Abstract
Ocean temperature rising drastically threatens the adaptation and survival of marine organisms, causing serious ecological impacts and economic losses. It is crucial to understand the adaptive mechanisms of marine organisms in response to high temperature. In this study, a novel regulatory mechanism that is mediated by hypoxia-inducible factor-1α (HIF-1α) was revealed in Pacific oyster (Crassostrea gigas) in response to heat stress. We identified a total of six HIF-1α genes in the C. gigas genome, of which HIF-1α and HIF-1α-like5 were highly induced under heat stress. We found that the HIF-1α and HIF-1α-like5 genes played critical roles in the heat shock response (HSR) through upregulating the expression of heat shock protein (HSP). Knocking down of HIF-1α via RNA interference (RNAi) inhibited the expression of heat shock factor 1 (HSF1) and HSP70 genes in C. gigas under heat stress. Both HIF-1α and HIF-1α-like5 promoted the transcriptional activity of HSF1 by binding to hypoxia response elements (HREs) within the promoter region. Furthermore, the survival of C. gigas under heat stress was significantly decreased after knocking down of HIF-1α. This work for the first time revealed the involvement of HIF-1α/HSF1/HSP70 pathway in response to heat stress in the oyster and provided an insight into adaptive mechanism of bivalves in the face of ocean warming.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Global climate change has driven environmental changes dramatically, including global temperature rise, shrinking ice sheets, and ocean warming. Global ocean warming is a critical indicator of the climate system. Surface temperature of the ocean has hit record high in 2020, and a continued increasing has been predicted (Cheng et al. 2021). The healthy and sustainable development of coastal ecosystem under ocean temperature rising is of substantial concern (McLeod et al. 2019; Solan and Whiteley 2016). Ocean temperature rising that linked to global warming was predicated to threaten the survival of marine animals, especially those distribute in coastal zones (Chen et al. 2021; Khan et al. 2020; Petes et al. 2007; Pinsky et al. 2019).
Marine bivalves play unique and essential roles in marine ecosystem with their worldwide distribution, ecological significance, economic benefits, and food resources (Guo et al. 2008; McLeod et al. 2019; Strehse and Maser 2020; Wijsman et al. 2019). However, the sessile bivalves are vulnerable to high temperature particularly during tidal fluctuations in summer since they are not able to escape from the habitats (Chapman 2006; Freitas et al. 2021). High temperature caused damages on the organs of marine bivalves, posing cascading effects on various physiological processes including immunity, metabolism, growth, reproduction, and development (Li et al. 2009, 2007; Mosca et al. 2013; Patrick et al. 2006; Zippay and Helmuth 2012). It is also worth noting that the heat stress is frequently accompanied by hypoxia stress, which together aggravates the damage to the marine organisms (Hu et al. 2022; Huo et al. 2019; McArley et al. 2020; Vaquer-Sunyer and Duarte 2011). Consequently, it is of great significance to discover the molecular mechanisms underlying the response of marine bivalves to high temperature, which will provide insights into understanding the adaptation of marine organisms to ocean warming.
The heat shock response (HSR) is an evolutionarily conserved cellular process, and it is particularly essential for the thermal tolerance of organisms under heat stress (Deka et al. 2016; Feder and Hofmann 1999; Somero 2020; Tomanek 2010). Although the HSR was firstly discovered in fruit flies under heat shock (Ritossa 1962), it was found to be a crucial mechanism to protect cells from various stimuli including hypoxia (Baird et al. 2006; Klumpen et al. 2017; Luo et al. 2021; Michaud et al. 2011). It was also reported that the heat shock transcription factor 1 (HSF1) was induced under hypoxia (Baird et al. 2006; Benjamin et al. 1990; Giaccia et al. 1992). The hypoxia-inducible factor-1α (HIF-1α) was an important transcription factor of hypoxia response (Semenza 2007). Previous studies suggested that it was also an evolutionarily conserved and a necessary component of heat acclimation (Treinin et al. 2003). HIF-1α regulated gene transcription by binding to the hypoxia response elements (HREs) within the promoter regions of target genes (Semenza 2007). Genes like HSP70 and HSF1 that involved in HSR were regulated via the HIF-1α pathway under hypoxic condition (Luo et al. 2021). Although it has been reported that HIF-1α activated the HSR via regulation of HSF1 (Agarwal and Ganesh 2020), the specific regulatory relationship between the two transcription factors is poorly understood in marine bivalves under heat stress.
The Pacific oyster (Crassostrea gigas) is widely distributed worldwide with important ecological and economic values which serve as an excellent model to study the adaption of marine invertebrates to costal environments (Dong et al. 2022; Guo 2009; Zhang et al. 2012). In this study, we found that the HIF-1α genes were induced in C. gigas in response to heat stress, resulting in the activation of HSR and contributing to thermal tolerance of C. gigas under heat stress. We for the first time revealed the role of HIF-1α/HSF1 pathway in oyster in response to high-temperature stress. Our findings suggested that HIF-1 could serve as a critical molecular integrator associated with thermal tolerance in a marine bivalve.
Materials and Methods
Gene Identification
The homologous sequence alignment was performed to identify the full set of the HIF-1α genes in C. gigas. We retrieved the HIF-1α protein sequences of representative species from NCBI database (https://www.ncbi.nlm.nih.gov/), including vertebrates (Homo sapiens, Mus musculus, Meleagris gallopavo, Gallus gallus, Anolis carolinensis, Chrysemys picta bellii, Xenopus laevis, Xenopus tropicalis, Danio rerio, Pangasianodon hypophthalmus, Branchiostoma belcheri) and invertebrates (Ciona intestinalis, Acanthaster planci, Strongylocentrotus purpuratus, Mizuhopecten yessoensis, Crassostrea virginica, Bombyx mori, Caenorhabditis elegans, Exaiptasia diaphana) (Supplementary Table 1). The downloaded sequences were then searched against the C. gigas genome (GenBank: GCA_902806645.1) using BLAST program with the E-value of 1E-10. The obtained sequences were finally confirmed by protein BLAST (BLASTP) (Altschul et al. 1997) against NCBI non-redundant (Nr) protein sequence database and manually curated using the Conserved Domain Search Service (https://www.ncbi.nlm.nih.gov/Structure/cdd/wrpsb.cgi). The gene location in the chromosome, the number of amino acids, and the base number of mRNA and UPR of the identified gene members were investigated according to the genome of C. gigas (GenBank: GCA_902806645.1).
Phylogenetic Analysis
The phylogenetic analysis of HIF-1α was performed using the protein sequences of C. gigas and representative species (H. sapiens, M. musculus, M. gallopavo, G. gallus, C. picta, P. sinensis, D. rerio, X. laevis, X. tropicalis, P. maximus, M. yessoensis, and C. virginica). The aryl hydrocarbon receptor nuclear translocator (ARNT, also known as HIF-1β) protein sequences of those species were also included in order to clearly distinguish the HIF-1α members. The phylogenetic tree was constructed using neighbor-joining (NJ) approach in MEGA7 (Kumar et al. 2016) with 1000 bootstraps.
Plasmid Construction
The AnimalTFDB 3.0 (http://bioinfo.life.hust.edu.cn/AnimalTFDB#!/) and JASPAR (http://jaspar.genereg.net/) were used to predict the putative binding sites of HIF-1α on promoter region of HSF1 (LOC105328117). The HSF1 promoter sequences containing different putative binding sites were amplified with specific primers and inserted into the NheI/HindIII site of pGL3-basic vector (Promega, USA) to construct the reporter plasmids (pGL3-HSF1). The open reading frame (ORF) of HIF-1α and HIF-1α-like5 was amplified and ligated to the pcDNA3.1( +) expression vector (Invitrogen, USA) to construct expression plasmids, respectively. The recombinant plasmids were constructed using the ClonExpress II One-Step Cloning Kit (Vazyme, Nanjing, China). Plasmids for transfection were prepared using EndoFree Max Plasmid Kit II (Tiangen, China) according to the manufacturer’s instructions. The primers used for plasmid construction were listed in Supplementary Table 2.
Cell Culture, Transfection, and Dual-Luciferase Reporter Assay
The HEK293T cells were cultured in DMEM high-glucose medium (Hyclone, USA) containing 10% FBS (Hyclone, USA) and 1% penicillin/streptomycin in incubator with 5% CO2 at 37 ℃. After 24 h culturing in 24-well plates, the cells were co-transfected with 0.5 µg of reporter plasmids and 0.25 µg of expression plasmids and 0.1 µg of pRL-TK Renilla luciferase plasmids (Promega, USA) using Xfect Transfection Reagent (Takara, Japan) according to the manufacturer’s protocol. After 48-h transfection, the Firefly luciferase and Renilla luciferase activities were determined using Dual-Luciferase Reporter Assay System (Promega, USA) according to the manufacturer’s instructions on a SYNERGY H1 microplate reader (BioTek, USA).
Long Double-Stranded RNA (dsRNA) Preparation
The HIF-1α and HIF-1α-like5 target cDNA fragment was amplified with primer pairs (Supplementary Table 3), respectively. The cDNA fragment was added T7 promoter by cloning to the pGEM®-T vector (Promega, USA). The linearized plasmid templates were used to synthesize dsRNA using T7 RiboMAX™ Express RNAi System (Promega, USA) according to the manufacturer’s instructions. The quality and concentration of dsRNA were assessed by 1.5% agarose gel electrophoresis and Nanodrop 2000 spectrophotometer (Thermo Scientific, USA).
Experiment Animals, RNAi, and Heat Shock Treatment
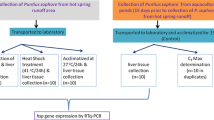
The healthy 1-year-old Pacific oysters (55 ± 6 mm in shell height) were collected from Weihai, China (July 2020), and transported to laboratory for experiment. The oysters were drilled with a small hole on shell near adductor muscle and were kept in filtered seawater (20 ± 0.5 ℃) for 1-week acclimation. During the acclimation period, oysters were fed with algae, and the water was changed daily. After acclimation, the oysters were randomly divided into three groups, the HIF-1α dsRNA injection group (n = 66), the HIF-1α-like5 dsRNA injection group (n = 66), and the PBS injection group (n = 66). The oysters were individually injected with 100 μL of 50 μg/100 μL HIF-1α dsRNA, 100 μL of 50 μg/100 μL HIF-1α-like5 dsRNA, or 100 μL 1 × PBS into the adductor muscle using micro syringe (Tian et al. 2021). The injected oysters were placed into tanks with filtered seawater at 35 ± 0.5 ℃ (Liu et al. 2019). The gills of six individuals were randomly sampled at 0, 3, 12, and 24 h from each group after injection. The gills were dissected, flash-frozen in liquid nitrogen, and stored at − 80 ℃ freezer until use for qRT-PCR analysis. Dead oysters were counted and removed from the tank during the experiment. Thirty-six individuals were used in each group.
Gene Expression Analysis
The mRNA expression levels of HIF-1α members and key genes in HSR pathway were determined in C. gigas under chronic heat stress (30 ℃) as previously reported (Fu et al. 2021). Briefly, the oysters were placed in tanks with a constant flow system, and the water was elevated at a rate of 1 °C/h to 30 °C. Total RNA was extracted from gills with TRIzol® reagent (Invitrogen, USA) and used for cDNA synthesis with PrimeScript™ RT Master Mix Perfect Real-Time Kit (Takara, Japan) according to the manufacturer’s introductions. The qRT-PCR was carried in triplicates (two individuals per replicate) on a LightCycler 480 real-time PCR machine (Roche, Switzerland) with a total volume of 10-μL reaction mix using SYBR Green PCR Master Mix (QIAGEN, Germany) with the following conditions: 95 °C for 2 min, 40 cycles at 95 °C for 5 s, and 60 °C for 10 s. The mRNA relative expression level of gene was calculated using the relative 2−ΔΔCT method (Livak and Schmittgen 2001). The EF-1α gene was used as control as previously reported (Liu et al. 2019). All the primers used for qRT-PCR analysis were listed in Supplementary Table 4.
Results
Identification of HIF-1α Genes in C. gigas Genome
A total of six HIF-1α homologous genes were identified in the C. gigas genome (Table 1). All of the members contain the classic conserved domains of HIF-1α, including basic-helix-loop-helix (bHLH) and Per-Arnt-Sim (PAS). The genes were renamed according to their location in the chromosome except for the HIF-1α (Gene ID: 105324565) which has been well annotated in NCBI. The phylogenetic analysis suggested that HIF-1α genes in C. gigas were clustered with those from invertebrates, and there was an obvious distinction between the HIF-1α and the ARNT (HIF-1β) clades (Fig. 1).
Phylogenetic analysis of HIF-1α genes in C. gigas. The phylogenetic tree was constructed using the neighbor-joining (NJ) method in MEGA 7 software with 1000 bootstraps. Abbreviations: Hsa, Homo sapiens; Mmu, Mus musculus; Mga, Meleagris gallopavo; Gga, Gallus gallus, Cpi, Chrysemys picta; Psi, Pelodiscus sinensis; Dre, Danio rerio; Xla, Xenopus laevis; Xtr, Xenopus tropicalis; Pma, Pecten maximus; Mye, Mizuhopecten yessoensis; Cvi, Crassostrea virginica; Cgi, Crassostrea gigas. The C. gigas HIF-1α were labeled by red spot
HIF-1α Genes Were Induced in C. gigas Under Heat Stress
In order to determine whether the HIF-1α genes were involved in response to heat stress, the mRNA expression levels of HIF-1α members were determined in C. gigas under chronic heat stress (30 ℃). The results showed that HIF-1α and HIF-1α-like5 were significantly induced in C. gigas at 22 h and 34 h post-heat stress (P < 0.05) (Fig. 2). Therefore, we focused on these two members for further analyses.
Knocking Down of HIF-1α Inhibited the Expression of HSF1 and HSP70 Genes in C. gigas Under Heat Stress
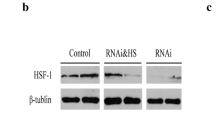
The expression of HSF1 and HSP70 genes was used to monitor the HSR. Apparently, both of HSF1 and HSP70 were significantly induced in C. gigas under chronic heat stress (P < 0.05) (Fig. 3). In order to confirm the involvement of HIF-1α in the HSR of C. gigas under heat stress, the RNAi treatment through dsRNA injection was used to knockdown the targeted HIF-1α homologs (Fig. 4). The mRNA expression level of HIF-1α was continuously decreased after HIF-1α dsRNA injection and significantly lower than that in PBS injection group. Notably, injection of HIF-1α dsRNA also significantly decreased the relative expression levels of HIF-1α-like5 after 24-h injection (P < 0.05) (Fig. 4A). Similarly, both mRNA expression levels of HIF-1α and the HIF-1α-like5 were significantly decreased after HIF-1α-like5 dsRNA injection, while there was no significant difference at 24 h post-injection compared with 0 h and PBS injection group (P < 0.05) (Fig. 4B). Moreover, the mRNA expression of HSF1 was significantly suppressed at 24 h after injection of HIF-1α dsRNA or HIF-1α-like5 dsRNA (P < 0.05) (Fig. 4C). Although the mRNA expression of HSP70 continued to decrease after dsRNA injection of HIF-1α or HIF-1α-like5, it was significantly higher in HIF-1α-like5 dsRNA injection group than that in HIF-1α dsRNA injection group at the early stage of the experiment (P < 0.05) (Fig. 4D).
Knocking down of HIF-1α- and HIF-1α-like5 altered expression of HIF-1α- and HSR-related genes. Relative expression of A HIF-1α, B HIF-1α-like5, C HSF1, and D HSP70 in C. gigas under heat stress after knocking down of HIF-1α genes. Different letters on the error bars represent significant differences (mean ± SD, n = 3, P < 0.05)
The HSF1 Was Directly Regulated by HIF-1α in C. gigas
To verify the regulatory role of HIF-1α on HSF1, a 2124 bp upstream sequence of the transcription start site of HSF1 was analyzed using dual-luciferase reporter assays. The results suggested that both of HIF-1α and HIF-1α-like5 significantly activated the transcriptional activity of HSF1 (P < 0.05) (Supplementary Fig. 1). To further determine the core promoter region, a series of plasmids with the truncated fragments of the HSF1 promoter containing different numbers of predicted HREs was constructed (Fig. 5A and Supplementary Fig. 2). The relative luciferase activity in the cells that was transfected with plasmids containing HREs of HSF1, and ORF of HIF-1α was significantly higher than that in negative controls (P < 0.05) (Fig. 5B). Similar result was observed in the cells containing HREs of HSF1 and ORF of HIF-1α-like5 (Fig. 5C). The position − 380 and − 180 of the HSF1 promoter with two binding sites was identified as the core region for HIF-1α and HIF-1α-like5 transcription regulation since the highest relative luciferase activity was observed (P < 0.05) (Fig. 5B and C).
Regulation of HIF-1α on HSF1 promoter in C gigas. A Schematic presentation of the putative binding sites of HIF-1α in HSF1 promoter region (2124 bp) in C. gigas. B Relative luciferase activity in HEK293T cells containing different deletion constructs of HSF1 promoter and HIF-1α. C Relative luciferase activity in HEK293T cells containing different deletion constructs of HSF1 promoter and HIF-1α-like5. Data were presented as means ± SD (n = 3). Different letters on the bar indicate significant difference (P < 0.05)
Knocking Down of HIF-1α Decreased the Survival of C. gigas Under Heat Stress
In order to investigate whether the expression of HIF-1α genes was associated with thermal tolerance of C. gigas under heat stress, we examined the survival of oysters under heat stress with in vivo RNAi to knock down the transcriptional levels of HIF-1α genes. The survival rate of C. gigas under heat stress at 24 h in PBS group, HIF-1α dsRNA group, and HIF-1α-like5 dsRNA group was 47%, 11% and 66%, respectively (Fig. 6). The HIF-1α dsRNA injection significantly reduced the survival rate of C. gigas under heat stress compared with the PBS and HIF-1α-like5 dsRNA group (P < 0.01). Although there was no significant difference between PBS and HIF-1α-like5 dsRNA group, the mortality was firstly observed in the HIF-1α-like5 dsRNA group after heat stress.
Discussion
Environmental temperature affects the physiological and biochemical processes of organisms (Tomanek 2014). Ocean temperature rising due to global warming has posed a serious threat to the marine animals, especially those widely distributed in tidal zones, causing ecological problems and economical losses (Barbosa Solomieu et al. 2015; Gunderson and Stillman 2015; Khan et al. 2020). The HSR is an evolutionarily conserved mechanism to protect cells from heat stress which contributes to thermal tolerance of marine organisms (Clark et al. 2008; Deka et al. 2016; Dong et al. 2011; Jeremias et al. 2018; Tomanek 2010). HSR and hypoxia response are interactive cellular processes during heat stress (Ely et al. 2014; Hu et al. 2022; Huo et al. 2019; Khan et al. 2020). The transcription factor, HIF-1α, coordinately activates multiple functional genes to promote cell survival under direct or indirect hypoxia (Bailey and Nathan 2018; Krejčová et al. 2019; Sun et al. 2016; Wang et al. 2016). Considering that, we investigated the involvement of HIF-1α in response to heat shock in C. gigas, which is a widely studied model of marine bivalves (Song et al. 2019; Zhang et al. 2012).
Previous studies have reported the involvement of HIF-1α in heat adaptation in invertebrates (Cai et al. 2014; Klumpen et al. 2017; Treinin et al. 2003). In this study, we identified a total of six HIF-1α genes in C. gigas genome, all of which contain classic conserved domains of HIF-1α family and have a closer homology to the clade of HIF-1α in vertebrates (Dengler et al. 2014; Wang et al. 1995). Since the gill is the respiratory and immune organ of bivalves and high temperature would cause damages on the gill tissues and affect its function (Tomanek 2012; Li et al. 2017), we further conducted the expression analysis of HIF-1α in gills of C. gigas under chronic heat stress. The results showed that HIF-1α and HIF-1α-like5 were greatly induced, which were consistent with previous observation in marine mollusks (Cai et al. 2014; Kawabe and Yokoyama 2012). These results suggested that the two members of HIF-1α could be functionally important for C. gigas in response to high temperature. Knocking down of HIF-1α or HIF-1α-like5 significantly decreased the expression levels of HSF1 and HSP70, which were the marker genes of HSR, indicating that the expression of HIF-1α was associated with activation of HSR in C. gigas under heat stress. This result was consistent with previous studies in which the HIF-1α was considered as crucial regulator for the activation of HSR (Agarwal and Ganesh 2020; Ali et al. 2011). The HIF-1α/HSF1 signaling pathway in large yellow croaker and oyster under hypoxia has been reported (Kawabe and Yokoyama 2011; Luo et al. 2021). Therefore, a potential regulation of HIF-1 on HSF1 in C. gigas in response to heat stress was speculated. As a transcription factor, HIF-1α activates gene transcription by binding to HREs in the promoter region of target genes (Dengler et al. 2014; Kaluz et al. 2008). In previous studies, several putative HREs (a core 5ʹ-RCGTG sequence) were identified in a region of − 1328 bp before the transcription start site of HSF1 in C. gigas (Kawabe and Yokoyama 2011). In this present study, the luciferase assay experiment revealed that two HREs located at position − 360 to − 180 of HSF1 promoter were both critical for the transcriptional regulation of HIF-1α and HIF-1α-like5 in C. gigas. It has been reported that functional HREs tended to localize at the proximal promoters of target genes and the proximal functional elements were necessary and sufficient to regulate the transcriptional activity (Ali et al. 2011; Dengler et al. 2014). These results supported the direct regulatory relationship between HIF-1α and HSF1 in C. gigas under heat stress.
The survival of C. gigas under heat stress was decreased after knocking down of HIF-1α gene, which provided further support on the regulation of HSP via HIF-1α-mediated signaling pathway. All of these results may imply the potential roles of the HIF-1α genes in the thermal tolerance of C. gigas. Heat stress could cause tissue damages and result in a series of physiological responses in oyster through induction of oxidative stress and energy trade-off (Li et al. 2021; Nash et al. 2019). Multiple cellular response pathways were involved in the protection processes from heat stress, in which the HSR and hypoxia response were the most studied interactive processes in organisms under heat stress (Ely et al. 2014; Hu et al. 2022; Huo et al. 2019). Moreover, the HIF-1α occupied an important position in the signaling pathways that regulated various physiological functions (Liang et al. 2022; Luo et al. 2021). Therefore, we reasoned that the HIF-1α served as one of the critical regulators for C. gigas in response to heat stress.
Conclusion
In this study, a total of six HIF-1α genes were identified in C. gigas genome and were investigated for their potential contribution in response to heat stress. The HIF-1α and HIF-1α-like5 were highly induced in C. gigas under chronic heat stress. The RNAi experiment in vivo and dual-luciferase reporter assay demonstrated the direct regulation of HIF-1α on the transcriptional activity of HSF1, suggesting the critical role of HIF-1α in contributing to thermal tolerance of C. gigas under heat stress. The present study for the first time revealed the involvement of HIF-1α/HSF1/HSP70 pathway in response to heat stress in the oyster and provided an insight into adaptive mechanism of bivalves in face of ocean warming.
References
Agarwal S, Ganesh S (2020) Perinuclear mitochondrial clustering, increased ROS levels, and HIF1 are required for the activation of HSF1 by heat stress. J Cell Sci 133:jcs245589
Ali YO, McCormack R, Darr A, Zhai RG (2011) Nicotinamide mononucleotide adenylyltransferase is a stress response protein regulated by the heat shock factor/hypoxia-inducible factor 1α pathway. J Biol Chem 286:19089–19099
Altschul SF, Madden TL, Schäffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–4340
Bailey PSJ, Nathan JA (2018) Metabolic regulation of hypoxia-inducible transcription factors: the role of small molecule metabolites and iron. Biomedicines 6:60
Baird NA, Turnbull DW, Johnson EA (2006) Induction of the heat shock pathway during hypoxia requires regulation of heat shock factor by hypoxia-inducible factor-1. J Biol Chem 281:38675–38681
Barbosa Solomieu V, Renault T, Travers M-A (2015) Mass mortality in bivalves and the intricate case of the Pacific oyster, Crassostrea gigas. J Invertebr Pathol 131:2–10
Benjamin IJ, Kröger B, Williams RS (1990) Activation of the heat shock transcription factor by hypoxia in mammalian cells. Proc Natl Acad Sci USA 87:6263–6267
Cai X, Huang Y, Zhang X, Wang S, Zou Z, Wang G, Wang Y, Zhang Z (2014) Cloning, characterization, hypoxia and heat shock response of hypoxia inducible factor-1 (HIF-1) from the small abalone Haliotis diversicolor. Gene 534:256–264
Chapman MG (2006) Intertidal seawalls as habitats for molluscs. J Molluscan Stud 72:247–257
Chen Y, Wang J, Liao M, Li X, Dong Y (2021) Temperature adaptations of the thermophilic snail Echinolittorina malaccana: insights from metabolomic analysis. J Exp Biol 224:jeb238659
Cheng L, Abraham J, Trenberth KE et al (2021) Upper ocean temperatures hit record high in 2020. Adv Atmos Sci 38:523–530
Clark MS, Fraser KPP, Peck LS (2008) Lack of an HSP70 heat shock response in two Antarctic marine invertebrates. Polar Biol 31:1059–1065
Deka K, Singh A, Chakraborty S, Mukhopadhyay R, Saha S (2016) Protein arginylation regulates cellular stress response by stabilizing HSP70 and HSP40 transcripts. Cell Death Discov 2:1–8
Dengler VL, Galbraith MD, Espinosa JM (2014) Transcriptional regulation by hypoxia inducible factors. Crit Rev Biochem Mol Biol 49:1–15
Dong Y, Yu S, Wang Q, Dong S (2011) Physiological responses in a variable environment: relationships between metabolism, hsp and thermotolerance in an intertidal-subtidal species. PLoS ONE 6:e26446
Dong Y, Liao M, Han G, Somero GN (2022) An integrated, multi-level analysis of thermal effects on intertidal molluscs for understanding species distribution patterns. Biol Rev 97:554–581
Ely BR, Lovering AT, Horowitz M, Minson CT (2014) Heat acclimation and cross tolerance to hypoxia: bridging the gap between cellular and systemic responses. Temperature (austin) 1:107–114
Feder ME, Hofmann GE (1999) Heat-shock proteins, molecular chaperones, and the stress response: evolutionary and ecological physiology. Ann Rev Physio 61:243–282
Freitas C, Villegas-Ríos D, Moland E, Olsen EM (2021) Sea temperature effects on depth use and habitat selection in a marine fish community. J Anim Ecol 90:1787–1800
Fu H, Jiao Z, Li Y, Tian J, Ren L, Zhang F, Li Q, Liu S (2021) Transient receptor potential (TRP) channels in the Pacific oyster (Crassostrea gigas): genome-wide identification and expression profiling after heat stress between C. gigas and C. angulata. Int J Mol Sci 22:3222
Giaccia AJ, Auger EA, Koong A, Terris DJ, Minchinton AI, Hahn GM, Brown JM (1992) Activation of the heat shock transcription factor by hypoxia in normal and tumor cell lines in vivo and in vitro. Int J Radiat Oncol Biol Phys 23:891–897
Gunderson AR, Stillman JH (2015) Plasticity in thermal tolerance has limited potential to buffer ectotherms from global warming. Proc Biol Sci 282:20150401
Guo X (2009) Use and exchange of genetic resources in molluscan aquaculture. Rev Aquac 1:251–259
Guo X, Wang Y, Wang L, Lee J-H (2008) Oysters. In: Kocher T, Kole C (eds) Genome mapping and genomics in fishes and aquatic animals. Springer, Berlin, Heidelberg, pp 163–175
Hu Z, Feng J, Song H, Zhou C, Yu Z-L, Yang M-J, Shi P, Guo Y-J, Li Y-R, Zhang T (2022) Mechanisms of heat and hypoxia defense in hard clam: insights from transcriptome analysis. Aquaculture 549:737792
Huo D, Sun L, Zhang L, Ru X, Liu S, Yang H (2019) Metabolome responses of the sea cucumber Apostichopus japonicus to multiple environmental stresses: heat and hypoxia. Mar Pollut Bull 138:407–420
Jeremias G, Barbosa J, Marques SM, Asselman J, Gonçalves FJM, Pereira JL (2018) Synthesizing the role of epigenetics in the response and adaptation of species to climate change in freshwater ecosystems. Mol Ecol 27:2790–2806
Kaluz S, Kaluzová M, Stanbridge EJ (2008) Regulation of gene expression by hypoxia: integration of the HIF-transduced hypoxic signal at the hypoxia-responsive element. Clin Chim Acta 395:6–13
Kawabe S, Yokoyama Y (2011) Novel isoforms of heat shock transcription factor 1 are induced by hypoxia in the Pacific oyster Crassostrea gigas. J Exp Zool 315A:394–407
Kawabe S, Yokoyama Y (2012) Role of hypoxia-inducible factor α in response to hypoxia and heat shock in the Pacific oyster Crassostrea gigas. Mar Biotechnol 14:106–119
Khan FU, Hu M, Kong H, Shang Y, Wang T, Wang X, Xu R, Lu W, Wang Y (2020) Ocean acidification, hypoxia and warming impair digestive parameters of marine mussels. Chemosphere 256:127096
Klumpen E, Hoffschröer N, Zeis B, Gigengack U, Dohmen E, Paul RJ (2017) Reactive oxygen species (ROS) and the heat stress response of Daphnia pulex: ROS-mediated activation of hypoxia-inducible factor 1 (HIF-1) and heat shock factor 1 (HSF-1) and the clustered expression of stress genes. Biol Cell 109:39–64
Krejčová G, Danielová A, Nedbalová P, Kazek M, Strych L, Chawla G, Tennessen JM, Lieskovská J, Jindra M, Doležal T, Bajgar A (2019) Drosophila macrophages switch to aerobic glycolysis to mount effective antibacterial defense. eLife 8:e50414
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874
Li A, Li L, Zhang Z, Li S, Wang W, Guo X, Zhang G (2021) Noncoding variation and transcriptional plasticity promote thermal adaptation in oysters by altering energy metabolism. Mol Biol Evol 38:5144–5155
Li Y, Qin JG, Abbott CA, Li X, Benkendorff K (2007) Synergistic impacts of heat shock and spawning on the physiology and immune health of Crassostrea gigas: an explanation for summer mortality in Pacific oysters. Am J Physiol Regul Integr Comp Physiol 293:R2353–R2362
Li Y, Qin JG, Li X, Benkendorff K (2009) Monthly variation of condition index, energy reserves and antibacterial activity in Pacific oysters, Crassostrea gigas, in Stansbury (South Australia). Aquaculture 286:64–71
Li Y, Song X, Wang W et al (2017) The hematopoiesis in gill and its role in the immune response of Pacific oyster Crassostrea gigas against secondary challenge with Vibrio splendidus. Dev Comp Immunol 71:59–69
Liang R, Liu N, Cao J, Liu T, Sun P, Cai X, Zhang L, Liu Y, Zou J, Wang L, Ding X, Zhang B, Shen Z, Yoshida S, Dou J, Wang S (2022) HIF-1α/FOXO1 axis regulated autophagy is protective for β cell survival under hypoxia in human islets. Biochim Biophys Acta Mol Basis Dis 1868(5):166356
Liu Y, Li L, Huang B, Wang W, Zhang G (2019) RNAi based transcriptome suggests genes potentially regulated by HSF1 in the Pacific oyster Crassostrea gigas under thermal stress. BMC Genom 20:639
Livak KJ, Schmittge TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25:402–408
Luo S-Y, Wang J-Q, Liu C, Gao X-M, Zhang Y-B, Ding J, Hou C-C, Zhu J-Q, Lou B, Shen W-L, Wu X-F, Zhang C-D, Tang D-J (2021) Hif-1α/Hsf1/Hsp70 signaling pathway regulates redox homeostasis and apoptosis in large yellow croaker (Larimichthys crocea) under environmental hypoxia. Zool Res 42:746–760
McArley TJ, Hickey AJR, Herbert NA (2020) Acute high temperature exposure impairs hypoxia tolerance in an intertidal fish. PLoS ONE 15:e0231091
McLeod IM, zu Ermgassen PSE, Gillies CL, Hancock B (2019) Chapter 25 - Can bivalve habitat restoration improve degraded estuaries? In: Wolanski E, Day JW, Elliott M, Ramachandran R (eds) Coasts and Estuaries. Elsevier, pp 427–442
Michaud MR, Teets NM, Peyton JT, Blobner BM, Denlinger DL (2011) Heat shock response to hypoxia and its attenuation during recovery in the flesh fly, Sarcophaga crassipalpis. J Insect Physiol 57:203–210
Mosca F, Narcisi V, Calzetta A, Gioia L, Finoia MG, Latini M, Tiscar PG (2013) Effects of high temperature and exposure to air on mussel (Mytilus galloprovincialis, Lmk 1819) hemocyte phagocytosis: modulation of spreading and oxidative response. Tissue Cell 45:198–203
Nash S, Johnstone J, Rahman MS (2019) Elevated temperature attenuates ovarian functions and induces apoptosis and oxidative stress in the American oyster, Crassostrea virginica: potential mechanisms and signaling pathways. Cell Stress Chaperones 24:957–967
Patrick S, Faury N, Goulletquer P (2006) Seasonal changes in carbohydrate metabolism and its relationship with summer mortality of Pacific oyster Crassostrea gigas (Thunberg) in Marennes-Oléron bay (France). Aquaculture 252:328–338
Petes LE, Menge BA, Murphy GD (2007) Environmental stress decreases survival, growth, and reproduction in New Zealand mussels. J Exp Mar Biol Ecol 351:83–91
Pinsky ML, Eikeset AM, McCauley DJ, Payne JL, Sunday JM (2019) Greater vulnerability to warming of marine versus terrestrial ectotherms. Nature 569:108–111
Ritossa F (1962) A new puffing pattern induced by temperature shock and DNP in drosophila. Experientia 18:571–573
Semenza GL (2007) Hypoxia-inducible factor 1 (HIF-1) pathway. Sci STKE 2007:cm8
Solan M, Whiteley N (2016) Stressors in the marine environment: physiological and ecological responses; societal implications. Oxford University Press
Somero GN (2020) The cellular stress response and temperature: function, regulation, and evolution. J Exp Zool A Ecol Integr Physiol 333:379–397
Song K, Wen S, Zhang G (2019) Adaptive evolution patterns in the Pacific oyster Crassostrea gigas. Mar Biotechnol 21:614–622
Strehse JS, Maser E (2020) Marine bivalves as bioindicators for environmental pollutants with focus on dumped munitions in the sea: a review. Mar Environ Res 158:105006
Sun S, Xuan F, Fu H, Ge X, Zhu J, Qiao H, Jin S, Zhang W (2016) Molecular characterization and mRNA expression of hypoxia inducible factor-1 and cognate inhibiting factor in Macrobrachium nipponense in response to hypoxia. Comp Biochem Physiol B Biochem Mol Biol 196–197:48–56
Tian J, Li Y, Fu H et al (2021) Physiological role of CYP17A1-like in cadmium detoxification and its transcriptional regulation in the Pacific oyster. Crassostrea Gigas Sci Total Environ 796:149039
Tomanek L (2010) Variation in the heat shock response and its implication for predicting the effect of global climate change on species’ biogeographical distribution ranges and metabolic costs. J Exp Biol 213:971–979
Tomanek L (2012) Environmental proteomics of the mussel Mytilus: implications for tolerance to stress and change in limits of biogeographic ranges in response to climate change. Integr Comp Biol 52:648–664
Tomanek L (2014) Proteomics to study adaptations in marine organisms to environmental stress. J Proteomics 105:92–106
Treinin M, Shliar J, Jiang H, Powell-Coffman JA, Bromberg Z, Horowitz M (2003) HIF-1 is required for heat acclimation in the nematode Caenorhabditis elegans. Physiol Genomics 14:17–24
Vaquer-Sunyer R, Duarte CM (2011) Temperature effects on oxygen thresholds for hypoxia in marine benthic organisms. Glob Chang Biol 17:1788–1797
Wang GL, Jiang BH, Rue EA, Semenza GL (1995) Hypoxia-inducible factor 1 is a basic-helix-loop-helix-PAS heterodimer regulated by cellular O2 tension. Proc Natl Acad Sci USA 92:5510–5514
Wang T, Meng J, Li L, Zhang G (2016) Characterization of CgHIFα-like, a novel bHLH-PAS transcription factor family member, and its role under hypoxia stress in the Pacific Oyster Crassostrea gigas. PLoS ONE 11:e0166057
Wijsman JWM, Troost K, Fang J, Roncarati A (2019) Global production of marine bivalves. Trends and challenges. In: Smaal AC, Ferreira JG, Grant J, Petersen JK, Strand Ø (eds) Goods and Services of Marine Bivalves. Springer International Publishing, Cham, pp 7–26
Zhang G, Fang X, Guo X, Li L, Luo R, Xu F, Yang P, Zhang L, Wang X, Qi H, Xiong Z, Que H, Xie Y, Holland PWH, Paps J, Zhu Y, Wu F, Chen Y, Wang Jiafeng, Peng C, Meng J, Yang L, Liu J, Wen B, Zhang N, Huang Z, Zhu Q, Feng Y, Mount A, Hedgecock D, Xu Z, Liu Y, Domazet-Lošo T, Du Y, Sun X, Zhang Shoudu, Liu B, Cheng P, Jiang X, Li J, Fan D, Wang W, Fu W, Wang T, Wang B, Zhang J, Peng Z, Li Yingxiang, Li Na, Wang Jinpeng, Chen M, He Y, Tan F, Song X, Zheng Q, Huang R, Yang Hailong, Du X, Chen L, Yang M, Gaffney P.M, Wang S, Luo L, She Z, Ming Y, Huang W, Zhang Shu, Huang B, Zhang Y, Qu T, Ni P, Miao G, Wang Junyi, Wang Q, Steinberg CEW, Wang H, Li Ning, Qian L, Zhang Guojie, Li Yingrui, Yang Huanming, Liu X, Wang Jian, Yin Y, Wang Jun (2012) The oyster genome reveals stress adaptation and complexity of shell formation. Nature 490:49–54
Zippay ML, Helmuth B (2012) Effects of temperature change on mussel, Mytilus. Integr Zool 7:312–327
Funding
This study was supported by grant from the National Key Research and Development Program of China (2022YFD2400300), the Key Research and Development Program of Shandong Province (No. 2021ZLGX03), the National Natural Science Foundation of China (Nos. 41976098 and 42276112) and the Agriculture Research System of China Project (CARS-49).
Author information
Authors and Affiliations
Contributions
S.L. conceived the study and obtained the funding. H.F., Y.L., and J.T. performed the experiment. H.F., B.Y., and Y.L. analyzed the data. F.H. drafted the manuscript, and S.L. revised the manuscript. Q.L. supervised the work. All authors reviewed manuscript.
Corresponding author
Ethics declarations
Ethics Approval
The C. gigas, C. angulata, and their reciprocal hybrids are neither an endangered nor protected species. All experiments in this study were conducted according to national and institutional guidelines.
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Fu, H., Li, Y., Tian, J. et al. Contribution of HIF-1α to Heat Shock Response by Transcriptional Regulation of HSF1/HSP70 Signaling Pathway in Pacific Oyster, Crassostrea gigas. Mar Biotechnol 25, 691–700 (2023). https://doi.org/10.1007/s10126-023-10231-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10126-023-10231-6