Abstract
The European wildcat is threatened by habitat fragmentation, illegal or incidental killings and hybridization with free-ranging domestic cats. Conservation projects should be based on a sound knowledge of the patterns of wildcat distribution, population size and dynamics. However, this information is scanty, mainly because of the species’ elusive behaviour. In this study, we tested the efficiency of a protocol that integrates the use of non-invasive genetic identifications and camera trapping for wildcat monitoring. The field work was carried out in the Foreste Casentinesi National Park, a protected area in the central Italian Apennines, where wildcat presence has been recently ascertained. DNA samples were extracted from scats collected during the survey and hair tufts trapped by valerian-treated sticks. Individual genotypes were identified using 10 autosomal microsatellites, mtDNA and Y chromosome markers. Additionally, we obtained 30-s long video clips from 20 camera trap stations associated to the hair traps. Our results confirmed the presence of wildcats in the study area. We identified six to nine wildcat individuals (respectively from non-invasive genetic sampling and camera trapping survey). Some of these showed anomalous coat colour patterns (one) or genetic signatures of hybridization (three). We further identified five domestic cats that were sharing parts of wildcats’ territories. We found individual variations in the response to valerian lure. We compared and evaluated the pros and cons of these monitoring methods. We concluded that, if used simultaneously, these methods may considerably increase the efficiency of wildcat detection and the quality of collected data.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The European wildcat (Felis silvestris silvestris Schreber, 1777) has a wide, yet partially fragmented, distribution throughout Europe (Driscoll and Nowell 2010; Lozano and Malo 2012). The species lives in a variety of habitat types. These habitats range from scrub pastures (Lozano et al. 2003; Monterroso et al. 2009; Lozano 2010) to forest patches intermixed with open fields (Klar et al. 2008, 2012) and, more marginally, to coniferous forests with rich undergrowth (Easterbee et al. 1991; Lozano et al. 2003). The European wildcat is a ‘strictly protected’ species included in ‘Annex IV’ of the European Habitats Directive (92/43/CEE). It is also included in ‘Annex II’ of the Bern Convention. The species is of ‘least concern’ for the IUCN (Driscoll and Nowell 2010), and it is ranked as ‘near threatened’ in the Red List of the Italian Vertebrates (Rondinini et al. 2013). The main threats to the European wildcat are the loss of suitable habitats (Klar et al. 2009, 2012), human-caused mortality—in particular, road kills (Nowell and Jackson 1996; Lüps et al. 2002; Schulenberg 2005; Krone et al. 2008), overgrazing by large game species (Lozano et al. 2007) and hybridization with the domestic cat (Felis silvestris catus) (Randi 2008; Oliveira et al. 2008a, b). Driscoll and Nowell (2010) reported a decreasing global population trend. However, despite poorly known distribution ranges, some local populations appear stable (Lozano et al. 2013). Projects for restoring ecological corridors are underway to counteract the consequences of habitat fragmentation and facilitate connections among isolated populations (Vogel and Mölich 2013). In the Italian peninsula, the species’ distribution range covers the entire southern and central Apennines (Cagnolaro et al. 1976; Ragni 1981) (Fig. 1a). Recent findings (Agostini et al. 2010; Tedaldi 2012; Ragni et al. 2014) suggest a northward wildcat expansion, sustained by suitable forested habitat corridors in protected areas along the Apennine ridge (Santolini et al. 2010). However, we cannot reject the alternative hypothesis, even if improbable (Ragni et al. 1994) that low-density populations have previously been missed in these regions. The European wildcat population in north-eastern Italy (Angelici and Genovesi 2003) is connected with the Dinaric-Balkan population (Mattucci et al. 2013). Recent observations (Bologna and Cristiani 2012) suggest the persistence of a north-western isolated population, even if the probable draining of the French source (Stahl and Artois 1994) makes its consistency uncertain. In Sardinia, the African wildcat Felis silvestris libyca Forster, 1780, is widely distributed. However, in Italy, both conservation status and population dynamics of the European wildcat are still poorly known.
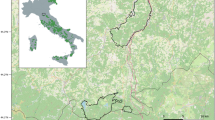
a Study area, located in the northern portion of the Foreste Casentinesi National Park, northern Apennines. b The sampling area in the Foreste Casentinesi National Park with the buffer area (grey), the minimum convex polygon and the additional buffer strip (dotted line) on which estimation of capture density was based. The legend shows the typologies of sampling sites (hair traps, camera stations and wildcat faecal depositions)
Reliable estimates of population abundance and trends are the key baseline data to assess the impact of threatening factors and outline sound conservation guidelines (see Council of Europe 1993). Wildcats have been monitored by direct sightings (Hartmann et al. 2013), live trapping (Bizzarri et al. 2010a), radio tracking (Monterroso et al. 2009; Bizzarri et al. 2010b) and non-invasive techniques such as camera trapping (Can et al. 2011; Kilshaw and Macdonald 2011; Anile et al. 2012a), scat surveys (Lozano et al. 2003; Lozano et al. 2013; Anile et al. 2014), hair trapping (Hupe and Simon 2007; Kéry et al. 2011; Steyer et al. 2013) and, opportunistically, through the collection of road kills (O’Brien et al. 2009; Hartmann et al. 2013). Each method has its own technical drawbacks. Even if morphological classification proved to be reliable for the wildcat (Randi et al. 2001; Lecis et al. 2006; Mattucci et al. 2013), the identification of individuals and hybrids by camera trapping can be difficult (Foster and Harmsen 2012; Kilshaw and MacDonald 2011). Scat surveys may fail to detect several individuals (Anile et al. 2014) and may be biased by identification mistakes in areas of sympatry of wild and free-ranging domestic cat (Monterroso et al. 2013). Genetic identifications can be flawed by variable genotyping success rates (Ruell and Crooks 2007; Anile et al. 2014), and responses to bait lures may be variable, depending on the individual (Kilshaw and Macdonald 2011; Monterroso et al. 2011; Anile et al. 2012b). Thus, it is advisable to integrate different methodologies as to balance the pros and cons of each. Recently, Anile et al. (2014) successfully assessed the population density of the wildcat population on the Etna volcano (Sicily) using both camera trapping and genetic analyses of faecal DNA. The integration of these methods has never been used to monitor wildcat populations in the Apennines. In this study, we planned to test and evaluate the integration of three non-invasive sampling techniques: individual identifications through camera trapping, genetic analyses of faecal DNA obtained from opportunistically collected scats and genotyping from hair trapping with valerian lure sticks in a systematic sampling design. We aimed to assess the feasibility and effectiveness of these methods for monitoring low-density wildcat populations.
Materials and methods
No animals were physically captured and manipulated during the work of this study. All the data were collected using non-invasive methods.
Study area
Based on the available information on wildcat presence, obtained from published records, forest rangers, wildlife technicians and preliminary field surveys (Cagnolaro et al. 1976; Agostini et al. 2010), we identified a 2,800-ha large study area within the northern sector of the Foreste Casentinesi National Park (Emilia Romagna, Italy; Fig. 1a). This area, which ranges from 600 to 1,150 m.a.s.l., is the northernmost Apennine location where the presence of the European wildcat has been recently confirmed (Ragni 2003; Ragni and Petruzzi 2010; Ragni et al. 2014). The Foreste Casentinesi National Park (36,000 ha) features over 29,000 ha of mixed woodlands (Fagus sylvatica, Quercus cerris, Quercus pubescens, Fraxinus excelsior, Castanea sativa, with introduced coniferous such as Pinus nigra, Picea abies, etc.) spaced out with wide pasturelands, grasslands and clearings. The fully protected centuries-old forests, offering a rich mammal community and low human density (about four residents per km2), represent a very suitable habitat for the European wildcat on the Apennine ridge (Santolini et al. 2010).
Genetic sampling protocol
We collected hair and scat samples within a 4 × 7-km grid, widely partitioned in 28 1 × 1-km large cells (Fig. 1b). We systematically placed a total of 45 raw pine sticks (60 × 4 × 4 cm) to uniformly cover the grid using, where possible, at least one lure in each cell (Hupe and Simon 2007; Weber et al. 2008; Kéry et al. 2011; Hartmann et al. 2013; Steyer et al. 2013). Each stick was identified with a code, geo-localized and drenched with valerian (Valeriana officinalis) hydroalcoholic tincture (70 %). Additionally, we made a hole of about 2 × 7 cm, longitudinally, at the top of each stick. We made two smaller holes, transversely, on each side and filled them with valerian root powder to obtain a stronger, uniform and longer lasting effect—even on rainy days. Valerian is a strong attractant that also elicits a rubbing behaviour (Monterroso et al. 2011). It has been used in several studies on wildcats (Hupe and Simon 2007; Weber et al. 2008; Kéry et al. 2011; Steyer et al. 2013). In order to catch as many hairs as possible, we scratched the surface of the sticks and applied a strip of bi-adhesive tape. We designed a ca. 25-km-long transect, linking the trap stations that were walked every 7–10 days to collect both hair and occasional scat samples, with a total effort of 17 two-day sampling sessions and 425 km of walked trails. To assess the capture density (number of individuals sampled per 10 km2), we traced a minimum convex polygon (MCP) considering the outermost pickets (22.4 km2, Fig. 1b). Considering the smallest home range used by a wildcat in central Italy (277.71 ha [95 % Kernel area] for an adult male, Bizzarri et al. 2010b), we further added a buffer strip of 939 m to the MCP (for a total of 42.3 km2, Fig. 1b).
Sampling was carried out between November 19, 2012 and June 24, 2013. However, heavy snowfalls between January and February 2013 forced us to stop the surveys for about 30 days. We inspected the lure sticks every 7–10 days. Attached hairs were removed with forceps and stored in an envelope with silica gel to keep the samples dry and to avoid the degradation of DNA. In order to prevent contamination, after each sampling event, the forceps were flamed and the lure sticks were scratched with an iron brush to remove any residual hairs. Finally, a new tape strip and an adequate quantity of valerian lure were applied. Scats were collected using sterile disposable gloves. The samples were stored in ethanol 96 % and frozen as soon as possible. We only collected samples that seemed to be recent and well hydrated.
Genetic analyses
DNA was extracted from both hair and faecal samples using the Blood & Tissue Kit® (Qiagen), following manufacturer’s instructions. Furthermore, hair samples were processed adding 20 μl of dithiothreitol to the digestion mix to efficiently degrade the keratin skeleton (McNevin et al. 2005).
We minimized contamination risks by using a laboratory dedicated to the pre-polymerase chain reaction (PCR) handling of non-invasively collected samples (Taberlet et al. 1999). Negative and positive controls were run alongside all reactions to monitor possible cross-contamination during extraction and amplification.
We sequenced and BLASTed a 719-bp portion of the mitochondrial DNA control region (mtDNA CR; sites 16,236–16,955, Tiedemann et al. 1996; Freeman et al. 2001, see supplementary material for the detailed sequencing protocol) to exclude samples that did not belong to Felis silvestris.
We further sequenced 877 bp (including the primers) of the mtDNA NADH dehydrogenase subunit 5 (ND5; nucleotides 13,131–14,007 mapped on the mitochondrial genome of the domestic cat; NCBI Reference Sequence NC001700, Lopez et al. 1996, see supplementary material), which, according to Driscoll et al. (2011), contains seven diagnostic single nucleotide polymorphisms (SNPs) discriminating European wildcats (F. s. silvestris) and domestic cats (F. s. catus). The sequences were aligned using SEQSCAPE 2.5 (Life Technologies) and trimmed using BIOEDIT 7.1.11 into equal sequences of 671 bp (positions 13,243–13,913). The complete sequence of mtDNA genome of the domestic cat (NCBI Reference Sequence: NC_001700, Lopez et al. 1996), trimmed at the positions mentioned above, was used as reference. The haplotype number and frequency were estimated using DNAsp 5.10.01. The ND5 alignment was used to obtain a network in NETWORK 4.6 (Fluxus Technology Ltd.) using a median-joining (MJ) algorithm (Bandelt et al. 1999) with ε = 10 and a transversions/transition weight of 3:1. We then cleaned up the resulting scheme using a maximum parsimony calculation (MP) (Polzin and Daneschmand 2003). The network was used to identify the mtDNA haplogroups and assign the haplogroups to the two cat subspecies.
The samples were then amplified at 10 autosomal microsatellite loci (STR; FCA23, FCA26, FCA43, FCA58, FCA77, FCA88, FCA96, FCA126, FCA132 and FCA149, Menotti-Raymond and Obrien 1995; Menotti-Raymond et al. 1997) five duplex reactions and one Y-linked STR (SMCY-7) that should show fixed alternative alleles in the two cat subspecies (Luo et al. 2007; Nussberger et al. 2013; see supplementary material).
The individual genotypes were determined in a multiple-tube approach with a mean of 4.68 (±0.79 SD) replicates per locus per individual. We determined the reliability (R) of each genotype and checked if further replicates were needed to reach R = 95 % in RELIOTYPE (Miller et al. 2002). The data set was used to assess the rate of allelic dropout (ADO) and false alleles (FA; Taberlet et al. 1999). Using the match function in GENALEX 6.501 (Peakall and Smouse 2006, 2012), we detected individuals that were sampled more than once. We used STRUCTURE 2.3.4 (Pritchard et al. 2000) to assign the genotypes to a domestic or wildcat reference population, assuming K = 2 genetic clusters (Oliveira et al. 2008a; O’Brien et al. 2009) and an individual proportion of membership threshold Qi = 0.8 (Pierpaoli et al. 2003; Oliveira et al. 2008a). Genotypes with intermediate Qi values were considered as putative hybrids. We used a panel of 77 free-living or house domestic cats, 235 putative European wildcats and 17 known silvestris × catus hybrids as a reference for the calculation of probability of identity among siblings (PIDsibs; Mills et al. 2000; Waits et al. 2001), chromosome Y subspecies assessment and mitochondrial and STRUCTURE analysis. These were collected in Italy from 2003 to 2010 and already analysed at 35 STRs (Mattucci et al. 2013). We ran 400,000 Markov Chain Monte Carlo (MCMC) steps after discarding the first 40,000 steps as burn-in, under the admixture model with correlated allele frequencies (Hertwig et al. 2009; Eckert et al. 2010). The PIDsib was calculated in GENALEX 6.501 (Peakall and Smouse 2006, 2012).
Camera trapping protocol
Ten camera traps (five Multipir12® and five Multipir12-HD®), with one passive infrared/motion front trigger sensor and two lateral preparation sensors, were tied to trees at about 2 m to the lured pickets. In accordance with the manufacturer’s instructions, the delay between the detection of the sensors and the triggering of the camera was about 1 s. Cameras were set on video mode with a video length of 30 s and an interval between consecutive shots of 60 s. Each camera was equipped with a 4-GB SDHC card and was powered by four rechargeable AA batteries. In order to avoid any interaction between animals and cameras, we chose an infrared flashlight of 940 nm. Due to technical issues, the cameras were placed on March 28 (one was placed on 8 April) and worked until 24 June with an average trapping effort of 43 nights each and a total of 819 trap days. After the first 45 days, all of the 10 cameras (except one that was withdrawn on 8 April due to malfunctioning) were simultaneously shifted to be associated to other hair traps in order to cover 20 sampling stations. According to the home range size estimated for and adult male wildcat (277.71 ha) considered above (see “Genetic sampling protocol”), we spaced the camera traps (mean distance = 1,039 m ± 368 m) so as to cover a continuous area, including non-monitored areas that were too small to host a whole wildcat home range. Consequently, the individuals with a home range greater than 277.7 ha were exposed to a greater number of traps (Otis et al. 1978; Karanth and Nichols 2002). Thus, to assess capture density (in this case, the number of individuals captured by camera traps per 10 km2), we considered a total monitored area of 30.4 km2 (the grey area in Fig. 1b).
We based the subspecies and the individual identification on coat colour patterns and body proportions of the animals (French et al. 1988; Ragni and Possenti 1996). In particular, according to the quality of the videos, we considered the number and distance of the tail rings and the proportional length of the dorsal stripe with respect to the body. We also considered the presence and shape of any additional sign on the pelage (Anile et al. 2012a). Furthermore, we considered the behaviour and body proportions to also infer sex and age.
To investigate the reactions towards the bait, we compiled an ethogram (Wells and Egli 2004; Ellis and Wells 2010). This included seven possible behaviours: indifference (I), curiosity (C), facial marking (FM), strong interaction (SI), spray marking (SM), diffidence (D) and fear (F) (Table 1). If two or more behaviours occurred during the same shooting, we only considered the strongest one (e.g. if a cat displayed curiosity followed by facial marking and strong interaction, we only considered the ‘strong interaction’ event).
To check for seasonal variation in the number of samples collected (from genetic sampling or from camera trapping), a Poisson regression model was used with the number of session elapsed as the independent variable and considering the logarithm as the link function.
Results
Genetic identifications
We collected a total of 63 non-invasive samples (30 hair and 33 scat samples). Out of these samples, 12 (36.7 %; one scat and 11 hair samples) were assigned to non-target species (one dog, six wild boars, one wolf, one badger, two beech martens and one brown hare), according to their mtDNA CR sequences.
Eighteen wildcat samples (eight hair and 10 scats) out of 51 (35.3 %) were successfully genotyped at the mtDNA CR and/or the STR panel. Furthermore, 17 yielded reliable ND5 mtDNA haplotypes (eight from hairs and nine from scats), while eight were successfully genotyped with STR markers (two and six, respectively). With regard to the sampling strategy, the capture success rate was 0.082 genotyped hair samples per 100 trap nights and one scat per 42.5 km. All confirmed wildcat biological samples were only found between March and June.
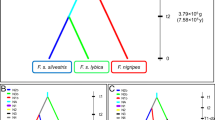
The rate of ADO was 12.1 % while the rate of FA was 1.6 % across all PCR reactions. The 10 autosomal STR loci yielded a value of PIDsib = 0.0001. None of the samples showed more than two alleles. This suggests that no contamination occurred among them. Microsatellite analyses allowed the detection of six individuals (three males and three females). The test performed with STRUCTURE assigned all individuals to the F. s. silvestris subspecies with Qi values >0.8 (Fig. 2). Using mitochondrial analyses of ND5 subunit, two main haplogroups (wildcat (W) and domestic cat (D), Fig. 3) were identified: the W haplogroup and the D haplogroup. We identified as pure wildcats only the individuals with concordant attributions at all the three markers (STR, mtDNA and SMCY-STR [for male], Table 2). Only one individual (individual 1, Table 2) met these requirements. Three individuals (3, 4 and 6, Table 2) were attributed to the wildcat according to their nuclear genotype (Qi > 0.9). However, their mtDNA haplotypes showed all the polymorphisms previously identified in domestic cats (H1 and H5 in Fig. 3). Furthermore, the samples belonging to one of these individuals (individual 3 in Table 2) were found about 1 km away from the nearest human settlement. Individual 6 was also captured by the camera traps and showed a wildcat phenotype (Fig. 4). Two individuals (2 and 5, Table 2) assigned to the wildcat cluster by microsatellite analyses did not yield reliable mitochondrial haplotypes.
Proportions of posterior probability assignments as inferred by microsatellite analysis in structure. Light grey corresponds to wildcat proportion membership, dark grey corresponds to domestic proportion membership. 1 = wildcat reference population; 2 = domestic cat reference population; 3 = admixed reference population; 4 = samples collected in this study
The network representing the phylogenetic relationships between the Italian reference samples (Mattucci et al. 2013), among which are those produced by our sampling campaign (this study). The two haplogroups (W and D) identify the samples, previously analysed with STR, through the presence of the diagnostic mutations indicated by Driscoll et al. (2011)
Considering the MCP (Fig. 1b), we found 2.6 captures/10 km2. Taking into account the added buffer, the rate changed to 1.41 captures/10 km2.
We found a slight, yet significant, positive dependency between the number of sessions elapsed since the first inspection and the number of samples (hairs and scats) collected per session (βk = 0.22; p < 0.01).
Camera trapping layer
We obtained a total of 570 animal captures. Out of these, 35 (6.1 %) were F. silvestris. Based on the coat colour marking patterns, size and proportion of the body and behaviour, 25 were referable to F. s. silvestris, five to F. s. catus and, at least, one to a putative hybrid. Out of the 35 videos, 20 (57.1 %) were useful to detect at least nine different individuals of wildcat - five males and four females (including the putative hybrid) and five different individuals of domestic cat (three males and two females). One wildcat female and the putative hybrid appeared in an advanced state of pregnancy. The total capture rate of wildcats was 3.1 captures/100 trap days. Considering the total area of 30.54 km2, we calculated 2.9 wildcats captures/10 km2 and 1.6 domestic cats capture/10 km2. The activity patterns of the wildcats in the study area were mainly nocturnal (76 % were captured between 9:00 pm and 5:00 am). Moreover, 92.3 % of the capture events occurred between May and June, with a significant positive dependency on the number of sessions elapsed (βk = 0.65; p < 0.01). Considering all of the wildcat captures, in 51.7 % of cases, the individuals showed no interest in the lures, while in 20.7 %, hairs were successfully trapped (Table 1). Regarding the single recognized individuals, six out of the nine detected individuals (four males and two females, individuals B, E, F, G, H, I) showed an interest in the lures. Four of them (two males and two females, individuals B, F, G, H) scratched on the picket leaving hair samples (FM or SI), while only one (male individual E) performed spray marking (Table 2).
Discussion
The concomitant use of three monitoring methods allowed us to detect between six and nine wildcat individuals in the study area (1.4–2.9 individuals sampled per 10 km2) and, at least, five domestic cats (1.6 individuals sampled per 10 km2).
Hair trapping proved to be the least efficient method to sample wildcat. These results are consistent with previous studies on carnivores (Long et al. 2007; Comer et al. 2011; Monterroso et al. 2014). The hair trapping success rate (0.08/100 trap days) was similar to that reported by recent surveys for felids (0.07/100 trap days, Steyer et al. 2013; 0.015/100 trap days, Garcia-Alaniz et al. 2010). The low capture efficiency of hair traps may depend on the variable response of wildcats to valerian lures. In a study by Monterroso et al. (2011), only 11.5 % of the wildcats detected showed an investigative behaviour towards the bait. Furthermore, Anile et al. (2012b) in Sicily and Kilshaw and MacDonald (2011) in Scotland found that none of the captured wildcats were interested in the valerian lures. Monterroso et al. (2013) obtained scarce results in collecting wildcat hairs in Spain, while surveys by Steyer et al. (2013) in Germany and Kéry et al. (2011) in Switzerland obtained better results. However, these last two studies did not include camera trapping. We found that four (44.4 %) of all of the detected wildcats reacted with the expected behaviour and left hair samples on the traps. These variable behaviours may have genetic bases. If individual reactions to valerian attractors are genetically inherited (Bradshaw 1992), wildcat populations, particularly isolated populations such as the Sicilian and Scottish ones, may exhibit different behaviours. Furthermore, only two (11 %) hair samples provided reliable individual genotypes. This may be due to a fast DNA degradation caused by the exposition of the pickets to environmental factors. Other studies reported similar genotyping rates (10 %, Ruell and Crooks 2007; Monterroso et al. 2014), although genotyping success can range between 0 and 100 % (Steyer et al. 2013). Despite these limitations, considering the difficulty in collecting good quality non-invasive wildcat samples, we suggest to use this method to compensate drawbacks of other techniques. Indeed, in our study, hair samples contributed to the characterization of the genetic profile of individuals. A strong point of this technique lies in its possible combination with the camera trapping. This makes it possible to associate the genetic data with the picture of an individual. However, in our study, only one individual (6-G) was both genotyped and camera trapped. Several hair trapping surveys reported that, in the breeding season (December—February), wildcat detection probability is at the highest (Weber et al. 2008; Kéry et al. 2011; Steyer et al. 2013). However, in our case, we found a different result. The positive relationship between the number of sessions elapsed since the first inspection and the number of samples collected per session (regarding both genetic and camera trapping surveys) indicated that the number of samples collected increased as the season approached the spring. This trend may have been influenced by seasonal variations in environmental conditions (food availability, snow cover, etc.) and/or a ‘trap-happy’ effect of the baited traps. Nevertheless, regarding genetic sampling, it cannot be excluded that part of the increasing performance in sample collection (especially scats) could be related to a more intense marking behaviour of wildcats or even a training effect of field researchers in finding wildcat scats.
Scat surveys are widely used in the non-invasive monitoring of elusive carnivores. They have proved to be a reliable source of samples for genetic analyses (Caniglia et al. 2012; Galaverni et al. 2012; Ruiz-González et al. 2013; Anile et al. 2014; DeMatteo et al. 2014). In our study, they provided the majority of biological samples that yielded reliable individual genotypes. The genotyping success rate of the faecal samples (1/42.5 km), as well as the proportion of samples that yielded reliable individual genotype (25 %), was lower compared with the most recent Italian study carried out in Sicily and based on a similar experimental design (1/27.9 km, 36 % Anile et al. 2014). The general low rate of success of genetic analyses using both mtDNA and microsatellites may be due to the considerable length of the amplicons of mtDNA and/or to environmental factors (temperature, UV, etc.) that may have acted on the samples in the time interval between sampling sessions (Broquet et al. 2007). Nevertheless, the average error rates (ADO = 12.1 %, FA = 1.6 %) and PIDsib for the used loci are consistent with the recommended values (Waits and Paetkau 2005; Broquet et al. 2007).
We identified at least three individuals (3, 4, 6 Table 2) showing apparent cytonuclear discordance. This is because they were assigned to the wildcat cluster using microsatellites (Qi > 0.9) but carried a ‘domestic’ mitochondrial haplotype. These findings could suggest the occurrence of past hybridization events and mtDNA introgression (Driscoll et al. 2011). However, the presence of mitochondrial haplotypes typical of the domestic population in some putative wildcats does not necessarily reveal this phenomenon. Indeed, the presence of domestic haplotypes in wildcat individuals has been found in several populations in Europe (Randi et al. 2001; Driscoll et al. 2007; Hertwig et al. 2009). Domestic cats share with F. s. libyca (from which it derived) the same diagnostic ND5 polymorphisms that distinguish African wildcat from the European wildcat (Driscoll et al. 2007). Then, shared haplotypes may represent ancient haplogroups, already present in the common ancestors of F. s. silvestris and F. s. libyca (Hertwig et al. 2009). Alternatively, a certain degree of gene flow between F. s. silvestris and F. s. libyca might have occurred as a result of undetected population dispersal before domestication. For these reasons, further investigation is required to better interpret these findings and allow the correct use of mitochondrial markers in conservation genetic studies.
Camera trapping results (3.1/100 trap days) highlight a higher capture rate compared with previous studies carried out in Turkey and Scotland (1.8/100 trap nights, Can et al. 2011; 2.3/100 trap nights, Kilshaw and Macdonald 2011), while comparable results were obtained by Anile et al. (2012a) in Sicily (2.9/100 trap nights). The quality of images provided by cameras using a 940-nm flashlight allowed an individual recognition in 57.1 % of the records. This result is comparable with those obtained by Can et al. (2009) (59.1 %), who used cameras with similar characteristics to ours. At the same time, Anile et al. (2012a) used a white flashlight and obtained 95.2 % pictures that were useful. Camera trapping is one of the most functional methods that allow the effective monitoring of several species (Silveira et al. 2003). It can provide estimates of population parameters (abundance, density, etc.), as well as valuable information about behaviour, circadian rhythms and species interactions (O’Connell et al. 2011). However, camera trapping can overestimate population size, particularly when capture–recapture methods are used to assess the abundance of low-density populations of elusive animals with few identification marks (Foster and Harmsen 2012). This should be taken into account, considering the higher number of wildcats we detected using camera traps compared with the genetic survey. However, compared with the genetic survey, camera trapping is also highly efficient in contacting the most elusive individuals. Indeed, the camera trapping survey counted a relatively high number of domestic cats, while the genetic survey did not show the presence of F. s. catus in the study area. It has been proved that the probability of finding faecal depositions is lower for the domestic cat than for the wildcat (Corbett 1979; Lozano and Urra 2007; Lozano et al. 2013). This fact could have affected the results of our scat survey. These aspects are of outermost importance when monitoring wildcat populations, as their management should not disregard the relationship between wild and domestic cats (Randi et al. 2001; Pierpaoli et al. 2003; Oliveira et al. 2008a, b; Hertwig et al. 2009).
Conclusions
In this study, we integrated three non-invasive monitoring methods to assess the presence and the abundance of the European wildcat in a small study area in the Italian Apennine. Comparison of the three data sets suggests how to offset the weaknesses of each method. However, some precautions are necessary to improve such an integrated approach. Our results highlight the low efficiency of valerian-treated lures in genetic surveys, so we recommend avoiding the exclusive use this technique that may lead to the underestimation of population size. When collecting either hair or scat samples for genetic analyses, we suggest to reduce, as much as possible, the sampling interval to prevent DNA degradation. Balestrieri et al. (2010) found that, in badger, an interval of 3–4 days yielded good genotyping rates. We also suggest performing the genetic analyses as soon as possible (Waits and Paetkau 2005). We recommend the use of two camera traps per station (Kilshaw and MacDonald 2011), one set in a high-resolution photo mode with a white flashlight for a better individual identification and the other in video- or multi-shot mode to investigate individual reactions to baits (O’Connell et al. 2011).
In agreement with a recent study carried out in other areas of the Foreste Casentinesi National Park (Ragni et al. 2014), our data suggest the presence of a well-established and stable European wildcat population. On the other hand, the widespread presence of domestic cats requires further studies to implement conservation measures.
The monitoring of the wildcat is very difficult as it is a solitary and secretive species. Knowledge about population parameters, such as the genetic status and ecology, often involves many years of sampling in the same area. However, it is important that wildlife management administrations obtain such information in order to effectively plan conservation measures, especially in areas where the presence of the species has only recently been confirmed. Our results suggest that by integrating some non-invasive techniques, monitoring performances can be improved, allowing the collection of sound data in a relatively short period of time.
References
Agostini N, Bottacci A, D’Amico C, Fabbri M, Mencucci M, Ragni B, Randi E, Tedaldi G (2010) il gatto selvatico nel Parco Nazionale delle Foreste Casentinesi. In: Randi E et al. (eds) atti del convegno Santa Sofia (FC) 7–8 Novembre 2008. Ente Parco Nazionale Foreste Casentinesi, pp 87–88
Angelici FM, Genovesi P (2003) Felis silvestris (Schreber, 1777). In: Boitani L, Lovari S, Vigna Taglianti A (eds.) Fauna d’Italia. Mammalia III. Carnivora - Artiodactyla. Calderini, Bologna, pp. 207–221
Anile S, Amico C, Ragni B (2012a) Population density estimation of the European wildcat (Felis silvestris silvestris) in Sicily using camera trapping. Wildl Biol Pract 8:1–12
Anile S, Arrabito C, Mazzamuto MV, Scornavacca D, Ragni B (2012b) A non-invasive monitoring on European wildcat (Felis silvestris silvestris Schreber, 1777) in Sicily using hair trapping and camera trapping: does scented lure work? Hystrix Ital J Mammal 23:45–50. doi:10.4404/hystrix-23.2-4657
Anile S, Ragni B, Randi E, Mattucci F, Rovero F (2014) Wildcat population density on the Etna volcano, Italy: a comparison of density estimation methods. J Zool 293:252–261. doi:10.1111/jzo.12141
Balestrieri A, Remonti L, Frantz AC, Capelli E, Zenato M, Dettori EE, Guidali F, Prigioni C (2010) Efficacy of passive hair-traps for the genetic sampling of a low-density badger population. Hystr Ital J Mammal 21:137–146
Bandelt HJ, Forster P, Röhl A (1999) Median-joining networks for inferring intraspecific phylogenies. Mol Biol Evol 16:37–48
Bizzarri L, Lacrimini M, Ragni B (2010a) Live capture and handling of the European wildcat in central Italy. Hystr Ital J Mammal 21:73–82
Bizzarri L, Capelletti P, Lacrimini M, Mariani M, Ragni B (2010b) A radio-telemetry study of the European wildcat in an area of the Umbrian Apennines. In: Randi E et al. (eds) atti del convegno Santa Sofia (FC) 7–8 Novembre 2008. Ente Parco Nazionale Foreste Casentinesi, pp 67–70
Bologna MA, Cristiani G (2012) Contributo alla teriofauna dell’Alta Val Tanaro, Alpi Liguri (CN-IM). Riv Piemont Stor Nat 33:295–319
Bradshaw JWS (1992) The behaviour of the domestic cat. C.A.B...I. International UK. BTO
Broquet T, Menard N, Petit E (2007) Noninvasive population genetics: a review of sample source, diet, fragment length and microsatellite motif effects on amplification success and genotyping error rates. Conserv Genet 8:249–260. doi:10.1007/s10592-006-9146-5
Cagnolaro L, Rosso D, Spagnesi M (1976) Inchiesta sulla distribuzione del Gatto selvatico (Felis silvestris Schreber) in Italia e nei Canton Ticino e Grigioni (Svizzera) e del Gatto selvatico sardo (Felis lyblca sarda Lataste) in Sardegna con notizie sulla Lince (Lynx lynx L.) 1971–1973. Ric Biol Selvag 64:1–109
Can OE, Kandemir I, Togan I (2011) The wildcat Felis silvestris in northern Turkey: assessment of status using camera trapping. Oryx 45:112–118. doi:10.1017/s0030605310001328
Caniglia R, Fabbri E, Cubaynes S, Gimenez O, Lebreton J-D, Randi E (2012) An improved procedure to estimate wolf abundance using non-invasive genetic sampling and capture-recapture mixture models. Conserv Genet 13:53–64. doi:10.1007/s10592-011-0266-1
Comer CE, Symmank ME, Kroll JC (2011) Bobcats exhibit low detection rates at hair collection stations in East Texas. Wildl Biol Pract 7:116–122. doi:10.2461/wbp.2011.7
Corbett LK (1979) Feeding ecology and social organization of wildcats (Felis silvestris) and domestic cats (Felis catus) in Scotland. Ph.D. dissertation. University of Aberdeen, Aberdeen
Council of Europe (1993) Seminar on the biology and conservation of the wildcat (Felis silvestris). Council of Europe, Strasbourg
DeMatteo KE et al (2014) Using detection dogs and genetic analyses of scat to expand knowledge and assist felid conservation in Misiones, Argentina. Integr Zool 9:623–639. doi:10.1111/1749-4877.12113
Driscoll C, Nowell K (2010) Felis silvestris. The IUCN Red List of Threatened Species. Version 2014.2. http://www.iucnredlist.org/details/8543/0. Accessed on 07 August 2014
Driscoll CA, Menotti-Raymond M, Roca AL, Hupe K, Johnson WE, Geffen E, Harley EH, Delibes M, Pontier D, Kitchener AC, Yamaguchi N, O’Brien SJ, Macdonald DW (2007) The Near Eastern origin of cat domestication. Science 317:519–523. doi:10.1126/science.1139518
Driscoll C, Yamaguchi N, O’Brien SJ, Macdonald DW (2011) A suite of genetic markers useful in assessing wildcat (Felis silvestris ssp.)—domestic cat (Felis silvestris catus) admixture. J Hered 102:S87–S90. doi:10.1093/jhered/esr047
Easterbee N, Hepburn LV, Jefferies DJ (1991) Survey of the status and distribution of the wildcat in Scotland, 1983–1987. Nature Conservancy Council for Scotland, Edinburgh
Eckert I, Suchentrunk F, Markov G, Hartl GB (2010) Genetic diversity and integrity of German wildcat (Felis silvestris) populations as revealed by microsatellites, allozymes, and mitochondrial DNA sequences. Mamm Biol 75:160–174. doi:10.1016/j.mambio.2009.07.005
Ellis SLH, Wells DL (2010) The influence of olfactory stimulation on the behaviour of cats housed in a rescue shelter. Appl Anim Behav Sci 123:56–62. doi:10.1016/j.applanim.2009.12.011
Foster RJ, Harmsen BJ (2012) A critique of density estimation from camera-trap data. J Wildl Manage 76:224–236. doi:10.1002/jwmg.275
Freeman AR, MacHugh DE, McKeown S, Walzer C, McConnell DJ, Bradley DG (2001) Sequence variation in the mitochondrial DNA control region of wild African cheetahs (Acinonyx jubatus). Heredity 86:355–362. doi:10.1046/j.1365-2540.2001.00840.x
French DD, Corbett LK, Easterbee N (1988) Morphological discriminants of Scottish wildcats Felis silvestris, domestic cats F. catus and their hybrids. J Zool Soc Lond 214:235–259
Galaverni M, Palumbo D, Fabbri E, Caniglia R, Greco C, Randi E (2012) Monitoring wolves (Canis lupus) by non-invasive genetics and camera trapping: a small-scale pilot study. Eur J Wildl Res 58:47–48
Garcia-Alaniz N, Naranjo EJ, Mallory FF (2010) Hair-snares: a non-invasive method for monitoring felid populations in the Selva Lacandona, Mexico. Trop Conserv Sci 3:403–411
Hartmann SA, Steyer K, Kraus RHS, Segelbacher G, Nowak C (2013) Potential barriers to gene flow in the endangered European wildcat (Felis silvestris). Conserv Genet 14:413–426. doi:10.1007/s10592-013-0468-9
Hertwig ST, Schweizer M, Stepanow S, Jungnickel A, Boehle UR, Fischer MS (2009) Regionally high rates of hybridization and introgression in German wildcat populations (Felis silvestris, Carnivora, Felidae). J Zool Syst Evol Res 47:283–297. doi:10.1111/j.1439-0469.2009.00536.x
Hupe K, Simon O (2007) Die Lockstockmethode-eine nicht invasive Methode zum Nachweis der Europäischen Wildkatze (Fels silvestris silvestris). Informat Natursch Niedersach 27:66–69
Karanth KU, Nichols JD (2002) Monitoring tigers and their prey: a manual for researchers, managers and conservationists in tropical Asia. Centre for Wildlife Studies, Bangalore, India
Kéry M, Gardner B, Stoeckle T, Weber D, Royle JA (2011) Use of spatial capture-recapture modeling and DNA data to estimate densities of elusive animals. Conserv Biol 25:356–364. doi:10.1111/j.1523-1739.2010.01616.x
Kilshaw K, Macdonald DW (2011) The use of camera trapping as a method to survey for the Scottish wildcat. Scott Nat Herit Commissioned Rep 479:1–32
Klar N, Fernandez N, Kramer-Schadt S, Herrmann M, Trinzen M, Buettner I, Niemitz C (2008) Habitat selection models for European wildcat conservation. Biol Conserv 141:308–319. doi:10.1016/j.biocon.2007.10.004
Klar N, Herrmann M, Kramer-Schadt S (2009) Effects and mitigation of road impacts on individual movement behavior of wildcats. J Wildl Managem 73:631–638. doi:10.2193/2007-574
Klar N, Herrmann M, Henning-Hahn M, Pott-Doerfer B, Hofer H, Kramer-Schadt S (2012) Between ecological theory and planning practice: (re-) connecting forest patches for the wildcat in Lower Saxony, Germany. Landsc Urban Plan 105:376–384. doi:10.1016/j.landurbplan.2012.01.007
Krone O, Guminsky O, Meinig H, Herrmann M, Trinzen M, Wibbelt G (2008) Endoparasite spectrum of wild cats (Felis silvestris Schreber, 1777) and domestic cats (Felis catus L.) from the Eifel, Pfalz region and Saarland, Germany. Eur J Wildl Res 54:95–100. doi:10.1007/s10344-007-0116-0
Lecis R, Pierpaoli M, Birò Z, Szemethy L, Ragni B, Vercillo F, Randi E (2006) Bayesian analyses of admixture in wild and domestic cats (Felis silvestris) using linked microsatellite loci. Mol Ecol 15:119–131
Long RA, Donovan TM, Mackay P, Zielinski WJ, Buzas JS (2007) Comparing scat detection dogs, cameras, and hair snares for surveying carnivores. J Wildl Managem 71:2018–2025. doi:10.2193/2006-292
Lopez JV, Cevario S, O’Brien SJ (1996) Complete nucleotide sequences of the domestic cat (Felis catus) mitochondrial genome and a transposed mtDNA tandem repeat (Numt) in the nuclear genome. Genomics 33:229–246
Lozano J (2010) Habitat use by European wildcats (Felis silvestris) in central Spain: what is the relative importance of forest variables? Anim Biodiv Conserv 33:143–150
Lozano J, Malo AF (2012) Conservation of European wildcat (Felis silvestris) in Mediterranean environments: a reassessment of current threats. In: Williams GS (ed) Mediterranean ecosystems: dynamics, management and conservation. Nova Science Publishers, Hauppauge, NY, pp 1–31
Lozano J, Urra F (2007) El gato doméstico, Felis silvestris catus. Galemys 19:35–38
Lozano J, Virgos E, Malo AF, Huertas DL, Casanovas JG (2003) Importance of scrub-pastureland mosaics for wild-living cats occurrence in a Mediterranean area: implications for the conservation of the wildcat (Felis silvestris). Biodiv Conserv 12:921–935. doi:10.1023/a:1022821708594
Lozano J, Virgos E, Cabezas-Diaz S, Mangas JG (2007) Increase of large game species in Mediterranean areas: is the European wildcat (Felis silvestris) facing a new threat? Biol Conserv 138:321–329
Lozano J, Virgos E, Cabezas-Diaz S (2013) Monitoring European wildcat Felis silvestris populations using scat surveys in central Spain: are population trends related to wild rabbit dynamics or to landscape features? Zool Stud 52 doi:10.1186/1810-522x-52-16
Luo S-J et al (2007) Development of Y chromosome intraspecific polymorphic markers in the Felidae. J Hered 98:400–413. doi:10.1093/jhered/esm063
Lüps P, Flückiger PF, Peier D, Schmidt P (2002) Fund einer Waldkatze Felis silvestris bei Oberbuchsiten. Mitt Nat forsch Ges Kanton Solothurn 39:41–45
Mattucci F, Oliveira R, Bizzarri L, Vercillo F, Anile S, Ragni B, Lapini L, Sforzi A, Alves PC, Lyons LA, Randi E (2013) Genetic structure of wildcat (Felis silvestris) populations in Italy. Ecol Evol 3:2443–2458. doi:10.1002/ece3.569
McNevin D, Wilson-Wilde L, Robertson J, Kyd J, Lennard C (2005) Short tandem repeat (STR) genotyping of keratinised hair—part 2. An optimised genomic DNA extraction procedure reveals donor dependence of STR profiles. Forensic Sci Int 153:247–259. doi:10.1016/j.forsciint.2005.05.005
Menotti-Raymond MA, Obrien SJ (1995) Evolutionary conservation of 10 microsatellite loci in 4 species of Felidae. J Hered 86:319–322
Menotti-Raymond M, David VA, Stephens JC, Lyons LA, Obrien SJ (1997) Genetic individualization of domestic cats using feline STR loci for forensic applications. J Forensic Sci 42:1039–1051
Miller CR, Joyce P, Waits LP (2002) Assessing allelic dropout and genotype reliability using maximum likelihood. Genetics 160:357–366
Mills LS, Citta JJ, Lair KP, Schwartz MK, Tallmon DA (2000) Estimating animal abundance using noninvasive DNA sampling: promise and pitfalls. Ecol Appl 10:283–294. doi:10.2307/2641002
Monterroso P, Brito JC, Ferreras P, Alves PC (2009) Spatial ecology of the European wildcat in a Mediterranean ecosystem: dealing with small radio-tracking datasets in species conservation. J Zool 279:27–35. doi:10.1111/j.1469-7998.2009.00585.x
Monterroso P, Alves PC, Ferreras P (2011) Evaluation of attractants for non-invasive studies of Iberian carnivore communities. Wildl Res 38:446–454. doi:10.1071/wr11060
Monterroso P, Castro D, Silva TL, Ferreras P, Godinho R, Alves PC (2013) Factors affecting the (in)accuracy of mammalian mesocarnivore scat identification in South-western Europe. J Zool 289:243–250. doi:10.1111/jzo.12000
Monterroso P, Rich LN, Serronha A, Ferreras P, Alves PC (2014) Efficiency of hair snares and camera traps to survey mesocarnivore populations. Eur J Wildl Res 60:279–289
Nowell K, Jackson P (1996) Wild cats. Status survey and conservation action plan. IUCN/SSC Cat Specialist Group, Gland, Switzerland and Cambridge, UK
Nussberger B, Greminger MP, Grossen C, Keller LF, Wandeler P (2013) Development of SNP markers identifying European wildcats, domestic cats, and their admixed progeny. Mol Ecol Resour 13:447–460. doi:10.1111/1755-0998.12075
O’Brien J, Devillard S, Say L, Vanthomme H, Leger F, Ruette S, Pontier D (2009) Preserving genetic integrity in a hybridising world: are European wildcats (Felis silvestris silvestris) in eastern France distinct from sympatric feral domestic cats? Biodiv Conserv 18:2351–2360. doi:10.1007/s10531-009-9592-8
O’Connell AF, Nichols JD, Karanth KU (2011) Camera traps in animal ecology. Methods Anal Springer. doi:10.1007/978-4-431-99495-4_14
Oliveira R, Godinho R, Randi E, Ferrand N, Alves PC (2008a) Molecular analysis of hybridisation between wild and domestic cats (Felis silvestris) in Portugal: implications for conservation. Conserv Genet 9:1–11. doi:10.1007/s10592-007-9297-z
Oliveira R, Godinho R, Randi E, Alves PC (2008b) Hybridization versus conservation: are domestic cats threatening the genetic integrity of wildcats (Felis silvestris silvestris) in Iberian Peninsula? Phil Trans R Soc B 363:2953–2961. doi:10.1098/rstb.2008.0052
Otis DL, Burnham KP, Anderson DR, White GC (1978) Statistical inference from capture data on closed animal populations. Wildl Monogr:1–135
Peakall R, Smouse PE (2006) GENALEX 6: genetic analysis in Excel. Population genetic software for teaching and research. Mol Ecol Notes 6:288–295. doi:10.1111/j.1471-8286.2005.01155.x
Peakall R, Smouse PE (2012) GenAlEx 6.5: genetic analysis in Excel. Population genetic software for teaching and research—an update. Bioinformatics 28:2537–2539. doi:10.1093/bioinformatics/bts460
Pierpaoli M, Birò ZS, Herrmann M, Hupe K, Fernandes M, Ragni B, Szemethy L, Randi E (2003) Genetic distinction of wildcat (Felis silvestris) populations in Europe, and hybridization with domestic cats in Hungary. Mol Ecol 12:2585–2598. doi:10.1046/j.1365-294X.2003.01939.x
Polzin T, Daneschmand SV (2003) On Steiner trees and minimum spanning trees in hypergraphs. Oper Res Lett 31:12–20
Pritchard JK, Stephens M, Donnelly PJ (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Ragni B (1981) Gatto selvatico. Felis silvestris Schreber, 1777. In: Distribuzione e biologia di 22 specie di Mammiferi in Italia. C.N.R., Roma: 105–113
Ragni B (2003) L’areale italiano del gatto selvatico Europeo (Felis silvestris silvestris): ancora un dilemma? Hystrix, supp. IV Congr. It. Teriologia PP:67
Ragni B, Petruzzi E (2010) Recent situation of the Old World wildcat (Felis silvestris Schreber, 1777) and Eurasian lynx (Lynx lynx Linnaeus, 1758) in the Apennines. In: Randi E et al. (eds) atti del convegno Santa Sofia (FC) 7–8 Novembre 2008. Ente Parco Nazionale Foreste Casentinesi pp 36–46
Ragni B, Possenti M (1996) Variability of coat-colour and markings system in Felis silvestris. Ital J Zool 63:285–292
Ragni B, Possenti M, Sforzi A, Zavalloni D, Ciani F (1994) The wildcat in central-northern Italian peninsula: a biogeographical dilemma. Biogeographia 17:553–566
Ragni B, Lucchesi M, Tedaldi G, Vercillo F, Fazzi P, Bottacci A, Quilghini G (2014). Il gatto selvatico Europeo nelle Riserve Naturali Casentinesi. Corpo Forestale dello Stato, UTB Pratovecchio
Randi E (2008) Detecting hybridization between wild species and their domesticated relatives. Mol Ecol 17:285–293. doi:10.1111/j.1365-294X.2007.03417.x
Randi E, Pierpaoli M, Beaumont M, Ragni B, Sforzi A (2001) Genetic identification of wild and domestic cats (Felis silvestris) and their hybrids using Bayesian clustering methods. Mol Biol Evol 18:1679–1693
Rondinini C, Battistoni A, Peronace V, Teofili C (2013) Lista Rossa IUCN dei Vertebrati Italiani. Comitato Italiano IUCN e Ministero dell’Ambiente e della Tutela del Territorio e del Mare, Roma. p 51
Ruell EW, Crooks KR (2007) Evaluation of noninvasive genetic sampling methods for felid and canid populations. J Wildl Managem 71:1690–1694
Ruiz-González A, Madeira MJ, Randi E, Urra F, Gómez-Moliner BJ (2013) Non-invasive genetic sampling of sympatric marten species (Martes martes and Martes foina): assessing species and individual identification success rates on faecal DNA genotyping. Eur J Wildl Res 59:371–386
Santolini R, Giuliani A, Tedaldi G, Morelli F, Ricci L, Moretti E, Savini C; (2010) Il gatto selvatico nell’appennino a nord dell’areale storico: analisi dell’offerta ambientale (dati preliminari) e indirizzi di conservazione. In: Randi E et al. (eds) atti del convegno Santa Sofia (FC) 7–8 Novembre 2008. Ente Parco Nazionale Foreste Casentinesi pp 83–86
Schulenberg J (2005) Säugetiere (Mammalia). In: A. Gunther, U. Nigmann, R. Achtziger (ed.) Analyse der Gefährdungsursachen von planungsrelevanten Tiergruppen in Deutschland zur Ergänzung der bestehenden Roten Listen gefährdeter Tiere. Schriftenr. Biol Vielfalt, BfN, Bonn-Bad Godesberg pp. 70–112
Silveira L, Jacomo ATA, Diniz JAF (2003) Camera trap, line transect census and track surveys: a comparative evaluation. Biol Conserv 114:351–355
Stahl P, Artois M (1994) Status and conservation of the wildcat (Felis silvestris) in Europe and around the Mediterranean rim. Convention on the Conservation of European Wildlife and Natural Habitats Standing Committee. Council Eur Nat Environ Ser 69:1–76
Steyer K, Simon O, Kraus RHS, Haase P, Nowak C (2013) Hair trapping with valerian-treated lure sticks as a tool for genetic wildcat monitoring in low-density habitats. Eur J Wildl Res 59:39–46. doi:10.1007/s10344-012-0644-0
Taberlet P, Waits LP, Luikart G (1999) Noninvasive genetic sampling: look before you leap. Trends Ecol Evol 14:323–327
Tedaldi G (2012) Rilevazione della presenza del gatto selvatico (Felis silvestris silvestris) in due siti di rete natura 2000 nella provincia di Bologna. Provincia di Bologna
Tiedemann R, Harder J, Gmeiner C, Haase E (1996) Mitochondrial DNA sequence patterns of harbour porpoises (Phocoena phocoena) from the North and the Baltic sea. Z Säugetierk 61:104–111
Vogel B, Mölich T (2013) Best practices for implementing biotope networks in highly fragmented landscapes: the safety net for the European wildcat. In: Proceedings of the 2013 International Conference on Ecology and Transportation (ICOET 2013)
Waits LP, Paetkau D (2005) Noninvasive genetic sampling tools for wildlife biologists: a review of applications and recommendations for accurate data collection. J Wildl Managem 69:1419–1433. doi:10.2193/0022-541x(2005)69[1419:ngstfw]2.0.co;2
Waits LP, Luikart G, Taberlet P (2001) Estimating the probability of identity among genotypes in natural populations: cautions and guidelines. Mol Ecol 10:249–256. doi:10.1046/j.1365-294X.2001.01185.x
Weber D, Stoeckle TM, Roth T (2008) Entwicklung und Anwendung einer neuen Wildkatzen-Nachweismethode (Development and application of a new method of detecting wildcats. In German). Internal report. Hintermann & Weber AG Rodersdorf
Wells DL, Egli JM (2004) The influence of olfactory enrichment on the behaviour of captive black-footed cats, Felis nigripes. Appl Anim Behav Sci 85:107–119. doi:10.1016/j.applanim.2003.08.013
Acknowledgments
We thank Nicole Marini for the irreplaceable help in the field and laboratory work. We are grateful to Nevio Agostini, Davide Alberti and the Research, Conservation and Development Department of the Foreste Casentinesi National Park (Parco Nazionale delle Foreste Casentinesi Monte Falterona e Campigna) and the Rangers of Premilcuore (commando stazione di Premilcuore, Corpo Forestale dello Stato) for the indispensable logistic support in the fieldwork. Thanks to Stefano Anile, Cristiano Tarantini, Marco Lucchesi, Giancarlo Tedaldi, Gabriele Cristiani, Katharina Steyer, Darius Weber and all the wildlife experts that helped us to develop the methodology applied in this work. Gratitude also goes to Federica Mattucci, Francesca Davoli, Romolo Caniglia, Elena Fabbri, Chiara Mengoni, Nadia Mucci, Aritz Ruiz-Gonzàles, Silvana Lapalombella, Marco Galaverni, Valentina Rovelli, Davide Nespoli, Alessandra Palladini and all the colleagues of the Conservation Genetics Group of the Institute for Environmental Protection and Research (ISPRA) for the assistance and the support in the laboratory. We also thank Martin Bennet for the linguistic revision. Part of the field equipment was provided by the Università di Roma Tre. The molecular analyses were funded by the Institute for Environmental Protection and Research (ISPRA, Istituto Superiore per la Protezione e la Ricerca Ambientale).
Conflict of interest
The authors declare no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by P. C. Alves
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(DOCX 18 kb)
Rights and permissions
About this article
Cite this article
Velli, E., Bologna, M.A., Silvia, C. et al. Non-invasive monitoring of the European wildcat (Felis silvestris silvestris Schreber, 1777): comparative analysis of three different monitoring techniques and evaluation of their integration. Eur J Wildl Res 61, 657–668 (2015). https://doi.org/10.1007/s10344-015-0936-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10344-015-0936-2