamide herbicides was isolated and named Y1T. In the present study, Sphingomonas starnbergensis 382T, S. sanxanigenens NX02T, S. haloaromaticamans A175T and S. fennica K101T, which were purchased from the culture collections of DMSZ and KACC, were used as reference strains for phenotypic characterization. Unless otherwise indicated, the morphological, physiological and biochemical characteristics of strain Y1T and the reference strains were observed after routine cultivation on R2A agar or in R2A broth at 30 °C.
Phenotypic characterization
The Gram reaction was determined using the non-staining method of Buck (1982). Cell morphology was determined through inverted microscopy (IX70; Olympus) and transmission electron microscopy (H-7650; Hitachi). The gliding motility was determined using the hanging-drop method (Bernardet et al. 2002). Growth at various temperatures (4, 10, 15, 20, 25, 30, 37, 40, 45 and 50 °C), salt concentrations (0.5–7 % NaCl with increments of 0.5 %, w/v), and pH values (pH 4.0–10.5 with increments of 0.5 pH units) was assessed in R2A broth and after incubation for up to 5 days. The pH was maintained using four different buffers: 100 mM citric acid-sodium citrate buffer (pH 4.0–6.0), 50 mM phosphate buffer (pH 5.5–8.0), 50 mM Tris–HCl buffer (pH 7.5–9.0) and 20 mM glycine-NaOH buffer (pH 8.5–10.5). Growth on nutrient agar, Trypticase Soy agar and MacConkey agar was evaluated after incubation at 28 °C for 5 days. Catalase and oxidase production was tested according to McCarthy and Cross (1984). Growth under anaerobic conditions was determined in R2A broth supplemented with or without 0.1 % (w/v) nitrate using the GasPak Anaerobic System (BBL) according to the manufacturer’s instructions. The degradation of DNA (in which DNase agar plates were flooded with 1 M HCl to detect DNase activity), casein, chitin, starch and CM-cellulose were investigated according to Smibert and Krieg (1994). Nitrate reduction, indole production, urease and gelatinase tests, and the assimilation and oxidation of various carbon compounds were performed using the API 20NE Kit (bioMérieux) and the Biolog GN2 System according to the manufacturer’s instructions. Sensitivity to antibiotics was tested on R2A agar plates using discs containing the following antibiotics: erythromycin (15 µg), clindamycin (2 µg), gentamicin (10 µg), chloramphenicol (30 µg), streptomycin (10 µg), roxithromycin (15 µg), lincomycin (2 µg), carbenicillin (100 µg), piperacillin (100 µg) and vancomycin (30 U). The ability to degrade butachlor, acetochlor, alachlor and metolachlor was determined according to the methods of Zhang et al. (2011). Each chloroacetamide herbicide was added at a final concentration of 100 mg/L.
16S rRNA gene phylogenetic analysis and genomic DNA G+C content determination
Genome DNA was purified in lysis solution containing 100 μg/mL proteinase K and 1 % (w/v) SDS, followed by phenol/chloroform extraction and 2-propanol precipitation according to standard procedures (Sambrook and Russell 2001). The nearly complete 16S rRNA gene sequence was obtained through PCR amplification using a set of universal primers, 5′-AGAGTTTGATCCTGGCTCAG-3′ (positions 8–27 in Escherichia coli 16S rRNA) and 5′-TACCTTGTTACGACTT-3′ (positions 1492-1507 in E. coli 16S rRNA), according to Lane (1991). The 16S rRNA gene sequence of strain Y1T was a continuous stretch of 1447 bp. Pairwise sequence similarity was calculated using a global alignment algorithm, implemented in the EzTaxon-e server (http://www.ezbiocloud.net/eztaxon; Kim et al. 2012). The 16S rRNA gene sequence alignment was performed using the CLUSTAL_X program (Thompson et al. 1997). Phylogenetic trees were constructed using the neighbor-joining method (Saitou and Nei 1987) and maximum-likelihood (Felsenstein 1981) with Kimura’s two-parameter calculation model in MEGA version 5.0 (Tamura et al. 2011). The bootstrap analysis of 1000 resamplings was used to evaluate the tree topology (Felsenstein 1985). The G+C content of the genomic DNA was determined through thermal denaturation (Mandel and Marmur 1968) using E. coli K-12 as a standard.
Determination of fatty acid and isoprenoid ubiquinone
The fatty acid profiles of strain Y1T and the related Sphingomonas species were determined according to the manufacturer’s instructions (Sherlock Microbial Identification System; MIDI Corporation) (Sasser 1990). All the strains were grown in R2A broth and harvested at the mid-exponential phase through centrifugation, washed with distilled water and freeze-dried. The fatty acid methyl esters were obtained from cells through saponification, methylation and extraction, and separated using gas chromatography (Agilent 6890N). The peaks were automatically integrated and fatty acid names and percentages were determined using the MIDI Sherlock MIS system (Library: TSBA6; Version, 6.0B). The polar lipid analysis of strain Y1T was performed at the Identification Service of the Deutsche Sammlung von Mikroorganismen und Zellkulturen (DSMZ, Braunschweig, Germany) as described by Tindall (1990). Quinone was extracted according to Collins et al. (1977) and separated through high performance liquid chromatography analysis (HPLC) (Tamaoka et al. 1983). Polyamines were extracted according to Busse and Auling (1988) and analyzed according to Su et al. (2006). For HPLC analysis, a Reverse-Phase C18 column (Agilent, USA, 100 mm × 2.1 mm; particle size 1.8 µm) was used; the mobile phase was methanol:acetonitrile:H2O (48:2:50, V:V), and the flow rate was 0.4 mL/min; the detection wavelength was 254 nm and the injection volume was 20 μL.
Results and discussion
Phenotypic characteristics
Strain Y1T was found to be Gram-negative, aerobic, non-spore-forming, and non-motile short rod (0.5–0.8 µm in width and 0.8–1.2 μm in length, Supplementary Fig. S2). Colonies grown on R2A agar plates for 3 days were observed to be smooth, circular, convex and yellow. The morphological, physiological and biochemical characteristics of strain Y1T are provided in Table 1, which lists the characteristics that differentiate strain Y1T from its closest phylogenetic relatives. Strain Y1T is able to degrade several commonly used chloroacetamide herbicides, such as butachlor, acetochlor and alachlor. When the inoculum was 5 % (v/v) and the initial concentration of different chloroacetamide herbicides was 100 mg/L, 88.4 ± 11.5 % of alachlor, 72.2 ± 6.3 % of acetochlor and 68.5 ± 8.7 % of butachlor were degraded after incubation for 7 days at 30 °C. Strain Y1T was found to be sensitive to erythromycin, gentamicin, chloramphenicol, roxithromycin, lincomycin and carbenicillin, while resistant to clindamycin, streptomycin, piperacillin and vancomycin.
16S rRNA gene sequence analysis
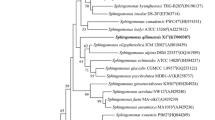
A nearly full-length 16S rRNA gene sequence of strain Y1T was determined (1450 nt, GenBank accession number JQ728997). In the neighbor-joining phylogenetic tree (Fig. 1), strain Y1T groups within Sphingomonas species and forms a subclade with Sphingomonas fennica K101T, S. formosensis CC-Nfb-2T, S. starnbergensis 382T, S. haloaromaticamans A175T, S. laterariae LNB2T, S. wittichii RW1T and S. histidinilytica UM2T. According to sequence similarity, strain Y1T is closely related to Sphingomonas starnbergensis 382T (95.7 %), S. sanxanigenens NX02T (95.7 %) and S. haloaromaticamans A175T (95.3 %), and shows <95 % similarity with other Sphingomonas species. Moreover, this clade was strongly supported by the maximum-likelihood tree (Supplementary Fig. S1). This branching pattern demonstrated that strain Y1T represents a novel species within the genus Sphingomonas. The DNA G+C content of strain Y1T was found to be 66 ± 0.4 mol%, consistent with the genus Sphingomonas (Takeuchi et al. 2001).
A neighbor-joining phylogenetic tree based on 16S rRNA gene sequences, showing the relationships between strain Y1T and members of the family Sphingomonadaceae. Bacillus subtilis DSM10T (AJ276351) was used as the outgroup. Percentage bootstrap values based on 1000 replicates are shown at branch nodes; only values at or above 50 % are shown. Bar, 0.02 substitutions per nucleotide position
Chemotaxonomic characteristics
Chemotaxonomic analysis showed that strain Y1T possesses Q-10 (99.1 %) as the major ubiquinone, but small amounts (0.9 %) of Q-11 could also be detected. This result is consistent with the genus Sphingomonas (Takeuchi et al. 2001). The fatty acid profiles of strain Y1T, S. starnbergensis 382T, S. sanxanigenens NX02T, S. haloaromaticamans A175T and S. fennica K101T are shown in Table 2. The major fatty acids (>3 %) of strain Y1T were determined to be C18:1 ω7c (38.2 %), C16:1 ω6c/C16:1 ω7c (28.5 %), C14:0 2-OH (14.3 %) and C16:0 (10.7 %). The fatty acid profile is characteristic for species of the genus Sphingomonas (Busse et al. 1999). However, some qualitative and quantitative differences in the proportions were observed between the isolate and the reference strains. C19:0 cyclo ω8c, C18:1 ω5c and C14:0 are not detected in strain Y1T, in contrast to other four tested strains. Moreover, compared to these four strains, strain Y1T possesses higher levels of C14:0 2-OH and C16:1 ω6c/C16:1 ω7c.
The polar lipids of strain Y1T were determined to be DPG, PG, PE, sphingoglycolipids (SGL1-SGL3), PDME, aminophospholipid (APL), and PC (Supplementary Fig. S3). The polyamine pattern of strain Y1T was determined to contain the major compound sym-homospermidine (14.35 μmol/g dry weight), accompanied by small amounts of spermidine and putrescine of (2.07 and 0.17 μmol/g dry weight, respectively). Both the polar lipids and polyamines of strain Y1T are in good agreement with the characteristics of the genus Sphingomonas (Busse et al. 1999; Takeuchi et al. 2001).
Taxonomic conclusion
The results of the phylogenetic analysis, phenotypic analysis and chemotaxonomic studies presented above support the conclusion that strain Y1T belongs to the genus Sphingomonas, while phylogenetic distinctiveness and some phenotypic differences (Table 1) confirmed that strain Y1T represents a species distinct from the recognized Sphingomonas species. Therefore, strain Y1T should be classified as representing a novel species of the genus Sphingomonas, for which the name Sphingomonas chloroacetimidivorans sp. nov. is proposed.
Description of Sphingomonas chloroacetimidivorans sp. nov
Sphingomonas chloroacetimidivorans (chloroacetamidivorans. N.L. n. chloroacetimidum, chloroacetimide herbicide; L. part. adj. vorans, devouring; N.L. part. adj. chloroacetimidivorans, chloroacetimide herbicide devouring, degrading).
Cells are Gram-negative, aerobic, non-spore-forming, and non-motile short rods. Colonies grown on R2A agar plates for 3 days are smooth, circular, convex and yellow. Growth occurs in 0–2.0 % (w/v) NaCl (optimum 0.5 %), at 20–40 °C (optimum 30 °C) and at pH 6.0–9.0 (optimum pH 7.0). No growth occurs on Trypticase Soy agar, nutrient agar or MacConkey agar. Gelatin, tyrosine, starch, casein, hydroxyethyl cellulose, aesculin, chitin and DNA are not hydrolyzed. These cells are positive to catalase and negative to oxidase, nitrate reduction, arginine dihydrolase, α-galactosidase, β-galactosidase, indole production, urease and N-acetyl-β-glucosaminidase. The following substrates are utilized for growth: propionate, caprate, 3-hydroxybutyrate, acetate. With the Biolog GN2 system, only the following four types of carbon sources are oxidized: dextrin, Tween 40, Tween 80 and acetic acid. The quinone system contains major amounts of Q-10 and lesser amounts of Q-11 and the major fatty acids are C18:1 ω7c, C16:1 ω6c/C16:1 ω7c, C14:0 2-OH and C16:0. The predominant polyamine is sym-homospermidine. The polar lipids comprise DPG, PG, PE, sphingoglycolipids (SGL1-SGL3), PDME, APL and PC. The DNA G+C content of the type strain is 66 ± 0.4 mol%. The type strain Y1T (=CCTCC AB 2011178T = KACC 16607T) was isolated from activated sludge in a chloroacetamide herbicides-manufacturing wastewater treatment facility in Kunshan City, Jiangsu Province, China.
Abbreviations
- KACC:
-
Korean agricultural culture collection
- CCTCC:
-
China center for type culture collection
References
Anzai Y, Kim H, Park JY, Wakabayashi H, Oyaizu H (2000) Phylogenetic affiliation of the pseudomonads based on 16S rRNA sequence. Int J Syst Evol Microbiol 50:1563–1589
Bernardet JF, Nakagawa Y, Holmes B (2002) Proposed minimal standards for describing new taxa of the family Flavobacteriaceae and emended description of the family. Int J Syst Evol Microbiol 52:1049–1070
Buck JD (1982) Nonstaining (KOH) method for determination of gram reactions of marine bacteria. Appl Environ Microbiol 44:992–993
Busse HJ, Auling G (1988) Polyamine pattern as a chemotaxonomic marker within the Proteobacteria. Syst Appl Microbiol 11:1–8
Busse HJ, Kämpfer P, Denner EBM (1999) Chemotaxonomic characterization of Sphingomonas. J Ind Microbiol Biotechnol 23:242–251
Chen C, Zheng Q, Wang YN, Yan XJ, Hao LK, Du X, Jian N (2010) Stakelama pacifica gen. nov., sp. nov., a new member of the family Sphingomonadaceae isolated from the Pacific Ocean. Int J Syst Evol Microbiol 60:2857–2861
Collins MD, Pirouz T, Goodfellow M, Minnikin DE (1977) Distribution of menaquinones in actinomycetes and corynebacteria. J Gen Microbiol 100:221–230
Felsenstein J (1981) Evolutionary trees from DNA sequences: a maximum likelihood approach. J Mol Evol 17:368–376
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791
Gich F, Overmann J (2006) Sandarakinorhabdus limnophila gen. nov., sp. nov., a novel bacteriochlorophyll a-containing, obligately aerobic bacterium isolated from freshwater lakes. Int J Syst Evol Microbiol 56:847–854
Jogler M, Chen H, Simon J, Rohde M, Busse HJ, Klenk HP, Tindall BJ, Overmann J (2013) Description of Sphingorhabdus planktonica gen. nov., sp. nov., and reclassification of three related Sphingopyxis species as members of the novel genus Sphingorhabdus. Int J Syst Evol Microbiol 63:1342–1349
Kämpfer P, Arun AB, Young CC, Busse HJ, Kassmannhuber J, Rosselló-Móra R, Geueke B, Rekha PD, Chen WM (2012) Sphingomicrobium lutaoense gen. nov., sp. nov., isolated from a coastal hot spring. Int J Syst Evol Microbiol 62:1326–1330
Kaur J, Kaur J, Niharika N, Lal R (2012) Sphingomonas laterariae sp. nov., isolated from a hexachlorocyclohexane-contaminated dump site. Int J Syst Evol Microbiol 62:2891–2896
Kim OS, Cho YJ, Lee K, Yoon SH, Kim M, Na H, Park SC, Jeon YS, Lee JH, Yi H, Won S, Chun J (2012) Introducing EzTaxon-e: a prokaryotic 16S rRNA gene sequence database with phylotypes that represent uncultured species. Int J Syst Evol Microbiol 62:716–721
Lane DL (1991) 16S/23S rRNA sequencing. In: Stackebrandt ER, Goodfellow M (eds) Nucleic acid techniques in bacterial systematics. Wiley, Chichester, pp 115–175
Lee KB, Liu CT, Anzai Y, Kim H, Aono T, Oyaizu H (2005) The hierarchical system of the ‘Alphaproteobacteria’: description of Hyphomonadaceae fam. nov., Xanthobacteraceae fam. nov. and Erythrobacteraceae fam. nov. Int J Syst Evol Microbiol 55:1907–1919
Lin SY, Shen FT, Lai WA, Zhu ZL, Chen WM, Chou JH, Lin ZY, Young CC (2012) Sphingomonas formosensis sp. nov., a polycyclic aromatic hydrocarbons degrading-bacterium isolated from agricultural soil. Int J Syst Evol Microbiol 62:1581–1586
Mandel M, Marmur J (1968) Use of ultraviolet absorbance-temperature profile for determining the guanine plus cytosine content of DNA. Methods Enzymol 12B:195–206
Maruyama T, Park HD, Ozawa K, Tanaka Y, Sumino T, Hamana K, Hiraishi A, Kato K (2006) Sphingosinicella microcystinivorans gen. nov., sp. nov., a microcystin-degrading bacterium. Int J Syst Evol Microbiol 56:85–89
Mccarthy AJ, Cross T (1984) A taxonomic study of Thermomonospora and other monosporic actinomycetes. Microbiol-SGM 130:5–25
Nigam A, Jit S, Lal R (2010) Sphingomonas histidinilytica sp. nov., isolated from a hexachlorocyclohexane dump site. Int J Syst Evol Microbiol 60:1038–1043
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Sambrook J, Russell DW (2001) Molecular cloning: a laboratory manual, 3rd edn. Cold Spring Harbor Laboratory, Laurel Hollow
Sasser M (1990) Identification of bacteria by gas chromatography of cellular fatty acids. MIDI Technical Note 101. Newark: MIDI
Smibert RM, Krieg NR (1994) Phenotypic characterization. In: Gerhardt P, Murray RGE, Wood WA, Krieg NR (eds) Methods for general and molecular bacteriology. American Society for Microbiology, Washington
Stamper DM, Tuovinen OH (1998) Biodegradation of the acetanilide herbicides alachlor, metolachlor, and propachlor. Crit Rev Microbiol 24:1–22
Su GX, Zhang WH, Liu YL (2006) Involvement of hydrogen peroxide generated by polyamine oxidative degradation in the development of lateral roots in soybean. J Integr Plant Biol 48:426–432
Takeuchi M, Hamana K, Hiraishi A (2001) Proposal of the genus Sphingomonas sensu stricto and three new genera, Sphingobium, Novosphingobium and Sphingopyxis, on the basis of phylogenetic and chemotaxonomic analyses. Int J Syst Evol Microbiol 51:1405–1417
Tamaoka J, Katayama-Fujimura Y, Kuraishi H (1983) Analysis of bacterial menaquinone mixtures by high performance liquid chromatography. J Appl Bacteriol 54:31–36
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL-X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882
Tindall BJ (1990) Lipid composition of Halobacterium lacusprofundi. FEMS Microbiol Lett 66:199–202
Uchida H, Hamana K, Miyazaki M, Yoshida T, Nogi Y (2012) Parasphingopyxis lamellibrachiae gen. nov., sp. nov., isolated from a marine annelid worm. Int J Syst Evol Microbiol 62:2224–2228
White D, Sutton S, Ringelberg D (1996) The genus Sphingomonas: physiology and ecology. Cur Opin Biotechnol 7:301–306
Wittich RM, BusseHJ Kämpfer P, Macedo AJ, Tiirola M, Wieser M, Abraham WR (2007) Sphingomonas fennica sp. nov. and Sphingomonas haloaromaticamans sp. nov., outliers of the genus Sphingomonas. Int J Syst Evol Microbiol 57:1740–1746
Yabuuchi E, Yano I, Oyaizu H, Hashimoto Y, Ezaki T, Yamamoto H (1990) Proposals of Sphingomonas paucimobilis gen. nov. and comb. nov., Sphingomonas parapaucimobilis sp. nov., Sphingomonas yanoikuyae sp. nov., Sphingomonas adhaesiva sp. nov., Sphingomonas capsulata comb. nov., and two genospecies of the genus Sphingomonas. Microbiol Immunol 34:99–119
Yabuuchi E, Yamamoto H, Terakubo S, Okamura N, Naka T, Fujiwara N, Kobayashi K, Kosako Y, Hiraishi A (2001) Proposal of Sphingomonas wittichii sp. nov. for strain RW1T, known as a dibenzo-p-dioxin metabolizer. Int J Syst Evol Microbiol 51:281–292
Zhang J, Zheng JW, Liang B, Wang CH, Ni YY, Cai S, He J, Li SP (2011) Biodegradation of chloroacetamide herbicides by Paracoccus sp. FLY-8. J Agric Food Chem 59:4614–4621
Acknowledgments
This work was financially supported by the National Science and Technology Support Plan (2012BAD15B03), the National Natural Science Foundation of China (Grant No. 31270157), the Program for New Century Excellent Talents in University (NCET-13-0861) and the National Natural Science Foundation of China (J1210056).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Kai Chen and Qing Chen have contributed equally to this work.
The GenBank/EMBL/DDBJ accession number for the 16S rRNA gene sequence of strain Y1T is JQ728997.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Chen, K., Chen, Q., Wang, GX. et al. Sphingomonas chloroacetimidivorans sp. nov., a chloroacetamide herbicide-degrading bacterium isolated from activated sludge. Antonie van Leeuwenhoek 108, 703–710 (2015). https://doi.org/10.1007/s10482-015-0526-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-015-0526-z