Abstract
Nowadays, breast cancer is one of the most widespread malignancies in women, and the second leading cause of cancer death among women. The progesterone receptor (PR) is one of the treatment targets in breast cancer, and can be blocked with selective progesterone receptor modulators (SPRMs). Since administration of chemical drugs can cause serious side effects, and patients, especially those undergoing long-term treatment, can suffer harmful consequences, there is an urgent need to discover novel potent drugs. Large-scale structural diversity is a feature of natural compounds. Accordingly, in the present study, we selected a library of 20,000 natural compounds from the ZINC database, and screened them against the PR for binding affinity and efficacy. In addition, we evaluated the pharmacodynamics and ADMET properties of the compounds and performed molecular docking. Moreover, molecular dynamics (MD) simulation was carried out in order to examine the stability of the protein. In addition, principal component analysis (PCA) was performed to study the motions of the protein. Finally, the MMPBSA method was applied in order to estimate the binding free energy. Our docking results reveal that compounds ZINC00936598, ZINC00869973 and ZINC01020370 have the highest binding energy into the PR binding site, comparable with that of Levonorgestrel (positive control). Moreover, RMSD, RMSF, Rg and H-bond analysis demonstrate that the lead compounds preserve stability in complex with PR during simulation. Our PCA analysis results were in accordance with MD results and the binding free energies support the docking results. This study paves the way for discovery of novel drugs from natural sources and with optimal efficacy, targeting the PR.

The binding mode of new progesterone receptor inhibitors.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Breast cancer (BC) is a deadly malignancy that affects the lives of millions women and their families worldwide, and is the second leading cause of cancer-related deaths in women [1]. BC has attracted growing scientific and clinical interest. A remarkable point is that BC is a clinically heterogeneous disease, associated with a large number of gene mutations. Detection of the mutated genes clarifies the genetic and molecular mechanisms of the disease; furthermore, it significantly increases the chances of finding a successful treatment [2]. In other words, recent findings on the molecular mechanisms of BC could lead to the discovery of new drug candidates [3]. The progesterone receptor (PR), also known as a member of the steroid/nuclear receptor (NR) superfamily, handles a composite network of distinct target genes. Ligand binding to PR leads to a conformational change in the receptor, which activates PR to bind to DNA and regulate transcription [4]. Loss of PR expression is associated with worse overall prognosis and survival among patients with BC.
Current studies have reported that the underlying PR molecular pathway is frequently dysregulated in BC; therefore, it can serve as an efficient therapeutic target [5].
Normally, PR has two isoforms in humans, which differ by structure and function [6]: PRA (~94 kDa) and PRB (~110 kDa). PRA and PRB exert opposite regulatory roles in gene transcription. Barring 164 amino acids at the N-terminus of PRB, the structures of the two isoforms are similar. Both isoforms contain one AF-1 domain with an important function, that of inhibiting the gene transcription. The AF-3 N-terminal domain prevents the function of the AF-1 domain in the PRB isoform [7]. Unlike PRA, PRB induces gene transcription [8]. Actually, PRA plays a role as the inhibitor of PRB and other members of the NR superfamily, such as estrogen receptor (ER) and androgen receptor (AR) [9]. In addition, two structural motifs have been found in PR isoforms: a DNA binding domain (DBD) and a ligand-binding domain (LBD). Tissue-specific expression of progesterone is induced by the function of more than 300 co-regulators that interact with PR.
The modulation of PR activity using a new class of synthetic steroids—selective progesterone receptor modulators (SPRMs)—has become an active area of research in BC treatment. SPRMs exert their effect as PR antagonists or PR agonists (progestins) [10]. Progestins, progesterone antagonists, and SPRMs bind to the PR on the nucleus, allowing a conformational change in the receptor structure. The shape of the receptor changes, enabling DNA to bind to the receptor. Agonists serve as co-activators of gene transcription and antagonists serve as co-repressors, inhibiting gene transcription [11]. Reported in 1981, Mifepristone (RU486) was the first glucocorticoid receptor antagonist and agonist [12]. Subsequently, numerous PR modulators have been described and synthesized [13]. Although considerable research has been devoted to BC, rather less attention has been paid to analysis of PR.
Since natural products have a large number of chiral centers, ring fusion and a higher density of functional groups, allowing for higher ligand affinity and better specificity for biological targets, they have therefore been considered for the treatment of nuclear receptor-mediated diseases such as cancers [14].
Hence, we report an extensive in silico analytical study was carried out in order to perform computational assessment of PR modulation by a large number of diverse ligands derived from natural compounds.
Materials and methods
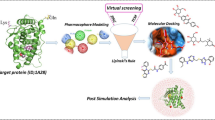
The strategy of the workflow is shown in Fig. 1.
Receptor preparation
The amino acid sequence of PR (Uniprot ID: P06401) was obtained from the UniProt Database (http://www.uniprot.org/) in FASTA format, and was used in BLAST analysis [15]. Then, BLASTP against the protein data bank (PDB) database was performed, in order to acquire a determined PDB structure with the best alignment scores. The X-ray crystal structure of PR (PDB ID: 1SQN, resolution: 1.45 Å) was downloaded from the RCSB website (http://www.rcsb.org). The PDB file had some missing residues, including amino acids 673–681, 706–707, 861, 933 in chain A and amino acids, 674–682, 861 in chain B. Therefore, the MODELLER 9.15 program [16] was utilized to fill these missing residues by treating the original structure as a template. The new protein model was then validated using the SAVES (https://services.mbi.ucla.edu/SAVES/) and RAMPAGE (http://mordred.bioc.cam.ac.uk/~rapper/rampage.php) servers. Subsequently, the protein was simulated through the AMBER03 force field, using GROMACS 5.1.3 software [17] for 100 ns.
Ligands and library preparation
In this study, several PR inhibitors, gathered from the literature, were selected as controls, and their structures were extracted from the PubChem database (https://pubchem.ncbi.nlm.nih.gov/). On the other hand, to explore new anti-cancer drugs, a massive collection of compounds was screened. Before initiating a virtual screening (VS) process, a library of potential ligands ought to be built, derived from the available compound databases. ZINC (http://zinc.docking.org/) is a free database, containing over 35 million commercially available structures. A library consisting of 20,000 compounds structures was selected randomly from the ZINC natural products database. All structures were saved in a mol2 file format, and converted to pdbqt format. Afterwards, all the ligands in pdbqt format were imported into openbabel software [18], and minimized in terms of energy using the MMFF94 force field.
Virtual screening
An integral component of the drug discovery process, VS relies on computer-based methods to design and discover novel drug-like compounds. In the present study, we used VS to determine the affinity of ligands, and appropriate binding mode, for the binding site of PR. All 20,000 selected compounds were docked into the binding cavity of the target PR molecule, through AutoDock Vina [19] in PyRx 0.8, in order to attain optimal complementarity of stereo- and physio-chemical properties. The configuration file was assigned to a grid size of x = 30, y = 30 and z = 30, and a grid Centre of x = 10.83, y = 10.16 and z = 17.3. The number of runs was set at 100 for each ligand. The Lamarckian genetic algorithm (LGA) was applied, whereas the remaining parameters were set to default. The 200 compounds with the best docking scores were selected for further assessment.
ADMET analysis
ADMET (absorption, distribution, metabolism, excretion and toxicological) properties, concomitant with determination of efficacy and clinical safety, perform a crucial role in the early steps of drug design and development. Forming a robust interaction between a drug and its target is necessary for effectiveness; however, the above-mentioned factors must be considered. A clinically effective drug must pass through the body in order to reach its target. ADMET factors identify the potency of the drug to reach its target in the body. However, determination of ADMET for all novel compounds using in vivo methods is a very challenging, expensive and time-consuming task. Hence, in order to forecast drug-like features of designed ligands, we utilized the FAF-Drugs 3 (a free ADME/tox filtering tool) web server [20], which can predict whether the physicochemical properties of drugs are acceptable by applying some filtering rules. The filtering rules for suitable physicochemical properties were set up using the well-known drug-like soft and Lipinski’s rule-of-five, with default values, respectively. All 200 compounds resulting from the previous VS step were subjected to this web server.
Molecular docking analysis
The output of ADMET analysis led to 56 drug-like molecules. In order to narrow down the number of lead compounds, all 56 ligands were subjected to the MTiOpenScreen web server (http://bioserv.rpbs.univ-paris-diderot.fr/services/MTiOpenScreen/), using the same parameters as for PyRx. This web service provides an automated platform, using AutoDock Vina [19], to perform VS. Furthermore, it yields detailed results of the docking between the target PR molecule and the most potent drug-like compounds. Hence, the top ten ligands resulting from the MTiOpenScreen were selected, and then subjected to the AutoDock 4.2 suite [21]. Moreover, the three-dimensional (3D) structures of PR control inhibitors were docked through Autodock 4.2 [21]. AutoDock is an automated docking software programmed for assessment of interactions amongst small molecules, such as ligands or drug-like compounds, and a receptor with known 3D structure. The PR molecule was also prepared by adding polar hydrogens and merging non-polar hydrogens. Kollman united atom charge, and atom type parameters were added. The dimensions of the grids were therefore 90 × 90 × 90 Å, with a spacing of 0.475 Å between the grid points. A grid Centre with x = 10.83, y = 10.16 and z = 17.3 was applied. The parameters were set as follows: a population size of 150, a maximum number of generations of 27,000 and a maximum number of 25,000,000 energy evaluations. Lamarckian genetic search algorithm was applied, and docking runs were set to 100 for each ligand.
Molecular dynamic simulation studies
In this study, molecular dynamic (MD) simulation was performed in two phases. In the first phase, the prepared conformation of PR was used as the initiating structure for MD simulation. The constituents of the simulation system were protein and water. In the second phase, the acquired conformation of PR from the first phase of MD, as well as the three top docked ligands (ZINC00936598, ZINC01020370 and ZINC00869973) and Levonorgestrel, were engaged in the MD simulation process. All MD simulations were performed by the GROMACS 5.1.3 software package, using the AMBER03 force field [17]. Water, ions, proteins and ligands were coupled to a temperature bath. Water molecules were characterized using a simple point charge (SPC216) model. Seven Cl− counter-ions were added, replacing water molecules, in order to guarantee total charge neutrality of the simulated system. The system was located in a box with dimensions 96 × 95 × 100 (all in Å). Periodic boundary conditions (PBC) were exerted in all three directions of the system. The force field parameters of the ligands were acquired from the antechamber module [22] of the Amber program, using the GAFF force field [23]. Partial charges of the ligands were assigned using the AM1-BCC model [24]. Initially, an energy minimization procedure was carried out. After energy minimization, a position restraint process was accomplished in association with NVT and NPT ensembles. An NVT ensemble was implemented at constant temperature of 300 K, using a coupling constant of 0.1 ps for 100 ps. After temperature equilibration, an NPT ensemble was carried out. In this step, a constant pressure of 1 bar was applied, with a coupling constant of 5.0 ps for 1 ns. NPT ensemble was performed after pressure equilibration. In both NVT and NPT ensembles, the coupling scheme of Berendsen was applied. The particle mesh Ewald (PME) method interaction was utilized. The Lincs algorithm for covalent and a 12 Å cutoff for long-range bond constraints were applied. In both phases, MD simulation was performed for 100 ns.
Principal component analysis
In order to reduce the complexity and dimensionality of the data and feature various forms of possible motions in the protein, principal component analysis (PCA) or essential dynamics (ED) was performed [25]. In this method, a covariance matrix is built, with a simple linear transformation in Cartesian coordinate space, after exclusion of translational and rotational movements. Thus, the backbone was selected and least squares fitted to the average structure, using g_covar and g_anaeig modules of GROMACS [26]. Diagonalization of the covariance matrix produced a set of eigenvectors. For every eigenvector, a specific eigenvalue defines the energetic impact of the component on the motion [27]. Porcupine plots were depicted using PyMol [28].
Free energy calculations
The molecular mechanics–Poisson Boltzmann surface area (MM-PBSA) method is pivotal to computing the binding free energy of protein–ligand complexes in the study of the molecular behavior of proteins [29]. In this study, the g_mmpbsa module of GROMACS was applied to estimate the binding free energy of selected complexes [30, 31]. Computation of the binding free energy using this method comprises three steps. In the first step, the potential energy in vacuum is calculated. Subsequently, polar and non-polar solvation energies were estimated, respectively. The non-polar solvation energy was calculated via the solvent-accessible surface area (SASA) model. For all calculations, the last 1000 frames of each trajectory were selected to assess the binding free energy.
Results and discussion
Receptor preparation
As shown in Table 1, the BLAST results represent the highest identity, query cover and score and the lowest E-value and resolution, between PR amino acid sequence and PDB ID: 1SQN. Hence, the PR ligand binding domain with bound norethindrone (PDB ID: 1SQN) was downloaded, from the RCSB website (http://www.rcsb.org). On the basis of the information in the PDB file, there are some missing residues, including amino acids 673–681, 706–707, 861, 933 in chain A and amino acids 674–682, 861 in chain B. Therefore, the missing residues were filled in using MODELLER 9.15 software [16] and specific scripts (Supplementary scripts). As the original structure of the 1SQN PDB file was changed, the prepared PDB file was validated using the SAVES (https://services.mbi.ucla.edu/SAVES/) and RAMPAGE (http://mordred.bioc.cam.ac.uk/~rapper/rampage.php) servers. The results show that the repaired PDB file has an overall quality factor of 89.683, in ERRAT test [32] (Fig. 2a), suggesting crystallographic reliability. An averaged 3D-1D score > = 0.2 in Verify3D test [33] was found for 89.42% of residues, confirming compatibility of the repaired model, and 99.6% of residues were in the favored allowed regions in Ramachandran plot, suggesting acceptable PHI–PSI angles (Fig. 2b), and 3D structure reliability.
Library preparation and virtual screening
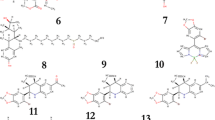
The two-dimensional (2D) structures of 12 PR inhibitors are illustrated in Fig. 3. In the present study, as natural compounds are known to exert preventive effects on nuclear receptor-mediated disease, such as different type of cancers [34], we selected a library of 20,000 natural compounds from the ZINC natural products database.
A structure-based virtual screening (SBVS) was carried out to investigate potential inhibitors amongst the selected library, and to narrow down the number of the possible drug-like candidates. SBVS was implemented over the set of natural compounds by molecular docking using AutoDock Vina [19], in PyRx 0.8. After screening, a set of 2500 compounds was selected by the estimated binding energy; the top 200 ligands were selected for further analysis.
ADMET analysis
The chosen top 200 compounds were scanned for their pharmacokinetics, pharmacodynamics descriptors, and toxicity, through the FAF-Drugs 3 (a free ADME/tox filtering tool) web server [20]. FAF-Drugs 3 predicts a set of descriptors, which are the ADMET and drug-like properties of compounds with probable medicinal properties (http://fafdrugs3.mti.univ-paris-diderot.fr/descriptors.html). Furthermore, the server employs some pre-defined filters, expressing the descriptors reference range, for considering the potency of the drug. In this study, we first applied the Drug-like soft filter to all the top 200 compounds. Among them, 92 compounds were able to successfully pass the filter (Supplementary Table 1). Subsequently, all 92 compounds were subjected to the Lipinski-RO5 filter, and 56 compounds were filtered successfully and selected for further analysis (Table 2).
The descriptors and reference range supplied in this test are explained in this link: http://fafdrugs3.mti.univ-paris-diderot.fr/filters.html.
The ADMET properties for final compounds reveal that these compounds meet drug-like criteria, and can be utilized as robust PR inhibitors.
Molecular docking analysis
Molecular docking analysis was conducted to investigate the drug candidates’ binding mode inside the PR structure. These analyses confirmed predictions suggested to date. In the first step, to narrow down the number of 56 compounds found in the previous stage, all 56 structures were subjected to MTiOpenScreen web server (http://bioserv.rpbs.univ-paris-diderot.fr/services/MTiOpenScreen/) using the same parameters as for PyRx. This web server is capable of performing docking analysis using both AutoDock and AutoDock Vina programs. The top ten compounds with highest estimated binding energy were chosen, and then subjected to AutoDock 4.2 software [21]. Moreover, structures of 12 PR control inhibitors were docked using the Autodock 4.2 program [21]. All 100 runs for each ligand were explored, and the best conformation, regarding binding energy and cluster rank, was considered. The interactions between ligand and PR structure were analyzed and visualized using the MOE (molecular operating environment) software package, version 2013.08 (Chemical Computing Group, Montreal, Canada).
The estimated binding energy, inhibitory constant (Ki) and the key residues involved in diverse interactions discussed below are listed in Tables 3 and 4. The estimated binding energy of these compounds varies between −5.82 and −9.03 kcal mol−1. Moreover, the 2D structures of the ten docked ligands are illustrated in Fig. 4. According to the results of molecular docking, the ZINC00936598 compound has the highest binding energy and the lowest Ki, with ZINC01020370 in second place, followed by ZINC00869973 (Table 4). The results suggest that these compounds have the greatest binding affinity to key PR binding site residues.
Compound ZINC00936598 has the highest negative binding energy (−9.03 kcal mol−1) in the PR binding cavity. The 2D and 3D interaction diagrams of this drug-like compound are illustrated in Fig. 5. As shown in Fig. 5a, the side chain guanidinium moiety of Arg 94 interacts with formamide oxygen through hydrogen binding interaction [35]. In addition, Pro24, Val26, and Phe146 residues have established hydrophobic interactions with an acetonaphthone group of the ligand (Fig. 5a).
The ZINC01020370 interaction study shows formation of hydrogen binding and hydrophobic interactions with the PR binding cleft residues (Fig. 6). The binding energy of this compound is −8.50 kcal mol−1. The side chain guanidinium moiety of Arg94 forms a hydrogen bond with the formamide oxygen of the ligand [36]. Moreover, Pro24, Val26, and Phe146 residues interact with the anisole group of the docked ligand through hydrophobic interactions (Fig. 6a).
As shown in Fig. 7a, the benzene ring of the ZINC00869973 compound was able to establish a Pi-cation (arene-cation) interaction, with the Lys150 side chain amine group, and, concurrently, another benzene ring of the ligand interacts with Trp93 through hydrophobic interaction [37]. The estimated binding energy of the ZINC00869973 compound equals to −8.48 kcal mol−1.
As listed in Table 3, the highest binding energy among the docked control drugs belongs to Levonorgestrel, with a value of −8.35 kcal mol−1. The 2D and 3D interaction diagrams of this drug, with PR binding sites are shown in Fig. 8. As illustrated in Fig. 8, the drug has not succeeded in establishing a hydrogen-binding interaction with any residues of the PR binding cavity. However, Leu110, Ile111, and Leu112 residues interact with Levonorgestrel through hydrophobic interactions (Fig. 8a).
Based on the results presented in Figs. 5–7, and Tables 3 and 4, the ZINC00936598, ZINC01020370, and ZINC00869973 compounds have the strongest affinity toward key PR binding site residues in comparison with the other natural compounds and the best-docked control drug (Levonorgestrel).
Furthermore, studies of the interactions of these natural compounds reveal not only that these compounds have the highest binding energy with key PR binding site residues, but also that the key residues interacting with these ligands were the same in all three cases. These data reflect the significant role of Glu23, Pro24, Asp25, Val26, Ile27, Gln53, Ser56, Val57, Trp60, Leu86, Gly90, Leu91, Trp93, Arg94, Gln143, Phe146 and Lys150 residues in PR inhibition.
Investigation of the interactions of the compound ZINC00057872 (binding energy of −8.41 kcal mol−1; Fig. 9a), reveals that the pyran ring oxygen of the ligand can interact with the side chain of Thr418 through hydrogen bonds. Simultaneously, hydrophobic interactions can be seen between Val473 and the ligand.
As shown in Fig. 10a, the aminobenzamide nitrogen of the ZINC00117321 compound interacts with the backbone of Pro24 through hydrogen bonds, and the aminobenzamide oxygen of the ligand interacts with the side chain of Arg94 via hydrogen bonds [35]. The binding energy of the ligand to the binding site of the PR is −8.20 kcal mol−1.
The 2D and 3D interaction diagrams of the other natural compounds and control drugs with PR binding site residues are depicted in Supplementary Figs. 1–16. As listed in Table 4, compounds ZINC00468904, ZINC00936596, ZINC00130087, ZINC00852626 and ZINC00057677 all have a binding energy less than −8 kcal mol−1. Moreover, the binding regions of these compounds are very far from those with a binding energy of more than −8 kcal mol−1.
Based on the results presented so far, it seems the amino acids Glu23, Pro24, Asp25, Val26, Ile27, Gln53, Ser56, Val57, Trp60, Leu86, Gly90, Leu91, Trp93, Gln143, Phe146, Lys150, and especially Arg94, play a key role in binding of inhibitors to the PR binding site. Our docking results are not consistent with those of previous studies [38, 39], and indicate that the PR structure has a potential ligand-binding site that could serve as an inhibitory pocket. In total, according to the molecular docking results, three top ranked natural compounds (ZINC00936598, ZINC01020370, and ZINC00869973) gave promising results as potential PR inhibitors and are likely to have a noteworthy role in the treatment of BC.
MD simulation studies
The simulation in this study was carried out in two phases. In the first phase, the goal was relaxation of the structure, and to study PR behavior, over a period of 100 ns. The first stage is necessary since, before docking and other stages of the study, the structure has to be energetically optimal and corrected in terms of internal clashes. Subsequently, screening and docking were performed on the output structure of the first phase. In the second phase, MD simulation was performed on the three complexes acquired from docking that had the most negative energy levels, i.e., PR-ZINC00936598, PR-ZINC01020370, PR-ZINC00869973, and the PR-Levonorgestrel complex as a control. The aim was to study the stability and other molecular behaviors of these complexes for 100 ns.
In order to study the stability and conformational changes of the free PR and above-mentioned complexes during MD simulation, root mean square deviation (RMSD) was calculated [35] for backbone atoms of both structures (Fig. 11) and each ligand separately (Fig. 12). As shown in Fig. 11, it can be said that all systems enjoy a level of RMSD between 0.100–0.415 nm during 100 ns of MD simulation. It is obvious that the RMSD of the free PR rises gradually in the first 3 ns; thereafter, no tangible rise in the RMSD rate of the free PR is seen until the end of the simulation, and the structure reaches stability. In the same way, the RMSD of the PR-ZINC00869973 complex enjoys a rise in first 1 ns, thereafter becoming stable until the end of the simulation. Moreover, the RMSD of the other complexes’ elevates for the first 2 ns, and continues almost similarly to the end of the simulation. Therefore, it is seen that all the systems equilibrate along the MD simulation, reflecting the notion that the presence of the ligand and various interactions with PR binding site residues cause no eminent deviation in complex structure during the MD simulation [40]. Moreover, Zheng et al. [41] reported that the PR structure preserves its stability during MD simulation, which is in accordance with our findings. Furthermore, as shown in Fig. 12, the RMSD values of the ligands in each complex are clearly stable from the beginning to the end of the simulation. From these results, it can be concluded that the protein–ligand complexes preserve stability throughout the 100 ns of MD simulation.
Also, in order to study the flexibility of the residues of these structures throughout the simulation, root mean square fluctuation (RMSF) of the apo and complex forms of PR backbone atoms was computed over the entire trajectory [42]. Residues with high RMSF values divulge more flexibility, whereas low RMSF values reflect restrictions on the movement of residues in 100 ns of MD simulation, and thus less flexibility. The rate of RMSF differences is shown in Fig. 13, showing that the great majority of free PR residues have RMSF values <0.25 nm. It can be seen that almost all systems have the same fluctuations in each residue. However, PR-Levonorgestrel and PR-ZINC00869973 complexes fluctuate more than the other complexes in some cases. Furthermore, again it seems that amino acids Glu23, Pro24, Asp25, Val26, Ile27, Gln53, Ser56, Val57, Trp60, Leu86, Gly90, Arg94, Leu91, Trp93, Gln143, Phe146 and Lys150 play roles in binding of ligands to the PR and enjoy less flexibility, suggesting stability of the active site. Our data also suggest that ligand binding to PR, and the bound formation, results in no major changes in flexibility of protein structure residues, confirming the RMSD results. This result supports the findings of a previous study [43].
The level of compactness or folding rate of the backbone atoms during the MD simulation was also evaluated using the radius of gyration (Rg) analysis [44]; Rg studies concentrate on protein structure compactness. As shown in Fig. 14, the free PR and PR-Levonorgestrel complex Rg values declined over the 100 ns MD simulation, whereas in the other complexes the Rg rate remains almost intact. Our data suggest that, while the inhibitor did not remarkably affect the flexibility and the deviation of the PR residues (Figs. 11 and 13), it was not able to induce a noticeable movement in PR structure, and hence protein compactness remains unchanged during MD simulation [45].
Intermolecular interactions of lead compounds with PR were evaluated during the simulation using g_hbond [46]. As depicted in Fig. 15, analysis of PR-Levonorgestrel complex revealed one hydrogen bond during the course of 100 ns simulation and a maximum of two hydrogen bonds. Similarly, in the case of the PR-ZINC01020370 and PR-ZINC00869973 complexes, the results revealed one hydrogen bond and a maximum of three hydrogen bonds during the 100 ns simulation. However, in the case of the PR-ZINC00936598 complex, the results showed at least one hydrogen bond and a maximum of four hydrogen bonds throughout 100 ns simulation. Comparison of H-bond analysis from MD simulations confirmed that the lead compound ZINC00936598 was more potent compared to ZINC01020370 and ZINC00869973.
Principal component analysis
In the present study, in order to better understand the conformational changes, the effect of ligand binding to the overall PR structure motion was investigated using PCA methods. The 2D projection of the first two major principal components (PC1 and PC2) was performed using ED, dividing conformational subspace to essential subspaces [47], for all lead compound-bound PR structures (Fig. 16). As illustrated in Fig. 16, it is obvious that the eigenvectors calculated from the MD trajectory for all systems are quite diverse, clearly demonstrating the difference in structural motion between the systems. It can be concluded that ligand binding to the PR structure resulted in conformational change. Therefore, the less occupied the conformational space in PCA analysis, the greater the inhibitory power of the bound ligand toward the protein structure. Our data are consistent with data reported by Singh et al. [48].
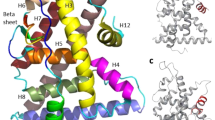
On the other hand, the first and the second principal modes of the lead compound-PR complexes were illustrated using porcupine plots [49]. As shown in Fig. 17, the protein loops and directions of domain motion have been plotted using spines. In the case of the PR-ZINC00936598 complex, it is obvious that chains A and B move downward-right and upward-left, respectively, and these movements in reverse direction were able to strike out the active site of the protein, and to establish more interactions between the ligand and active site residues. Similarly, in the case of PR-ZINC00869973, such motions, the same as PR-ZINC00936598, are more visible and with a sharper angle. Concerning the PR-ZINC01020370 complex, with a slight difference, chain A and chain B move downward and upward, respectively. However, in the case of PR-Levonorgestrel complex, chain A and chain B move counter-clockwise and clockwise, respectively, showing a different pattern of movement than the other complexes. All these motions demonstrate the conformational changes induced by ligand binding to the PR structure [50].
Free energy calculations
In order to investigate the stability and binding of lead compounds after MD simulation, binding free energy of the compounds was estimated by the MM-PBSA method. MM-PBSA is a powerful tool used to study the binding free energy of protein–ligand complexes. As listed in Table 5, Levonorgestrel (−163.041 kJ mol−1) and ZINC00869973 (−105.943 kJ mol−1) compounds, enjoy less binding energy than ZINC01020370 (−62.175 kJ mol−1) and ZINC00936598 (−59.002 kJ mol−1). Similarly, calculated van der Waal energies for Levonorgestrel (−215.132 kJ mol−1) and ZINC00869973 (−201.224 kJ mol−1) compounds were less than those of the other hits, whereas, electrostatic energy values in ZINC00869973 (−60.966 kJ mol−1) and ZINC00936598 (−59.195 kJ mol−1) were lower than the others. In the case of SASA energy, all four compounds have almost the same value. However, in contrast to the other energies, polar solvation energy contributes positively to the total binding free energy value. The strong computed binding energies support our docking results [51].
Conclusions
In this study, we used a wide range of computational methods to examine the effect of plant-derived drug compounds on the structure of PR. VS, ADMET study, molecular docking, MD simulation, PCA and MMPBSA analyses led us to findings that could explain the important effects of these compounds on the structure of PR. Among the 20,000 compounds studied after VS, ADMET and docking, three compounds (ZINC00936598, ZINC01020370 and ZINC00869973) were selected based on docking energies.
The MD simulation for 100 ns of the complex of each of these three compounds with the PR structure showed that, in the RMSD analysis, each of the three complexes was equilibrated and stable. RMSF analysis showed less flexibility in the residues during simulation, and confirmed the stability of common amino acids in the interactions (Glu23, Pro24, Asp25, Val26, Ile27, Gln53, Ser56, Val57, Trp60, Leu86, Gly90, Arg94, Leu91, Trp93, Gln143, Phe146 and Lys150). However, the Rg analysis of all complexes remained intact, indicating the stability of PR compactness during simulation. Additionally, H-bond analysis revealed the greater power of ZINC00936598 in PR inhibition. Furthermore, PCA analysis confirmed that ligand binding to PR structure resulted in conformational changes, and also changes in the direction of different domain movements, in each structure. Finally, the MMPBSA analysis indicated strong ligand binding energies with PR after simulation, confirming the docking results.
References
DeSantis C, Siegel R, Bandi P, Jemal A (2011) Breast cancer statistics, 2011. CA Cancer J Clin. 61(6):408–418. https://doi.org/10.3322/caac.20134
Toss A, Cristofanilli M (2015) Molecular characterization and targeted therapeutic approaches in breast cancer. Breast Cancer Res 17(1):60. https://doi.org/10.1186/s13058-015-0560-9
Gatza ML, Lucas JE, Barry WT, Kim JW, Wang Q, D. Crawford M, B. Datto M, Kelley M, Mathey-Prevot B, Potti A, Nevins JR (2010) A pathway-based classification of human breast cancer. Proc Natl Acad Sci USA 107(15):6994–6999. https://doi.org/10.1073/pnas.0912708107
Grimm SL, Hartig SM, Edwards DP (2016) Progesterone receptor signaling mechanisms. J Mol Biol 428(19):3831–3849. https://doi.org/10.1016/j.jmb.2016.06.020
Abdel-Hafiz HA, Horwitz KB (2012) Control of progesterone receptor transcriptional synergy by SUMOylation and deSUMOylation. BMC Mol Biol 13(1):10. https://doi.org/10.1186/1471-2199-13-10
Anderson E (2002) Progesterone receptors - animal models and cell signaling in breast cancer: the role of oestrogen and progesterone receptors in human mammary development and tumorigenesis. Breast Cancer Res 4(5):197. https://doi.org/10.1186/bcr452
Bain DL, Franden MA, McManaman JL, Takimoto GS, Horwitz KB (2000) The N-terminal region of the human progesterone A-receptor: structural analysis and the influence of the dna binding domain. J Biol Chem 275(10):7313–7320. https://doi.org/10.1074/jbc.275.10.7313
Wetendorf M, Demayo FJ (2014) Progesterone receptor signaling in the initiation of pregnancy and preservation of a healthy uterus. Int J Dev Biol 58(0):95–106. https://doi.org/10.1387/ijdb.140069mw
Li H, Fidler ML, Lim CS (2005) Effect of initial subcellular localization of progesterone receptor on import kinetics and transcriptional activity. Mol Pharm 2(6):509–518. https://doi.org/10.1021/mp0500418
Wagenfeld A, Saunders PTK, Whitaker L, Critchley HOD (2016) Selective progesterone receptor modulators (SPRMs): progesterone receptor action, mode of action on the endometrium and treatment options in gynecological therapies. Expert Opin Ther Targets 20(9):1045–1054. https://doi.org/10.1080/14728222.2016.1180368
Girish C, Jayanthi M, Sivaraman G (2005) Asoprisnil: A selective progesterone receptor modulator. Indian J Pharmacol 37(4):266
Robbins A, Spitz IM (1996) Mifepristone: clinical pharmacology. Clin Obstet Gynecol 39(2):436–450
Spitz IM (2003) Progesterone antagonists and progesterone receptor modulators: an overview. Steroids 68(10–13):981–993. https://doi.org/10.1016/j.steroids.2003.08.007
Buss A (2010) Chiral centers. In: Natural product chemistry for drug discovery. RSC, London, p 37
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215(3):403–410. https://doi.org/10.1016/S0022-2836(05)80360-2
Sanchez R, Sali A (1997) Evaluation of comparative protein structure modeling by MODELLER-3. Proteins Suppl 1:50–58
Berendsen HJC, van der Spoel D, van Drunen R (1995) GROMACS: a message-passing parallel molecular dynamics implementation. Comput Phys Commun 91(1):43–56. https://doi.org/10.1016/0010-4655(95)00042-E
O'Boyle NM, Banck M, James CA, Morley C, Vandermeersch T, Hutchison GR (2011) Open babel: an open chemical toolbox. Journal of Cheminformatics 3(1):33. https://doi.org/10.1186/1758-2946-3-33
Trott O, Olson AJ (2010) AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J Comb Chem 31(2):455–461. https://doi.org/10.1002/jcc.21334
Miteva MA, Violas S, Montes M, Gomez D, Tuffery P, Villoutreix BO (2006) FAF-drugs: free ADME/tox filtering of compound collections. Nucleic Acids Res 34(suppl_2):W738–W744. https://doi.org/10.1093/nar/gkl065
Morris GM, Huey R, Lindstrom W, Sanner MF, Belew RK, Goodsell DS, Olson AJ (2009) AutoDock4 and AutoDockTools4: automated docking with selective receptor flexibility. J Comput Chem 30(16):2785–2791. https://doi.org/10.1002/jcc.21256
Wang J, Wang W, Kollman PA, Case DA (2006) Automatic atom type and bond type perception in molecular mechanical calculations. J Mol Graph Model 25(2):247–260. https://doi.org/10.1016/j.jmgm.2005.12.005
Wang J, Wolf RM, Caldwell JW, Kollman PA, Case DA (2004) Development and testing of a general amber force field. J Comput Chem 25(9):1157–1174. https://doi.org/10.1002/jcc.20035
Jakalian A, Jack DB, Bayly CI (2002) Fast, efficient generation of high-quality atomic charges. AM1-BCC model: II. Parameterization and validation. J Comput Chem 23(16):1623–1641. https://doi.org/10.1002/jcc.10128
David CC, Jacobs DJ (2014) Principal component analysis: a method for determining the essential dynamics of proteins. Methods Mol Biol 1084:193–226. https://doi.org/10.1007/978-1-62703-658-0_11
Kumar A, Rajendran V, Sethumadhavan R, Purohit R (2013) Molecular dynamic simulation reveals damaging impact of RAC1 F28L mutation in the switch I region. PLoS One 8(10):e77453. https://doi.org/10.1371/journal.pone.0077453
C GPD, B R, Chakraborty C, N N, Ali SK, Zhu H (2014) Structural signature of the G719S-T790M double mutation in the EGFR kinase domain and its response to inhibitors. Sci Rep 4:5868. https://doi.org/10.1038/srep05868
The PyMOL Molecular Graphics System, Version 2.0, Schrödinger, LLC
Genheden S, Ryde U (2015) The MM/PBSA and MM/GBSA methods to estimate ligand-binding affinities. Expert Opin Drug Discovery 10(5):449–461. https://doi.org/10.1517/17460441.2015.1032936
Kumari R, Kumar R, Lynn A (2014) g_mmpbsa—a GROMACS tool for high-throughput MM-PBSA calculations. J Chem Inf Model 54(7):1951–1962. https://doi.org/10.1021/ci500020m
Baker NA, Sept D, Joseph S, Holst MJ, McCammon JA (2001) Electrostatics of nanosystems: application to microtubules and the ribosome. Proc Natl Acad Sci USA 98(18):10037–10041. https://doi.org/10.1073/pnas.181342398
Colovos C, Yeates TO (1993) Verification of protein structures: patterns of nonbonded atomic interactions. Protein Sci 2(9):1511–1519. https://doi.org/10.1002/pro.5560020916
Bowie J, Luthy R, Eisenberg D (1991) A method to identify protein sequences that fold into a known three-dimensional structure. Science 253(5016):164–170. https://doi.org/10.1126/science.1853201
Nobili S, Lippi D, Witort E, Donnini M, Bausi L, Mini E, Capaccioli S (2009) Natural compounds for cancer treatment and prevention. Pharmacol Res 59(6):365–378. https://doi.org/10.1016/j.phrs.2009.01.017
Karami M, Jalali C, Mirzaie S (2017) Combined virtual screening, MMPBSA, molecular docking and dynamics studies against deadly anthrax: an in silico effort to inhibit bacillus anthracis nucleoside hydrolase. J Theor Biol 420(Supplement C):180–189. https://doi.org/10.1016/j.jtbi.2017.03.010
Park H, Park SY, Ryu SE (2013) Homology modeling and virtual screening approaches to identify potent inhibitors of slingshot phosphatase 1. J Mol Graph Model 39(Supplement C):65–70. https://doi.org/10.1016/j.jmgm.2012.10.008
Manivannan P, Muralitharan G (2014) Molecular modeling of abc transporter system—permease proteins from Microcoleus chthonoplastes PCC 7420 for effective binding against secreted aspartyl proteinases in Candida albicans—a therapeutic intervention. Interdisciplinary Sciences: Computational Life Sciences 6(1):63–70. https://doi.org/10.1007/s12539-014-0189-x
Sheikh IA (2016) Stereoselectivity and the potential endocrine disrupting activity of di-(2-ethylhexyl)phthalate (DEHP) against human progesterone receptor: a computational perspective. J Appl Toxicol 36(5):741–747. https://doi.org/10.1002/jat.3302
Sarath Josh MK, Pradeep S, Vijayalekshmy Amma KS, Sudha Devi R, Balachandran S, Sreejith MN, Benjamin S (2016) Human ketosteroid receptors interact with hazardous phthalate plasticizers and their metabolites: an in silico study. J Appl Toxicol 36(6):836–843. https://doi.org/10.1002/jat.3221
Jadhav A, Dash R, Hirwani R, Abdin M (2017) Sequence and structure insights of kazal type thrombin inhibitor protein: studied with phylogeny, homology modeling and dynamic MM/GBSA studies. Int J Biol Macromol. https://doi.org/10.1016/j.ijbiomac.2017.11.020
Zheng L, Lin VC, Mu Y (2016) Exploring flexibility of progesterone receptor ligand binding domain using molecular dynamics. PLoS One 11(11):e0165824. https://doi.org/10.1371/journal.pone.0165824
John A, Sivashanmugam M, Umashankar V, Natarajan SK (2016) Virtual screening, molecular dynamics, and binding free energy calculations on human carbonic anhydrase IX catalytic domain for deciphering potential leads. J Biomol Struct Dyn 1–14. https://doi.org/10.1080/07391102.2016.1207565
Shahlaei M, Madadkar-Sobhani A, Mahnam K, Fassihi A, Saghaie L, Mansourian M (2011) Homology modeling of human CCR5 and analysis of its binding properties through molecular docking and molecular dynamics simulation. Biochim Biophys Acta Biomembr 1808(3):802–817. https://doi.org/10.1016/j.bbamem.2010.12.004
Sepehri S, Saghaie L, Fassihi A (2017) Anti-HIV-1 activity prediction of novel Gp41 inhibitors using structure-based virtual screening and molecular dynamics simulation. Molecular Informatics 36(3):1600060. https://doi.org/10.1002/minf.201600060
Fakhar Z, Naiker S, Alves CN, Govender T, Maguire GEM, Lameira J, Lamichhane G, Kruger HG, Honarparvar B (2016) A comparative modeling and molecular docking study on Mycobacterium tuberculosis targets involved in peptidoglycan biosynthesis. J Biomol Struct Dyn 34(11):2399–2417. https://doi.org/10.1080/07391102.2015.1117397
Yang X, Lu J, Ying M, Mu J, Li P, Liu Y (2017) Docking and molecular dynamics studies on triclosan derivatives binding to FabI. J Mol Model 23(1):25. https://doi.org/10.1007/s00894-016-3192-9
Verma S, Singh A, Kumari A, Tyagi C, Goyal S, Jamal S, Grover A (2017) Natural polyphenolic inhibitors against the antiapoptotic BCL-2. J Recept Signal Transduct Res 37(4):391–400. https://doi.org/10.1080/10799893.2017.1298129
Singh SP, Gupta D (2017) Discovery of potential inhibitor against human acetylcholinesterase: a molecular docking and molecular dynamics investigation. Comput Biol Chem 68(Supplement C):224–230. https://doi.org/10.1016/j.compbiolchem.2017.04.002
Zobnina V, Lambreva MD, Rea G, Campi G, Antonacci A, Scognamiglio V, Giardi MT, Polticelli F (2017) The plastoquinol–plastoquinone exchange mechanism in photosystem II: insight from molecular dynamics simulations. Photosynth Res 131(1):15–30. https://doi.org/10.1007/s11120-016-0292-4
Aguayo-Ortiz R, Chavez-Garcia C, Straub JE, Dominguez L (2017) Characterizing the structural ensemble of [gamma]-secretase using a multiscale molecular dynamics approach. Chem Sci 8(8):5576–5584. https://doi.org/10.1039/C7SC00980A
Saadhali SA, Hassan S, Hanna LE, Ranganathan UD, Kumar V (2016) Homology modeling, substrate docking, and molecular simulation studies of mycobacteriophage Che12 lysin A. J Mol Model 22(8):180. https://doi.org/10.1007/s00894-016-3056-3
Acknowledgment
We thank Behbahan Faculty of Medical Sciences for financial support (grant number: 9523).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
None Declared.
Electronic supplementary material
ESM 1
(PDF 5746 kb)
Rights and permissions
About this article
Cite this article
Zarezade, V., Abolghasemi, M., Rahim, F. et al. In silico assessment of new progesterone receptor inhibitors using molecular dynamics: a new insight into breast cancer treatment. J Mol Model 24, 337 (2018). https://doi.org/10.1007/s00894-018-3858-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00894-018-3858-6