Abstract
Wildcats are among the most elusive and least investigated carnivores in Central Europe. Here, we propose a hair-trapping method that allows reliable detection of wildcat presence even in low-density habitats. The trap is simple, consisting of a wooden stick with valerian as cat attractant. We performed non-invasive genetic wildcat monitoring in the Kellerwald-Edersee National Park, Germany, between 2007 and 2011. Our results provide the first evidence of wildcat presence in this region. Microsatellite analysis and mtDNA sequencing of hair samples furthermore confirm the existence of at least six individuals (males and females) in the study region. Four individuals were detected over two consecutive years, suggesting the resident status of wildcats in this area. Our results show that the lure stick method releases its full potential when combined with genetic analysis and is a sensitive tool which not only enables the detection of wildcat presence but also provides individual identification, even in recently colonised low-density areas.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In the first half of the last century, the European wildcat (Felis silvestris silvestris Schreber 1777) was close to extinction over large parts of its range. In Germany, wildcat populations persisted only in few refugia, such as the low mountain ranges of the Harz, the Eifel and the Taunus mountains (Raimer 1988). Additionally, the German wildcat population today faces threats like habitat fragmentation and hybridisation with domestic cats. Especially in North Rhine-Westphalia, high rates of hybridisation were observed (Hertwig et al. 2009).
The wildcat is protected in Germany since 1934 and has to be monitored periodically based on the European Council Directive 92/43/EEC (Appendix IV). Wildcat monitoring is largely based on sightings, camera and live trapping, radio tracking, scat and track surveys and, opportunistically, through the occurrence of roadkills. However, due to their similarity, wildcats and domestic cats (Felis silvestris catus) can hardly be distinguished by pelage characteristics (Krüger et al. 2009), even under favourable weather and light conditions. Distinguishing field signs from scat surveys and tracks is difficult, too, due to the similarity of these signs in wildcats and domestic cats (Lozano et al. 2003; Okarma et al. 2002). Live trapping of wildcats with cage traps combined with radio telemetry allows detailed examination of individuals in terms of spatial distribution, social organisation and physical condition and provides high-quality blood, urine and tissue samples. Nevertheless, live trapping is intrusive, logistically difficult and a laborious task for large study areas. Therefore, the development of a new economic and easy-to-use monitoring system for the wildcat was required. Hupe and Simon (2007) tested a non-invasive method to obtain hair samples for morphological identification of free-living wildcats in the Solling low mountain range. The method uses rough-sawn wooden sticks treated with valerian (Valeriana officinalis) root extract. Valerian is known to be an effective olfactory attractant for feline species, especially for cats (Jerosch et al. 2010; Monterroso et al. 2011).
Purely morphological identification of hairs obtained with the lure stick method is time-consuming, tedious and limited only to the detection of wildcat presence; no further information such as individualisation or family relationships can be inferred. Therefore, we combined for the first time the hair-trapping method of Hupe and Simon (2007) with subsequent genetic analysis of hair samples, allowing for an effective and reliable differentiation between wild and domestic cats. Preliminary trials of the combination of hair trapping and genetic finger printing resulted in the detection of wildcat presence in several areas where the species was considered extinct (Nowak and Steyer 2009). In order to test the suitability of valerian-based hair trapping for wildcat monitoring in recently colonised low-density habitats, we chose the Kellerwald-Edersee National Park in central Germany as a study region. In the area of the national park, which recently became an UNESCO world natural heritage, the wildcat population underwent a severe population decline at the beginning of the twentieth century through hunting, and the last occasional shootings of wildcats are documented until the 1950s (collection material, Research Institute and Natural History Museum Frankfurt). Between 1950 and 2004, there was no evidence for wildcat occurrence in the Kellerwald region, suggesting the obvious lack of wildcats. In the year 2000, a cage-trap monitoring of wildcats was not successful in detecting any wildcat individuals, either (Semrau 2000). In 2004, a dead wildcat was found 20 km near the national park border, accompanied by sightings of cats with wildcat pelage characters. This initially indicated a return of wildcats to one of their former habitats, the Kellerwald region.
The aim of this study was to evaluate the use of hair trapping with valerian lure (Fig. 1) for the detection of wildcat presence. We collected hair samples regularly and studied them by genetic means, i.e. mitochondrial and microsatellite analysis, documenting the feasibility and effectiveness of genetic wildcat monitoring even in low-density areas of the species.
Material and methods
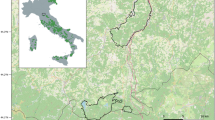
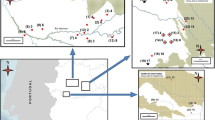
The Kellerwald-Edersee National Park has a size of 5,700 ha and is located in the north of Germany's federal state of Hesse (51°10′ N, 9°00′ E). Its altitude varies from 200 to 600 m above sea level. The mean annual temperature is 7 °C, and the annual rainfall is between 600 and 800 mm. There are no public roads or human settlements in the national park. The dominant vegetation is Luzulo-Fagetum Meusel 1937 beech grove with scattered glades and meadows.
Samples were collected in the mating period of wildcat from December to May (Piechocki 1990). Sampling periods varied across years due to snowfall, effectively being from February to April. Rough wooden sticks with a dimension of 100 × 4.8 × 2.4 cm were set up along hiking trails which are accessible by vehicles. Most parts of the world natural heritage site were not accessible for sampling by reasons of missing trails and/or to avoid disturbance. Sampling strategy was further based on known habitat preferences of the wildcat (Klar et al. 2008). Lure sticks were wetted with valerian oil (Madaus GmbH, Cologne, Germany) to attract wildcats (see Fig. 1).
Inspection of lure sticks was carried out in 7–10-day intervals. Attached hairs were removed with forceps and stored in plastic bags with silica gel to keep samples dry and to avoid degradation of DNA. In order to prevent contamination from remaining hair material, sticks were flamed after collection of hairs. After each sampling event, lure sticks were treated again with valerian oil. Prior to genetic analysis, all hair samples were inspected under a microscope in order to exclude hairs obviously not stemming from carnivores as evidenced by their morphology using an identification key (Teerink 1991). For separating similar hair from red fox (Vulpes vulpes) and wildcat, cross sections are usually required (Toth 2002), but to avoid possible negative effects in downstream processes, hairs were not cut or treated with oil. Hairs of potential carnivores were only subjected to mitochondrial DNA (mtDNA) analysis, and felid hairs, additionally, to microsatellite analysis.
DNA extraction was carried out following the QIAamp DNA Investigator kit (Qiagen, Hilden) protocols for hair. Contamination risks were minimised using a laboratory dedicated to the pre-polymerase chain reaction (PCR) handling of non-invasively collected samples (Taberlet et al. 1999). To minimise genotyping errors based on allelic dropout and false alleles, a minimum of ten hairs with roots from the same hair cluster were used for microsatellite analysis (Goossens et al. 1998). Negative controls were run alongside all reactions to monitor for possible cross contamination during extraction and amplification.
For mtDNA sequence analysis, PCR reactions were carried out in a total volume of 15 μl, including 3 μl DNA isolate, 3 mM MgCl2, 1X standard Taq (Mg-free) reaction buffer, 0.2 mM of each dNTP, 0.3 μM of each primer and 0.66 units Taq polymerase (New England BioLabs). Primers for the control region (LF4 5′-GACATAATAGTGCTTAATCGTGC-3′, Eckert et al. 2009; and H16498 5′-CCTGAAGTAAGAACCAGATG-3′, Kocher et al. 1989) were used to identify cat haplotypes, following Eckert et al. (2009). An initial denaturation step at 94 °C was followed by 41 cycles (30 s at 94 °C, 30 s at 54 °C and 30 s at 72 °C) with a final extension step for 10 min at 72 °C. PCR products were purified with ExoSAP-IT (Affymetrix). Sequencing was carried out using the BigDye Terminator 3.1 sequencing kit (Applied Biosystems) with an initial denaturation step of 60 s at 95 °C, followed by 30 cycles of 10 s at 96 °C, 10 s at 50 °C and 120 s at 60 °C. Products were purified with ABI-XTerminator beads (Applied Biosystems) and separated on an ABI 3730 DNA Analyzer (Applied Biosystems). Sequences were aligned with the ClustalW algorithm (Thompson et al. 1994) in BioEdit 7.0.9.0 (Hall 1999) and compared to wildcat and domestic cat reference samples from the collection of the Senckenberg Research Institute and Natural History Museum.
For nuclear DNA analysis, one sex marker (Zn-finger, Pilgrim et al. 2005) and 14 microsatellite markers (Menotti-Raymond et al. 1999) were amplified in four multiplex reactions: FCA8, FCA171, FCA571 and FCA124; FCA149, FCA170, FCA88 and FCA275; FCA364, FCA132 and FCA576; and FCA232, FCA347 and FCA567. To avoid genotyping errors, a multiple tube approach with three replicates was implemented (Navidi et al. 1992). After initial denaturation (15 min at 95 °C), 46 cycles (30 s at 94 °C, 90 s at 50 °C and 60 s at 72 °C) were run with a final extension step for 30 min at 72 °C. Products were sized together with a LIZ size standard on an ABI 3730 DNA Analyzer (Applied Biosystems). Fragment length was scored using GeneMarker 1.9 (SoftGenetics); consensus genotypes and the probability of identity for siblings (PIDsibs) were calculated with GIMLET 1.3.3 (Valière 2002). Individualisation was done by using the match function in GENALEX 6 (Peakall and Smouse 2006). Subspecies identification was performed in Structure (Pritchard et al. 2000) version 2.3.3. The number of genetic clusters was set to K = 2 in order to differentiate between wild and domestic cats (Oliveira et al. 2007; O'Brien et al. 2009). Analyses were based on 100,000 MCMC steps after discarding the first 100,000 steps as burn-in, under the admixture model with correlated allele frequencies (Eckert et al. 2009; Hertwig et al. 2009). A panel of 35 wildcats (from Hesse, Rhineland-Palatinate, Thuringia and Luxembourg, described as wildcats on basis of pelage characters, intestine length and mitochondrial analysis) and 35 domestic cats from Hesse (internal database) with an assignment index of q i > 0.98 for their respective cluster was used as reference data set for calculation of PIDsibs (the probability to discriminate two siblings) and for Structure analysis. Assignment threshold was set to q i > 0.8 for (sub-)species identification (Oliveira et al. 2007; Pierpaoli et al. 2003).
Results
Between 2007 and 2011, lure sticks were placed in the region for a cumulative total of 35,300 days. Wildcats were trapped 25 times (based on genetic data), representing a total capture success of 0.07/100 trap days. Capture success rate was lowest in 2008 (0.3/100 trap days) and highest in 2007 (0.15/100 trap days, Table 1). The number of positive lure stick stations was between 2 in 2008 and 11 in 2010 (Table 1). Wild boar hair was found once in 2009, twice in 2008 and repeatedly in 2011 at different lure sticks.
Out of a total of 37 samples, 24 yielded reliable mitochondrial sequences (success rate 65 % across sampling years and samples). Lowest success rates were observed in 2008 and 2009 with 40 %, and the highest mtDNA success rate, in 2007 (80 %). Altogether, four different Felis mtDNA haplotypes could be amplified, three belonging to wildcats and one observed only in domestic cats (GenBank accession numbers JX045658-JX045661). None of the three wildcat haplotypes has previously been observed in hybrids (unpublished data). In 2007, haplotype FS03 was detected twice. In 2008 and 2009, only haplotype FS06 appeared in our samples. In 2010, haplotype diversity was greatest with FS03, FS06 and FS22 being detected in our study area. Two different haplotypes (FS03 and FS22) could be observed in the final sampling year 2011. Two further hair samples from 2011 were identified as red fox (V. vulpes) by mtDNA analysis (Table 1).
Hair quantity was not sufficiently large for reliable microsatellite analyses in 2007 and was also not successful for samples collected in 2008. In later study years, however, success rates were 70 % (2010) and 100 % (2009 and 2010) (Table 1). In total, 18 of 24 hair samples with a minimum of ten hairs with roots could be individualised successfully (75 %). Rates of allelic dropout were 11 %, and false alleles occurred at 4.8 % across all PCR reactions. In no reaction, we detected more than two alleles at each locus, confirming that no hairs from multiple individuals, potentially visiting the same lure stick between two samplings, were combined. PIDsibs calculation showed that the five most variable microsatellite loci were sufficient to discriminate siblings with a probability of 0.007 (data not shown) as recommended in Waits and Paetkau (2005). For individualisation, mitochondrial haplotype information and all 14 microsatellite loci were used. In the case of assignment of hair samples to individuals A and C, a minimum of 11 loci were identical among samples. Samples that were assigned to individuals B, D and F show identical genetic consensus profiles at a minimum of 13 loci. Mismatches in the assignment of consensus genotypes to individuals were solely caused by allelic dropout.
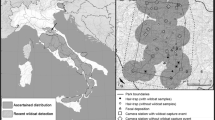
Due to the lack of microsatellite data, individual identification was not possible in the years 2007 and 2008. Between 2009 and 2011, however, six different wildcat individuals were detected (five males and one female, Fig. 2). Individual wildcats were re-sampled up to five times, with an average of 2.67 detections per individual. Four individuals could be detected in two consecutive years. The mean maximum distance between lure sticks where the same individual was detected ranged from 0 (individual C) to 4.7 km (individual F), with an overall mean maximum distance of 2.5 km. Based on the minimum convex polygon, we estimated an average of 11.7 captures/100 km² with a maximum in 2007 (17.4 captures/100 km²) and a minimum in 2009 (5.5 captures/100 km²). Due to the observed mean, maximum distance between lure sticks visited by the same individual, a buffer area of 2.5 km² was placed around lure sticks. With respect to that buffer zone, the average was 7.4 captures/100 km² (minimum 3.4 captures/100 km² in 2008 and maximum in 2010 with 15.3 captures/100 km²).
All detected wildcat individuals could clearly be assigned as belonging to the central German wildcat population, with no evidence for hybridisation with domestic cats (q i > 0.98, Fig. 3). The only evidence of domestic cat was obtained in 2009 at a lure stick which was situated 2 km away from the nearest human settlement.
Discussion
Comparison of sampling methods
The detection of wildcats is much more effective during mating season (Hupe 2007). Thus, we decided that a restriction of hair trapping from December to March should lead to the best results with regard to wildcat detection even in low-density areas where capture success rates are generally expected to be rather low. Studies of live trapping detected between 0.3 and 0.5 wildcats/100 trap nights (Bizzarri et al. 2010; Daniels et al. 2001; Sarmento et al. 2009) and 1.7 and 1.8 wildcats/100 trap nights (Monterroso et al. 2009; Potocnik et al. 2002); however, these data were not from recently colonised areas. Also, the logistics involved in live-trapping studies are substantial—in sharp contrast to camera trapping or the lure stick method. Cameras or lure sticks are operated largely unsupervised compared to live trapping and do not require the handling of live animals, which usually necessitates trained personnel and special permissions. Camera trapping as detection method for wildcats showed capture rates between 0.016 (Sarmento et al. 2009) and 4 captures/100 trap nights (Anile et al. 2010). However, such camera-trap studies provide no information regarding an individual's sex, nor the possibility to obtain genetic material for further analysis of, e.g. genetic diversity, inbreeding rate, pedigree reconstruction, hybridisation and individual discrimination.
The detection rates of wildcats with the lure stick method in the Kellerwald-Edersee National Park in central Germany were 0.03–0.15 captures/100 trap nights. These rates seem to be lower than the above-mentioned examples but were conducted in an area known for its very low wildcat densities. Considering the trapping effort in contrast to live trapping, this is a cost effective and easy-to-use non-invasive method for the detection of wildcats. To systematically compare the capture success rates of the lure stick method with live trapping and/or camera trapping, comparative studies in areas with varying wildcat densities are required. The increased information gain due to the genetic analysis of hairs collected with the lure stick method is a significant advantage in comparison with camera trapping. To our experience, one trained person can easily handle between 20 and 50 sticks during a sampling period. Assuming weekly inspections over two consecutive months, this adds to ~200–400 inspections (1,200–3,000 trap days), indicating a very high chance of wildcat detection if wildcats are present in the area. Thus, it should be possible for a single person to obtain analysable wildcat hairs in the framework of a short-term field project in a low-density area.
Constraints and possible improvements of the lure stick method
Non-invasively collected samples, like hairs from valerian-treated lure sticks, show increased rates of DNA degradation, resulting in lowered amplification success and higher allelic dropout and false allele rates. Moreover, they are particularly sensitive in terms of cross contamination (Beja-Pereira et al. 2009). Broquet et al. (2007) reviewed success rates of mtDNA and microsatellite amplifications from hairs. In that study, the mean values for mtDNA and microsatellite amplification success rate based on hairs were reported to lie between 55 and 95 % and between 45 and 100 %, respectively. The amplification success rates in our study (65 % for mtDNA and 75 % for microsatellite analysis) were in line with these values. Also, allelic dropout rate was comparable to that of other studies, with a rate of 11 % across all PCR reactions (Broquet et al. 2007). The occurrence of allelic dropout and false alleles can typically be further minimised by performing PCR reactions in more replicates (Taberlet et al. 1999).
The false allele rate of 4.8 % in our study and the often low rates of mtDNA and microsatellite amplification success in our and other studies (Broquet et al. 2007) using hair samples as source of DNA can be caused by several factors. Ruell and Crooks (2007) suggested that the fine structure of cat hair harbours much less DNA than hairs of species with coarser hair. Additionally, hairs sampled from lure sticks are usually shed hairs, which contain lower DNA concentrations as plucked hairs, too (Gagneux et al. 1997). Next to hair structure, the low amplification success rates are caused by several strong snow fall events and high intervals of inspection, which results in long-term exposure of hair samples to humidity, temperature fluctuation and UV radiation (Bonin et al. 2004). In future studies, an improvement of success rates will likely be achieved by increasing the frequency of inspections to avoid exposure to environmental conditions as much as possible.
Conclusions
Although the effectiveness of hair trapping and amplification success rates was comparatively low, a total of five male wildcats and one female wildcat were detected over the sampling period in this low-density region. Even though our study was not designed to allow for reliable population size estimation, it seems highly plausible to assume that only a portion of the individuals present in this area were detected. The higher detection rates of males could be due to the ongoing colonisation process or is a bias of the lure which may attract more males than females. For future studies with the aim to estimate population sizes, it will be important to further investigate this possible bias and account for it appropriately.
The obtained data not only confirmed the presence of wildcat in the national park but also suggested the potential establishment of a wildcat population. Male wildcats leave their native territory when reaching sexual maturity and disperse across wide areas in order to find new territories (Piechocki 1990). Thus, the detection of only male wildcats in an area does not provide support for the idea that a resident wildcat population is present. The detection of a female wildcat and three male wildcat individuals over two consecutive years, however, provides substantial support for the existence of a resident population in the sampling area. This finding is supported by the recent observation of juvenile wildcats in the national park area (Uwe Liehr, personal communication).
Currently performed large-scale application of this method shows that success rates in known high-density wildcat regions are usually much higher compared to the detection rates obtained in this study (unpublished data). Thus, we presume that population densities in this recently established population are still low.
Our study shows that the approach of combining hair trapping and subsequent genetic analysis leads to a successful detection of wildcats in a low-density area. The method proved to be an ideal tool for monitoring wildcats, and its application in future studies in other areas will provide more information on potential ways of improving it. The assessment of a potential sex bias of this method will need specific attention. Based on a systematic grid-based hair-trapping approach and capture-mark-recapture methods, even population density estimations of this elusive predator and conservation flagship species should be possible in the near future.
References
Anile S, Bizzarri L, Ragni B (2010) Estimation of European wildcat population size in Sicily (Italy) using camera trapping and capture-recapture analyses. Ital J Zool 77:241–246
Beja-Pereira A, Oliveira R, Alves PC et al (2009) Advancing ecological understandings through technological transformations in noninvasive genetics. Mol Ecol Resour 9:1279–1301
Bizzarri L, Lacrimini M, Ragni B (2010) Live capture and handling of the European wildcat in central Italy. Hystrix 21:73–82
Bonin A, Bellemain E, Eidesen PB et al (2004) How to track and assess genotyping errors in population genetics studies. Mol Ecol 13:3261–3273
Broquet T, Ménard N, Petit E (2007) Noninvasive population genetics: a review of sample source, diet, fragment length and microsatellite motif effects on amplification success and genotyping error rates. Conserv Genet 8:249–260
Daniels MJ, Beaumont MA, Johnson PJ et al (2001) Ecology and genetics of wild-living cats in the north-east of Scotland and the implications for the conservation of the wildcat. J Appl Ecol 38:146–161
Eckert I, Suchentrunk F, Markov G, Hartl GB (2009) Genetic diversity and integrity of German wildcat (Felis silvestris) populations as revealed by microsatellites, allozymes, and mitochondrial DNA sequences. Mamm Biol 75:160–174
Gagneux P, Boesch C, Woodruff DS (1997) Microsatellite scoring errors associated with noninvasive genotyping based on nuclear DNA amplified from shed hair. Mol Ecol 6:861–868
Goossens B, Waits LP, Taberlet P (1998) Plucked hair samples as a source of DNA: reliability of dinucleotide microsatellite genotyping. Mol Ecol 7:1237–1241
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucl Acids Symp Ser 41:95–98
Hertwig ST, Schweizer M, Stepanow S et al (2009) Regionally high rates of hybridization and introgression in German wildcat populations (Felis silvestris, Carnivora, Felidae). J Zool Syst Evol Res 47:283–297
Hupe K (2007) Untersuchung zum Vorkommen der Wildkatze (Felis silvestris silvestris) in Wäldern und bewaldeten Höhenzügen zwischen Solling und Hainberg im Hinblick auf eine mögliche Vernetzung der Harz- und Sollingpopulation. Inform.d. Naturschutz Niedersachs 27:38–45
Hupe K, Simon O (2007) Die Lockstockmethode - eine nicht invasive Methode zum Nachweis der Europäischen Wildkatze (Felis s. silvestris). Inform d Naturschutz Niedersachs 27:66–69
Jerosch S, Götz M, Klar N, Roth M (2010) Characteristics of diurnal resting sites of the endangered European wildcat (Felis silvestris silvestris): implications for its conservation. J Nat Conserv 18:45–54
Klar N, Fernández N, Kramer-Schadt S et al (2008) Habitat selection models for European wildcat conservation. Biol Conserv 141:308–319
Kocher TD, Thomas WK, Meyer A et al (1989) Dynamics of mitochondrial DNA evolution in animals: amplification and sequencing with conserved primers. P Natl Acad Sci USA 86:6196–6200
Krüger M, Hertwig ST, Jetschke G, Fischer MS (2009) Evaluation of anatomical characters and the question of hybridization with domestic cats in the wildcat population of Thuringia, Germany. J Zool Syst Evol Res 47:268–282
Lozano J, Virgos E, Malo AF et al (2003) Importance of scrub-pastureland mosaics for wild-living cats occurrence in a Mediterranean area: implications for the conservation of the wildcat (Felis silvestris). Biodivers Conserv 12:921–935
Menotti-Raymond M, David VA, Lyons LA et al (1999) A genetic linkage map of microsatellites in the domestic cat (Felis catus). Genomics 57:9–23
Monterroso P, Brito JC, Ferreras P, Alves PC (2009) Spatial ecology of the European wildcat in a Mediterranean ecosystem: dealing with small radio-tracking datasets in species conservation. J Zool 279:27–35
Monterroso P, Alves PC, Ferreras P (2011) Evaluation of attractants for non-invasive studies of Iberian carnivore communities. Wildl Res 38:446
Navidi W, Arnheim N, Waterman MS (1992) A multiple-tubes approach for accurate genotyping of very small DNA samples by using PCR: statistical considerations. Am J Hum Genet 50:347–359
Nowak C, Steyer K (2009) Abschlußbericht Genetisches Monitoring der Wildkatze im Rahmen des Projektes “Ein Rettungsnetz für die Wildkatze”- Teilbereich Kontrolle. Conservation Genetics Group, Senckenberg Gelnhausen
Oliveira R, Godinho R, Randi E et al (2007) Molecular analysis of hybridisation between wild and domestic cats (Felis silvestris) in Portugal: implications for conservation. Conserv Genet 9:1–11
Okarma H, Sniezko S, Olszanska A (2002) The occurrence of wildcat in the Polish Carpathian Mountains. Acta Theriol 47:499–504
O'Brien J, Devillard S, Say L et al (2009) Preserving genetic integrity in a hybridising world: are European Wildcats (Felis silvestris silvestris) in eastern France distinct from sympatric feral domestic cats? Biodivers Conserv 18:2351–2360
Peakall R, Smouse PE (2006) GENALEX 6: genetic analysis in Excel. Population genetic software for teaching and research. Mol Ecol Notes 6:288–295
Piechocki R (1990) Die Wildkatze. Die Neue Brehm-Bücherei, Wittenberg-Lutherstadt
Pierpaoli M, Biro ZS, Herrmann M et al (2003) Genetic distinction of wildcat (Felis silvestris) populations in Europe, and hybridization with domestic cats in Hungary. Mol Ecol 12:2585–2598
Pilgrim KL, McKelvey KS, Riddle AE, Schwartz MK (2005) Felid sex identification based on noninvasive genetic samples. Mol Ecol Notes 5:60–61
Pritchard JK, Stephensa M, Donnellya P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Raimer F (1988) Die Wildkatze in Hessen und Niedersachsen. University of Kassel, Diploma thesis
Semrau M (2000) Studie zur Situation der Wildkatze im Kellerwald. University of Göttingen, Diploma thesis
Potocnik H, Kljun F, Racnik J et al (2002) Experience obtained from box trapping and handling wildcats in Slovenia. Acta Theriol 47:211–219
Ruell EW, Crooks KR (2007) Evaluation of noninvasive genetic sampling methods for felid and canid populations. J Wildl Manage 71:1690–1694
Sarmento P, Cruz J, Eira C, Fonseca C (2009) Spatial colonization by feral domestic cats Felis catus of former wildcat Felis silvestris silvestris home ranges. Acta Theriol 54:31–38
Taberlet P, Waits LP, Luikart G (1999) Noninvasive genetic sampling: look before you leap. Trends Ecol Evol 14:323–327
Teerink BJ (1991) Atlas and identification key hair of West-European mammals. Cambridge University Press, Cambridge
Thompson JD, Higgins DG, Gibson TJ (1994) Clustal-W—improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680
Toth AM (2002) Identification of Hungarian Mustelidae and other small carnivores using guard hair analysis. Acta Zool Acad Sci Hung 48:237–250
Valière N (2002) GIMLET: a computer program for analysing genetic individual identification data. Mol Ecol Notes 10:1046–1046
Waits LP, Paetkau D (2005) Noninvasive genetic sampling tools for wildlife biologists: a review of applications and recommendations for accurate data collection. J Wildl Manage 69:1419–1433
Acknowledgements
We thank Achim Frede and the team from the Nationalpark Kellerwald-Edersee and Karsten Wittern (Förderverein für den Nationalpark Kellerwald-Edersee e.V.). We are also grateful to numerous wildcat experts in Germany and Switzerland who helped to develop the methodology shown in this manuscript, including Thomas Mölich, Burkhard Vogel and Thomas Norgall (Bund für Umwelt- und Naturschutz Deutschland, BUND); Jürgen Thein, Karsten Hupe, Malte Götz, Martina Denk and Tabea Stoeckle. Parts of the applied molecular methods were originally developed by ECOGENICS, CH. Wildcat tissue samples were kindly provided by Mathias Herrmann, Matthias Krüger, Franz Müller, Uwe Müller and Jacques Pir. Technical assistance in the laboratory by Mascha Siemund and João Barateiro Diogo is gratefully acknowledged. We appreciate the work of Armin Bürgel, who was an encouraged supporter of this project. This study was funded by the Kellerwald-Edersee National Park. Additional funding comes from the BUND Hessen and the Landesoffensive zur Entwicklung wissenschaftlich-ökonomischer Exzellenz of the state of Hesse. RHSK was funded by grant SAW-2011-SGN-3 from the Leibniz Association.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by P. C. Alves
Rights and permissions
About this article
Cite this article
Steyer, K., Simon, O., Kraus, R.H.S. et al. Hair trapping with valerian-treated lure sticks as a tool for genetic wildcat monitoring in low-density habitats. Eur J Wildl Res 59, 39–46 (2013). https://doi.org/10.1007/s10344-012-0644-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10344-012-0644-0