Abstract
Only a few cold-adapted halophilic proteases have been reported. Here, the gene mcp03 encoding a cold-adapted halophilic protease MCP-03 was cloned from deep-sea psychrotolerant bacterium Pseudoalteromonas sp. SM9913, which contains a 2,130-bp ORF encoding a novel subtilase precursor. The recombinant MCP-03, expressed in Escherichia coli BL21 and purified from fermented broth, is a multi-domain protein with a catalytic domain and two PPC domains. Compared to mesophilic subtilisin Carlsberg, MCP-03 had characteristics of a typical cold-adapted enzyme (e.g., higher activity at low temperatures, lower optimum temperature and higher thermolability). MCP-03 also exhibited good halophilic ability with maximal activity at 3 M NaCl/KCl and good stability in 3 M NaCl. Deletion mutagenesis showed that the C-terminal PPC domains were unnecessary for enzyme secretion but had an inhibitory effect on MCP-03 catalytic efficiency and were essential for keeping MCP-03 thermostable.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The average depth of the oceans is about 3,800 m and almost 60% of the earth’s surface is made up of the deep-sea floor (water depths greater than 2,000 m) (Brunnegard et al. 2004). The estimated total input of particulate organic nitrogen to the deep-sea sediment is 24–80 μmol/m2/day and the recycling efficiency of sedimentary particulate organic nitrogen is 94% (Brunnegard et al. 2004). It has been shown that high proteolysis rate exists in deep-sea sediments (Bianchi et al. 2003; Huston and Deming 2002), suggesting that there are abundant bacterial proteases. However, the bacterial proteases in deep-sea sediments are largely unknown. Although the proteases from some deep-sea Pseudoalteromonas (Chen et al. 2003, 2007; Xiong et al. 2007; Kurata et al. 2007) and Pseudomonas strains (Zeng et al. 2003) have been studied, more knowledge of the bacterial proteases involved in sedimentary organic nitrogen degradation is needed to understand the mechanism of organic nitrogen degradation and marine nitrogen recycling. In addition, since deep-sea bacterial proteases are relatively seldom exploited, their further study may find new proteases for future technological applications.
Based on catalytic type, proteases include aspartic, cysteine, metallo, serine, threonine and yet unclassified proteases (Barrett et al. 2004). Peptidase family S8, also known as the subtilisin family or subtilase family, is the second largest family of serine proteases. Subtilases are all characterized by a D/H/S catalytic triad and an alpha/beta-fold catalytic center containing seven-stranded parallel beta-sheets (Rawlings et al. 2006). Although subtilase precursors are usually mosaic proteins, the mature form of subtilases is usually mono-domain (Barrett et al. 2004; Siezen and Leunissen 1997). Some members from Vibrio and its close relatives are mosaic proteins and possess a special C-terminal (C-t) domain required for secretion, which is proteolytically removed upon extracellular activation (Barrett et al. 2004). However, it has been found that some mature subtilases are mosaic proteins with C-t domains, and the function of some C-t domains has been elucidated (Chen et al. 2007; Itoi et al. 2006; Tsujbo et al. 1996; Zhao et al. 2008). For example, the C-t domains of some subtilases with collagenolytic activity were able to bind collagen (Itoi et al. 2006; Zhao et al. 2008).
Pseudoalteromonas sp. SM9913 was isolated from deep-sea sediment as an efficient producer of extracellular proteases. We have been studying the extracellular proteolytic system of strain SM9913 to clarify the role of the strain and its proteases in sedimentary organic nitrogen degradation, the structure and function of the proteases and their adaptation to the deep-sea environment. The proteolytic system of strain SM9913 consists of at least three extracellular proteases (MCP-01, MCP-02 and MCP-03). MCP-01, the most abundant protease secreted by strain SM9913, is a cold-adapted protease (Chen et al. 2003). It is a new type of multi-domain subtilase (named deseasin) with a polycystic kidney disease (PKD) domain at its C-terminus, which can bind insoluble protein, such as collagen (Chen et al. 2007; Zhao et al. 2008). MCP-02 is a cold-adapted metalloprotease belonging to M4 family, and its cold adaptation mechanism was recently studied in detail (Xie et al. 2009). In this study, the gene encoding protease MCP-03 was cloned, sequenced and expressed in E. coli. The recombinant protease was purified and characterized, which showed that it is a cold-adapted multi-domain subtilase with high activity under extreme saline conditions. Moreover, the function of the C-t PPC domains of MCP-03 was studied by deletion mutagenesis.
Materials and methods
Cloning of the gene mcp-03 encoding serine protease MCP-03
The genomic DNA of Pseudoalteromonas sp. SM9913 was prepared according to a previously described method (Chen et al. 2007). Two primers were designed according to the conserved sequences of the serine protease active sites (Maciver et al. 1994), and then a 565-bp DNA fragment was amplified by PCR from the genomic DNA of strain SM9913. With the 565-bp DNA fragment as a probe, one positive clone containing 83.5% of the total protease gene was observed by Southern blotting and colony hybridization using the method described by Taguchi et al (1995). Using TAIL PCR (Thermal Asymmetric Interlaced PCR) (Liu and Whittier 1995), the absent sequence of this gene and the 374 upstream base-pairs were cloned. Through assembly, a 2,504-bp sequence containing a 2,130-bp open reading frame (ORF) encoding MCP-03 was obtained. This gene, named mcp-03, was submitted to GenBank under Accession No. DQ422814.
Plasmids and mutagenesis
The expression plasmid pET22b-MCP03 was constructed by ligation of gene mcp-03 into the EcoRI–XhoI restriction sites of the pET22b plasmid (Invitrogen). Using the pET22b-MCP03 plasmid as a template, the DNA fragments of progressive C-t truncation mutants, BQ1, BQ2, BQ3 and BQ4, were generated by PCR, respectively, using the method described by Pues et al (1997). The generated DNA fragments were subcloned into the vector pET22b for mutant expression.
Expression and purification
All the expression plasmids constructed above were transformed into E. coli BL21-(DE3) competent cells. All the proteins were expressed as C-terminally His6-tagged proteins and purified with a His•Bind metal chelating column. To express active MCP-03 and the mutants from the transformed E. coli BL21, the transformed strain was grown in LB medium containing 50 μg/mL ampicillin at 37°C for 12 h. The culture was diluted 100-fold and grown in LB-amp at 37°C to an OD600 = 1.0. Expression was induced with 0.25 mM IPTG with agitation at 150 rpm at 15°C. After 50 h (when the protease activity in the fermented broth peaked), the fermented broth was centrifuged at 10,000g at 4°C to remove the E. coli BL21 cells. The supernatant was incubated for another 4 h at 15°C to promote protease precursor maturation. The supernatant proteins were precipitated by 65% ammonium sulfate saturation. The dissolved proteins, after dialysis, were added to a His Bind metal chelating column (Novagen) and were eluted with 1 M imidazole. Proteins were pooled, and their purity was analyzed by SDS-PAGE using the Laemmli method (Laemmli 1970). All purification procedures were performed at 0–4°C.
Enzyme activity assay and characterization of MCP-03 and its mutants
The proteolytic activity of recombinant protease MCP-03 and its truncated mutants was assayed using synthetic peptide N-Succinyl-Ala-Ala-Pro-Phe-p-nitroanilide (AAPF, 0.5 mg/mL) as substrate at 45°C for 10 min with Peek’s method (Peek et al. 1993). The optimum pH of MCP-03 was determined using a previously described method (Chen et al. 2003), and the optimum temperature was determined by monitoring MCP-03 activity over 10 min at optimum pH between 0 and 60°C. The thermostability was studied by incubating the enzyme at 50°C and the residual enzyme activity was measured every 5 min. The effects of phenylmethylsulfonyl flouride (PMSF, 10 mM), urea (3 M and 6 M), SDS (1%, w/v), Triton X-100 (1%, v/v), dithiotreitol (10 mM), NaCl and KCl (0–5 M) on MCP-03 activity were investigated by measuring MCP-03 activity at 45°C after MCP-03 was incubated with every agent in 50 mM tris–HCl (pH 8.0) containing 10 mM CaCl2 at 0°C for 30 min. To analyze the substrate specificity of MCP-03, the ability to hydrolyze 10 synthetic peptides (0.5 mg/mL), FAAF (N-Succinyl-Phe-Ala-Ala-Phe-p-nitroanilide, the following peptides were abbreviated in the same way), AAPF, AAPR, AAPL, FVR, AAPK, AAA, AAV, GGG and GGF, was measured at 45°C using Peek’s method (Peek et al. 1993). K m values of MCP-03 and its mutants were determined by Lineweaver–Burk plots, which were made by linear regression with initial rates between 0 and 1 mg/mL of AAPF at 45°C. k cat values of MCP-03 and its mutants were calculated with the formula k cat = V/[E], where V is the reaction rate measured at 45°C and [E] is the enzyme concentration in the reaction mixture. Protein concentration was assessed using the Bradford method (Bradford 1976). The N-terminal (N-t) amino acid sequence of the active MCP-03 protease was determined as previously described (Chen et al. 2007).
Results
Cloning and sequence analysis of gene mcp-03 encoding protease MCP-03
The gene mcp-03 encoding protease MCP-03 from Ps. sp. SM9913 was cloned and sequenced. As shown in Fig. 1, the gene contains a single complete ORF composed of 2,130 bp with an initiation codon of ATG and a termination codon of TAA. A possible TATA-like promoter site (5′-TATAAA-3′) is located 14 bp upstream of the initiation codon for transcription. The gene encodes a polypeptide consisting of 709 amino acids with a calculated molecular mass of 72.6 kDa. BLAST amino acid sequence similarity searches showed that mcp-03 is translated as a subtilisin-like protease precursor.
Nucleotide and deduced amino acid sequences of the MCP-03 precursor. A possible TATA-like promoter site (5′-TATAAA-3′) located 14-bp upstream of the initiation codon. The first 27 amino acid residues underlined are the predicted signal peptide. The first amino acid residue of mature MCP-03 is written in bold and indicated by +1. Three amino acid residues—D43, H102 and S283—forming the catalytic triad are written in bold and boxed. The sequences of two C-t PPC domains are boxed, respectively
The protease precursor deduced from mcp-03 consists of four domains: a signal peptide sequence, an N-t prosequence, a catalytic domain and a C-t extension (Fig. 1). The first 27 N-terminal (N-t) amino acids is a signal peptide sequence identified by the SignalP program (Bendtsen et al. 2004). The five N-t amino acid residues of the purified enzyme were determined to be P1–F2–A3–T4–P5 by Edman degradation sequencing analysis. Based on the C-t sequence of the signal peptide and the N-t sequence of mature MCP-03 protease, the MCP-03 N-t prosequence is composed of 118 amino acid residues (N-118–K-1) and has high identity to the N-t prosequences of other subtilisins, especially in the N1 and N2 motifs that were speculated to be critical for nucleation of the folding process (Shinde et al. 1999). Therefore, the MCP-03 N-t prosequence may function as an intramolecular chaperone to guide correct folding, like in other subtilisins (Shinde et al. 1997, 1999). Amino acid sequence homology analysis of the MCP-03 catalytic domain (P1-S341) with other subtilisin-like proteases indicated that the three amino acid residues (D43, H102 and S280) that likely form the catalytic triad and their surrounding residues are fully conserved (data not shown). Even though MCP-03 has the highest identity (82%) and similar precursor structure with protease AprI secreted by the marine bacterium Alteromonas sp. strain O-7 (Tsujbo et al. 1996), the MCP-03 catalytic domain exhibits low identity (35%–44%) to other subtilisin family members, such as subtilisins BPN′ (44%) (Wells et al. 1983), subtilisins E (42%) (Stahl and Ferrari 1984), subtilisin Carlsberg (41%) (Jacobs et al. 1985) and the psychrophilic subtilisins, S41 (35%) (Davail et al. 1994) and S39 (36%) (Narinx et al. 1992). The C-t extension of MCP-03 precursor is characterized by a repeat of PPC domain (Bacterial Pre-peptidase C-terminal domain), PPC1 + PPC2, which are each composed of 85 amino acid residues. Two PPC domains show 96.4–97.6% and 87.3–92.9% similarity, respectively, to those in the C-t pro-region of several known Gram-negative bacterial proteases in NCBI database (P70765, Q00971, Q9LCJ5 and Q60106).
Characterization of protease MCP-03
Mcp-03 was cloned into pET22b and expressed in E. coli BL21 (DE3). The active form of recombinant MCP-03 protease was purified from the fermented broth with a molecular weight of approximately 58 kDa analyzed by SDS-PAGE (Fig. S1). Based on its N-t sequence and molecular weight, active MCP-03 is composed of the catalytic domain and the two C-t PPC domains.
Since MCP-03 sequence analysis showed it is probably a subtilase of serine proteases, 10 synthetic substrates of subtilases and other serine proteases were selected to analyze MCP-03 specificity. The specific activities of MCP-03 to these substrates were measured and compared with those of subtilisin Carlsberg, the subtilase family archetype. As shown in Table 1, both enzymes very effectively hydrolyzed AAPF. AAPL, AAPK, AAPR, FAAF and FVR were hydrolyzed less effectively; and AAA, AAV, GGF and GGG had hydrolysis below the detection limit of our assay. These results indicated that MCP-03 has broad substrate specificity.
With AAPF as a substrate, MCP-03 had the highest activity at 45°C and retained 14% of its maximal activity at 0°C. In contrast, subtilisin Carlsberg, a mesophilic subtilase, only had ~1% of its maximal activity at 0°C (Fig. 2a). As such, MCP-03 had higher relative activity at low temperatures than subtilisin Carlsberg. In addition, the optimal temperature of MCP-03 against AAPF was about 15°C lower than that of subtilisin Carlsberg. MCP-03 had high thermolability, in which activity dropped rapidly at 50°C (Fig. 2b). These MCP-03 results, compared with subtilisin Carlsberg, indicated that MCP-03 has cold-adapted enzyme characteristics. MCP-03 displayed an alkaline pH activity profile with an optimal pH of 8.0 (Fig. 2c), approximately the pH of seawater, indicating its adaptation to the sea environment.
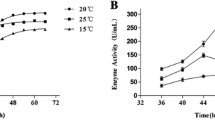
a Effect of temperature on MCP-03 and subtilisin Carlsberg activity. Enzyme activities toward AAPF were measured at pH 8.0, 0–65°C. The specific activity of MCP-03 (4,300 μ/mg) at 45°C and the specific activity of subtilisin Carlsberg (63,484 μ/mg) at 60°C correspond to 100% activity, respectively. b Effect of temperature on the stability of MCP-03 and subtilisin Carlsberg. Enzymes were incubated at 50°C, and the residual activity toward AAPF was measured every 5 min at pH 8.0, 45°C for MCP-03 and 60°C for subtilisin Carlsberg. The specific activities of MCP-03 (4,300 μ/mg) and subtilisin Carlsberg (63,484 μ/mg) without incubation correspond to 100% activity, respectively. c Effect of pH on the activity of MCP-03. The activity of MCP-03 toward AAPF was measured at 45°C in broad pH buffers ranging from pH 3 to pH 11 as previously described (Chen et al. 2003). The specific activity of MCP-03 (4,300 μ/mg) at pH 8.0 corresponds to 100% activity. d Effect of NaCl and KCl on MCP-03 and subtilisin Carlsberg activity. The enzyme activity was measured with AAPF as substrate at 45°C for MCP-03 and 60°C for subtilisin Carlsberg in 20 mM Tris–HCl (pH 8.0) containing NaCl (or KCl) and 10 mM CaCl2. The specific activity of MCP-03 (7,697 μ/mg) in 3 M NaCl and the specific activity of subtilisin Carlsberg (63,484 μ/mg) in 0 M NaCl correspond to 100% activity, respectively
MCP-03 exhibited the highest activity in 3 M NaCl/KCl and retained higher activity in 5 M NaCl than in 0 M NaCl (Fig. 2d). Moreover, MCP-03 was stable for at least 1 month at 4°C in 20 mM Tris–HCl (pH 8.0) containing 3 M NaCl and 10 mM CaCl2 (data not shown). These results showed that MCP-03 is a halophilic protease.
The activity of MCP-03 was completely inhibited by 10 mM PMSF. It was sensitive to urea (3–6 M), 1% SDS and 1% Triton X-100, indicating that these protein denaturants could cause partial denaturation of MCP-03. As much as 10 mM dithiotreitol, a thiol-reducing agent, inhibited MCP-03 activity by 50% (Table 2).
Analysis of the MCP-03 C-t PPC domain function by deletion mutagenesis
In order to study the function of the C-t extension of MCP-03, an extensive C-t truncation analysis by deletion mutagenesis was undertaken. As shown in Fig. 3a, the mutant BQ1 was lacking the PPC2 domain at the C-terminus; BQ2 and BQ3 were lacking PPC2 and a part of PPC1 domain; BQ4 was lacking both PPC1 and PPC2. MCP-03 and the mutants, BQ1 to pBQ4, were expressed as C-terminally His6-tagged proteins and purified directly from fermented broth (Fig. 3b), suggesting that the PPC domains in the C-t extension of MCP-03 are unnecessary for its secretion through the E. coli membrane. With AAPF as a substrate, the K m values of the recombinant enzyme and all mutants were similar (Table 3). However, the PPC domains had inhibitory effects on MCP-03 catalytic efficiency, because the k cat/K m value of MCP-03, which has two PPC domains in its C terminus, was much lower than that of the mutants containing one or no PPC domain (Table 3). The thermal stability experiment revealed that MCP-03 had higher thermostability than any mutant at 50°C, and the thermostability of pBQ1 with one PPC domain was higher than any other mutant with a partial or no PPC domain (Table 3). These results suggested that the MCP-03 PPC domains may play a role in enzyme thermostability.
a Schematic diagram of MCP-03 and its C-terminal truncated mutants. The number at the left and the right of the schematic diagram represents the location of the first and the last amino acid residue of the MCP-03 precursor sequence shown in Fig. S1. b The purified active enzymes of MCP-03 and its C-terminal truncated mutants of MCP-03 analyzed by 12.5% SDS-PAGE. Lane 1, MCP-03; lane 2, BQ1; lane 3, BQ2; lane 4, BQ3; lane 5, BQ4
Discussion
Most deep-sea environments are extreme, influenced by low temperature, low nutrient concentration, high hydrostatic pressure, moderate salinity and fluidity. In order to adapt to such a harsh environment, bacterial proteases may have special structures and functions to carry out protein degradation. For example, the protease secreted by deep-sea bacterium Pseudomonas strain DY-A and protease MCP-01 from deep-sea strain SM9913 are cold-adapted enzymes (Zeng et al. 2003; Chen et al. 2003). Protease MCP-01 has a C-t PKD domain, which can bind to insoluble proteins to facilitate protein degradation by MCP-01 (Zhao et al. 2008). Exploration of the deep-sea proteases may not only find novel proteases with special structures and functions, but may also be helpful in understanding deep-sea nitrogen recycling.
In this study, the gene mcp03 encoding protease MCP-03 from deep-sea cold-adapted bacterium Pseudoalteromonas sp. SM9913 was cloned and expressed in E. coli, and the active enzyme of recombinant MCP-03 was purified and characterized. Sequence analysis of MCP-03 showed that it is a novel subtilase. The recombinant MCP-03 is a multi-domain subtilase with two PPC domains at its C-terminus. Compared to mesophilic subtilisin Carlsberg, MCP-03 had typical characteristics of a cold-adapted enzyme including higher activity at low temperatures, lower optimum temperature and higher thermolability, indicating that it is a cold-adapted enzyme. Subtilisin Carlsberg is the archetype of subtilase family (Barrett et al. 2004). Most subtilases are nonspecific peptidases with a preference for an aromatic amino acid residue at the P1 position (Siezen and Leunissen 1997). MCP-03 has broad substrate specificity. The broad substrate specificity of MCP-03 is beneficial for strain SM9913 to adequately utilize the surrounding deep-sea proteins. MCP-03 displayed alkaline pH activity profile (pH 7.0 to 9.0) with an optimal pH of 8.0, approximately the pH of the seawater. Moreover, MCP-03 is a halophilic protease with the highest activity in 3 M NaCl/KCl. These results suggest adaptation of MCP-03 to the alkaline, cold, saline and low-nutrient deep-sea environment. Although many halophilic proteases from archaea and bacteria have been reported (Mellado et al. 2005), only a few are cold-adapted halophilic proteases (Xiong et al. 2007). These cold-adapted halophilic proteases may aid in processing of saline food, such as seafood.
The PPC domain is found in some members of metalloprotease families M4, M9 and M28 as well as serine protease family S8 (Barrett et al. 2004). The PPC domains are usually cleaved after secretion but prior to protease activation (Yeats et al. 2003). There are few mature proteases containing PPC domains. While their actual function is not clear, they may aid secretion/localization or inhibit the protease until needed (Yeats et al. 2003). The PPC domain of protease XCEXPR is reportedly required for secretion through the E. coli JM109 outer membrane (Liu et al. 1990). Proteases AprI and AprII with C-t PPC domains show lower activity than the matured enzymes without PPC domains (Tsujbo et al. 1996). The homologous C-t PPC domain of V. vulnificus metalloprotease is essential for efficient attachment to insoluble protein substrates and erythrocyte membranes (Miyoshi et al. 1997). In this paper, the mature recombinant MCP-03 enzyme has been shown to have a catalytic domain and two PPC domains at its C-terminus. In order to study the effect of these PPC domains on MCP-03, a series of deletion mutants were designed and expressed. The MCP-03 C-t PPC domains are unnecessary for secretion through the E. coli membrane, similar to V. vulnificus metalloprotease (Miyoshi et al. 1997). Moreover, the MCP-03 C-t PPC domains had inhibitory effect on catalytic efficiency since the mutant without any PPC had higher catalytic efficiency than MCP-03 and the mutant with one PPC. This is consistent with the effect of PPC on the activity of proteases AprI and AprII (Tsujbo et al. 1996). In addition, the MCP-03 PPC domains might aid enzyme thermostability because MCP-03 and the mutant with one PPC are more thermostable than the mutants lacking PPC.
Abbreviations
- C-t:
-
C-terminal
- N-t:
-
N-terminal
- PPC domain:
-
Bacterial pre-peptidase C-terminal domain
- PKD domain:
-
Polycystic kidney disease domain
- FAAF:
-
N-Succinyl-Phe-Ala-Ala-Phe-p-nitroanilide
- AAPF:
-
N-Succinyl-Ala-Ala-Pro-Phe-p-nitroanilide
- AAPR:
-
N-Succinyl-Ala-Ala-Pro-Arg-p-nitroanilide
- AAPL:
-
N-Succinyl-Ala-Ala-Pro-Leu-p-nitroanilide
- FVR:
-
N-Benzoyl-Phe-Val-Arg-p-nitroanilide
- AAPK:
-
N-Succinyl-Ala-Ala-Pro-Lys-p-nitroanilide
- AAA:
-
N-Succinyl-Ala-Ala-Ala-p-nitroanilide
- AAV:
-
N-Succinyl-Ala-Ala-Val-p-nitroanilide
- GGG:
-
N-Succinyl-Gly-Gly-Gly-p-nitroanilide
- GGF:
-
N-Succinyl-Gly-Gly-Phe-p-nitroanilide
References
Barrett AJ, Rawlings ND, Woessner JF (2004) Handbook of proteolytic enzymes, 2nd edn. Elsevier press, London
Bendtsen JD, Nielsen H, vonHeijne G, Brunak S (2004) Improved prediction of signal peptides: SignalP 3.0. J Mol Biol 340:783–795
Bianchi A, Calafat A, Wit RD, Garcin J, Tholosan O, Cacho I, Canals M, Fabrés J, Grout H, Masqué P, Sanchez-Cabeza J-A, Sempéré R (2003) Microbial activity at the deep water sediment boundary layer in two highly productive systems in the Western Mediterranean: the Almeria-Oran front and the Malaga upwelling. Oceanologica Acta 25:315–324
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254
Brunnegard J, Grandel S, Stahl H, Tengberg A, Hall POJ (2004) Nitrogen cycling in deep-sea sediments of the Porcupine Abyssal Plain, NE Atlantic. Prog Oceanogr 63:159–181
Chen X-L, Zhang Y-Z, Gao P-J, Luan X-W (2003) Two different proteases produced by a deep-sea psychrotrophic strain Pseudoaltermonas sp. SM9913. Mar Biol 143:989–993
Chen X-L, Xie B-B, Lu J-T, He H-L, Zhang Y-Z (2007) A novel type of subtilase from the psychrophilic bacterium Pseudoalteromonas sp. SM9913: catalytic and structural properties of deseasin MCP-01. Microbiol-SGM 153:2116–2125
Davail S, Feller G, Narinx E, Gerday C (1994) Cold adaptation of proteins. Purification, characterization, and sequence of the heat-labile subtilisin from the Antarctic psychrophile bacillus TA41. J Biol Chem 269:17448–17453
Huston AL, Deming JW (2002) Relationships between microbial extracellular enzymatic activity and suspended and sinking particulate organic matter: seasonal transformations in the North Water. Deep-Sea Res II 49:5211–5225
Itoi Y, Horinaka M, Tsujimoto Y, Matsui H, Watanabe K (2006) Characteristic features in the structure and collagen-binding ability of a thermophilic collagenolytic protease from the thermophile Geobacillus collagenovorans MO-1. J Bacteriol 188:6572–6579
Jacobs M, Eliasson M, Uhlen M, Flock JI (1985) Cloning, sequencing and expression of subtilisin Carlsberg from Bacillus licheniformis. Nucleic Acids Res 13:8913–8926
Kurata A, Miyazaki M, Kobayashi T, Nogi Y, Horikoshi K (2007) Alkalimonas collagenimarina sp. nov., a psychrotolerant, obligate alkaliphile isolated from deep-sea sediment. Int J Syst Evol Microbiol 57:1549–1553
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227:680–685
Liu YG, Whittier RF (1995) Thermal asymmetric interlaced PCR: automatable amplification and sequencing of insert end fragments from P1 and YAC clones for chromosome walking. Genomics 25:674–681
Liu Y-N, Tang J-L, Clark BR, Dow JM, Danniels MJ (1990) A multipurpose broad host range cloning vector and its use to characterise an extracellular protease gene of Xanthomonas campestris pathovar campestris. Mol Gen Genet 220:433–440
Maciver B, Mchale RH, Saul DJ, Bergquist PL (1994) Cloning and sequencing of a serine proteinase gene from a thermophilic bacillus species and its expression in Escherichia coli. Appl Environ Microbiol 60:3981–3988
Mellado E, Sánchez-Porro C, Ventosa A (2005) Proteases produced by halophilic bacteria and archaea. Microbial Enz Biotrans 17:181–190
Miyoshi S, Wakae H, Tomochika K, Shinoda S (1997) Functional domains of a zinc metalloprotease from Vibrio vulnificu. J Bacteriol 23:7606–7609
Narinx E, Davail S, Feller G, Gerdy C (1992) Nucleotide and derived amino acid sequence of the subtilisin from the Antarctic psychrotroph Bacillus TA39. Biochim Biophys Acta 113:111–113
Peek K, Veitch DP, Prescott M, Daniel RM, MacIver B, Bergquist PL (1993) Some characteristics of a proteinase from a thermophilic Bacillus sp. expressed in Escherichia coli: comparison with the native enzyme and its processing in E. coli and in vitro. Appl Environ Microbiol 59:1168–1175
Pues H, Holz B, Weinhold E (1997) Construction of a deletion library using a mixture of 5′-truncated primers for inverse PCR (IPCR). Nucleic Acids Res 25:1303–1304
Rawlings ND, Morton FR, Barrett AJ (2006) MEROPS: the peptidase database. Nucleic Acids Res 34(Database Issue):D270–D272
Shinde UP, Liu JJ, Inouye M (1997) Protein memory through altered folding mediated by intramolecular chaperones. Nature 389:520–522
Shinde UP, Fu X, Inouye M (1999) A pathway for conformational diversity in proteins mediated by intramolecular chaperones. J Biol Chem 274:15615–15621
Siezen RJ, Leunissen JAM (1997) Subtilases: the superfamily of subtilisin-like proteases. Protein Sci 6:501–523
Stahl ML, Ferrari E (1984) Replacement of the Bacillus subtilis subtilisin structural gene with an in vitro-derived deletion mutation. J Bacteriol 158:411–418
Taguchi S, Odaka A, Watanabe Y, Momose H (1995) Molecular characterization of a gene encoding extracellular serine protease isolated from a subtilisin inhibitor-deficient mutant of Streptomyces albogriseolus S-3253. Appl Environ Microbiol 61(1):180–186
Tsujbo H, Miyamoto K, Tanaka K, Kaidzu Y, Imada C, Okami Y, Inamori Y (1996) Cloning and sequence analysis of a protease-encoding gene from the marine bacterium Alteromonas sp. strain O-7. Biosci Biotechnol Biochem 60:1284–1288
Wells JA, Ferrari E, Henner DJ, Estell DA, Chen EY (1983) Cloning, sequencing, and secretion of Bacillus amyloliquefaciens subtilisin in Bacillus subtilis. Nucleic Acids Res 11:7911–7925
Xie BB, Bian F, Chen XL, He HL, Guo J, Gao X, Zeng YX, Chen B, Zhou BC, Zhang YZ (2009) Cold adaptation of zinc metalloproteases in the thermolysin family from deep sea and Arctic Sea ice bacteria revealed by catalytic and structural properties and molecular dynamics: new insights into relationship between conformational flexibility and hydrogen bonding. J Biol Chem 284:9257–9269
Xiong H, Song L, Xu Y, Tsoi M-Y, Dobretsov S, Qian P-Y (2007) Characterization of proteolytic bacteria from the Aleutian deep-sea and their proteases. J Ind Microbiol Biotechnol 34:63–71
Yeats C, Bentley S, Bateman A (2003) New knowledge from old: In silico discovery of novel protein domains in Streptomyces coelicolo. BMC Microbiol 3:1–20
Zeng R, Zhang R, Zhao J, Lin N (2003) Cold-active serine alkaline protease from the psychrophilic bacterium Pseudomonas strain DY-A: enzyme purification and characterization. Extremophiles 7:335–337
Zhao G-Y, Chen X-L, Zhao H-L, Xie B-B, Zhou B-C, Zhang Y-Z (2008) Hydrolysis of insoluble collagen by deseasin MCP-01 from deep-sea Pseudoalteromonas sp. SM9913. J Biol Chem 283:36100–36107
Acknowledgments
The work was supported by the Hi-Tech Research and Development program of China (2006AA09Z414, 2007AA091903), the National Natural Science Foundation of China (30770040), the Program for New Century Excellent Talents in University (NCET-06-0578), the Foundation for Young Excellent Scientists in Shandong Province (2006BS02002), and the COMRA Program (DYXM-115-02-2-6).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by F. Robb.
X.-L. Chen and B.-Q. Yan contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Yan, BQ., Chen, XL., Hou, XY. et al. Molecular analysis of the gene encoding a cold-adapted halophilic subtilase from deep-sea psychrotolerant bacterium Pseudoalteromonas sp. SM9913: cloning, expression, characterization and function analysis of the C-terminal PPC domains. Extremophiles 13, 725–733 (2009). https://doi.org/10.1007/s00792-009-0263-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00792-009-0263-1