Abstract
Haloferax alexandrinus Strain TM JCM 10717T = IFO 16590T is an extreme halophilic archaeon able to produce significant amounts of canthaxanthin. Its genome sequence has been analysed in this work using bioinformatics tools available at Expasy in order to look for genes encoding nitrate reductase-like proteins: respiratory nitrate reductase (Nar) and/or assimilatory nitrate reductase (Nas). The ability of the cells to reduce nitrate under aerobic conditions was tested. The enzyme in charge of nitrate reduction under aerobic conditions (Nas) has been purified and characterised. It is a monomeric enzyme (72 ± 1.8 kDa) that requires high salt concentration for stability and activity. The optimum pH value for activity was 9.5. Effectiveness of different substrates, electron donors, cofactors and inhibitors was also reported. High nitrite concentrations were detected within the culture media during aerobic/microaerobic cells growth. The main conclusion from the results is that this haloarchaeon reduces nitrate aerobically thanks to Nas and may induce denitrification under anaerobic/microaerobic conditions using nitrate as electron acceptor. The study sheds light on the role played by haloarchaea in the biogeochemical cycle of nitrogen, paying special attention to nitrate reduction processes. Besides, it provides useful information for future attempts on microecological and biotechnological implications of haloarchaeal nitrate reductases.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction.
Haloferax alexandrinus was described in 2002 as an extreme halophilic archaea able to produce significant amounts of canthaxanthin (Asker and Ohta 2002a, b), a carotenoid of high interest for several biotechnological uses (Rodrigo-Baños et al. 2015). Shortly after that, lipidic characterisation of the Hfx. alexandrinus strain TM JCM 10717T = IFO 16590T was also reported (Asker et al. 2002). Even taking into account the potential use of this haloarchaea as carotenoid producer, studies about this strain are scarce.
During the last decade, several haloarchaeal genomes have been fully sequenced and annotated. The sequence of the Hfx. alexandrinus strain TM JCM 10717T = IFO 16590T was reported first in 2013 and later modified in 2015 (http://www.ncbi.nlm.nih.gov/genome/16378?genome_assembly_id=176792). More recently, the genome sequence of Hfx. alexandrinus strain Arc-Hr has been published (http://www.ncbi.nlm.nih.gov/genome/16378?genome_assembly_id=204114). Although genomic “era” for archaea started late compared to other organisms, currently it is possible to carry out genomics in parallel to biochemical studies for many of the most representative species of the class Halobacteria, commonly named haloarchaea (Gupta et al. 2015, 2016).
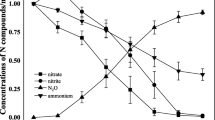
Haloarchaea constitute the main microbial populations in salty environments, and consequently, they play an important role in the main biogeochemical cycles. Nitrogen is a basic element for life, and it accounts for approximately 6% of the dry mass on average. The biogeochemical cycle of nitrogen (N-cycle) makes possible nitrogen interconversions from the most strongly reduced state, as [NH3], in the −3 oxidation state, to the most highly oxidised state, nitrate ion, [NO3]−, in the +5 oxidation state (Richardson and Watmough 1999; Thomson et al. 2012). This cycle is constituted by several pathways with bacteria and archaea playing an important role. Nitrate can be used as nitrogen source for growth under aerobic conditions (assimilatory nitrate reduction) or as final electron acceptor under anaerobic conditions (denitrification) (Bothe et al. 2006).
In nitrate assimilation, first NO3 − is incorporated into the cells by high-/low-affinity transporters and further reduced to NH4 +, via NO2 −, by two sequential reduction reactions catalysed by assimilatory nitrate reductase (Nas; EC 1.6.6.2) and assimilatory nitrite reductase (Nir; EC 1.7.7.1). These two enzymes are located within the cytoplasm. The NH4 + produced is further incorporated into carbon skeletons by the glutamine synthetase/glutamate synthase pathway (GS-GOGAT; EC 6.3.1.2, EC 1.4.7.1, respectively) or via glutamate dehydrogenase (GDH; EC 1.4.1.2) (Martínez-Espinosa et al. 2006; Pire et al. 2014).
Two classes of assimilatory nitrate reductases (Nas) have been described from microorganisms: the ferredoxin- or flavodoxin-dependent Nas and the NADH-dependent enzyme (Moreno-Vivián et al. 1999). The Fd-Nas are usually monomers with a molecular mass between 75 and 85 kDa (Mikami and Ida 1984; Rubio et al. 1996), whilst NADH-Nas proteins are heterodimers of 45 kDa FAD-containing diaphorase and 95 kDa catalytic subunit with molybdenum cofactor and a putative N-terminal [4Fe-4S] centre (Richardson et al. 2001).
Apart from assimilatory nitrate reductases, there are two other types of nitrate reductases-like proteins (Richardson et al. 2001; Sparacino-Watkins et al. 2014): respiratory nitrate reductases (Nar) and dissimilatory nitrate reductases (usually termed Nap). These reductases differ in their cellular location and function: respiratory membrane-bound enzyme (Nar) plays a key role in the generation of metabolic energy by using nitrate as a terminal electron acceptor (nitrate respiration/denitrification) (Richardson et al. 2001; Torregrosa-Crespo et al. 2016). This enzyme is an heterotrimer as well as the periplasmic nitrate reductase (Nap), which participates in the dissipation of excess of reducing power for redox balancing (nitrate dissimilation) (Richardson et al. 2001).
In silico studies revealed that genes encoding the main proteins involved in nitrogen cycle have been found in archaeal genomes (Cabello et al. 2004). However, physiological and biochemical characterisation of such as kind of proteins is still poor in Archaea domain. Particularly, proteins involved in NO3 − reduction to NO2 − related to both, assimilation or denitrification, have only been studied in members of the Haloferax and Haloarcula genera (Yoshimatsu et al. 2000, 2002, 2007; Torregrosa-Crespo et al. 2016; Hattori et al. 2016). Besides, assimilatory nitrate reduction pathway has only been explored in the haloarchaea Hfx. mediterranei at the time of writing this work (Martínez-Espinosa et al. 2001a, b, 2006; Pire et al. 2014; Esclapez et al. 2015).
This work summarises the in silico analysis of the Hfx. alexandrinus strain TM JCM 10717T = IFO 16590T genome looking for the sequences encoding nitrate reductases-like proteins. Biochemical characterisation of the enzyme catalysing nitrate reduction to nitrite (Nas) under aerobic conditions is also reported. This is the second study about Nas (and consequently about assimilatory nitrate reduction) in haloarchaea. The results show that Hfx. alexandrinus is able to use nitrate as sole nitrogen source for growth under aerobic conditions. Potential capability to use nitrate as final electron acceptor (under anaerobic/microaerobic conditions) is also expected.
Materials and methods
Genome analysis
Haloferax alexandrius strain TM JCM 10717T = IFO 16590T genome available at NCBI (http://www.ncbi.nlm.nih.gov/genome/16378?genome_assembly_id=176792) was used to perform in silico analysis with the aim to identify genes coding for nitrate reductase-like proteins. Standard bioinformatics tools available at Expasy portal were used (http://www.expasy.org/) (Gasteiger et al. 2003). Genomics were carried out using ClustalW software for multiple sequence alignment (http://embnet.vital-it.ch/software/ClustalW.html) (Thompson et al. 1997) and BLAST software for biological sequence similarity search and search on protein sequence database (http://blast.ncbi.nlm.nih.gov/Blast.cgi) (Altschul et al. 1990). Protparam (http://web.expasy.org/protparam/) was used to get physical–chemical parameters of the nitrate reductases predicted-like proteins.
Growth conditions
Hfx. alexandrinus strain TM JCM 10717T = IFO 16590T from Japan Collection of Microorganisms was used (RIKEN BioResource Center). The cells were grown in culture media containing the following mixture of salts: (gl−1) 250 NaCl, 20 MgSO4 × 7H2O, 2 KCl, 3 Na3C6H5O7, 0.05 FeSO4 × 7H2O, 0.0002 MnSO4 × H2O and 0.5 KH2PO4 (Asker and Ohta 2002a). This medium also contained glucose and KNO3, 5 and 10 g l−1, respectively. The pH value of the culture medium was adjusted to pH 7.4 using 1 M KOH. Hfx. alexandrinus was grown aerobically at 37 °C in 500-ml batch cultures in 1 L × 20 erlenmeyer flasks using a rotary shaker (New Brunswick innova44) at 180 rpm. Growth was monitored for 10 days measuring the optical density at 600 nm. Nitrite excreted within the media by the cells was quantified using diazo coupling method (Snell and Snell 1949).
Assimilatory nitrate reductase purification
In order to purify Nas, cells were harvested at mid-exponential phase of growth (100 h of incubation) by centrifugation at 30,000g for 20 min in a Beckman Avanti J 30 centrifuge. All the purification steps were carried out at room temperature following the protocol previously described by Martínez-Espinosa et al. (2001b) with some minor changes.
Step 1 Preparation of crude extract. The freshly harvested cells were washed using the mixture of salts previously described and centrifuged at 30,000g for 20 min at room temperature. After that, the cells were resuspended in 50 mM phosphate buffer pH 7.4, containing 2.5 M (NH4)2SO4 (buffer A). The cells were disrupted by sonication (3′ × 8 pulses in ice), and the suspension was centrifuged at 105,000g for 1.5 h at 4 °C. The supernatant was collected and used as the source of enzyme.
Step 2 Sepharose 4B chromatography. The supernatant from the previous step was chromatographed on a Sepharose 4B column (2.5 × 30 cm) equilibrated with buffer A. After introducing the sample, the column was washed with two volumes of buffer A. Elution was carried out with a decreasing linear gradient of 2.5–0.5 M (NH4)2SO4 in 50 mM phosphate buffer pH 7.4 at a flow rate of 48 ml h−1. The total volume of the gradient was 1.5 L. Fractions containing Nas activity were pooled and applied to a DEAE cellulose column.
Step 3 DEAE cellulose chromatography. A DEAE cellulose column (1 × 6 cm) was equilibrated with two column volumes of buffer A. The column was washed using the same buffer at a flow rate of 30 ml/h. The enzyme was eluted with 50 mM phosphate buffer pH 7.4 (buffer B), containing 4.3 M NaCl at a flow rate of 30 ml h−1. Fractions containing Nas activity were pooled and applied to a gel filtration column.
Step 4 Sephacryl S-300 chromatography. Fractions containing Nas activity were loaded on a Sephacryl S-300 column (Pharmacia HiPrep 16/60), previously equilibrated with buffer B containing 2 M NaCl. Buffer B was also used for protein elution (flow rate of 30 ml h−1). After elution, the fractions containing Nas activity (15 ml in total) were immediately dialysed against 100 volumes of 50 mM phosphate buffer pH 7.4, containing 4.3 M NaCl to stabilise the Nas protein (Martínez-Espinosa et al. 2001b).
Protein determination, nitrate reductase assay and enzymatic activity characterisation
The protein content was determined by the Bradford method, with bovine serum albumin (fraction V) as a standard.
Nitrate reductase activity was measured by colorimetric determination of nitrite as previously described. The appearance of nitrite was followed using the diazo coupling method (Snell and Snell 1949; Martínez-Espinosa et al. 2001b).
Nas-specific activity is expressed as nmol of NO2 − appearing per min per mg of protein. Enzymatic activities were explored at different pHs (using phosphate, TRIS–HCl or carbonate/bicarbonate buffers), temperatures ranging from 20 to 90 °C and in the presence of different salt concentrations (0–2 M NaCl or KCl). All the assays were carried out in triplicate and against a control assay without enzyme.
The kinetic results were processed using the Michaelis–Menten equation. The values of V max and K m were determined from the analysis of the corresponding Michaelis–Menten curves using Excel software.
To analyse the effect of several electron donors and inhibitors on the Nas activity, NADH, NADPH, azide, cyanide, EDTA and sulphite were added to the reaction mixture at 1 mM final concentration.
UV–visible spectra from pure protein sample were obtained to identify signals from metal cofactors. The oxidised spectrum was obtained first and the reduced by re-running the same sample after addition of a few crystals of sodium DT (which was used as a reductant reactive).
Gel electrophoresis and estimation of Nas Mr value
The Mr of Nas was estimated by SDS-PAGE taking into account that molecular masses of halophilic proteins are overestimated in SDS-PAGE (around 13–17%) (Johnsen and Schönheit 2004). Molecular mass markers were proved by Sigma (marker M4038).
Results
Haloferax alexandrius strain TM JCM 10717T = IFO 16590T genome is available at NCBI (http://www.ncbi.nlm.nih.gov/genome/16378?genome_assembly_id=176792). This genome is fully sequenced and annotated. Recently, it has been stated that annotation errors are quite common in haloarchaeal genomes (Pfeiffer and Oesterhelt 2015) and nomenclature used is usually confusing. In order to explore potential capability of Hfx. alexandrinus to reduce nitrate, the genome previously mentioned was analysed. Two different sequences encoding nitrate reductases-like proteins were located (Table 1). Both of them are annotated as “nitrate reductases”. Similarities search using Blast and sequences alignments using ClustalW from Expasy stated that one of the sequences (Accession No. ELZ94752.1) corresponds to the respiratory nitrate reductase beta subunit (in charge of the electron transfer during nitrate reduction under anoxic conditions), whilst the other sequence (Accession No. ELZ88427.1) shows the highest similarity to the assimilatory nitrate reductases (in charge of the nitrate reduction to nitrite under aerobic conditions). Sequences coding for the respiratory nitrate reductase alpha subunit (catalytic subunit) were not identified.
Figure 1 displays sequence alignments of the Hfx. alexandrinus ELZ88427.1 sequence and other halophilic assimilatory nitrate reductase-like proteins. It has the best scores with Nas from Hfx. volcanii (99% identity) and with Nas from Hfx. mediterranei (83% identity). Hfx. mediterranei is the only haloarchaea from where assimilatory and respiratory nitrate reductases have been isolated and biochemically characterised up to now (Martínez-Espinosa et al. 2001b; Torregrosa-Crespo et al. 2016).
The N-terminal of the Hfx. alexandrinus protein ELZ88427.1 contains a twin arginine “-RR-”. The twin arginine (“RR”) motif (also termed Tat signal peptide) is involved in proteins translocation to the outside of the cytoplasmic membrane (Maillard et al. 2007). The conserved consensus sequence for this motif (S/T-RR-X-FLK) has been identified in few archaeal respiratory nitrate reductases (Torregrosa-Crespo et al. 2016). However, the N-terminal of the protein ELZ88427.1 is not similar to the consensus Tat signal peptide. Consequently, this protein may be the assimilatory nitrate reductase, a cytoplasmic enzyme reducing nitrate to nitrite aerobically to allow cells growth.
Protparam was use to get physical–chemical parameters of the Hfx. alexandrinus’ ELZ88427.1 sequence, finding that it has 713 amino acidic residues (predicted Mr = 76049.8 Da) from which the total number of negatively charged residues (Asp + Glu) reach 110 against a total number of positively charged residues (Arg + Lys) of 57. Other predicted parameters were: pI: 4.52; instability index (II): 35.15; and aliphatic index: 72.95.
Once it was verified that the genome contains a gene encoding a putative assimilatory nitrate reductase (Nas), cells were grown aerobically in minimal culture media containing 100 mM KNO3 as sole nitrogen source for growth, in order to explore Hfx. alexandrinus capacity to reduce nitrate aerobically.
As it is displayed in Fig. 2, cells were able to grow aerobically using nitrate. Nas activity was detected between 72 and 168 h of incubation, and it reached the maximum value when the absorbance of the culture was around 0.47 (at 600 nm). This maximum activity value was observed shortly after the beginning of the exponential phase of growth, and in that moment, high nitrite concentration within the media was quantified (up to 18.8 mM). This growth phase is characterised by oxygen depletion (culture medium is initially aerobic, but it becomes microaerobic as soon as the biomass increases shortly before the stationary phase of growth) (Hochstein and Lang 1991; Torregrosa-Crespo et al. 2016). Consequently, under these circumstances, the respiratory pathway could also be induced as it has been previously described in Hfx. mediterranei, which is known as a denitrifier (Mancinelli and Hochstein 1986; Torregrosa-Crespo et al. 2016). The nitrite excretion here detected as well as the presence of genes coding for at least three of the four enzymes involved in denitrification indirectly suggests that Hfx. alexandrinus could induce denitrification under microaerobic conditions (see Sect. “Discussion”). The growth rate calculated under these growth conditions was 0.010 ± 0.002 (h−1).
To purify Nas, cells were harvested at the beginning of the exponential phase of growth (where maximum Nas activity was detected under aerobic conditions). The purification scheme is summarised in Table 2. It involves Sepharose 4B, DEAE cellulose, Sephacryl S-300 chromatographies. These protocols were previously tested to purify Nas from Hfx. mediterranei, and they allow successful purifications of halophilic proteins (pure concentrated protein samples in a short period of time with low cost). Nas from Hfx. alexandrinus was purified 70-fold, and the specific activity of purified enzyme was 0.23 U/mg protein. These values are lower than those obtained from Hfx. mediterranei Nas purification (the enzyme was purified 177-fold, and specific activity was 0.55 U/mg protein) (Martínez-Espinosa et al. 2001b). Hfx. alexandrinus Nas activity decreased about 40% in 1 week when the crude extract was stored at temperatures around 4 °C. At temperatures higher than 4 °C, the activity depletion in the crude extract was even more dramatic (60–80%). However, the activity of the pure sample was more stable (2–3 weeks stored at 4 °C). Consequently, it was necessary to start the purification process immediately after getting the crude extract. This pattern was also observed during the Hfx. mediterranei Nas purification, and it could be due to: (1) the action of different proteases, (2) interactions between Nas and other enzymes in the crude extract or (3) the instability of the iron–sulphur clusters and other metallocofactors (MoCo, for instance) in the presence of oxygen.
SDS-PAGE of the purified enzyme showed one band of Mr 72 ± 1.8 kDa (Fig. 3). It is important to highlight that molecular masses of halophilic proteins are usually overestimated by SDS-PAGE. This effect is due to the presence of large amounts of negatively charged amino acids (Johnsen and Schönheit 2004). Taking into account the magnitude of the Mr overestimation (13–17%), a molecular mass of around 70 kDa for Hfx. alexandrinus Nas would be expected. This value correlates with the molecular mass predicted from the protein sequence (Table 1).
Fractions containing Nas activity from DEAE cellulose chromatography were combined and used for the characterisation assays. After DEAE cellulose column, Nas sample was bright brown colour which agrees with those results obtained from other assimilatory nitrate reductases. This colour is mainly due to the presence of Fe-S clusters in the Nas. To confirm the presence of Fe-S clusters in the protein, protein samples from DEAE cellulose as well as pure protein fractions from Sephacryl S-300 were used to get UV–Vis spectra in the fully oxidised and fully reduced forms. In addition to the expected absorbance maximum at 280 nm (due to protein), there was a broad band showing a maximum peak at 404 nm in the fully oxidised protein sample, which is consistent with the presence of Fe-S clusters. These clusters usually exhibit a maximum between 400 and 460 nm. This peak shifted up to 450 nm in the fully reduced protein. These results are similar to those obtained from Hfx. mediterranei Nas (Martínez-Espinosa et al. 2001b).
The effect of several electron donors such as NADH, NADPH or MV on Nas activity was tested. Reduced methylviologen (MV) was the best electron donor (in vitro) for Hfx. alexandrinus Nas, as it was previously described for its homolog from Hfx. mediterranei (Martínez-Espinosa et al. 2001b). Nas from Hfx. alexandrinus did not use electrons from either NADH (1 mM) or NADPH (1 mM mM) (in the presence or absence of DT within the reaction mixture). Dithionite (DT) was not able to reduce nitrate in the absence of MV. These results suggest that Nas from Hfx. alexandrinus could be a ferredoxin-dependent enzyme (Martínez-Espinosa et al. 2001b). Conserved Cys residues that may serve as ligands to Fe atoms (Fe-S clusters) are highlighted in Fig. 1.
Several inhibitors of nitrate reductases were also tested. Dithiothreitol (DTT 1 mM) was not effective as Nas inhibitor (only 2% inhibition was determined compared to the control). Sulphite and EDTA caused 30 and 52% inhibition respectively, at 1 mM final concentration. Azide (1 mM) and cyanide (1 mM) strongly inhibited the enzyme (90 and 98% inhibition, respectively).
pH dependence of enzymatic activity (Fig. 4) as well as the effect of salt concentration (Table 3) on enzymatic activity was also analysed. Optimum pH for activity was slightly alkaline (9.5). The effect of NaCl and KCl at different concentrations (up to 2 M) was studied finding that the highest the salt concentration, the highest the activity value. However, Nas activity was significantly higher in the presence of KCl than in the presence of NaCl (Table 3). Like other halophilic nitrate reductases from genus Haloferax Martínez-Espinosa et al. 2001b), Nas from Hfx. alexandrinus showed a remarkable thermophilicity and worked well up to 50 °C in the presence of high salt concentrations.
Kinetic parameters of halophilic Nas were determined varying the concentration of one substrate (MV) at several fixed concentrations of the other substrate (nitrate), in the presence of 120 mM bicarbonate/carbonate buffer (pH 9.0) containing 1 M NaCl. The halophilic enzyme followed a Michaelis–Menten kinetic. K m values for nitrate and MV were 45 ± 5.2 and 6.46 ± 0.74 µM, respectively. V max values for nitrate and MV were 61.1 ± 3.4 and 19.01 ± 1.7 U/mg prot., respectively. The value of K m for nitrate is under the range of the values obtained from other nitrate reductases (reported K m values are between 0.1 and 1.6 mM) (Alvarez Ossorio et al. 1992; Martínez-Espinosa et al. 2001b).
Discussion
Nitrate cycle in archaea, and in particular in haloarchaea, has been poorly described up to now. Taking into account that haloarchaea constitute the major microbial populations in salty environments, it is worthy to explore how relevant is their contribution in the main biogeochemical cycles. Nevertheless, the nature of the archaeal cells in terms of cell membranes composition, molecular biology machineries, etc, makes it difficult (but at the same time interesting) to study haloarchaeal metabolic pathways from biochemical and molecular biology points of view.
New efforts have been made to sequence haloarchaeal genomes and to improve genome annotations, thus improving current knowledge about this group of extremophiles. The in silico analysis of the Hfx. alexandrinus strain TM JCM 10717T = IFO 16590T genome (which annotation is not completely detailed at the time of writing this work) revealed that there are two genes coding for nitrate reductases-like proteins: assimilatory nitrate reductase (which catalyses the nitrate reduction to nitrite under aerobic conditions) and the beta subunit (also termed NarH) of the respiratory nitrate reductases (which catalyses the reduction of nitrate to nitrite under anaerobic conditions). It was impossible to identify genes coding for the large subunit of the respiratory nitrate reductases (NarG, also called alpha subunit) in Hfx. alexandrinus genome. Potential capacities of Hfx. alexandrinus to carry out nitrate assimilation and nitrate respiration were checked first by in silico searches looking for genes encoding the structural enzymes catalysing both pathways. Genes coding for all the enzymes required to assimilate nitrate (ferredoxin-dependent nitrite reductase: ELZ88359.1; glutamine synthetase: ELZ90622.1; glutamate synthase: ELZ92264.1; glutamate dehydrogenase: ELZ95726.1), as well as most of the enzymes involved in denitrification (copper-containing nitrite reductase: ELZ87995.1; nitric oxide reductase: ELZ88003.1; nitrous oxide reductase accessory protein: WP_006600978.1) have been identified. The presence of genes encoding structural enzymes of denitrification as well as nitrite excretion within the media at the end of the exponential phase of growth under the culture conditions used in this study suggests that Hfx. alexandrinus could be denitrifier. Consequently, this haloarchaea could potentially use nitrate in both senses, as nitrogen source for growth or as final electron acceptor to respire. Regarding denitrification, it remains unclear whether or not there are genes coding for the catalytic subunit of the respiratory nitrate reductase as well as the nitrous oxide reductase (the last enzyme in the denitrification pathway). Genome annotation errors are quite common in haloarchaeal genomes (Pfeiffer and Oesterhelt 2015). Several aspects such as start codon misassignments, disrupted genes as well as poor knowledge based on experimental characterisation of the genes/proteins functions contribute to this persistent problem hampering research in the biosciences related to extreme microbes.
This in silico analysis was the starting point to study assimilatory nitrate reduction in Hfx. alexandrinus. The cells were able to grow aerobically in minimal culture media in the presence of 100 mM nitrate as sole nitrogen source (Fig. 2). These culture conditions were used to purify and characterise the assimilatory nitrate reductase (Nas) from Hfx alexandrinus, which is the first enzyme of the pathway. Nas has been purified as a monomer showing similar biochemical characteristics than those reported from Hfx. mediterranei Nas (Martínez-Espinosa et al. 2001b), in terms of molecular mass, optimal pH for activity and effect of high salt concentration on activity at stability (even in the presence of high temperature). Like other non-halophilic and halophilic nitrate and nitrite reductases, cyanide and azide were strong inhibitors for Hfx. alexandrinus Nas (Alvarez Ossorio et al. 1992; Hochstein and Lang 1991; Ken-Ichi and Hochstein 1996; Moreno-Vivián et al. 1999; Martínez Espinosa et al. 2001a, b). These compounds are thought to be inhibited by metal chelation, and the primary site of action is probably the molybdenum (McDonald and Coddington 1974). Nas activity from Hfx. alexandrinus showed a remarkable thermophilicity working well up to 50 °C in the presence of high salt concentrations (2 M NaCl or KCl), as it was expected taking into account the environmental conditions of the ecosystems inhabited by this haloarchaea. Nas activity was higher in the presence of KCl than in the presence of NaCl under all the conditions assayed, which makes sense taking into account that KCl is the salt that haloarchaea accumulated intracellularly to be isotonic with their environment (Oren 2013). One important feature to be highlighted is that Hfx. alexandrinus Nas has greater affinity for its substrate than Hfx. mediterranei Nas (Martínez-Espinosa et al. 2001b). K m value for nitrate in the case of Hfx. alexandrinus Nas was 0.045 mM, which is approximately 1/21 of the value reported for nitrate from Hfx. mediterranei Nas (Martínez-Espinosa et al. 2001b). However, Hfx. mediterranei grows much better aerobically in the presence of nitrate than Hfx. alexandrinus, as it can be concluded comparing the growth rates of Hfx. alexandrinus (µ = 0.010 ± 0.002 h−1) and Hfx. mediterranei grown aerobically in the presence of nitrate as sole nitrogen source (Martínez-Espinosa et al. 2001a, b).
In conclusion, Hfx. alexandrinus strain TM JCM 10717T = IFO 16590T is able to use nitrate as sole nitrogen source for growth under aerobic conditions thanks to Nas. In 2002, Asker and Ohta (Asker and Ohta 2002a) described this species as a strict aerobe unable to grow anaerobically by using alternative electron acceptors such as nitrate or DMSO, or by fermenting l-arginine. Results from genomics here presented as well as nitrite excretion during the cells growth in the presence of nitrate confirm that Hfx. alexandrinus strain TM JCM 10717T = IFO 16590T may induce denitrification under anaerobic/microaerobic conditions using nitrate as electron acceptor. On the other hand, the same authors detected aerobic reduction of nitrate and nitrite without gas production (Asker and Ohta 2002a), which is consistent with the induction of the assimilatory nitrate pathway. These results about Hfx. alexandrinus strain TM JCM 10717T = IFO 16590T Nas constitute the second study about assimilatory nitrate reduction in haloarchaea providing useful information about haloarchaeal Nas.
Abbreviations
- Nas:
-
Assimilatory nitrate reductase
- Fd-Nas:
-
Ferredoxin assimilatory nitrate reductase dependent
- Nar:
-
Respiratory nitrate reductase
- DT:
-
Dithionite
- DTT:
-
Dithiothreitol
- MV:
-
Methylviologen
References
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Alvarez Ossorio M, Muriana FJG, De La Rosa FF, Relimpio AM (1992) Purification and characterization of nitrate reductase from the halophile archaebacterium Haloferax mediterranei. Z Naturforsch 47c:670–676
Asker D, Ohta Y (2002a) Haloferax alexandrinus sp. nov., an extremely halophilic canthaxanthin-producing archaeon from a solar saltern in Alexandria (Egypt). Int J Syst Evol Microbio 52:729–738
Asker D, Ohta Y (2002b) Production of canthaxanthin by Haloferax alexandrinus under non-aseptic conditions and a simple, rapid method for its extraction. Appl Microbiol Biotechnol 58:743–750
Asker D, Awad T, Ohta Y (2002) Lipids of Haloferax alexandrinus strain TM(T): an extremely halophilic canthaxanthin-producing archaeon. J Biosci Bioeng 93:37–43
Bothe H, Ferguson SJ, Newton WE (2006) Biology of the nitrogen cycle. Elsevier, Amsterdam
Cabello P, Roldán MD, Moreno-Vivián C (2004) Nitrate reduction and the nitrogen cycle in archaea. Microbiology 150:3527–3546
Esclapez J, Pire C, Camacho M, Bautista V, Martínez-Espinosa RM, Zafrilla B et al (2015) Transcriptional profiles of Haloferax mediterranei based on nitrogen availability. J Biotechnol 193:100–107. doi:10.1016/j.jbiotec.2014.11.018
Gasteiger E, Gattiker A, Hoogland C, Ivanyi I, Appel RD, Bairoch A (2003) ExPASy: The proteomics server for in-depth protein knowledge and analysis. Nucleic Acids Res 31:3784–3788
Gupta RS, Naushad S, Baker S (2015) Phylogenomic analyses and molecular signatures for the class Halobacteria and its two major clades: a proposal for division of the class Halobacteria into an emended order Halobacteriales and two new orders, Haloferacales ord. nov. and Natrialbales ord. nov., containing the novel families Haloferacaceae fam. nov. and Natrialbaceae fam. nov. Int J Syst Evol Microbiol 65:1050–1069. doi:10.1099/ijs.0.070136-0
Gupta RS, Naushad S, Fabros R, Adeolu M (2016) A phylogenomic reappraisal of family-level divisions within the class Halobacteria: proposal to divide the order Halobacteriales into the families Halobacteriaceae, Haloarculaceae fam. nov., and Halococcaceae fam. nov., and the order Haloferacales into the families, Haloferacaceae and Halorubraceae fam nov. Antonie Van Leeuwenhoek 109:565–587. doi:10.1007/s10482-016-0660-2
Hattori T, Shiba H, Ashiki K, Araki T, Nagashima YK, Yoshimatsu K et al (2016) Anaerobic Growth of Haloarchaeon Haloferax volcanii by Denitrification Is Controlled by the Transcription Regulator NarO. J Bacteriol 198:1077–1086
Hochstein LI, Lang F (1991) Purification and properties of a dissimilatory nitrate reductase from Haloferax denitrificans. Arch Biochem Biophys 288:380–385
Johnsen U, Schönheit P (2004) Novel xylose dehydrogenase in the halophilic archaeon Haloarcula marismortui. J Bacteriol 186:6198–6207
Ken-Ichi I, Hochstein LI (1996) The purification and properties of a copper nitrite reductase from Haloferax denitrificans. Curr Microbiol 32:72–76
Maillard J, Spronk CA, Buchanan G, Lyall V, Richardson DJ, Palmer T, Vuister GW, Sargent F (2007) Structural diversity in twin-arginine signal peptide-binding proteins. Proc Natl Acad Sci USA 104:15641–15646
Mancinelli RL, Hochstein LI (1986) The occurrence of denitrification in extremely halophilic bacteria. FEMS Microbiol Lett 35:55–58
Martínez-Espinosa RM, Marhuenda-Egea FC, Bonete MJ (2001a) Purification and characterisation of a possible assimilatory nitrite reductase from the halophile archaeon Haloferax mediterranei. FEMS Microbiol Lett 196:113–118
Martínez-Espinosa RM, Marhuenda-Egea FC, Bonete MJ (2001b) Assimilatory nitrate reductase from the haloarchaeon Haloferax mediterranei: purification and characterisation. FEMS Microbiol Lett 204:381–385
Martínez-Espinosa RM, Esclapez J, Bautista V, Bonete MJ (2006) An octameric prokaryotic glutamine synthetase from the haloarchaeon Haloferax mediterranei. FEMS Microbiol Lett 264:110–116
McDonald DW, Coddington A (1974) Properties of assimilatory nitrate reductase from Aspergillus nidulans. Eur J Biochem 46:169–178
Mikami B, Ida S (1984) Purification and properties of ferredoxin-nitrate reductase from the cyanobacterium Plectonema boryanum. Biochim Biophys Acta 791:294–304
Moreno-Vivián C, Cabello P, Martínez-Luque M, Blasco R, Castillo F (1999) Prokaryotic nitrate reduction: molecular properties and functional distinction among bacterial nitrate reductases. J Bacteriol 181:6573–6584
Oren A. (2013) Life at high salt concentrations, intracellular KCl concentrations, and acidic proteomes. Front Microbiol 4:315. doi:10.3389/fmicb.2013.00315
Pfeiffer F, Oesterhelt D (2015) A manual curation strategy to improve genome annotation: application to a set of haloarchael genomes. Life (Basel) 5:1427–1444. doi:10.3390/life5021427
Pire C, Martínez-Espinosa RM, Pérez-Pomares F, Esclapez J, Bonete MJ (2014) Ferredoxin-dependent glutamate synthase: involvement in ammonium assimilation in Haloferax mediterranei. Extremophiles 18:147–159. doi:10.1007/s00792-013-0606-9
Richardson DJ, Watmough NJ (1999) Inorganic nitrogen metabolism in bacteria. Curr Opin Microbiol 3:207–219
Richardson DJ, Berks BC, Russell DA, Spiro S, Taylor C (2001) Functional, biochemical and genetic diversity of prokaryotic nitrate reductases. Cell Mol Life Sci 58:165–178
Rodrigo-Baños M, Garbayo I, Vílchez C, Bonete MJ, Martínez-Espinosa RM (2015) Carotenoids from Haloarchaea and Their Potential in Biotechnology. Mar Drugs 13:5508–5532. doi:10.3390/md13095508
Rubio LM, Herrero A, Flores E (1996) A cyanobacterial narB gene encodes a ferredoxin-dependent nitrate reductase. Plant Mol Biol 30:845–850
Snell CD, Snell CT (1949) Colorimetric Methods of Analysis, 2. Van Nostrand, New York, pp 802–807
Sparacino-Watkins C, Stolz JF, Basu P (2014) Nitrate and periplasmic nitrate reductases. Chem Soc Rev 43:676–706. doi:10.1039/c3cs60249d
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882
Thomson AJ, Giannopoulos G, Pretty J, Baggs EM, Richardson DJ (2012) Biological sources and sinks of nitrous oxide and strategies to mitigate emissions. Philos Trans R Soc Lond B Biol Sci 367:1157–1168. doi:10.1098/rstb.2011.0415
Torregrosa-Crespo J, Martínez-Espinosa RM, Esclapez J, Bautista V, Pire C, Camacho M, Richardson DJ, Bonete MJ (2016) Anaerobic Metabolism in Haloferax Genus: Denitrification as Case of Study. Adv Microb Physiol 68:41–85
Yoshimatsu K, Sakurai T, Fujiwara T (2000) Purification and characterization of dissimilatory nitrate reductase from denitrifying halophilic archaeon Haloarcula marismortui. FEBS Lett 470:216–220
Yoshimatsu K, Iwasaki T, Fujiwara T (2002) Sequence and electron paramagnetic resonance analyses of nitrate reductase NarGH from a denitrifying halophilic euryarchaeote Haloarcula marismortui. FEBS Lett 516:145–150
Yoshimatsu K, Araya O, Fujiwara T (2007) Haloarcula marismortui cytochrome b-561 is encoded by the narC gene in the dissimilatory nitrate reductase operon. Extremophiles 11:41–47
Acknowledgements
This work was funded by research grant from the MINECO Spain (CTM2013-43147-R) and by funds from the Department of Biology, Faculty of Science, Anadolu University (Turkey).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there is no conflict of interest regarding the publication of this paper.
Additional information
Communicated by M. da Costa.
Rights and permissions
About this article
Cite this article
Kilic, V., Kilic, G.A., Kutlu, H.M. et al. Nitrate reduction in Haloferax alexandrinus: the case of assimilatory nitrate reductase. Extremophiles 21, 551–561 (2017). https://doi.org/10.1007/s00792-017-0924-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00792-017-0924-4