Abstract
DNM1 developmental and epileptic encephalopathy (DEE) is characterized by severe to profound intellectual disability, hypotonia, movement disorder, and refractory epilepsy, typically presenting with infantile spasms. Most of the affected individuals had de novo missense variants in DNM1. DNM1 undergoes alternative splicing that results in expression of six different transcript variants. One alternatively spliced region affects the tandemly arranged exons 10a and 10b, producing isoforms DNM1A and DNM1B, respectively. Pathogenic variants in the DNM1 coding region affect all transcript variants. Recently, a de novo DNM1 NM_001288739.1:c.1197-8G > A variant located in intron 9 has been reported in several unrelated individuals with DEE that causes in-frame insertion of two amino acids and leads to disease through a dominant-negative mechanism. We report on a patient with DEE and a de novo DNM1 variant NM_001288739.2:c.1197-46C > G in intron 9, upstream of exon 10a. By RT-PCR and Sanger sequencing using fibroblast-derived cDNA of the patient, we identified aberrantly spliced DNM1 mRNAs with exon 9 spliced to the last 45 nucleotides of intron 9 followed by exon 10a (NM_001288739.2:r.1196_1197ins[1197-1_1197-45]). The encoded DNM1A mutant is predicted to contain 15 novel amino acids between Ile398 and Arg399 [NP_001275668.1:p.(Ile398_Arg399ins15)] and likely functions in a dominant-negative manner, similar to other DNM1 mutants. Our data confirm the importance of the DNM1 isoform A for normal human brain function that is underscored by previously reported predominant expression of DMN1A transcripts in pediatric brain, functional differences of the mouse Dnm1a and Dnm1b isoforms, and the Dnm1 fitful mouse, an epilepsy mouse model.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
DNM1 encodes dynamin 1, one of several members of the dynamin-like family. Dynamin 1 is a neuron-specific GTPase implicated in endocytosis and mediates uptake of synaptic vesicles in presynaptic terminals [1,2,3]. Dynamins are well known for their self-oligomerization into contractile helical polymers that surround a membrane tube, nucleotide-driven conformational changes leading to constriction of the polymer and the membrane, and finally GTP hydrolysis-dependent fission of membrane tubes. Dnm1 shows a predominant and developmentally regulated expression in the brain [4, 5]. In mammals, Dnm1 undergoes extensive alternative splicing resulting in the expression of multiple transcript variants (Fig. 1a). The first alternatively spliced region of Dnm1 affects the two tandemly arranged exons 10a and 10b encoding 46 amino acid residues of the middle region that vary in 11 residues in human DNM1. The expression of these mutually exclusive mRNAs results in the production of isoforms Dnm1a and Dnm1b. The second alternative splicing region affects the 3′ region of the gene. Alternative splicing in this region leads to the generation of different C-terminal ends in dynamin 1 isoforms [6,7,8,9]. Dynamin 1 is composed of the N-terminal GTPase domain, the middle domain or stalk region, a bundle signaling element, a phosphoinositide-4,5-bisphosphate-binding pleckstrin homology (PH) domain, and a proline-rich domain (Fig. 1b) [10, 11].
DNM1 transcript variants, domain structure, and pathogenic variants. a Schematic representation of the exon–intron structure of exons 7–14 in the six alternatively spliced DNM1 transcript variants encoding the DNM1 central region present in NCBI and Ensemble databases (last accessed 12/2022). Exons are given by boxes and are numbered. Location of the intronic variants NM_001288739.2:c.1197-46C > G and NM_001288739.2:c.1197-8G > A identified in this study and in several independent patients are marked by red and blue lines, respectively. Relative DNM1 expression in cortex and cerebellum according to the GTEx portal (last accessed 12/2022) is shown next to the transcript variants on the right. Length of the bars represents the rate of expression (violet, strong expression; grey, no expression). b Schematic representation of the DNM1 domain structure (NP_004399.2 and NP_001275668.1). Amino acid numbering is given. The polypeptide within the central region encoded by the alternatively spliced exon 10a or 10b is highlighted in dark blue. DMN1 de novo pathogenic variants associated with developmental and epileptic encephalopathy according to HGMD professional database (last accessed 12/2022) are given below the domain structure (black). De novo variants associated with a milder neurodevelopmental disorder are given in dark grey. Recurrent variants are indicated by the number of identified unrelated patients in brackets. If a variant only affects one DNM1 isoform, this is indicated by the suffix [a] and [b] for DNM1 isoforms NP_001275668.1 and NP_004399.2, respectively. Homozygous loss-of-function variants in patients with severe neurodevelopmental delay and early-onset epilepsy are underlined. C, C-terminus; GED, GTPase effector domain; N, N-terminus, PH, pleckstrin homology; PRD, prolin-rich domain
In 2014, de novo missense variants in DNM1 were reported in five individuals who had infantile spasms in the first year of life and severe to profound intellectual disability [12]. Up to date, at least 60 individuals with DNM1 encephalopathy were described. Most of them carry de novo missense variants or in-frame insertions affecting the GTPase or middle domain, including a few recurrent variants that alter codons 43, 45, 65, 177, 206, 237, and 359. These variants affect all transcript variants of DNM1 (Fig. 1). The majority of affected individuals had a relatively homogeneous phenotype that can be subsumed under developmental and epileptic encephalopathies (DEE) (Table 1) [12,13,14,15,16,17,18,19,20,21,22,23,24,25,26,27,28,29,30,31,32,33,34,35,36,37,38,39,40,41,42,43,44,45]. The phenotypic spectrum has recently been broadened by reports of a few patients with mild-to-moderate developmental delay and/or intellectual disability with self-limited or no seizures. Affected individuals carried missense variants located in the PH domain of dynamin 1, while a 4-year-old girl had a de novo DNM1 variant c.139G > A/p.(Val47Met) in the GTPase domain, previously reported in more severely affected individuals, and a 5-year-old girl had a de novo c.1214C > T/p.(Pro405Leu) variant, exclusively affecting exon 10b (GenBank: NM_004408.4) (Fig. 1b; Table 1) [44, 46,47,48]. Interestingly, several individuals with severe DEE carried the recurrent de novo DNM1 non-coding variant, NM_001288739.1:c.1197-8G > A (Table 1) [44, 49, 50]. This transition is located in intron 9, just 8 bp upstream of the alternatively spliced exon 10a, and affects only three out of six DNM1 transcript variants (GenBank: NM_001288737.2, NM_001288738.2, and NM_001288739.2; Fig. 1a). A splicing minigene assay revealed insertion of 6 bp in the DNM1A mRNA that is predicted to cause an insertion of two amino acids in the dynamin-1 middle domain (GenBank: NM_001288739.2:r.1196_1197[1197-1_1197-6]; p.(Arg399_Thr400insCysArg)) [44].
All disease-associated missense and in-frame insertion variants in DNM1 are supposed to exert a dominant-negative effect of the DNM1 mutant on wild-type DNM1 by (i) defective GTP binding, (ii) impaired GTP hydrolysis, (iii) interfering with self-assembly, or (iv) the inability to interact with phosphoinositide-4,5-bisphosphate, collectively causing impaired synaptic vesicle endocytosis [11, 17, 31, 44, 51,52,53,54,55,56,57]. Altogether, DNM1 encephalopathy associated with specific autosomal dominant variants belongs to the group of synaptic vesicle cycling disorders in which synaptic transmission and plasticity are impaired [13, 17].
Here, we report a patient with developmental and epileptic encephalopathy who carried a de novo deep intronic DMN1 variant NM_001288739.2:c.1197-46C > G predicted to create a new splice acceptor site. DNM1 transcript analysis revealed aberrantly spliced DNM1 mRNAs with an inclusion of 45 nucleotides between exons 9 and 10a in the patient. The encoded DNM1 isoform A mutant is predicted to contain 15 novel amino acids between Ile398 and Arg399 [NP_001275668.1:p.(Ile398_Arg399ins15)], while DNM1 isoform B is likely left intact. Our data and previously published data confirm the importance of the DNM1A isoform for normal brain function.
Material and methods
Editorial policies and ethical considerations
The parents of the proband provided written informed consent for participation in the study, clinical data and specimen collection, genetic analysis, and publication of relevant findings under a protocol approved by the Ethics Committee of the Hamburg Medical Chamber (PV7038-4438-BO-ff).
Exome sequencing and variant filtering
Trio exome sequencing was performed on genomic DNA extracted from leukocytes of the patient and his healthy parents by CeGaT. Enrichment was carried out using the SureSelect Human All Exon V6 kit (Agilent Technologies). Each captured library was then loaded and sequenced on the HiSeq platform (Illumina, San Diego, CA). fastp (v0.21.0) [58] was used to remove artificial and low-quality (Phred quality score below 15) sequences from the 3′ end of sequence reads. Putative base calling errors located in regions where two reads of a read pair overlap were corrected (fastp option: “–correction”). The sequences were then aligned to the human reference assembly (NCBI GRCh38 (GCA_000001405.15)) with the Burrows-Wheeler Aligner (BWA mem, v0.7.17-r1188) [59]. Strelka2 (v.2.9.10) [60] and GATK4 (v.4.1.9.0) [61] were used to detect genetic variation. Variants were annotated using the Ensembl Variant Effect Predictor (v.103.0) [62]. Only exonic and intronic variants that were de novo (absent from public databases) or rare (with a minor allele frequency [MAF] < 0.5% and no homo- and hemizygotes in public databases) were retained. Variants with poor depth of sequencing coverage (total read depth < 10) and in low-quality regions (checked in IGV) were discarded.
Variant validation
Sequence validation was performed by Sanger sequencing. Primers designed to amplify the selected region of DNM1 intron 9 (NM_001288739.2) are described in Supplementary Table 1. Amplicons were directly sequenced using the ABI BigDye Terminator Sequencing Kit (Applied Biosystems) and an automated capillary sequencer (ABI 3500, Applied Biosystems). Sequence electropherograms were analyzed using the Sequence Pilot software (JSI Medical Systems).
The DNM1 variant NM_001288739.2:c.1197-46C > G was submitted to the LOVD database (https://databases.lovd.nl/shared/genes/DNM1), with the LOVD Variant ID #0000908423.
Transcript analysis
Total RNA was extracted from cultured primary fibroblasts of the patient and three healthy individuals (Monarch Total RNA Miniprep Kit, New England Biolabs). RNA concentration and purity of the samples were assessed by use of the Epoch™ Microplate Spectrophotometer (BioTek). One microgram of total RNA was reverse transcribed (LunaScript® RT SuperMix Kit, New England Biolabs). Primers designed to amplify DNM1 cDNA fragments from fibroblast-derived cDNA of the patient and healthy controls are described in Supplementary Table 1. PCR products were cloned into the pCR2.1 TOPO TA Cloning Vector (ThermoFisherScientific). Individual Escherichia coli clones were subjected to colony PCR followed by Sanger sequencing.
Results
Clinical findings
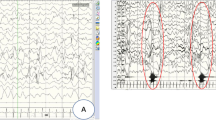
The patient is a 2-year-old boy and the second child of healthy non-consanguineous parents of Caucasian origin. Family history was unremarkable, except for a developmental disorder without seizures in a 20-year-old maternal first cousin-once-removed. Pregnancy was uneventful and delivery was at 42 weeks of gestation. Birth measurements were normal, with weight of 3480 g (− 0.28 z), length of 54 cm (0.75 z), and occipitofrontal circumference (OFC) of 36 cm (0.55 z). During the first days, breastfeeding was difficult, but he was bottle fed without any problems. In the neonatal phase, the mother described him as hypotonic and to be easily startled. From 4 weeks of age, the parents noticed twitching of the right leg and more frequent crying. About 10 days later, a first prolonged seizure occurred and the patient was admitted to our hospital. At the age of 6 weeks, body measurements were within the normal range, with weight of 4200 g (0.26 z), length of 57 cm (0.83 z), and OFC of 37.5 cm (0.2 z). Clinical examination was normal and he did not have any dysmorphic features. EEG showed an immature irregular activity with multifocal and generalized epileptic discharges. Brain MRI was normal, aside from a right choroidal fissure cyst. Metabolic workup revealed normal results, including enzyme activities for CLN1 and CLN2. There was no history of a congenital infection and TORCH serology was negative. Echocardiogram and abdominal ultrasound revealed patent foramen ovale without any other abnormalities.
After the first prolonged seizure at 6 weeks of age, the boy developed frequent seizures of more than 100 per day, including myoclonic jerks, tonic clonic seizures, and tonic seizures. He was treated with various medications, but these resulted in only a slight improvement in seizure frequency: pyridoxine, pyridoxal phosphate, folic acid, topiramate, levetiracetam, brivaracetam, clobazam, vigabatrin, ethosuximide, and zonisamide. He also received ketogenic diet. Due to hypsarrhythmia on EEG, a treatment with prednisolone according to the ICISS scheme was initiated. He later received high-dose methylprednisolone, but this resulted in only temporary improvement. At the age of 2 years, he still had > 50 myoclonic and tonic seizures per day, including prolonged seizures of > 15 min several times a day. EEG showed a general slowing with multifocal epileptic discharges.
At last follow-up at 2 years of age, the patient had normal growth parameters with weight of 10.8 kg (− 1.30 z), length of 89 cm (− 0.13 z), and OFC of 47.5 cm (− 1.88 z). In addition to the therapy-resistant epilepsy, he attained no motor and cognitive developmental milestones. He was severely hypotonic and had not attained head control. Except for subtle movements of the feet, he did not show any motor activity. He was not able to roll, crawl, sit, or walk. He did not show any reaction to visual or auditory stimuli and his responses to tactile stimuli were rare. His gaze did not fix on objects. There was no speech development. Due to frequent vomiting, a percutaneous jejunostomy was performed at the age of 20 months. Together, the patient showed the typical clinical picture of a developmental and epileptic encephalopathy with therapy-resistant epilepsy and no achievement of developmental milestones.
Genetic and transcript findings
We performed trio exome sequencing in the patient and parents and did not detect any biallelic or X-chromosomal variant in a known disease gene or disease gene candidate at the time of analysis that could underlie his severe developmental and epileptic encephalopathy. We identified three de novo variants in the patient (Supplementary Table 2), among them a deep intronic variant in the DEE-associated gene DNM1, NM_004408.4:c.1335 + 1600C > G, located in the 9631-bp large intron 10 (Fig. 1a). NM_004408.4 represents the longest DNM1 transcript and contains the alternatively spliced exon 10b. The NM_004408.4:c.1335 + 1600C > G transversion was absent from the gnomAD databases v.2.1.1 and 3.2.1, while a C-to-T transition at the same position, NM_004408.4:c.1335 + 1600C > T, was present in 1 of 151,934 alleles (worldwide allele frequency of 0.000006582; gnomAD v.3.2.1). We validated the DNM1 intronic variant in leukocyte- and fibroblast-derived DNA of the patient and confirmed its absence in parental DNA samples by Sanger sequencing (Fig. 2a). The C-to-G change is at the identical intronic position according to the two other DNM1 transcript variants NM_001005336.3 and NM_001374269.1 that all contain exon 10b spliced to exon 11 (Fig. 1a). The three other DNM1 transcript variants NM_001288737.2, NM_001288738.2, and NM_001288739.2 contain exon 10a instead of exon 10b (Fig. 1a). According to the three latter transcript variants, the transversion is located in intron 9 at position − 46 upstream of the alternatively spliced exon 10a: NM_001288739.2:c.1197-46C > G. To predict the effect of the non-coding change on splicing of DNM1 pre-mRNAs, we used four in silico programs that all predicted creation of a new splice acceptor site in intron 9 (Supplementary Table 3).
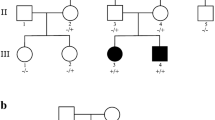
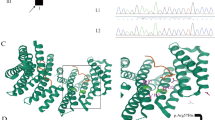
Validation of the DNM1 c.1197-46C > G variant and DNM1 transcript analysis. a Partial sequence electropherograms demonstrating a DNM1 NM_001288739.2:c.1197-46C > G variant in the heterozygous state in leukocyte- and fibroblast-derived DNA of the patient. The healthy parents (mother and father) do not carry the intronic DNM1 variant in leukocyte-derived DNA. An arrow points to the heterozygous variant. b 2% agarose gel picture showing amplicons of RT-PCR using fibroblast-derived cDNA of the patient and three controls with primers located in exons 9 (forward) and 10a (reverse; orange arrows). Two amplicons, one of the expected size (168 bp) and one larger product (~ 220 bp), were present in the patient. One amplicon (168 bp) was observed in the three controls. Schematics of the exon-exon and exon-intron-exon junctions as well as the size of the amplicons (after sequencing) are shown on the right. c Partial sequence electropherogram of the aberrantly spliced DMN1 transcript in the patient. Cloning of patient-derived RT-PCR amplicons into pCR2.1 TOPO TA cloning vector followed by colony PCR and Sanger sequencing of individual amplicons identified the larger amplicon (213 bp) to represent DNM1 transcripts harboring the last 45 bp of intron 9 between exons 9 and 10a (r.1196_1197ins[1197-1_1197-45]). Triplets and encoded amino acid residues in the three-letter code are shown below the sequence. Residues in blue indicate 15 novel amino acids located between Ile398 and Arg399. In, intron
To analyze potential aberrant splicing of DNM1 pre-mRNAs, we performed qualitative RT-PCR using fibroblast-derived cDNA of the patient and controls. Cultured fibroblasts predominantly express the exon 10b containing transcript variant NM_001374269.1 (Supplementary Fig. 1a according to GTeX). RT-PCR using primers located in exons 9 and 11 resulted in a single amplicon of 219 bp in the patient and controls (Supplementary Fig. 1b), which represents DNM1 transcripts in which exon 9 was spliced to exon 10b (Supplementary Fig. 1c). To specifically amplify exon 10a including transcript variants, we combined the forward primer in exon 9 with a reverse primer in exon 10a and obtained the expected wild-type amplicon of 168 bp in controls and the 168-bp amplicon in addition to a larger PCR product (~ 220 bp) in the patient (Fig. 2b). Cloning of patient-derived amplicons followed by colony PCR and Sanger sequencing revealed the presence of wild-type DNM1 transcripts in which exon 9 was spliced to exon 10a as well as aberrantly spliced DNM1 mRNAs that contain exon 9, the last 45 bp of intron 9, and exon 10a in the patient (Fig. 2c). Inclusion of 45 nucleotides of intron 9 in the DNM1 mRNA is in accordance with usage of the newly generated splice acceptor site in intron 9: NM_001288738.2:r.1196_1197ins [1197-1_1197-45]. On protein level, the 45-nucleotide in-frame insertion in DNM1 mRNAs including exon 10a predicts insertion of 15 DNM1-unrelated amino acid residues between isoleucine 398 and arginine 399, but only in the encoded DNM1A isoforms [NP_001275668.1:p.(Ile398_Arg399ins15)] and not in DNM1B isoforms produced from alternatively spliced DNM1 transcripts with the divergent exon 10b. The 15 inserted amino acids are located before the 46 residues of the middle domain encoded by exon 10a in the patient (Figs. 1b and 2c).
Discussion
We report on a 2-year-old male patient with a de novo DNM1 pathogenic variant in intron 9 who had the clinical findings of DEE with therapy-resistant epilepsy. The phenotype of the severely affected boy fits the clinical spectrum of DNM1 encephalopathy [17, 44]. In particular, our patient had a similar severe neurological phenotype as individuals with heterozygous DNM1 variants affecting both DNM1A and DNM1B isoforms [17], individuals with heterozygous DNM1 variants affecting only DNM1A isoform A [44, 49, 50], and the recently reported individuals with homozygous DNM1 loss-of-function variants [63, 64] (Table 1). The data highlights (i) the importance of DNM1 isoform A in brain development and cognitive function and (ii) a similar severe neurodevelopmental disorder in patients with heterozygous DNM1 variants causing a dominant-negative effect and patients with biallelic loss-of-function DNM1 variants. A few individuals with heterozygous DNM1 pathogenic variants were reported to show milder neurodevelopmental phenotypes, including three patients with a variant affecting the PH domain of dynamin 1 (Fig. 1b; Table 1) [46,47,48].
The intronic variant NM_001288739.2:c.1197-46C > G affects three out of six DNM1 transcript variants that include exon 10a instead of 10b, while almost all other DNM1 pathogenic variants affect all six DNM1 transcript variants (Fig. 1). The c.1197-46C > G variant caused aberrant splicing of DNM1 pre-mRNAs by inclusion of the last 45 nucleotides of intron 9 between exon 9 and exon 10a in mature DNM1 transcripts: NM_001288739.2:r.1196_1197ins[1197-1_1197-45]. The aberrantly spliced DNM1 transcripts encode DNM1 mutant proteins with 15 DNM1-unrelated amino acids (SHGCSSSCPHLLPGC) inserted before a peptide of 46 residues in the middle domain that is encoded by exon 10a. Accordingly, cells of the patient expressing DNM1A transcript variants are supposed to produce 50% wild-type and 50% mutant DNM1A [NP_001275668.1:p.(Ile398_Arg399ins15)]. The disease relevance of DNM1 mRNAs including exon 10a is underscored by a previously reported de novo DNM1 pathogenic variant NM_001288739.1:c.1197-8G > A in several unrelated patients with DEE [44, 49, 50]. This G-to-A transition in intron 9 is predicted to create a new splice acceptor site (Supplementary Table 3) and results in an in-frame insertion of two novel amino acids between arginine 399 and threonine 400 [p.(Arg399_Thr400insCysArg)] [44]. We assume that production of DNM1B isoforms encoded by DNM1 mRNAs with the divergent exon 10b is left intact in patients carrying a DNM1 sequence change located directly upstream of the splice acceptor preceding exon 10a.
Dimerization of dynamin is mediated by the middle domain. Interaction between dynamin dimers drives the assembly into the helical polymer at the neck of the clathrin-coated endocytic pit [10, 11]. Several amino acid substitutions affecting the middle domain impair the higher-order self-assembly process of dynamin, including Arg399Ala [65] and the disease-associated amino acid substitutions Gly359Ala and Gly397Asp [31, 53]. These data demonstrate that residues 397–399 of the dynamin middle domain are important for dynamin dimerization followed by coordinated polymerization [53, 65]. Based on this data, any amino acid insertion between Ile398 and Thr400 likely leads to a dominant-negative effect of the DNM1A mutant on dynamin 1 dimerization, polymerization into oligomers, or, as proposed previously, oligomerization-induced GTPase activation [44].
A spontaneous heterozygous mutation in Dnm1, only affecting exon 10a, has been identified in the “fitful” (allele symbol: Ftfl) mouse which develops tonic-clonic seizures from 2 to 3 months of age. The point mutation in exon 10a results in the substitution p.Ala408Thr leading to expression of the Dnm1aFtfl mutant protein, while production of wild-type Dnm1b isoforms is intact. The Dnm1aFtfl mutant protein does not assemble into higher-order dynamin complexes and interferes with endocytosis. Thus, the Dnm1 p.Ala408Thr variant has a dominant-negative effect, possibly by binding of Dnm1 mutant with wild type that results in non-functional heterodimers [6]. These data show the importance of the Dnm1a isoform for normal brain function in mice.
Interestingly, DNM1 transcript variants NM_001288737.2 and NM_001288739.2 including exon 10a show higher expression in the human cortex than the alternatively spliced mRNAs including exon 10b (NM_001005336.3, NM_001374269.1, and NM_004408.4) (Fig. 1a according to GTeX). This finding was confirmed by RNA-seq data in human pediatric brain samples that show a 5.7-fold higher expression of DNM1A compared with DNM1B mRNAs in the cortex [44]. Developmental neuronal expression of Dnm1 transcript variants with exon 10a or 10b and encoded isoforms has been studied in the mouse. Dnm1b expression is high during embryonic and early postnatal development, while Dnm1a expression increases postnatally with synaptic maturation. Differential developmental regulation of DNM1 exon10a and 10b alternative splicing in the brain suggests potential functional differences between the two Dnm1 isoforms [6, 66]. Subcellular localization studies demonstrated co-localization of Dnm1a with clathrin at the plasma membrane, while Dnm1b isoform preferentially localizes at the Golgi apparatus [7, 66]. Furthermore, Dnm1a and b have different capacities to bind the interaction partner amphiphysin 1 and to rescue epileptic pathology in the Dnm1Ftfl mouse. While Dnm1a is able to prevent the seizure phenotype in the mutant mouse, this is not the case for Dnm1b. Overall, functional differences between Dnm1 isoforms a and b with varying amino acid sequences in a portion of the middle domain have been postulated to reflect differential endocytic requirements during early and adult brain functions [66]. These data together with the identification of pathogenic variants that specifically affect DNM1 transcripts including exon 10a and that cause pathology in mice and DEE in humans suggest that particularly the DNM1A isoform plays a critical role in human brain development.
In conclusion, we show here that a de novo DNM1 intronic variant NM_001288739.2:c.1197-46C > G in a patient with DEE specifically affects DNM1 transcripts including the alternatively spliced exon 10a and causes inclusion of 45 intronic nucleotides in DNM1A mRNAs. The encoded DNM1 isoform A mutant is predicted to contain 15 novel amino acids between Ile398 and Arg399 [NP_001275668.1:p.(Ile398_Arg399ins15)] and is supposed to act in a dominant-negative manner, potentially by affecting the higher-order oligomerization of dynamin 1. Functional differences between the two Dnm1a and Dnm1b isoforms and the incapability of Dnm1b to functionally compensate in the presence of a pathogenic variant only affecting Dnm1 isoform a in the Dnm1Ftfl mouse [6, 66] can be recapitulated in patients with DEE and DNM1 pathogenic variants affecting exon 10a and not exon 10b.
References
Ferguson SM, Brasnjo G, Hayashi M, Wolfel M, Collesi C, Giovedi S, Raimondi A, Gong LW, Ariel P, Paradise S, O’Toole E, Flavell R, Cremona O, Miesenbock G, Ryan TA, De Camilli P (2007) A selective activity-dependent requirement for dynamin 1 in synaptic vesicle endocytosis. Science 316(5824):570–574. https://doi.org/10.1126/science.1140621
Robinson PJ, Sontag JM, Liu JP, Fykse EM, Slaughter C, McMahon H, Sudhof TC (1993) Dynamin GTPase regulated by protein kinase C phosphorylation in nerve terminals. Nature 365(6442):163–166. https://doi.org/10.1038/365163a0
van der Bliek AM, Meyerowitz EM (1991) Dynamin-like protein encoded by the Drosophila shibire gene associated with vesicular traffic. Nature 351(6325):411–414. https://doi.org/10.1038/351411a0
Nakata T, Iwamoto A, Noda Y, Takemura R, Yoshikura H, Hirokawa N (1991) Predominant and developmentally regulated expression of dynamin in neurons. Neuron 7(3):461–469. https://doi.org/10.1016/0896-6273(91)90298-e
Scaife R, Margolis RL (1990) Biochemical and immunochemical analysis of rat brain dynamin interaction with microtubules and organelles in vivo and in vitro. J Cell Biol 111(6 Pt 2):3023–3033. https://doi.org/10.1083/jcb.111.6.3023
RM Boumil VA Letts MC Roberts C Lenz CL Mahaffey ZW Zhang T Moser WN Frankel 2010 A missense mutation in a highly conserved alternate exon of dynamin 1 causes epilepsy in fitful mice PLoS Genet 6 8. https://doi.org/10.1371/journal.pgen.1001046
Cao H, Garcia F, McNiven MA (1998) Differential distribution of dynamin isoforms in mammalian cells. Mol Biol Cell 9(9):2595–2609. https://doi.org/10.1091/mbc.9.9.2595
Sontag JM, Fykse EM, Ushkaryov Y, Liu JP, Robinson PJ, Sudhof TC (1994) Differential expression and regulation of multiple dynamins. J Biol Chem 269(6):4547–4554
Urrutia R, Henley JR, Cook T, McNiven MA (1997) The dynamins: redundant or distinct functions for an expanding family of related GTPases? Proc Natl Acad Sci USA 94(2):377–384. https://doi.org/10.1073/pnas.94.2.377
Antonny B, Burd C, De Camilli P, Chen E, Daumke O, Faelber K, Ford M, Frolov VA, Frost A, Hinshaw JE, Kirchhausen T, Kozlov MM, Lenz M, Low HH, McMahon H, Merrifield C, Pollard TD, Robinson PJ, Roux A, Schmid S (2016) Membrane fission by dynamin: what we know and what we need to know. EMBO J 35(21):2270–84. https://doi.org/10.15252/embj.201694613
Ferguson SM, De Camilli P (2012) Dynamin, a membrane-remodelling GTPase. Nat Rev Mol Cell Biol 13(2):75–88. https://doi.org/10.1038/nrm3266
Euro E-RESC, Phenome E, Genome P, Epi KC (2014) De novo mutations in synaptic transmission genes including DNM1 cause epileptic encephalopathies. Am J Hum Genet 95(4):360–370. https://doi.org/10.1016/j.ajhg.2014.08.013
John A, Ng-Cordell E, Hanna N, Brkic D, Baker K (2021) The neurodevelopmental spectrum of synaptic vesicle cycling disorders. J Neurochem 157(2):208–228. https://doi.org/10.1111/jnc.15135
Fernandez-Marmiesse A, Roca I, Diaz-Flores F, Cantarin V, Perez-Poyato MS, Fontalba A, Laranjeira F, Quintans S, Moldovan O, Felgueroso B, Rodriguez-Pedreira M, Simon R, Camacho A, Quijada P, Ibanez-Mico S, Domingno MR, Benito C, Calvo R, Perez-Cejas A, Carrasco ML, Ramos F, Couce ML, Ruiz-Falco ML, Gutierrez-Solana L, Martinez-Atienza M (2019) Rare variants in 48 genes account for 42% of cases of epilepsy with or without neurodevelopmental delay in 246 pediatric patients. Front Neurosci 13:1135. https://doi.org/10.3389/fnins.2019.01135
Investigators GPP, Smedley D, Smith KR, Martin A, Thomas EA, McDonagh EM, Cipriani V, Ellingford JM, Arno G, Tucci A, Vandrovcova J, Chan G, Williams HJ, Ratnaike T, Wei W, Stirrups K, Ibanez K, Moutsianas L, Wielscher M, Need A, Barnes MR, Vestito L, Buchanan J, Wordsworth S, Ashford S, Rehmstrom K, Li E, Fuller G, Twiss P, Spasic-Boskovic O, Halsall S, Floto RA, Poole K, Wagner A, Mehta SG, Gurnell M, Burrows N, James R, Penkett C, Dewhurst E, Graf S, Mapeta R, Kasanicki M, Haworth A, Savage H, Babcock M, Reese MG, Bale M, Baple E, Boustred C, Brittain H, de Burca A, Bleda M, Devereau A, Halai D, Haraldsdottir E, Hyder Z, Kasperaviciute D, Patch C, Polychronopoulos D, Matchan A, Sultana R, Ryten M, Tavares ALT, Tregidgo C, Turnbull C, Welland M, Wood S, Snow C, Williams E, Leigh S, Foulger RE, Daugherty LC, Niblock O, Leong IUS, Wright CF, Davies J, Crichton C, Welch J, Woods K, Abulhoul L, Aurora P, Bockenhauer D, Broomfield A, Cleary MA, Lam T, Dattani M, Footitt E, Ganesan V, Grunewald S, Compeyrot-Lacassagne S, Muntoni F, Pilkington C, Quinlivan R, Thapar N, Wallis C, Wedderburn LR, Worth A, Bueser T, Compton C, Deshpande C, Fassihi H, Haque E, Izatt L, Josifova D, Mohammed S, Robert L, Rose S, Ruddy D, Sarkany R, Say G, Shaw AC, Wolejko A, Habib B, Burns G, Hunter S, Grocock RJ, Humphray SJ, Robinson PN, Haendel M, Simpson MA, Banka S, Clayton-Smith J, Douzgou S, Hall G, Thomas HB, O’Keefe RT, Michaelides M, Moore AT, Malka S, Pontikos N, Browning AC, Straub V, Gorman GS, Horvath R, Quinton R, Schaefer AM, Yu-Wai-Man P, Turnbull DM, McFarland R, Taylor RW, O’Connor E, Yip J, Newland K, Morris HR, Polke J, Wood NW, Campbell C, Camps C, Gibson K, Koelling N, Lester T, Nemeth AH, Palles C, Patel S, Roy NBA, Sen A, Taylor J, Cacheiro P, Jacobsen JO, Seaby EG, Davison V, Chitty L, Douglas A, Naresh K, McMullan D, Ellard S, Temple IK, Mumford AD, Wilson G, Beales P, Bitner-Glindzicz M, Black G, Bradley JR, Brennan P, Burn J, Chinnery PF, Elliott P, Flinter F, Houlden H, Irving M, Newman W, Rahman S, Sayer JA, Taylor JC, Webster AR, Wilkie AOM, Ouwehand WH, Raymond FL, Chisholm J, Hill S, Bentley D, Scott RH, Fowler T, Rendon A, Caulfield M (2021) 100,000 genomes pilot on rare-disease diagnosis in health care - preliminary report. N Engl J Med 385(20):1868–1880. https://doi.org/10.1056/NEJMoa2035790
Nakashima M, Kouga T, Lourenco CM, Shiina M, Goto T, Tsurusaki Y, Miyatake S, Miyake N, Saitsu H, Ogata K, Osaka H, Matsumoto N (2016) De novo DNM1 mutations in two cases of epileptic encephalopathy. Epilepsia 57(1):e18-23. https://doi.org/10.1111/epi.13257
von Spiczak S, Helbig KL, Shinde DN, Huether R, Pendziwiat M, Lourenco C, Nunes ME, Sarco DP, Kaplan RA, Dlugos DJ, Kirsch H, Slavotinek A, Cilio MR, Cervenka MC, Cohen JS, McClellan R, Fatemi A, Yuen A, Sagawa Y, Littlejohn R, McLean SD, Hernandez-Hernandez L, Maher B, Moller RS, Palmer E, Lawson JA, Campbell CA, Joshi CN, Kolbe DL, Hollingsworth G, Neubauer BA, Muhle H, Stephani U, Scheffer IE, Pena SDJ, Sisodiya SM, Helbig I, Epi KC, Euro E-RESNWG (2017) DNM1 encephalopathy: a new disease of vesicle fission. Neurology 89(4):385–394. https://doi.org/10.1212/WNL.0000000000004152
Li H, Fang F, Xu M, Liu Z, Zhou J, Wang X, Wang X, Han T (2019) Clinical assessments and EEG analyses of encephalopathies associated with dynamin-1 mutation. Front Pharmacol 10:1454. https://doi.org/10.3389/fphar.2019.01454
Lin L, Zhang Y, Pan H, Wang J, Qi Y, Ma Y (2020) Clinical and genetic characteristics and prenatal diagnosis of patients presented GDD/ID with rare monogenic causes. Orphanet J Rare Dis 15(1):317. https://doi.org/10.1186/s13023-020-01599-y
Takata A, Nakashima M, Saitsu H, Mizuguchi T, Mitsuhashi S, Takahashi Y, Okamoto N, Osaka H, Nakamura K, Tohyama J, Haginoya K, Takeshita S, Kuki I, Okanishi T, Goto T, Sasaki M, Sakai Y, Miyake N, Miyatake S, Tsuchida N, Iwama K, Minase G, Sekiguchi F, Fujita A, Imagawa E, Koshimizu E, Uchiyama Y, Hamanaka K, Ohba C, Itai T, Aoi H, Saida K, Sakaguchi T, Den K, Takahashi R, Ikeda H, Yamaguchi T, Tsukamoto K, Yoshitomi S, Oboshi T, Imai K, Kimizu T, Kobayashi Y, Kubota M, Kashii H, Baba S, Iai M, Kira R, Hara M, Ohta M, Miyata Y, Miyata R, Takanashi JI, Matsui J, Yokochi K, Shimono M, Amamoto M, Takayama R, Hirabayashi S, Aiba K, Matsumoto H, Nabatame S, Shiihara T, Kato M, Matsumoto N (2019) Comprehensive analysis of coding variants highlights genetic complexity in developmental and epileptic encephalopathy. Nat Commun 10(1):2506. https://doi.org/10.1038/s41467-019-10482-9
Bowling KM, Thompson ML, Amaral MD, Finnila CR, Hiatt SM, Engel KL, Cochran JN, Brothers KB, East KM, Gray DE, Kelley WV, Lamb NE, Lose EJ, Rich CA, Simmons S, Whittle JS, Weaver BT, Nesmith AS, Myers RM, Barsh GS, Bebin EM, Cooper GM (2017) Genomic diagnosis for children with intellectual disability and/or developmental delay. Genome Med 9(1):43. https://doi.org/10.1186/s13073-017-0433-1
Zhu X, Padmanabhan R, Copeland B, Bridgers J, Ren Z, Kamalakaran S, O’Driscoll-Collins A, Berkovic SF, Scheffer IE, Poduri A, Mei D, Guerrini R, Lowenstein DH, Allen AS, Heinzen EL, Goldstein DB (2017) A case-control collapsing analysis identifies epilepsy genes implicated in trio sequencing studies focused on de novo mutations. PLoS Genet 13(11):e1007104. https://doi.org/10.1371/journal.pgen.1007104
Deng XL, Yin F, Zhang CL, Ma YP, He F, Wu LW, Peng J (2016) Dynamin-1-related infantile spasms: a case report and review of literature. Zhonghua Er Ke Za Zhi 54(11):856–859. https://doi.org/10.3760/cma.j.issn.0578-1310.2016.11.014
Fung CW, Kwong AK, Wong VC (2017) Gene panel analysis for nonsyndromic cryptogenic neonatal/infantile epileptic encephalopathy. Epilepsia Open 2(2):236–243. https://doi.org/10.1002/epi4.12055
Yao R, Zhou Y, Tang J, Li N, Yu T, He Y, Wang C, Wang J, Wang J (2021) Genetic diagnosis spectrum and multigenic burden of exome-level rare variants in a childhood epilepsy cohort. Front Genet 12:782419. https://doi.org/10.3389/fgene.2021.782419
Allen NM, Conroy J, Shahwan A, Lynch B, Correa RG, Pena SD, McCreary D, Magalhaes TR, Ennis S, Lynch SA, King MD (2016) Unexplained early onset epileptic encephalopathy: exome screening and phenotype expansion. Epilepsia 57(1):e12–e17. https://doi.org/10.1111/epi.13250
Epi KC, Phenome E, Genome P, Allen AS, Berkovic SF, Cossette P, Delanty N, Dlugos D, Eichler EE, Epstein MP, Glauser T, Goldstein DB, Han Y, Heinzen EL, Hitomi Y, Howell KB, Johnson MR, Kuzniecky R, Lowenstein DH, Lu YF, Madou MR, Marson AG, Mefford HC, Esmaeeli Nieh S, O’Brien TJ, Ottman R, Petrovski S, Poduri A, Ruzzo EK, Scheffer IE, Sherr EH, Yuskaitis CJ, Abou-Khalil B, Alldredge BK, Bautista JF, Berkovic SF, Boro A, Cascino GD, Consalvo D, Crumrine P, Devinsky O, Dlugos D, Epstein MP, Fiol M, Fountain NB, French J, Friedman D, Geller EB, Glauser T, Glynn S, Haut SR, Hayward J, Helmers SL, Joshi S, Kanner A, Kirsch HE, Knowlton RC, Kossoff EH, Kuperman R, Kuzniecky R, Lowenstein DH, McGuire SM, Motika PV, Novotny EJ, Ottman R, Paolicchi JM, Parent JM, Park K, Poduri A, Scheffer IE, Shellhaas RA, Sherr EH, Shih JJ, Singh R, Sirven J, Smith MC, Sullivan J, Lin Thio L, Venkat A, Vining EP, Von Allmen GK, Weisenberg JL, Widdess-Walsh P, Winawer MR (2013) De novo mutations in epileptic encephalopathies. Nature 501(7466):217–221. https://doi.org/10.1038/nature12439
Ko A, Youn SE, Kim SH, Lee JS, Kim S, Choi JR, Kim HD, Lee ST, Kang HC (2018) Targeted gene panel and genotype-phenotype correlation in children with developmental and epileptic encephalopathy. Epilepsy Res 141:48–55. https://doi.org/10.1016/j.eplepsyres.2018.02.003
Snoeijen-Schouwenaars FM, van Ool JS, Verhoeven JS, van Mierlo P, Braakman HMH, Smeets EE, Nicolai J, Schoots J, Teunissen MWA, Rouhl RPW, Tan IY, Yntema HG, Brunner HG, Pfundt R, Stegmann AP, Kamsteeg EJ, Schelhaas HJ, Willemsen MH (2019) Diagnostic exome sequencing in 100 consecutive patients with both epilepsy and intellectual disability. Epilepsia 60(1):155–164. https://doi.org/10.1111/epi.14618
Yuskaitis CJ, Ruzhnikov MRZ, Howell KB, Allen IE, Kapur K, Dlugos DJ, Scheffer IE, Poduri A, Sherr EH (2018) Infantile spasms of unknown cause: predictors of outcome and genotype-phenotype correlation. Pediatr Neurol 87:48–56. https://doi.org/10.1016/j.pediatrneurol.2018.04.012
Dhindsa RS, Bradrick SS, Yao X, Heinzen EL, Petrovski S, Krueger BJ, Johnson MR, Frankel WN, Petrou S, Boumil RM, Goldstein DB (2015) Epileptic encephalopathy-causing mutations in DNM1 impair synaptic vesicle endocytosis. Neurol Genet 1(1):e4. https://doi.org/10.1212/01.NXG.0000464295.65736.da
Epi KC (2016) De novo mutations in SLC1A2 and CACNA1A are important causes of epileptic encephalopathies. Am J Hum Genet 99(2):287–298. https://doi.org/10.1016/j.ajhg.2016.06.003
Lazzara A, Asghar S, Zacharia T, Byler D (2018) DNM1 mutation in a child associated with progressive bilateral mesial temporal sclerosis. Clin Case Rep 6(11):2037–2039. https://doi.org/10.1002/ccr3.1793
Naess K, Bruhn H, Stranneheim H, Freyer C, Wibom R, Mourier A, Engvall M, Nennesmo I, Lesko N, Wredenberg A, Wedell A, von Dobeln U (2021) Clinical presentation, genetic etiology, and coenzyme Q10 levels in 55 children with combined enzyme deficiencies of the mitochondrial respiratory chain J Pediatr 228:240–51 e2. https://doi.org/10.1016/j.jpeds.2020.08.025
Palmer EE, Schofield D, Shrestha R, Kandula T, Macintosh R, Lawson JA, Andrews I, Sampaio H, Johnson AM, Farrar MA, Cardamone M, Mowat D, Elakis G, Lo W, Zhu Y, Ying K, Morris P, Tao J, Dias KR, Buckley M, Dinger ME, Cowley MJ, Roscioli T, Kirk EP, Bye A, Sachdev RK (2018) Integrating exome sequencing into a diagnostic pathway for epileptic encephalopathy: evidence of clinical utility and cost effectiveness. Mol Genet Genomic Med 6(2):186–199. https://doi.org/10.1002/mgg3.355
Hamdan FF, Myers CT, Cossette P, Lemay P, Spiegelman D, Laporte AD, Nassif C, Diallo O, Monlong J, Cadieux-Dion M, Dobrzeniecka S, Meloche C, Retterer K, Cho MT, Rosenfeld JA, Bi W, Massicotte C, Miguet M, Brunga L, Regan BM, Mo K, Tam C, Schneider A, Hollingsworth G, Deciphering Developmental Disorders S, FitzPatrick DR, Donaldson A, Canham N, Blair E, Kerr B, Fry AE, Thomas RH, Shelagh J, Hurst JA, Brittain H, Blyth M, Lebel RR, Gerkes EH, Davis-Keppen L, Stein Q, Chung WK, Dorison SJ, Benke PJ, Fassi E, Corsten-Janssen N, Kamsteeg EJ, Mau-Them FT, Bruel AL, Verloes A, Ounap K, Wojcik MH, Albert DVF, Venkateswaran S, Ware T, Jones D, Liu YC, Mohammad SS, Bizargity P, Bacino CA, Leuzzi V, Martinelli S, Dallapiccola B, Tartaglia M, Blumkin L, Wierenga KJ, Purcarin G, O’Byrne JJ, Stockler S, Lehman A, Keren B, Nougues MC, Mignot C, Auvin S, Nava C, Hiatt SM, Bebin M, Shao Y, Scaglia F, Lalani SR, Frye RE, Jarjour IT, Jacques S, Boucher RM, Riou E, Srour M, Carmant L, Lortie A, Major P, Diadori P, Dubeau F, D’Anjou G, Bourque G, Berkovic SF, Sadleir LG, Campeau PM, Kibar Z, Lafreniere RG, Girard SL, Mercimek-Mahmutoglu S, Boelman C, Rouleau GA, Scheffer IE, Mefford HC, Andrade DM, Rossignol E, Minassian BA, Michaud JL (2017) High rate of recurrent de novo mutations in developmental and epileptic encephalopathies. Am J Hum Genet 101(5):664–685. https://doi.org/10.1016/j.ajhg.2017.09.008
Helbig KL, Farwell Hagman KD, Shinde DN, Mroske C, Powis Z, Li S, Tang S, Helbig I (2016) Diagnostic exome sequencing provides a molecular diagnosis for a significant proportion of patients with epilepsy. Genet Med 18(9):898–905. https://doi.org/10.1038/gim.2015.186
Joshi C, Kolbe DL, Mansilla MA, Mason SO, Smith RJ, Campbell CA (2016) Reducing the cost of the diagnostic odyssey in early onset epileptic encephalopathies. Biomed Res Int 2016:6421039. https://doi.org/10.1155/2016/6421039
Moller RS, Larsen LH, Johannesen KM, Talvik I, Talvik T, Vaher U, Miranda MJ, Farooq M, Nielsen JE, Svendsen LL, Kjelgaard DB, Linnet KM, Hao Q, Uldall P, Frangu M, Tommerup N, Baig SM, Abdullah U, Born AP, Gellert P, Nikanorova M, Olofsson K, Jepsen B, Marjanovic D, Al-Zehhawi LI, Penalva SJ, Krag-Olsen B, Brusgaard K, Hjalgrim H, Rubboli G, Pal DK, Dahl HA (2016) Gene panel testing in epileptic encephalopathies and familial epilepsies. Mol Syndromol 7(4):210–219. https://doi.org/10.1159/000448369
Rim JH, Kim SH, Hwang IS, Kwon SS, Kim J, Kim HW, Cho MJ, Ko A, Youn SE, Kim J, Lee YM, Chung HJ, Lee JS, Kim HD, Choi JR, Lee ST, Kang HC (2018) Efficient strategy for the molecular diagnosis of intractable early-onset epilepsy using targeted gene sequencing. BMC Med Genomics 11(1):6. https://doi.org/10.1186/s12920-018-0320-7
Kolnikova M, Skopkova M, Ilencikova D, Foltan T, Payerova J, Danis D, Klimes I, Stanik J, Gasperikova D (2018) DNM1 encephalopathy - atypical phenotype with hypomyelination due to a novel de novo variant in the DNM1 gene. Seizure 56:31–33. https://doi.org/10.1016/j.seizure.2018.01.020
Posey JE, Harel T, Liu P, Rosenfeld JA, James RA, Coban Akdemir ZH, Walkiewicz M, Bi W, Xiao R, Ding Y, Xia F, Beaudet AL, Muzny DM, Gibbs RA, Boerwinkle E, Eng CM, Sutton VR, Shaw CA, Plon SE, Yang Y, Lupski JR (2017) Resolution of disease phenotypes resulting from multilocus genomic variation. N Engl J Med 376(1):21–31. https://doi.org/10.1056/NEJMoa1516767
French CE, Dolling H, Megy K, Sanchis-Juan A, Kumar A, Delon I, Wakeling M, Mallin L, Agrawal S, Austin T, Walston F, Park SM, Parker A, Piyasena C, Bradbury K, Next Generation Children's Project C, Ellard S, Rowitch DH, Raymond FL (2022) Refinements and considerations for trio whole-genome sequence analysis when investigating Mendelian diseases presenting in early childhood HGG Adv 3 3 100113. https://doi.org/10.1016/j.xhgg.2022.100113
Parthasarathy S, Ruggiero SM, Gelot A, Soardi FC, Ribeiro BFR, Pires DEV, Ascher DB, Schmitt A, Rambaud C, Represa A, Xie HM, Lusk L, Wilmarth O, McDonnell PP, Juarez OA, Grace AN, Buratti J, Mignot C, Gras D, Nava C, Pierce SR, Keren B, Kennedy BC, Pena SDJ, Helbig I, Cuddapah VA (2022) A recurrent de novo splice site variant involving DNM1 exon 10a causes developmental and epileptic encephalopathy through a dominant-negative mechanism. Am J Hum Genet 109(12):2253–2269. https://doi.org/10.1016/j.ajhg.2022.11.002
Truty R, Patil N, Sankar R, Sullivan J, Millichap J, Carvill G, Entezam A, Esplin ED, Fuller A, Hogue M, Johnson B, Khouzam A, Kobayashi Y, Lewis R, Nykamp K, Riethmaier D, Westbrook J, Zeman M, Nussbaum RL, Aradhya S (2019) Possible precision medicine implications from genetic testing using combined detection of sequence and intragenic copy number variants in a large cohort with childhood epilepsy. Epilepsia Open 4(3):397–408. https://doi.org/10.1002/epi4.12348
Brereton E, Fassi E, Araujo GC, Dodd J, Telegrafi A, Pathak SJ, Shinawi M (2018) Mutations in the PH Domain of DNM1 are associated with a nonepileptic phenotype characterized by developmental delay and neurobehavioral abnormalities. Mol Genet Genomic Med 6(2):294–300. https://doi.org/10.1002/mgg3.362
Choi E, Dale B, RamachandranNair R, Ejaz R (2021) Pathogenic DNM1 gene variant presenting with unusually nonsevere neurodevelopmental phenotype a case report. Neurol Genet 7(5):e618. https://doi.org/10.1212/NXG.0000000000000618
van der Ven AT, Johannsen J, Kortum F, Wagner M, Tsiakas K, Bierhals T, Lessel D, Herget T, Kloth K, Lisfeld J, Scholz T, Obi N, Wortmann S, Prokisch H, Kubisch C, Denecke J, Santer R, Hempel M (2021) Prevalence and clinical prediction of mitochondrial disorders in a large neuropediatric cohort. Clin Genet 100(6):766–770. https://doi.org/10.1111/cge.14061
Devanna P, van de Vorst M, Pfundt R, Gilissen C, Vernes SC (2018) Genome-wide investigation of an ID cohort reveals de novo 3’UTR variants affecting gene expression. Hum Genet 137(9):717–721. https://doi.org/10.1007/s00439-018-1925-9
Sahly AN, Krochmalnek E, St-Onge J, Srour M, Myers KA (2020) Severe DNM1 encephalopathy with dysmyelination due to recurrent splice site pathogenic variant. Hum Genet 139(12):1575–1578. https://doi.org/10.1007/s00439-020-02224-5
Chappie JS, Acharya S, Leonard M, Schmid SL, Dyda F (2010) G domain dimerization controls dynamin’s assembly-stimulated GTPase activity. Nature 465(7297):435–440. https://doi.org/10.1038/nature09032
Damke H, Binns DD, Ueda H, Schmid SL, Baba T (2001) Dynamin GTPase domain mutants block endocytic vesicle formation at morphologically distinct stages. Mol Biol Cell 12(9):2578–2589. https://doi.org/10.1091/mbc.12.9.2578
Ford MG, Jenni S, Nunnari J (2011) The crystal structure of dynamin. Nature 477(7366):561–566. https://doi.org/10.1038/nature10441
Marks B, Stowell MH, Vallis Y, Mills IG, Gibson A, Hopkins CR, McMahon HT (2001) GTPase activity of dynamin and resulting conformation change are essential for endocytosis. Nature 410(6825):231–235. https://doi.org/10.1038/35065645
Pucadyil TJ, Schmid SL (2008) Real-time visualization of dynamin-catalyzed membrane fission and vesicle release. Cell 135(7):1263–1275. https://doi.org/10.1016/j.cell.2008.11.020
Vallis Y, Wigge P, Marks B, Evans PR, McMahon HT (1999) Importance of the pleckstrin homology domain of dynamin in clathrin-mediated endocytosis. Curr Biol 9(5):257–260. https://doi.org/10.1016/s0960-9822(99)80114-6
van der Bliek AM, Redelmeier TE, Damke H, Tisdale EJ, Meyerowitz EM, Schmid SL (1993) Mutations in human dynamin block an intermediate stage in coated vesicle formation. J Cell Biol 122(3):553–563. https://doi.org/10.1083/jcb.122.3.553
Chen S, Zhou Y, Chen Y, Gu J (2018) fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34(17):i884–i890. https://doi.org/10.1093/bioinformatics/bty560
Li H, Durbin R (2010) Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics 26(5):589–595. https://doi.org/10.1093/bioinformatics/btp698
Kim S, Scheffler K, Halpern AL, Bekritsky MA, Noh E, Kallberg M, Chen X, Kim Y, Beyter D, Krusche P, Saunders CT (2018) Strelka2: fast and accurate calling of germline and somatic variants. Nat Methods 15(8):591–594. https://doi.org/10.1038/s41592-018-0051-x
McKenna A, Hanna M, Banks E, Sivachenko A, Cibulskis K, Kernytsky A, Garimella K, Altshuler D, Gabriel S, Daly M, DePristo MA (2010) The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res 20(9):1297–1303. https://doi.org/10.1101/gr.107524.110
McLaren W, Gil L, Hunt SE, Riat HS, Ritchie GR, Thormann A, Flicek P, Cunningham F (2016) The ensembl variant effect predictor. Genome Biol 17(1):122. https://doi.org/10.1186/s13059-016-0974-4
AlTassan R, AlQudairy H, Alromayan R, Alfalah A, AlHarbi OA, Gonzalez-Alvarez AC, Arold ST, Kaya N (2022) Clinical radiological and genetic characterization of a patient with a novel homoallelic loss-of-function variant in DNM1 Genes (Basel) 13 12 https://doi.org/10.3390/genes13122252
Yigit G, Sheffer R, Daana M, Li Y, Kaygusuz E, Mor-Shakad H, Altmuller J, Nurnberg P, Douiev L, Kaulfuss S, Burfeind P, Wollnik B, Brockmann K (2022) Loss-of-function variants in DNM1 cause a specific form of developmental and epileptic encephalopathy only in biallelic state. J Med Genet 59(6):549–553. https://doi.org/10.1136/jmedgenet-2021-107769
Ramachandran R, Surka M, Chappie JS, Fowler DM, Foss TR, Song BD, Schmid SL (2007) The dynamin middle domain is critical for tetramerization and higher-order self-assembly. EMBO J 26(2):559–566. https://doi.org/10.1038/sj.emboj.7601491
Asinof S, Mahaffey C, Beyer B, Frankel WN, Boumil R (2016) Dynamin 1 isoform roles in a mouse model of severe childhood epileptic encephalopathy. Neurobiol Dis 95:1–11. https://doi.org/10.1016/j.nbd.2016.06.014
Acknowledgements
We thank the patient’s family for participation in this study. We further thank Henrike Wilshusen and Elena Hamdorf for skilful technical assistance.
Funding
This study was supported by a grant from the Deutsche Forschungsgemeinschaft (KU 1240/13–1 to K.K.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Harms, F.L., Weiss, D., Lisfeld, J. et al. A deep intronic variant in DNM1 in a patient with developmental and epileptic encephalopathy creates a splice acceptor site and affects only transcript variants including exon 10a. Neurogenetics 24, 171–180 (2023). https://doi.org/10.1007/s10048-023-00716-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10048-023-00716-w