Abstract
Two halophilic archaeal strains TBN4T and TBN5 were isolated from Taibei marine solar saltern in Jiangsu, China. Both strains showed light red-pigmented colonies and their cells were rod, motile and Gram-stain-negative. They were able to grow at 25–50°C (optimum 37°C), at 1.4–4.3 M NaCl (optimum 2.1 M NaCl), at 0–1.0 M MgCl2 (optimum 0.005 M MgCl2) and at pH 6.0–9.0 (optimum pH 7.0). Their cells lyse in distilled water and minimal NaCl concentration to prevent cell lysis is 8% (w/v). The major polar lipids of the two strains were PG (phosphatidylglycerol), PGP-Me (phosphatidylglycerol phosphate methyl ester), PGS (phosphatidylglycerol sulfate) and five glycolipids chromatographically identical to S-TGD-1 (sulfated galactosyl mannosyl glucosyl diether), S-DGD-1 (sulfated mannosyl glucosyl diether), TGD-1 (galactosyl mannosyl glucosyl diether), DGD-1 (mannosyl glucosyl diether) and DGD-2 (an unknown diglycosyl diether). Phylogenetic analysis revealed that TBN4T and strain TBN5 formed a distinct clade with genus Haladaptatus (showing 90.0–90.9% 16S rRNA gene similarities). The DNA G + C content of strain TBN4T and strain TBN5 are 66.1 and 65.4 mol%, respectively. The DNA–DNA hybridization value between strain TBN4T and strain TBN5 was 94.3%. The phenotypic, chemotaxonomic and phylogenetic properties suggest that strain TBN4T and strain TBN5 represent a novel species in a new genus within the family Halobacteriaceae, for which the name Halorussus rarus gen. nov., sp. nov. is proposed. The type strain is TBN4T (=CGMCC 1.10122T = JCM 16429T).
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Extremely halophilic archaea, members of the order Halobacteriales, thrive in diverse hypersaline biotopes such as salt lakes, artificial crystallizer ponds of marine solar salterns, salty fermented food and salted hides (Oren 2006). Some special members can also be found in low-salt habitats such as low-salt, sulfide-rich springs and sediments of seashore marshes (Purdy et al. 2004; Savage et al. 2007, 2008). Other than these aerobic members, one species preference for anaerobic conditions was isolated from a deep-sea, hypersaline, anoxic basin (Antunes et al. 2008). The family Halobacteriaceae, the single family described within the order Halobacteriales, currently contains 110 species classified in 29 genera (as of 27 July 2010) (Oren et al. 2009; Cui et al. 2010; Euzéby, 2010). In addition, currently there are 3 new genera and 17 new species as a paper in press online into the International Journal of Systematic and Evolutionary Microbiology. Some of these halophilic archaea were first isolated from thalassohaline hypersaline environments and the others were obtained from athalassohaline environments, showing that both kinds of saline ecosystems are harbor-rich communities of halophilic archaea. In addition to many inland salt lakes, soda lakes and salt mines, more than 100 marine solar salterns, artificial shallow ponds for the production of sea salt from seawater, are located in tropical and subtropical offshore areas along 18,000 km coast of marginal sea at Eastern China (Xu et al. 1999, 2007; Xin et al. 2000; Castillo et al. 2006; Cui et al. 2007; Wang et al. 2007). These solar salterns mainly include three kinds of ponds, evaporation ponds, concentration ponds and crystallizer ponds, in which diverse halotolerant and halophilic microorganisms thrive (Antón et al. 2002). To better understand halophilic archaeal diversity of marine solar salterns of Eastern China, brines and saline soils were collected from marine solar salterns and a medium with low concentration of nutrients was used in cultivating assay. In this study, we characterize two strains, TBN4T and TBN5, as a novel species in a novel genus of the family Halobacteriaceae.
Materials and methods
Origin of the strains and culture condition
Strain TBN4T and strain TBN5 were isolated from brine sampling from Taibei marine solar saltern (34°43′38″N, 119°17′48″E) near Lianyungang city of Jiangsu province, China on 28 August 2008. The brine had a temperature of 25°C, a pH of 7.2 and a total salinity of 285 g/l. The neutral oligotrophic haloarchaeal medium (NOM) used for the isolation procedure contained the following ingredients (g/l): yeast extract 0.05, fish peptone 0.25, sodium pyruvate 1.0, KCl 5.4, K2HPO4 0.3, CaCl2 0.25, NH4Cl 0.25, MgSO4·7H2O 26.8, MgCl2·6H2O 23.0, NaCl 184.0 (pH adjusted to 7.0–7.2 with 1 M NaOH solution) (Cui et al. 2010). The medium was solidified with 2.0% agar. One milliliter brine was added into 9 ml liquid NOM medium, serially diluted with the same medium and then plated onto the NOM agar plates. The plates are incubated in the dark at 37°C for at least 8 weeks in sealed plastic bags. A plate with 50–100 colonies was selected for picking single colonies. To ensure purity, a single colony of each strain was re-streaked twice onto NOM plates. The strains were routinely grown aerobically at 37°C in NOM-3 medium (NOM series medium) with the following modifications (g/l): yeast extract 1.0, fish peptone 0.25, sodium formate 0.25, sodium acetate 0.25, sodium lactate 0.25, sodium pyruvate 0.25.
Phenotypic characterization
Phenotypic tests were performed according to the proposed minimal standards for the description of novel taxa in the order Halobacteriales (Oren et al. 1997). The type strains Haladaptatus paucihalophilus JCM 13897T, Haladaptatus litoreus CGMCC 1.7737T, Halalkalicoccus tibetensis CGMCC 1.3240T, Halalkalicoccus jeotgali JCM 14584T, Halococcus morrhuae CGMCC 1.2153T, Haloferax elongans JCM 14791T were selected as reference strains in positive and negative testing. Cell morphology and motility in exponentially growing liquid cultures were examined using a Nikon microscope equipped with phase-contrast optics (model: E400). Minimal salt concentration to prevent cell lysis was tested by suspending washed cells in serial sterile saline solutions containing NaCl ranging from 0 to 15% (w/v) and the stability of the cells was detected by light microscopic examination.
The Gram stain was performed by following the method outlined by Dussault (1955). Most miscellaneous biochemical tests and nutritional tests were performed as described and proposed by Oren et al. (1997). Briefly, growth and gas formation with nitrate as electron acceptor were tested in 9-ml stoppered tubes, completely filled with liquid NOM medium to which NaNO3 (5 g/l) had been added, and containing an inverted Durham tube. The formation of gas from nitrate was detected by the presence of gas bubbles in Durham tubes and the formation of nitrite was monitored colorimetrically. Anaerobic growth in the presence of l-arginine and DMSO (5 g/l) were tested in completely filled 9-ml stoppered tubes. Starch hydrolysis was determined on NOM agar plates supplemented with 2 g/l soluble starch and detected by flooding the plates with Lugol’s iodine solution. Gelatin hydrolysis was performed by growing colonies on NOM agar plates amended with 1% (w/v) gelatin and flooding the plates with Frazier’s reagent (McDade and Weaver 1959) after growth was established. Esterase activity was detected as outlined by Gutiérrez and González (1972). Tests for catalase and oxidase activities were performed as described by Gonzalez et al. (1978). Production of H2S was tested by growing the isolates and reference strains in a tube with the NOM liquid medium supplemented with 0.5% (w/v) sodium thiosulfate; a filter-paper strip impregnated with lead acetate was used for H2S detection (Cui et al. 2007). To test for growth on single carbon sources, fish peptone and sodium pyruvate were omitted from the NOM medium and the compound to be tested was added at a concentration of 5 g/l. Antibiotic susceptibilities were determined by the Gutiérrez (2008) method on NOM agar plates with antibiotic discs containing the following concentrations (μg per disc, unless otherwise indicated): ampicillin (10), anisomycin (20), aphidicolin (20), bacitracin (0.04 IU per disc), chloramphenicol (30), ciprofloxacin (5), erythromycin (15), gentamicin (10), kanamycin (30), nalidixic acid (30), neomycin (30), norfloxacin (10), novobiocin (30), penicillin G (10 IU per disc), rifampin (5), streptomycin (10), tetracycline (30) and vancomycin (30).
Polar lipids analysis
Tested halophilic archaeal strains were extracted in the wet state directly after harvesting by a modification of the Kates (1986) procedure using a chloroform/methanol system. The fresh cell paste (more than 2.0 g wet weight) was diluted to 20 ml with pure water, and to the suspension in a 250-ml glass-stoppered flask was added 60 ml of chloroform–methanol (1:2, v/v); the mixture was shaken at 150 rpm for 2–3 h at room temperature. To the mixture, 10 ml of water was added, with gentle mixing to form a two-phased system. The lower chloroform phase was withdrawn and brought to dryness in a rotary evaporator (28–33°C). The lipid residue was dissolved in chloroform and made to a known volume with chloroform. The polar lipid was purified out of the above lipid solution by adding about ten volumes of cold acetone and mixing well on the vortexer. The precipitate of polar lipid was centrifuged down and re-washed several times by suspending it in 10-ml portions of cold acetone and centrifuge until the washings were only faintly colored. The acetone-washed precipitate of polar lipid was dried in a desiccator in vacuum. The purified polar lipids were analyzed using one- and two-dimensional TLC, as described previously (Kates 1986). Merck silica gel 60 F254 aluminum-backed thin-layer plates were used in TLC analysis. In two-dimensional TLC, the first solvent was chloroform–methanol-water (65:25:4, by vol.) and the second solvent was chloroform–methanol–acetic acid–water (80:12:15:4, by vol.) which was also used in one-dimensional TLC. Both one- and two-dimensional TLC plates were sprayed with sulfuric acid–ethanol (1:2, by vol.) followed by heating at 150°C for 3 min to detect phospholipids and glycolipids.
Phylogenetic analysis of 16S rRNA genes and determination of G + C content
Genomic DNAs from halophilic archaeal strains were prepared as described by Ng et al. (1995). The 16S rRNA gene was amplified via PCR using primers 0018F and 1518R (Cui et al. 2009). PCR was performed in a thermal cycler (MJ Research PTC-150, USA) for 30 cycles (5 min denaturing step at 95°C in the first cycle, 1 min denaturing at 95°C, 1 min annealing at 60°C and 1.5 min elongation at 72°C, with a final extension step at 72°C for 10 min). The PCR products were examined on a 1.0% (w/v) agarose gel and then cloned into the pEASY-T vector (TransGen Biotech, China) and transformed into Escherichia coli Mach1. Sixteen transformants were randomly picked and sequenced at the Sino-GenoMax Company Limited (Beijing, China) to determine whether the two strains possessed multiple distinct 16S rRNA gene sequences. Multiple sequence alignments were performed using the ClustalW program integrated in MEGA 5 software (Kumar et al. 2008). Phylogenetic trees were reconstructed using the Neighbor-Joining (Saitou and Nei 1987), Maximum-Parsimony (Fitch 1971) and Maximum-Likelihood (Felsenstein 1981) algorithms in MEGA 5 software (http://www.megasoftware.net/). The percentages of replicate trees in which the associated taxa clustered together in the bootstrap test (1,000 replicates) are shown next to the branches. 16S rRNA gene sequence similarity was calculated by comparison with those of related halophilic archaea from the on-line EzTaxon server (Chun et al. 2007). The DNA G + C content was determined by the HPLC method (Mesbah et al. 1989). DNA–DNA hybridization analyses were performed according to the thermal denaturation and renaturation method of De Ley et al. (1970) as modified by Huß et al. (1983), using a Beckman Coulter DU800 spectrophotometer equipped with a high performance temperature controller. The DNA–DNA hybridizations were carried out in 2× SSC at 80°C and the experiment was carried out in triplicate.
Results and discussion
Cells of both strains TBN4T and TBN5 were motile and rod shaped (1–5 by 0.4–0.5 μm) when grown in NOM liquid medium (Fig. 1). Their cells stained Gram-negative and their colonies were light red-pigmented. Strains TBN4T and TBN5 were able to grow at 25–50°C (optimum 37°C both), at 1.4–4.3 M NaCl (optimum 2.1 M NaCl), at pH 6.0–9.0 (optimum pH 7.0) and both strains did not require Mg2+ for growth. Cells lyse in distilled water and the minimal NaCl concentration to prevent cell lysis is 8% (w/v). Both strains produced indole from tryptone and produce H2S from sodium thiosulfate. They hydrolyzed Tween 80, gelatin and casein and weakly hydrolyzed starch. They were able to grow in defined and complex media; d-glucose, d-mannose, d-galactose, maltose, sucrose, lactose, starch, glycerol, acetate, pyruvate, dl-lactate and l-malate as single carbon sources yielded the best growth. Acid was produced during growth on carbohydrates. Strain TBN4 could utilize d-mannitol as single carbon sources, but strain TBN5 could not. More detailed results of phenotypic tests and nutritional features of the two strains are given in the species descriptions.
The polar lipids of the two strains were PG (phosphatidylglycerol), PGP-Me (phosphatidylglycerol phosphate methyl ester), PGS (phosphatidylglycerol sulfate) and five glycolipids chromatographically identical to S-TGD-1 (sulfated galactosyl mannosyl glucosyl diether), S-DGD-1 (sulfated mannosyl glucosyl diether), TGD-1 (galactosyl mannosyl glucosyl diether), DGD-1 (mannosyl glucosyl diether) and DGD-2 (an unknown diglycosyl diether) (Fig. 2; Supplementary Figure S1). The presence of phosphatidylglycerol sulfate (PGS) within our two strains, the members of Haladaptatus (Savage et al. 2007; Cui et al. 2010) and Halalkalicoccus jeotgali JCM 14584T (Table 1) help to differentiate them from related members of Halococcus (Namwong et al. 2007; Oren et al. 2009) and Halalkalicoccus tibetensis CGMCC 1.3240T (Roh et al. 2007; Xue et al. 2005), which do not contain PGS. The glycolipid profile sets strains TBN4T and TBN5 apart from the members of Haladaptatus which contain S-DGD-1, TGD-1 and one unidentified glycolipid (Cui et al. 2010), Halalkalicoccus tibetensis CGMCC 1.3240T which does not contain any glycolipid (Xue et al. 2005), Halalkalicoccus jeotgali JCM14584T which contain S2-DGD and minor unidentified glycolipids (Table 1).
Analysis of the lipid composition of strain TBN4T using two-dimensional TLC, showing the presence of five glycolipids in strain TBN4T. PG phosphatidylglycerol, PGP-Me phosphatidylglycerol phosphate methyl ester, PGS phosphatidylglycerol sulfate, DGD-1 mannosyl glucosyl diether, DGD-2 an unknown diglycosyl diether, S-DGD-1 sulfated mannosyl glucosyl diether, TGD-1 galactosyl mannosyl glucosyl diether, S-TGD-1 sulfated galactosyl mannosyl glucosyl diether, F first dimension of TLC, S second dimension of TLC
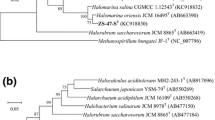
Sixteen complete 16S rRNA gene sequences of strain TBN4T and strain TBN5 were obtained. Sequence comparisons indicated that both strains have one kind of 16S rRNA gene sequence; they are 99.9% similar to each other. Both strains showed low levels of 16S rRNA gene sequence similarity to other members of the family Halobacteriaceae. The closest related recognized species were the members of Halalkalicoccus (90.0–91.3% similarity) and Haladaptatus (90.4–90.9% similarity). Phylogenetic analysis using the Neighbor-Joining (NJ) algorithm revealed that TBN4T and strain TBN5 formed a distinct clade with the members of genus Haladaptatus (Fig. 3). The phylogenetic position was also confirmed in a tree generated using the Maximum-Parsimony and Maximum-Likelihood algorithms (Supplementary Figures S2 and S3). The result based on phylogenetic analysis of 16S rRNA gene reveals that both strains represent a novel phylogenetic taxon.
Neighbor-Joining phylogenetic tree based on 16S rRNA gene sequences showing the relationships between strain TBN4T, strain TBN5 and other close relatives within the family Halobacteriaceae. Bootstrap values (%) are based on 1,000 replicates and are shown for branches with more 70% bootstrap support. Bar 0.02 substitutions per nucleotide position
The DNA G + C content of strain TBN4T and strain TBN5 is 66.1 and 65.4 mol%, respectively. These values are within the range of those of Halococcus (59.5–67.0%) (Namwong et al. 2007; Oren et al. 2009), but higher than the values reported for Haladaptatus (54.0–59.2%) (Savage et al. 2007; Cui et al. 2010) and Halalkalicoccus (61.5–63.2%) (Roh et al. 2007; Xue et al. 2005). The DNA–DNA hybridization value between strain TBN4T and strain TBN5 was 94.3%. The data show that the two strains should be classified in the same species, since the generally accepted threshold value to separate two species is 70% (Stackebrandt and Goebel 1994).
This polyphasic taxonomic study provides evidence that strains TBN4T and TBN5 represent a novel species of a new genus within the family Halobacteriaceae, for which the name Halorussus rarus gen. nov., sp. nov. is proposed. The type strain is TBN4T (=CGMCC 1.10122T = JCM 16429T). Characteristics that distinguish strains TBN4T and TBN5 from other genera within Halobacteriaceae are shown in Table 1.
Description of Halorussus gen. nov
Halorussus (Ha.lo.rus’sus. Gr. masc. n. hals, halos, salt; L. masc. adj. russus red; N.L. masc. n. Halorussus red salt organism)
Cells are rods under optimal growth conditions and stain Gram-negative. Aerobic heterotrophs. Cells lyse in distilled water. Oxidase and catalase tests are positive. Extremely halophilic, with growth occurring in media containing 1.4–4.3 M NaCl. Temperatures between 25 and 50°C and pH between 6.0 and 9.0 may support growth. Sugars are metabolized, in some cases with formation of acids. The polar lipids are PG (phosphatidylglycerol), PGP-Me (phosphatidylglycerol phosphate methyl ester), PGS (phosphatidylglycerol sulfate) and five glycolipids chromatographically identical to S-TGD-1 (sulfated galactosyl mannosyl glucosyl diether), S-DGD-1 (sulfated mannosyl glucosyl diether), TGD-1 (galactosyl mannosyl glucosyl diether), DGD-1 (mannosyl glucosyl diether) and DGD-2 (an unknown diglycosyl diether). The genomic DNA G + C content is between 65.4 and 66.1 mol%. Isolated from marine solar salterns. The type species is Halorussus rarus. Recommended three-letter abbreviation: Hrs.
Description of Halorussus rarus sp. nov
Halorussus rarus (ra’rus. L. masc. adj. rare)
Cells are motile and rod shaped (1.0–5.0 by 0.4–0.5 μm) under optimal growth conditions and stain Gram-negative. Colonies on agar plates containing 2.1 M NaCl are light red, elevated and round. Chemoorganotrophic and aerobic. Growth occurs at 25–50°C (optimum 37°C), at 1.4–4.3 M NaCl (optimum 2.1 M NaCl), at 0–1.0 M MgCl2 (optimum 0.005 M MgCl2) and at pH 6.0–9.0 (optimum pH 7.0). Cells lyse in distilled water and minimal NaCl concentration to prevent cell lysis is 8% (w/v). Catalase- and oxidase-positive. Do not grow under anaerobic conditions with nitrate, arginine and DMSO. Nitrate reduction to nitrite is observed and no gas formation from nitrate. H2S is produced from sodium thiosulfate and positive for indole formation. Tween 80, casein and gelatin are hydrolyzed, starch is hydrolyzed weakly. Urease activity is detected but lysine decarboxylase and ornithine decarboxylase are not produced. PHA (Poly-β-hydroxyalkanoate) is not produced. The following substrates are utilized as single carbon and energy sources for growth: d-glucose, d-mannose, d-galactose, maltose, sucrose, lactose, starch, glycerol, d-mannitol, acetate, pyruvate, dl-lactate, succinate, l-malate and citrate. The following substrates are utilized as single carbon, nitrogen or energy sources for growth: l-alanine, l-aspartate, l-glutamate and l-ornithine. No growth occurred on d-fructose, l-sorbose, d-ribose, d-xylose, d-sorbitol, fumarate, glycine, l-arginine and l-lysine. Acid is produced from d-glucose, d-mannose, d-galactose, sucrose and lactose. Sensitive to the following antibiotics (μg per disc, unless otherwise indicated): novobiocin (30), bacitracin (0.04 IU per disc), anisomycin (20), aphidicolin (20) and rifampin (5). Resistant to the following antibiotics: erythromycin (15), penicillin G (10 IU per disc), ampicillin (10), chloramphenicol (30), neomycin (30), norfloxacin (10), ciprofloxacin (5), streptomycin (10), kanamycin (30), tetracycline (30), vancomycin (30), gentamicin (10) and nalidixic acid (30). The polar lipids are PG (phosphatidylglycerol), PGP-Me (phosphatidylglycerol phosphate methyl ester), PGS (phosphatidylglycerol sulfate) and five glycolipids chromatographically identical to S-TGD-1 (sulfated galactosyl mannosyl glucosyl diether), S-DGD-1 (sulfated mannosyl glucosyl diether), TGD-1 (galactosyl mannosyl glucosyl diether), DGD-1 (mannosyl glucosyl diether) and DGD-2 (an unknown diglycosyl diether). The DNA G + C content of TBN4T is 66.1 mol% (as determined by HPLC).
The type strain is TBN4T (=CGMCC 1.10122T = JCM 16429T), and was isolated from Taibei marine solar saltern near Lianyungang city of Jiangsu province, China.
References
Antón J, Oren A, Benlloch S, Rodríguez-Valera F, Amann R, Rosselló-Mora R (2002) Salinibacter ruber gen. nov., sp. nov., a novel, extremely halophilic member of the Bacteria from saltern crystallizer ponds. Int J Syst Evol Microbiol 52:485–491
Antunes A, Taborda M, Huber R, Moissl C, Nobre MF, da Costa MS (2008) Halorhabdus tiamatea sp. nov., a non-pigmented, extremely halophilic archaeon from a deep-sea, hypersaline anoxic basin of the Red Sea, and emended description of the genus Halorhabdus. Int J Syst Evol Microbiol 58:215–220
Castillo AM, Gutiérrez MC, Kamekura M, Ma YH, Cowan DA, Jones BE, Grant WD, Ventosa A (2006) Halovivax asiaticus gen. nov., sp. nov., a novel extremely halophilic archaeon isolated from Inner Mongolia, China. Int J Syst Evol Microbiol 56:765–770
Chun J, Lee JH, Jung Y, Kim M, Kim S, Kim BK, Lim YW (2007) EzTaxon: a web-based tool for the identification of prokaryotes based on 16S ribosomal RNA gene sequences. Int J Syst Evol Microbiol 57:2259–2261
Cui HL, Lin ZY, Dong Y, Zhou PJ, Liu SJ (2007) Halorubrum litoreum sp. nov., an extremely halophilic archaeon from a solar saltern. Int J Syst Evol Microbiol 57:2204–2206
Cui HL, Zhou PJ, Oren A, Liu SJ (2009) Intraspecific polymorphism of 16S rRNA genes in two halophilic archaeal genera, Haloarcula and Halomicrobium. Extremophiles 13:31–37
Cui HL, Sun FF, Gao X, Dong Y, Xu XW, Zhou YG, Liu HC, Oren A, Zhou PJ (2010) Haladaptatus litoreus sp. nov., an extremely halophilic archaeon from a marine solar saltern, and emended description of the genus Haladaptatus. Int J Syst Evol Microbiol 60:1085–1089
De Ley J, Cattoir H, Reynaerts A (1970) The quantitative measurement of DNA hybridization from renaturation rates. Eur J Biochem 12:133–142
Dussault HP (1955) An improved technique for staining red halophilic bacteria. J Bacteriol 70:484–485
Euzéby J (2010) Notification that new names and new combinations have appeared in volume 60, part 5, of the IJSEM. Int J Syst Evol Microbiol 60:1715–1716
Felsenstein J (1981) Evolutionary trees from DNA sequences: a maximum likelihood approach. J Mol Evol 17:368–376
Fitch WM (1971) Toward defining the course of evolution: minimum change for a specific tree topology. Syst Zool 20:406–416
Gonzalez C, Gutierrez C, Ramirez C (1978) Halobacterium vallismortis sp. nov. an amylolytic and carbohydrate-metabolizing, extremely halophilic bacterium. Can J Microbiol 24:710–715
Gutiérrez C, González C (1972) Method for simultaneous detection of proteinase and esterase activities in extremely halophilic bacteria. Appl Microbiol 24:516–517
Gutiérrez MC, Castillo AM, Kamekura M, Ventosa A (2008) Haloterrigena salina sp. nov., an extremely halophilic archaeon isolated from a salt lake. Int J Syst Evol Microbiol 58:2880–2884
Huß VAR, Festl H, Schleifer KH (1983) Studies on the spectrophotometric determination of DNA hybridization from renaturation rates. Syst Appl Microbiol 4:184–192
Kates M (1986) Techniques of lipidology, 2nd edn. Elsevier, Amsterdam, pp 106–107, 241–246
Kumar S, Dudley J, Nei M, Tamura K (2008) MEGA: a biologist-centric software for evolutionary analysis of DNA and protein sequences. Briefings Bioinf 9:299–306
McDade JJ, Weaver RH (1959) Rapid methods for the detection of gelatin hydrolysis. J Bacteriol 77:60–64
Mesbah M, Premachandran U, Whitman WB (1989) Precise measurement of the G + C content of deoxyribonucleic acid by high-performance liquid chromatography. Int J Syst Evol Bacteriol 39:159–167
Namwong S, Tanasupawat S, Visessanguan W, Kudo T, Itoh T (2007) Halococcus thailandensis sp. nov., from fish sauce in Thailand. Int J Syst Evol Microbiol 57:2199–2203
Ng WL, Yang CF, Halladay JT, Arora P, DasSarma S (1995) Protocol 25: isolation of genomic and plasmid DNAs from Halobacterium halobium. In: DasSarma S, Fleischmann EM (eds) Archaea: a laboratory manual: halophiles. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, pp 179–180
Oren A (2006) The order Halobacteriales. In: Dworkin M, Falkow S, Rosenberg E, Schleifer KH, Stackebrandt E (eds) The prokaryotes, vol 3, 3rd edn. Springer, New York, pp 113–164
Oren A, Ventosa A, Grant WD (1997) Proposed minimal standards for description of new taxa in the order Halobacteriales. Int J Syst Bacteriol 47:233–238
Oren A, Arahal DR, Ventosa A (2009) Emended descriptions of genera of the family Halobacteriaceae. Int J Syst Evol Microbiol 59:637–642
Purdy KJ, Cresswell-Maynard TD, Nedwell DB, McGenity TJ, Grant WD, Timmis KN, Embley TM (2004) Isolation of haloarchaea that grow at low salinities. Environ Microbiol 6:591–595
Roh SW, Nam YD, Chang HW, Sung Y, Kim KH, Lee HJ, Oh HM, Bae JW (2007) Halalkalicoccus jeotgali sp. nov., a halophilic archaeon from shrimp jeotgal, a traditional Korean fermented seafood. Int J Syst Evol Microbiol 57:2296–2298
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Savage KN, Krumholz LR, Oren A, Elshahed MS (2007) Haladaptatus paucihalophilus gen. nov., sp. nov., a halophilic archaeon isolated from a low-salt, sulfide-rich spring. Int J Syst Evol Microbiol 57:19–24
Savage KN, Krumholz LR, Oren A, Elshahed MS (2008) Halosarcina pallida gen. nov., sp. nov., a halophilic archaeon from a low-salt, sulfide-rich spring. Int J Syst Evol Microbiol 58:856–860
Stackebrandt E, Goebel BM (1994) Taxonomic note: a place for DNA–DNA reassociation and 16S rRNA sequence analysis in the present species definition in bacteriology. Int J Syst Bacteriol 44:846–849
Wang QF, Li W, Yang H, Liu YL, Cao HH, Dornmayr-Phaffenhuemer M, Stan-Lotter H, Guo GQ (2007) Halococcus qingdaonensis sp. nov., a halophilic archaeon isolated from a crude sea-salt sample. Int J Syst Evol Microbiol 57:600–604
Xin H, Itoh T, Zhou P, Suzuki K, Kamekura M, Nakase T (2000) Natrinema versiforme sp. nov., an extremely halophilic archaeon from Aibi salt lake, Xinjiang, China. Int J Syst Evol Microbiol 50:1297–1303
Xu Y, Zhou P, Tian X (1999) Characterization of two novel haloalkaliphilic archaea Natronorubrum bangense gen. nov., sp. nov. and Natronorubrum tibetense gen. nov., sp. nov. Int J Syst Bacteriol 49:261–266
Xu XW, Wu YH, Wang CS, Oren A, Zhou PJ, Wu M (2007) Haloferax larsenii sp. nov., an extremely halophilic archaeon from a solar saltern. Int J Syst Evol Microbiol 57:717–720
Xue YF, Fan HP, Ventosa A, Grant WD, Jones BE, Cowan DA, Ma YH (2005) Halalkalicoccus tibetensis gen. nov., sp. nov., representing a novel genus of haloalkaliphilic archaea. Int J Syst Evol Microbiol 55:2501–2505
Acknowledgments
This work was supported by the National Natural Science Foundation of China (No. 30970006) and the opening project of State key Laboratory of Microbial Resources, Institute of Microbiology, Chinese Academy of Sciences (No. SKLMR-20100604).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by A. Oren.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Cui, HL., Gao, X., Yang, X. et al. Halorussus rarus gen. nov., sp. nov., a new member of the family Halobacteriaceae isolated from a marine solar saltern. Extremophiles 14, 493–499 (2010). https://doi.org/10.1007/s00792-010-0329-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00792-010-0329-0