Abstract
Cryptosporidiosis is an important though underreported public health concern. Molecular tools might be helpful in improving its diagnosis. In this study, ZR Fecal DNA MiniPrep™ Kit (ZR) and NucliSens® easyMAG® (EM) were compared using four Cryptosporidium-seeded feces and 29 Cryptosporidium-positive stools. Thereafter, ZR was selected for prospective evaluation of Cryptosporidium detection by 18S rDNA and LAXER quantitative PCR (qPCR) in 69 stools from 56 patients after Cryptosporidium detection by glycerin, modified Ziehl–Neelsen (ZN) and auramine–phenol (AP) stainings. The combination of any of the two extraction methods with 18S qPCR yielded adequate detection of Cryptosporidium in seeded stools, but the ZR kit showed the best performance. All 29 Cryptosporidium-positive samples were positive with 18S qPCR, after both ZR and EM extraction. However, false-negative results were found with LAXER qPCR or nested PCR. Cryptosporidiosis was diagnosed in 7/56 patients. All the microscopic methods enabled the initial diagnosis, but Cryptosporidium was detected in 12, 13, and 14 samples from these seven patients after glycerin, ZN, and AP staining respectively. Among these samples, 14 and 12 were positive with 18S and LAXER qPCR respectively. In two patients, Cryptosporidium DNA loads were found to be correlated with clinical evolution. Although little known, glycerin is a sensitive method for the initial detection of Cryptosporidium. When combined with 18S qPCR, ZR extraction, which had not been evaluated so far for Cryptosporidium, was an accurate tool for detecting Cryptosporidium and estimating the oocyst shedding in the course of infection.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cryptosporidiosis is an emerging disease caused by apicomplexan parasites belonging to the genus Cryptosporidium. Twenty-seven recognized species and more than 40 genotypes have been described in the literature [1–3], but C. parvum and C. hominis represent the most prevalent agents infecting humans [4, 5]. Some other species are considered to be pathogens in both immunocompetent and immunocompromised individuals, but their prevalence remains much lower. Infection is most commonly spread by the fecal–oral route, through oocyst-contaminated water or food or by direct contact with infected humans or animals as well as contaminated non-living objects (fomites). Transmission by aerosol inhalation of oocysts has also been reported, but it remains very uncommon [6]. The severity of illness depends on both host and parasite factors, which include immune and nutritional status, age, and species/genotype involved. In healthy subjects, cryptosporidiosis is a transient self-limiting gastroenteritis-like syndrome that usually lasts 1–2 weeks [4]. But immunocompromised patients (especially with HIV/AIDS) can suffer from profuse, watery diarrhea, resulting in malabsorption, wasting, and reduced life expectancy [6, 7]. Extra-intestinal manifestations, such as biliary involvement, pancreatitis, and respiratory tract disease, may also occur in deeply immunosuppressed individuals with severe cryptosporidiosis [8]. Nevertheless, the treatment options are limited, and no antiparasitic drug has been proven to possess clear activity against Cryptosporidium [9].
The lessons learnt from the numerous waterborne and foodborne outbreaks that have occurred worldwide during the 2 past decades [10, 11] have elevated Cryptosporidium spp. to the rank of parasites of major public health importance. In 2012, a joint Food and Agriculture Organization (FAO) and WHO expert committee ranked Cryptosporidium in fifth place among the 24 most important food-borne parasites [2]. In addition, the parasite has been recently identified as one of the main causes of mild-to-severe diarrhea in children <5 years old in developing countries [12]. Information regarding the overall incidence of cryptosporidiosis is limited. In France, cryptosporidiosis cases have been notified to the ANOFEL Cryptosporidium National Network (ACNN) since 2006, and ≈ 100 cases per year (≈0.15/100,000 population/year) have been reported [13]. However, cryptosporidiosis probably continues to be underestimated, primarily because sampling from patients with gastrointestinal impairments and specific requests for diagnostic tests is insufficient.
Biological diagnosis relies on the detection of Cryptosporidium oocysts in stool samples by microscopic examination of smears prepared from formalin–ether concentrates and stained with the historical modified Ziehl–Neelsen (ZN) method [14], but sensitivity is low (~75 %) [15, 16]. Increased sensitivities of 90–100 % can be obtained with fluorescent auramine O and immunofluorescent assays using monoclonal antibodies [16–18]. But the sensitivity of these microscopic methods remains dependent on the clinical status of the patient, yielding false-negative results among poorly symptomatic or asymptomatic subjects with low parasitic loads and light infections. Immunochromatographic assays (ICA) have also been developed to detect Cryptosporidium antigens. However, a recent French blind multicenter study demonstrated limited interest, since the sensitivity of the four tested ICAs ranged from 50.1 % to 86.7 % for C. parvum and C. hominis, and was <35 % for the other species [19].
More recently, molecular methods, which are required for species identification and furthermore allow DNA quantification when real-time assays are used, have been developed. The higher sensitivity reported for Cryptosporidium PCR in several studies suggests that molecular tools could improve the diagnosis of cryptosporidiosis [20, 21], and they are increasingly used as first-line in clinical diagnostics [22]. 18S rDNA is the most frequently used target [23]. PCRs targeting the Cryptosporidium outer wall protein (COWP) [24], 60-kDa glycoprotein (gp60) [25], actin [26], LAXER [27], beta-tubulin [28] or HSP-70 [29] have also been developed. However, the small amount of target DNA combined with the low efficiency of total DNA extraction due to oocyst cell wall sturdiness, and the presence of PCR inhibitors in the samples to be amplified, may result in detection of false negatives. Moreover, only a few publications have focused on the optimization of Cryptosporidium DNA extraction in stools [30–34]. In this context, we aimed to (i) develop an optimal procedure for Cryptosporidium DNA isolation, detection and quantification, by comparing the performance of two commercial DNA extraction kits and the accuracy of two real-time quantitative PCR (qPCR) methods targeting 18S rDNA or LAXER locus, and (ii) evaluate the contribution of qPCR to the diagnosis and follow-up of cryptosporidiosis by comparing their performances to those of conventional microscopic techniques [ZN, auramine–phenol (AP), and glycerin] in a prospective study enrolling patients at risk for Cryptosporidium infection.
Materials and methods
Fecal sample collection
The fecal samples analyzed in this study included: (i) four human negative fecal samples of distinct origin (one child and three adults) and consistency (two soft, one liquid and one greasy stools, that were identified as S1, S2, S3, and S4 respectively), selected on the basis of negative results for cryptosporidia after ZN staining, and for other parasite cysts or ova after formalin–ether concentration, that were spiked with C. hominis oocysts isolated from a human diarrheal stool containing 2.105 oocysts per milliliter (P1) for comparison of two DNA extraction methods, (ii) 29 Cryptosporidium-positive fecal specimens from diarrhoeal patients (P2 to P30) stored at +4 °C in tubes containing 2.5 % potassium dichromate, and selected from the French ACNN collection [13], which included 13 C. hominis (P2 to P14), 12 C. parvum (P15 to P26) and 4 C. felis (P27 to P30) isolates, according to molecular identification performed as described previously [13], and (iii) 69 fecal specimens from 56 patients presenting gastro-intestinal symptoms, prospectively collected in the Parasitology–Mycology Laboratory of Lille University Hospital between September 2012 and April 2013. Fecal samples from subjects hospitalized in Lille University Hospital, considered at risk for cryptosporidiosis, were included in the study based on the following criteria: (i) liquid consistency (all adult or pediatric immunocompetent and immunosuppressed patients), or (ii) soft consistency if age was < 5 years, or if the patient had undergone a transplant, suffered from hematological malignancy, or an infectious disease. Stools submitted to the Parasitology–Mycology Laboratory of Lille University Hospital by other hospital centers for specific Cryptosporidium detection were also included.
Authorization for utilization of the stool isolates that were collected in Lille University Hospital was obtained from the French Ministry of Research (N°DC-2008-642). Furthermore, as required by French regulations, the ACNN collection was declared to the French Consultative Committee on Information Treatment of the Ministry of Research.
Light microscopic detection of Cryptosporidium
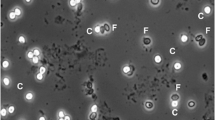
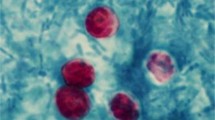
Microscopic detection of Cryptosporidium was performed after diphasic concentration for all samples, except those from the ACNN collection, for which the diagnosis of cryptosporidiosis had previously been established by microscopy in the ACNN laboratory of origin. The samples were subjected to formalin–ether concentration using a Parasep Fecal Parasite Concentrator system (DiaSys, Wokingham, UK) and were extemporaneously examined after glycerin preparation [35]. Two air-dried smears were prepared from stool concentrate, fixed in methanol, and stained by ZN (RAL, Paris, France) and AP (Sigma–Aldrich, Saint-Quentin Fallavier, France) techniques. Glycerin preparations and ZN-stained smears were observed at 400x magnification and under oil immersion at 1,000x under a light microscope respectively, whereas AP-stained smears were screened at 400x magnification and checked at 1,000x under the oil immersion lens of a fluorescent microscope.
Comparison of DNA extraction methods
Firstly, two extraction methods using the ZR Fecal DNA MiniPrep™ Kit (ZR) and the automated NucliSens® easyMAG® method (EM) were compared using aliquots from four Cryptosporidium-negative stools (S1 to S4) that were seeded with 104 C. hominis oocysts (S5 suspension) provided by P1 stool. DNA extractions were performed in triplicate from 150 mg of stool accordingly to the maximal starting amount of material compatible with ZR extraction kit. The automated EM method was adapted from the protocol previously defined by Mary et al. [32] [lower initial quantity of stool, i.e., 150 mg instead of 400 mg, and mechanical lysis using the MagNA Lyser instrument (Roche Diagnostics, Meylan, France) instead of the FastPrep-24 grinder, at 7,000 rpm for 70 sec with 425–600 μm beads (Sigma–Aldrich)], while the manual ZR extractions were carried out following strictly the steps in the manufacturer’s instructions.
Thereafter, the ZR and EM methods were further compared using positive stool samples from the ACNN collection. For each DNA extraction method, 1 ml of the stool that had been conserved in potassium dichromate was first washed with sterile water, and then centrifuged six times at 2,000 g for 10 min. DNA extraction was then performed on the pellet, as described above.
Lastly, the ZR kit was used for DNA extraction from the 69 stool specimens that had been prospectively collected between September 2012 and April 2013, and stored at −20 °C before DNA extraction.
For each DNA extract, the DNA yield (ng/μl) was determined using the Thermo ScientificTM NanoDropTM 3300 fluorospectrometer (Thermo Fischer Scientific, Inc., Illkirch, France).
Quantification of Cryptosporidium DNA using real-time qPCR
Two qPCR assays were performed on all DNA extracts. The first assay (18S qPCR), which was developed by Mary et al. [32], targets 18S rDNA, and is a pan-Cryptosporidium assay. The second assay (LAXER qPCR), which can detect C. parvum, C. hominis, and C. meleagridis, was designed to detect a 138-bp fragment positioned inside a specific 452-bp fragment sequenced by Laxer et al. [27, 36]. The qPCR technical conditions used in this study were those described by Mary et al. [32] and Fontaine & Guillot [36], except for the following points: (i) the 18S qPCR DNA template and mixture final volume were 2 μl and 20 μl respectively, instead of 1 μl and 25 μl, and (ii) the LAXER qPCR primers and probe concentrations were 0.4 μM and 0.1 μM respectively, instead of 0.3 μM and 0.2 μM, whereas the DNA template and mixture final volume were half of those described in the original publication.
18S and LAXER amplifications were performed with the LightCycler® 480 Thermal Cycling System (Roche Diagnostics) and ABI7500 Real-Time PCR System (Applied Biosystems®, Life Technologies, Saint Aubin, France) respectively. The detection limit of each qPCR method, which was determined using 10-fold serial dilutions of plasmid DNA carrying the target genes, reached 1 and 0.37 copies per microliter of DNA extract, i.e., 2.0 and 1.85 copies per reaction for the 18S and LAXER qPCRs respectively. Considering five copies of 18S [37] and one copy of LAXER [27] per sporozoite, i.e., 20 copies of 18S and four copies of LAXER per oocyst, these quantities were equivalent to 0.1 and 0.46 oocysts per reaction respectively, which corresponded to five and 23 oocysts in the total DNA extract (taking into account a 100-μl elution volume), or in the 150-mg initial amount of stool respectively. The qPCR efficiencies reached 94.5 % and 100 % respectively. In order to quantify Cryptosporidium DNA, plasmid suspensions containing 103 to 105 copies/μl or 3.68.103 to 3.68.105 copies/μl were included as standards in each 18S or LAXER qPCR series respectively. PCR inhibitor detection was performed using the TaqMan Exogenous Internal Positive Control reagent kit (Applied Biosystems). A negative control without a DNA template was included in each qPCR run. Each DNA extract was tested pure and diluted 10 times. Cryptosporidium detection was considered positive if at least one replicate was positive. The parasitic loads that are presented in tables and figures were calculated from the undiluted replicate, except when this replicate was negative. This exception only concerned one sample (see Table 1, stool P25), for which DNA quantification was obtained by multiplying by 10 the detected quantity.
In order to check the coherence between the Cryptosporidium DNA amounts detected with the 18S (5 gene copies [37]) and LAXER (single copy gene [27]) qPCRs, a 18S/LAXER ratio was calculated from Cryptosporidium DNA quantities (in copies/μl). These ratios were calculated for the 29 Cryptosporidium positive clinical specimens (P2 to P30, Table 1).
Nested-PCR detection and 18S rDNA genotyping of Cryptosporidium
Nested PCR was performed for all Cryptosporidium-positive stool specimens (ACNN collection and samples obtained from the prospective study), according to the protocol developed by Xiao et al. [23]. The isolates obtained from the prospective study were further identified by 18S rDNA sequencing using the 3500 Dx Genetic Analyser (Applied Biosystems). The sequences were compared to the GenBank database, using the BLAST program [38]. The DNA extracts from the prospective study were further tested with a typing 18S qPCR using C. parvum and C. hominis-specific probes, as described previously [32].
Statistical analysis
The mean total DNA and Cryptosporidium DNA quantities were compared by the Student's t- test, using GraphPad Prism software (version 6.01, GraphPad Software Inc., California, USA). Cryptosporidium DNA quantities in positive stool samples were compared using a Wilcoxon signed rank test. A p-value ≤ 0.05 was considered to be statistically significant.
Results
Comparison of ZR Fecal DNA MiniPrep™ Kit (ZR) and NucliSens® easyMAG® (EM) methods for molecular detection of Cryptosporidium oocysts in spiked human stools
The total DNA quantification in DNA extracts obtained from the suspension containing 104 oocysts of C. hominis (S5) and from the four spiked stool samples (S1 to S4) revealed significantly higher DNA quantities with the ZR kit for S3, S4, and S5 samples (p ≤ 0.01) (Fig. 1a). Nevertheless, the DNA yields were similar for the EM and ZR extraction methods performed on S1 and S2.
Comparison of absolute DNA and Cryptosporidium DNA yields obtained from four spiked stool samples (S1-S4) and a suspension (S5) using two extraction methods. Seeded samples included two softs (S1-S2), one liquid (S3) and one greasy (S4) stools from one child and three adults. Absolute DNA quantities (a) were measured using NanoDropTM. Cryptosporidium DNA was quantified using 18S (b) or LAXER (c) qPCR. ** p ≤ 0.01. ZR, ZR Fecal DNA MiniPrep™ Kit (Zymo Research); EM, NucliSens® easyMAG® (bioMérieux)
Detection of Cryptosporidium DNA by 18S qPCR was positive for all the samples. Except for greasy stool S4, Cryptosporidium DNA quantities detected were significantly higher with ZR than EM (Fig. 1b). Quantities were higher in the liquid stool S3 and in the oocyst suspension S5, with up to 57,800 copies/μl being detected with ZR (Fig. 1b). The weakest signals were obtained in the child soft stool S1 (where 110 copies/μl and 458 copies/μl were detected for the EM and ZR methods respectively), and in the greasy stool S4 (33 and 45 copies/μl respectively). Detection of PCR inhibitors was positive in the three DNA extracts from the paediatric soft stool S1 with the EM method. However, despite low DNA quantities were found in the greasy stool S4, no PCR inhibitors were detected in this stool. Furthermore, none of the ZR DNA extracts presented PCR inhibition.
When the number of Cryptosporidium LAXER copies was determined by qPCR, the ZR kit still yielded the best performance for stools S2, S3, and S5, for which 824, 7,600, and 5,540 copies/μl were detected with ZR and 0, 2,110, and 1,600 copies/μl with EM extracts respectively (Fig. 1c). ZR performances were also better on stool S1, for which a very weak signal was observed for one ZR replicate, whereas the three EM replicates were negative. However, the LAXER qPCR was negative in S4 ZR extracts, while Cryptosporidium DNA was detected in only one out of three EM replicates (five copies/μl) for this stool. Globally, fewer samples were positive using LAXER when compared to 18S qPCR for Cryptosporidium detection. Furthermore, around 10-fold lower DNA yields were systematically found with LAXER qPCR, and no amplification was observed in the S1 EM extracts containing PCR inhibitors (Fig. 1c).
Overall, the combination of any of the two DNA extraction methods with 18S qPCR yielded adequate detection of Cryptosporidium. Although only four seeded samples were tested, our data indicate that the performances of DNA extraction could be influenced by the origin and consistency of the stool. Since higher Cryptosporidium DNA yields and lower frequency of PCR inhibitors were obtained using the ZR kit, this method was selected for further evaluation of Cryptosporidium positive stool samples and comparison with automated extraction using EM.
Performance of ZR and EM methods for Cryptosporidium DNA extraction from positive stool samples
The total DNA quantities obtained from the 29 Cryptosporidium-positive human stool samples from the ACNN collection reached 0 to 70.7 ng/μl and 5.0 to 111.7 ng/μl with the ZR and EM methods respectively (Table 1). PCR inhibition was detected in no ZR or EM extracts. All the samples were positive with 18S qPCR for both DNA extraction methods, but Cryptosporidium DNA amounts were significantly higher in ZR than EM extracts (p = 0.002). Nested PCR was also positive for all ZR extracts, but surprisingly, four EM extracts from P2, P4, P15, and P27 stools, corresponding to two C. hominis, one C. parvum, and one C. felis isolates respectively, were negative. The Cryptosporidium LAXER qPCR was positive for all C. hominis and C. parvum samples using the EM method, but ZR extraction yielded two negative C. hominis samples (P13, P14), which corresponded to samples with very low DNA amounts using 18S qPCR (42 and 45 copies/μl respectively). Despite these two false negative samples with ZR, when DNA quantities were compared between ZR and EM extracts in the 23 samples that were positive with LAXER qPCRs, the Cryptosporidium DNA amounts that were detected in ZR extracts were found to be significantly higher than the ones detected in EM extracts (p = 0.004). The LAXER qPCR was negative for the four C. felis isolates (P27 to P30) with both the EM and ZR extracts. Moreover, when both LAXER and 18S qPCRs were positive, the latter yielded higher Cryptosporidium DNA amounts (Table 1), with an \( \frac{18S}{LAXER} \) median value reaching 5.8 and 6.6 for the ZR and EM kits respectively.
Prospective comparison of microscopy and molecular tools for the detection of Cryptosporidium in patients at risk
The stool samples collected for parasitological examination between September 2012 and April 2013 included 58 specimens from 47 patients hospitalized at Lille University Hospital, and 11 specimens from nine patients at other hospitals. The M:F gender ratio was 1.15, and underlying diseases included hematological malignancy (three stem-cell transplants, two myelodysplastic syndromes, one acute lymphoid leukemia, one acute myeloid leukemia, one multiple myeloma), kidney transplantation (n = 6), human immunodeficiency virus infection (n = 2), primary immunodeficiency (hyper-IgM syndrome, n = 1), auto-immune hepatitis (n = 1), and pancreatic adenocarcinoma (n = 1). Thirty-three patients (15 children and 18 adults) were immunocompetent. The immune status of four persons was unknown.
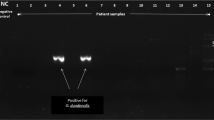
Cryptosporidiosis was diagnosed by microscopic detection in 12, 13, and 14 samples from 7/56 (12.5 %) patients after glycerin, ZN, and AP staining respectively. Among patients with cryptosporidiosis, two were hospitalized at Lille University Hospital and five at other hospitals. Therefore, the incidence of cryptosporidiosis in patients at risk reached 4.3 % (2/47) at Lille University Hospital. Taking into account all Lille University Hospital patients who underwent parasitological stool examinations between September 2012 and April 2013, the incidence of cryptosporidiosis was 0.3 % (2/651). The patients included three immunocompromised (two kidney transplants and one primary immunodeficiency) and four immunocompetent patients (Table 2). The molecular identification yielded five C. parvum and two C. hominis isolates, with full agreement between the DNA sequencing and typing qPCR.
In comparing the microscopic and molecular methods, 48 samples were negative with all techniques. All samples from patients with cryptosporidiosis were positive with both AP staining and 18S qPCR (n = 14, Table 3). For the latter, six supplementary samples were later positive (Ct > 40). The LAXER qPCR detected only 12 of the 14 microscopically positive samples. One supplementary sample was positive for a patient previously diagnosed with cryptosporidiosis (patient no. 3, Table 2).
Contribution of qPCR to the follow-up of cryptosporidiosis
When evaluating qPCR for the follow-up of cryptosporidiosis in two patients with biliary dissemination (patient no. 1, Table 2) or acute gastroenteritis (patient no. 2, Table 2), both the 18S and LAXER Cryptosporidium DNA loads were found to be correlated to disease severity. Firstly, for patient no. 1, an 18-year-old male with hyper-IgM syndrome suffering from sclerosing cholangitis, a dramatic increase in parasitic DNA loads was detected simultaneously with bioclinical worsening, revealed by strong elevation of hepatic enzymes combined with reinforcement of abdominal pains, despite treatment with nitazoxanide (Fig. 2a). Identically, for patient no. 2, a 12-year-old kidney transplant child with choleriform diarrhea, the Cryptosporidium DNA quantities rose simultaneously with both hepatic and pancreatic involvement despite nitazoxanide administration. Clinical improvement and a gradual decrease in the parasite DNA loads (until negativation by LAXER qPCR) were observed after treatment with intravenous immunoglobulins (IVIg), which was prescribed to treat a hypogammaglobulinemia at 3.3 g/l (Fig. 2b).
Follow-up of Cryptosporidium oocysts and DNA loads in two patients with chronic sclerosing cholangitis (a) and abrupt watery diarrhea secondarily complicated by acute pancreatitis (b). Considering the date of the first stool diagnosing cryptosporidiosis as day 0 (D0), nitazoxanide treatment (arrowhead) was initiated at D45 and D2 for patients nos. 1 and 2 respectively, while IVIg injections (arrow) were used only for patient no. 2 (D25 and D26). ALP, alkaline phosphatase (n = 35-–30 IU/l); GGT, γ-glutamyl-transpeptidase (n = 5–45 IU/l); LIP, lipase (n = 13–60 IU/l). Cryptosporidium oocysts semi-quantification using microscopy (glycerin staining): (+++), abundant (>1 oocysts/field at 400× magnification); (++), moderately abundant (>10 oocysts/slide); (+), scarce (1–5 oocysts/slide); (−), absent
Discussion
When evaluating microscopy and molecular tools for the diagnosis of cryptosporidiosis, our prospective study confirmed the higher sensitivity of AP [16, 17]. Although two samples that were positive with AP were not detected with glycerin (yielding a lower sensitivity of 85.7 %), glycerin was positive in all the fecal specimens at the time of diagnosis, and only became negative in the course of cryptosporidiosis. Like the negative staining technique of Heine [39], glycerin is poorly known by medical biologists, and few laboratories use it for the routine diagnosis of cryptosporidiosis. However, it is a fast, reliable, and cost-effective method compared to ZN or AP. Thus, it could be widely used in patients with diarrhea in non-specialized laboratories. The high sensitivity of 18S qPCR confirmed the usefulness of this method for cryptosporidiosis diagnosis. As reported in previous publications revealing a significant increase in cryptosporidiosis cases (up to 2.5-fold) detected with 18S nested PCR in HIV-positive patients, or children with diarrhea [17, 20, 40], 18S qPCR positivity in six supplementary samples could correspond to those that were misdiagnosed using microscopic methods. By contrast, Calderaro et al. showed that real-time PCR, ICA, and fluorescent microscopy offered similar performance despite a large cohort (1,040 stools from 533 patients with a suspected intestinal parasitosis) [21]. However, this study was conducted in a low-prevalence area where an elevated proportion of light infection is expected, and the sensitivity of the molecular method that was used (multiplex qPCR with Cryptosporidium hominis/parvum detection using a LAXER targeting assay) could have been too low to increase the diagnosis of cryptosporidiosis in such a setting. In all, this study suggests that repeated sequences such as 18S rDNA could be a more appropriate target, but further large prospective studies are needed to confirm our data.
Our evaluation of the ZR Fecal DNA MiniPrep™ Kit and NucliSens® easyMAG® performed on spiked stools confirmed the usefulness of the EM automated method for Cryptosporidium DNA extraction [30, 32]. Furthermore, even if high performance has been reported for Blastocystis DNA extraction from stools with the ZR kit (94 % vs 48 % sensitivity for QIAamp® DNA Stool Mini Kit (QIA) in [41]), our data provides the first evidence of the efficiency of this method for Cryptosporidium DNA extraction from human stools. The significantly higher Cryptosporidium DNA quantities we obtained using ZR in most spiked stools and in stools from the ACNN collection could result from the mechanical pre-treatment step (bead-beating) which is systematically used prior to DNA isolation in this kit, and allow both optimal stool homogeneisation and oocyst disruption, whereas it was specifically added for EM extraction in our study, accordingly to Mary et al. who reported significantly enhanced DNA yields when adding bead-beating (by 2.17-fold, p < 0.0001) [32]. Bead-beating was also found by Halstead et al. [31] to improve yields for parasitic targets. Thermal treatment has also been reported to increase the sensitivity of Cryptosporidium detection from 60 % to 100 % in stools positive by microscopy after extraction with QIA [30]. These results underline the importance of mechanical or thermal pre-treatment for lysis of Cryptosporidium oocysts and optimal DNA extraction. If not included in the DNA extraction method provided by the supplier (such as for EM), it should be included as a supplementary step. Moreover, our results on S1 stool (Cryptosporidium-negative, seeded with oocysts) confirm that an efficient removal of PCR inhibitors, which was optimal with ZR kit, is essential. Although the presence of inhibitors in all S1 EM replicates did not completely prevent DNA amplification by 18S qPCR in this stool, it probably led to an underestimation of the quantities that were detected. Furthermore, the greater impact of PCR inhibitors on LAXER qPCR (which was negative for all EM DNA extracts) is consistent with its lower sensitivity (5-fold fewer target genes per sporozoite than 18S). It could also result from differences in inhibition strength according to the targeted locus.
Although Cryptosporidium DNA quantities were higher with ZR, both the EM and ZR kits allowed the efficient detection (100 %) of the parasite with 18S qPCR in Cryptosporidium-positive stool samples from the ACNN collection. 18S nested PCR was also 100 % sensitive with the ZR kit, but was not able to detect the parasite in four of the 29 EM extracts in our study. Such false-negative results have been reported previously by several authors [32, 42]. In our study, since nested PCR targets long DNA fragments (820 bp), the false negatives we observed could be related to DNA fragmentation resulting from mechanical pretreatment, which could have a lower impact on 18S qPCR since this assay targets a smaller DNA fragment (178 bp). As expected, the LAXER qPCR yielded lower sensitivity. These results were consistent with the intrinsic performance of this assay, which is specific for the detection of C. parvum and C. hominis, and does not cross-react with C. felis [36]. Additionally, two ZR DNA extracts from C. hominis isolates, which were found to be very weakly positive with 18S qPCR, were negative with the LAXER qPCR (Table 3). Once again, these data are consistent with the higher sensitivity of 18S qPCR, resulting from the higher number of 18S copies than LAXER copies (five vs one) in the sporozoite genome [27, 37]. Our results are in line with this statement, since the proportion of these two targets were respected, with an \( \frac{18S}{LAXER} \) median value reaching 5.8 and 6.6 for the ZR and EM extraction performed on Cryptosporidium-positive samples.
Beyond the differences in performance for DNA recovery and co-purification of PCR inhibitors, the choice of a DNA extraction method depends on several features including processing/hands-on time, cost, and the variety of validated sample types (Table 4). The NucliSens® easyMAG® is an IVD-labeled system allowing DNA extraction from a wide panel of specimens and volumes, well adapted to high-functioning laboratories. Moreover, automation enables up to 24 extractions simultaneously, releasing non-negligible hands-on time. The system also incorporates sample and reagent traceability, improving the safety of the process analysis. However, the main disadvantage of the EM platform remains the price of the robot, inadequate for small laboratories. The overall cost of the manual ZR Fecal DNA MiniPrep™ Kit is lower and provides high quality/quantity DNA from stools, particularly since bead-beating is processed prior to DNA extraction and inhibitors are efficiently removed. However, it requires a higher hands-on time and achieves lower standardization.
Our prospective study revealed a 4.3 % incidence of cryptosporidiosis in patients at risk who were hospitalized at Lille University Hospital between September 2012 and April 2013, and a 0.3 % incidence among patients who underwent parasitological stool examination in Lille during this period. Taking into account all the patients with cryptosporidiosis who were diagnosed at Lille University Hospital (including the five patients staying at other hospitals), the incidence reached 1.1 % and was similar to the 1.1-2.3 % national prevalence estimated by the ACNN [13]. But our data shows an unexpected excess of Cryptosporidium cases in the North Region in this period (six, versus two or one cases reported in 2010 and 2011 respectively). Such an observation was also reported at the national level. Indeed, a 2-fold increase in notification of cryptosporidiosis cases to the ACNN was observed between September and December 2012, with 55 cases versus 29 and 22 cases notified in the same period in 2010 and 2011 respectively. Interestingly, this situation emerged simultaneously with a dramatic (1.8- to 4.9-fold) increase in cryptosporidiosis cases in three European countries, i.e., the Netherlands, the United Kingdom and Germany [43]. Increased reporting in 2012 compared with 2011 was also observed in Spain (268 %), Finland (127 %), Belgium (103 %), Ireland (35 %) and Slovenia (20 %) [44]. These observations support the value of systematic Cryptosporidium screening for patients presenting gastrointestinal symptoms, which is still insufficiently implemented in French laboratories, and leads to an underestimation of the cryptosporidiosis burden [13].
Although real-time PCR was found to be a reliable tool for the quantification of microsporidia and the monitoring of treatment efficacy [45], reports on Cryptosporidium infection follow-up with qPCR were lacking. In the present study, the Cryptosporidium DNA loads were found to be correlated with the disease severity in two cases of persistent cryptosporidiosis. The increase in DNA loads despite treatment with nitazoxanide in the two patients was consistent with the insufficient anti-Cryptosporidium activity of this drug among severely immunosuppressed patients [9]. For the kidney transplant child, the dramatic decrease of DNA loads after initiation of immunotherapy suggested that IVIg, which has already been reported for the treatment of several bacterial or viral infections [46], could also be useful in treating cryptosporidiosis in immunocompromised patients. These new data open the way for larger studies to evaluate the efficacy of IVIg for the treatment of cryptosporidiosis.
Altogether, these results indicate that, although little known, glycerin is a sensitive method for Cryptosporidium detection. Furthermore, the combination of the ZR Fecal DNA Kit and 18S qPCR was found to be an adequate procedure for Cryptosporidium DNA detection in human stools. qPCR was more sensitive than microscopy in patients at risk, and was also more sensitive and accurate in estimating the oocyst shedding in the course of infection. Furthermore, the 18S qPCR typing method was an accurate tool in providing rapid identification of the two most prevalent Cryptosporidium species. Using this procedure in patients at risk (i.e., children and immunocompromised individuals) could be helpful in preventing the misdiagnosis of cryptosporidiosis and quickly identifying the sources of infection in the case of outbreaks.
References
Ryan U, Paparini A, Tong K, Yang R, Gibson-Kueh S, O’Hara A et al (2015) Cryptosporidium huwi n. sp. (Apicomplexa: Eimeriidae) from the guppy (Poecilia reticulata). Exp Parasitol 150:31–35. doi:10.1016/j.exppara.2015.01.009
Ryan U, Fayer R, Xiao L (2014) Cryptosporidium species in humans and animals: current understanding and research needs. Parasitology 141:1667–1685. doi:10.1017/S0031182014001085
Šlapeta J (2013) Cryptosporidiosis and Cryptosporidium species in animals and humans: a thirty-colour rainbow? Int J Parasitol 43:957–970. doi:10.1016/j.ijpara.2013.07.005
Chalmers RM, Katzer F (2013) Looking for Cryptosporidium: the application of advances in detection and diagnosis. Trends Parasitol 29:237–251. doi:10.1016/j.pt.2013.03.001
Cacciò SM, Thompson RCA, McLauchlin J, Smith HV (2005) Unravelling Cryptosporidium and Giardia epidemiology. Trends Parasitol 21:430–437. doi:10.1016/j.pt.2005.06.013
Bouzid M, Hunter PR, Chalmers RM, Tyler KM (2013) Cryptosporidium pathogenicity and virulence. Clin Microbiol Rev 26:115–134. doi:10.1128/CMR.00076-12
Chalmers RM, Davies AP (2010) Minireview: clinical cryptosporidiosis. Exp Parasitol 124:138–146. doi:10.1016/j.exppara.2009.02.003
Hunter PR, Nichols G (2002) Epidemiology and clinical features of Cryptosporidium infection in immunocompromised patients. Clin Microbiol Rev 15:145–154
Checkley W, White AC, Jaganath D, Arrowood MJ, Chalmers RM, Chen X-M et al (2015) A review of the global burden, novel diagnostics, therapeutics, and vaccine targets for Cryptosporidium. Lancet Infect Dis 15:85–94. doi:10.1016/S1473-3099(14)70772-8
Baldursson S, Karanis P (2011) Waterborne transmission of protozoan parasites: review of worldwide outbreaks — an update 2004–2010. Water Res 45:6603–6614. doi:10.1016/j.watres.2011.10.013
Robertson LJ, Chalmers RM (2013) Foodborne cryptosporidiosis: is there really more in Nordic countries? Trends Parasitol 29:3–9. doi:10.1016/j.pt.2012.10.003
Kotloff KL, Nataro JP, Blackwelder WC, Nasrin D, Farag TH, Panchalingam S et al (2013) Burden and aetiology of diarrhoeal disease in infants and young children in developing countries (the Global Enteric Multicenter Study, GEMS): a prospective, case–control study. Lancet 382:209–222. doi:10.1016/S0140-6736(13)60844-2
ANOFEL Cryptosporidium National Network (2010) Laboratory-based surveillance for Cryptosporidium in France, 2006–2009. Euro Surveill 15:19642
Henriksen SA, Pohlenz JF (1981) Staining of cryptosporidia by a modified Ziehl–Neelsen technique. Acta Vet Scand 22:594–596
Cacciò SM, Pozio E (2006) Advances in the epidemiology, diagnosis and treatment of cryptosporidiosis. Expert Rev Anti Infect Ther 4:429–443. doi:10.1586/14787210.4.3.429
Chalmers RM, Campbell BM, Crouch N, Charlett A, Davies AP (2011) Comparison of diagnostic sensitivity and specificity of seven Cryptosporidium assays used in the UK. J Med Microbiol 60:1598–1604. doi:10.1099/jmm.0.034181-0
Khurana S, Sharma P, Sharma A, Malla N (2012) Evaluation of Ziehl–Neelsen staining, auramine phenol staining, antigen detection enzyme linked immunosorbent assay and polymerase chain reaction, for the diagnosis of intestinal cryptosporidiosis. Trop Parasitol 2:20–23. doi:10.4103/2229-5070.97234
Jex AR, Smith HV, Monis PT, Campbell BE, Gasser RB (2008) Cryptosporidium—biotechnological advances in the detection, diagnosis and analysis of genetic variation. Biotechnol Adv 26:304–317. doi:10.1016/j.biotechadv.2008.02.003
Agnamey P, Sarfati C, Pinel C, Rabodoniriina M, Kapel N, Dutoit E et al (2011) Evaluation of four commercial rapid immunochromatographic assays for detection of Cryptosporidium antigens in stool samples: a blind multicenter trial. J Clin Microbiol 49:1605–1607. doi:10.1128/JCM.02074-10
Martín-Ampudia M, Mariscal A, Lopez-Gigosos RM, Mora L, Fernandez-Crehuet J (2012) Under-notification of cryptosporidiosis by routine clinical and laboratory practices among non-hospitalised children with acute diarrhoea in Southern Spain. Infection 40:113–119. doi:10.1007/s15010-011-0188-3
Calderaro A, Montecchini S, Gorrini C, Dettori G, Chezzi C (2011) Similar diagnostic performances of antigen detection and nucleic acid detection of Cryptosporidium spp. in a low-prevalence setting. Diagn Microbiol Infect Dis 70:72–77. doi:10.1016/j.diagmicrobio.2010.11.017
van Lieshout L, Roestenberg M (2015) Clinical consequences of new diagnostic tools for intestinal parasites. Clin Microbiol Infect 21:520–528. doi:10.1016/j.cmi.2015.03.015
Xiao L, Escalante L, Yang C, Sulaiman I, Escalante AA, Montali RJ et al (1999) Phylogenetic analysis of Cryptosporidium parasites based on the small-subunit rRNA gene locus. Appl Environ Microbiol 65:1578–1583
Hawash Y, Dorgham LS, Al-Hazmi AS, Al-Ghamdi MS (2014) Prevalence of Cryptosporidium-associated diarrhea in a high-altitude community of Saudi Arabia detected by conventional and molecular methods. Korean J Parasitol 52:479–485. doi:10.3347/kjp.2014.52.5.479
Abe N, Matsubayashi M, Kimata I, Iseki M (2006) Subgenotype analysis of Cryptosporidium parvum isolates from humans and animals in Japan using the 60-kDa glycoprotein gene sequences. Parasitol Res 99:303–305. doi:10.1007/s00436-006-0140-0
Homem CG, Nakamura AA, Silva DC, Teixeira WFP, Coelho WMD, Meireles MV (2012) Real-time PCR assay targeting the actin gene for the detection of Cryptosporidium parvum in calf fecal samples. Parasitol Res 110:1741–1745. doi:10.1007/s00436-011-2694-8
Laxer MA, Timblin BK, Patel RJ (1991) DNA sequences for the specific detection of Cryptosporidium parvum by the polymerase chain reaction. Am J Trop Med Hyg 45:688–694
Tanriverdi S, Tanyeli A, Başlamişli F, Köksal F, Kilinç Y, Feng X et al (2002) Detection and genotyping of oocysts of Cryptosporidium parvum by real-time PCR and melting curve analysis. J Clin Microbiol 40:3237–3244
Sulaiman IM, Morgan UM, Thompson RC, Lal AA, Xiao L (2000) Phylogenetic relationships of Cryptosporidium parasites based on the 70-kilodalton heat shock protein (HSP70) gene. Appl Environ Microbiol 66:2385–2391
Hawash Y (2014) DNA extraction from protozoan oocysts/cysts in feces for diagnostic PCR. Korean J Parasitol 52:263–271. doi:10.3347/kjp.2014.52.3.263
Halstead FD, Lee AV, Couto-Parada X, Polley SD, Ling C, Jenkins C et al (2013) Universal extraction method for gastrointestinal pathogens. J Med Microbiol 62:1535–1539. doi:10.1099/jmm.0.058743-0
Mary C, Chapey E, Dutoit E, Guyot K, Hasseine L, Jeddi F et al (2013) Multicentric evaluation of a new real-time PCR assay for quantification of Cryptosporidium spp. and identification of Cryptosporidium parvum and Cryptosporidium hominis. J Clin Microbiol 51:2556–2563. doi:10.1128/JCM.03458-12
Elwin K, Robinson G, Hadfield SJ, Fairclough HV, Iturriza-Gómara M, Chalmers RM (2012) A comparison of two approaches to extracting Cryptosporidium DNA from human stools as measured by a real-time PCR assay. J Microbiol Methods 89:38–40. doi:10.1016/j.mimet.2012.02.006
Masny A, Rozej W, Gołab E (2009) Development of efficient DNA isolation procedures for Cryptosporidium and Trichinella PCR detection in fecal samples. Med Dosw Mikrobiol 61:259–265
Dutoit E, Dewitte J-M, Dei-Cas E, Camus D (1988) Rapid method for detection of Cryptosporidium oocysts in stools. Le Biologiste XXII:159–160
Fontaine M, Guillot E (2002) Development of a TaqMan quantitative PCR assay specific for Cryptosporidium parvum. FEMS Microbiol Lett 214:13–17
Le Blancq SM, Khramtsov NV, Zamani F, Upton SJ, Wu TW (1997) Ribosomal RNA gene organization in Cryptosporidium parvum. Mol Biochem Parasitol 90:463–478
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410. doi:10.1016/S0022-2836(05)80360-2
Potters I, Van Esbroeck M (2010) Negative staining technique of Heine for the detection of Cryptosporidium spp.: a fast and simple screening technique. Open Parasitol J 4:1–4
Kaushik K, Khurana S, Wanchu A, Malla N (2008) Evaluation of staining techniques, antigen detection and nested PCR for the diagnosis of cryptosporidiosis in HIV seropositive and seronegative patients. Acta Trop 107:1–7. doi:10.1016/j.actatropica.2008.02.007
Yoshikawa H, Dogruman-Al F, Dogruman-Ai F, Turk S, Kustimur S, Balaban N et al (2011) Evaluation of DNA extraction kits for molecular diagnosis of human Blastocystis subtypes from fecal samples. Parasitol Res 109:1045–1050. doi:10.1007/s00436-011-2342-3
Jothikumar N, da Silva AJ, Moura I, Qvarnstrom Y, Hill VR (2008) Detection and differentiation of Cryptosporidium hominis and Cryptosporidium parvum by dual TaqMan assays. J Med Microbiol 57:1099–1105. doi:10.1099/jmm.0.2008/001461-0
Fournet N, Deege MP, Urbanus AT, Nichols G, Rosner BM, Chalmers RM et al (2012) Simultaneous increase of Cryptosporidium infections in the Netherlands, the United Kingdom and Germany in late summer season, 2012. Euro Surveill 18:1–5
European Center for Disease Prevention and Control (ECDC) (2014) Annual epidemiological report 2014. Food- and waterborne diseases and zoonoses
Menotti J, Cassinat B, Porcher R, Sarfati C, Derouin F, Molina J-M (2003) Development of a real-time polymerase-chain-reaction assay for quantitative detection of Enterocytozoon bieneusi DNA in stool specimens from immunocompromised patients with intestinal microsporidiosis. J Infect Dis 187:1469–1474. doi:10.1086/374620
Mouthon L, Lortholary O (2003) Intravenous immunoglobulins in infectious diseases: where do we stand? Clin Microbiol Infect 9:333–338. doi:10.1046/j.1469-0691.2003.00694.x
Acknowledgments
We wish to thank Michèle Wauquier and Filoména Naji (Parasitology–Mycology Laboratory of Lille University Hospital Center) for their technical assistance, and the members of the French ANOFEL Cryptosporidium National Network: Ahmed Abou Bacar CHU Strasbourg, Isabelle Accoceberry CHU Bordeaux, Patrice Agnamey CHU Amiens, Adela Angoulvant CHU Bicêtre Paris, Dominique Aubert CHU Amiens, Belkhadi Ghania CHU St Antoine, Paris, Antoine Berry CHU Toulouse, Denis Blanchet CHU Cayenne, Julie Bonhomme CHU Caen, Françoise Botterel CHU Créteil, Marie-Elisabeth Bougnoux CHU Necker, Paris, Pierre Buffet CHU Pitié, Paris, Frédéric Dalle CHU Dijon, Eric Dannaoui HEGP, Paris, Marie-Laure Dardé CHU Limoges, Ludovic De Gentile CHU Angers, Anne Debourgogne CHU Nancy, Monique Debruyne (Cerba, Paris), Brigitte Degeilh CHU Rennes, Magalie Demar CHU Cayenne, Nicole Desbois CHU Fort de France, Guillaume Desoubeaux CHU Tours, Pascal Delaunay CHU Nice, Pierre Flori CHU St Etienne, Gilles Gargala CHU Rouen, Agathe Goubard Biomnis Paris, Frédéric Grenouillet CHU Besançon, Samia Hamane CHU St Louis, Paris, Sandrine Houzé CHU Bichat, Paris, Jamet Deborah CHU Brest, Nathalie Kapel CHU Pitié, Paris, Franck Labbe CH Le Havre, Denis Leméteil Lab. St Valéry en Caux, Denis Magne CHU St Antoine Paris, Pierre Marty CHU Nice, Jean Menotti CHU St Louis, Paris, Laurence Millon CHU Besançon, Christelle Morelle CHU Montpellier, Florent Morio CHU Nantes, Jean-Benjamin Murat CHU Grenoble, Gilles Nevez CHU Brest, Muriel Nicolas CHU Guadeloupe, Philippe Poirier CHU Clermont-Ferrand, Méja Rabodonirina CHU Lyon, Marie-Hélène Rodier CHU Poitiers, Marc Sautour CHU Dijon, Marc Thellier CHU Pitié, Paris, Anne Totet CHU Amiens, Alexis Valentin CHU Toulouse, Isabelle Villena CHU Reims, Hélène Yera CHU Cochin, Paris.
This work was supported by grants from the Lille University Hospital Center, from the University of Lille, Pasteur Institute of Lille, “Centre National de la Recherche Scientifique” (CNRS), and “Institut National de la Santé et de la Recherche Médicale” (Inserm). It was presented at the First French North African Parasitology and Mycology meeting in Rabat, Morocco, in October 2013, and at the European Congress of Clinical Microbiology and Infectious Diseases (ECCMID) in Barcelona, Spain, in May 2014.
Author information
Authors and Affiliations
Consortia
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Eduardo Dei-Cas is deceased.
E. Fréalle and E. Dutoit contributed equally to this work.
Rights and permissions
About this article
Cite this article
Le Govic, Y., Guyot, K., Certad, G. et al. Assessment of microscopic and molecular tools for the diagnosis and follow-up of cryptosporidiosis in patients at risk. Eur J Clin Microbiol Infect Dis 35, 137–148 (2016). https://doi.org/10.1007/s10096-015-2519-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10096-015-2519-2