Abstract
There is a great interest in xylanases due to the wide variety of industrial applications for these enzymes. We cloned a xylanase gene (xyn8) from an environmental genomic DNA library. The encoded enzyme was predicted to be 399 amino acids with a molecular weight of 45.9 kD. The enzyme was categorized as a glycosyl hydrolase family 8 member based on sequence analysis of the putative catalytic domain. The purified enzyme was thermolabile, had an activity temperature optimum of 20°C on native xylan substrate, and retained significant activity at lower temperatures. At 4°C, the apparent K m was 3.7 mg/ml, and the apparent k cat was 123/s.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Hemicellulose is the second most common component of biomass and thus represents a large source of renewable substrate (Ward and Moo-Young 1989). By developing technologies, such as enzymatic hydrolysis, to better utilize this material, the cost efficiency of transitioning to a more bio-based economy will be improved. Hemicellulose is composed primarily of xylan, a β-1,4-linked polymer of xylose residues, that is frequently decorated with a wide variety of chemical moieties that can limit enzymatic access to the main chain (Biely et al. 1986; Poutanen et al. 1991; Bajpai 1997). As a result, a suite of enzymes is generally required to ensure complete enzymatic hydrolysis of native xylan.
The most critical enzymatic components are the endo-xylanases (EC 3.2.1.8). These enzymes hydrolyze the internal β-1,4 bonds of the main xylan chain (Saha and Bothast 1999). Based on sequence analysis of the catalytic domain, these xylanases are categorized into glycosyl hydrolase (GH) families (Henrissat and Bairoch 1996). Members of a GH family share a common structural motif and catalytic mechanism. Of the xylanase gene sequences in the public domain, the vast majority is categorized as GH10 or GH11 family members (Collins et al. 2005). GH11 xylanases generally have a lower molecular weight and have higher specific activities than the GH10 enzymes. However, GH10 enzymes can better hydrolyze substrates that are highly branched. Besides these two families, a few xylanases have also been discovered in GH families 5, 7, 8, and 43 (Collins et al. 2005).
There is a great interest in discovering xylanase enzymes for various applications. Xylanases are needed to more fully hydrolyze lignocellulosic biomass into simple sugars that can then be fermented to products, such as liquid fuel and chemical feedstocks. The enzymes are also used in the production processes of other industries, such as animal feed, baking, and paper (Beg et al. 2001; Subramaniyan and Prema 2002). Most of the focus has been to develop thermophilic enzymes that can tolerate high temperatures (Andrews et al. 2004; Lee and Lim 2004; Sun et al. 2005). By elevating the temperature, the rates of the reaction are increased. However, there are situations when using psychrophilic enzymes might be preferred (Gerday et al. 2000; Collins et al. 2005). For instance, colder reaction conditions would preserve the properties of heat labile products. Alternatively, the extent of enzymatic digestion could be easily controlled by heat denaturing the cold-active enzymes. The use of colder reaction conditions would also lead to energy savings.
To discover new enzymes, the traditional strategy has been to culture and screen individual strains and then clone the unknown genes of interest. However, less than 1% of the microbiological organisms from the environment can be cultured using standard laboratory techniques, thus narrowing the spectrum of organisms that can be studied (Amann et al. 1995). To address this issue, a more recent approach has been to create libraries from genetic material that is isolated directly from an environmental sample that has not been subjected to any laboratory culturing (Lorenz et al. 2002; Streit et al. 2004). This “metagenomic” approach enables the exploration of a much wider sequence space than would otherwise be possible.
We screened a genomic DNA library created from the population of microorganisms collected from the waste lagoon of a dairy farm. A gene encoding a xylanase enzyme (Xyn8) was isolated and cloned. The encoded enzyme was classified as a GH8 family member based on sequence analysis of the catalytic domain. The enzyme was active against native xylan substrates but had only negligible cellulolytic activity. In addition, the enzyme retained significant activity even at 4°C.
Materials and methods
Sample collection and genomic DNA preparation
Samples were collected from a lagoon used to contain the manure wastewater from a dairy farm in Modesto, California. This lagoon was 14°C at the time of sampling. Microorganisms from 3 ml aliquots of the wastewater were pelleted by centrifugation. The cells were disrupted using the CLS-TC buffer and two 0.25 in. ceramic spheres of the FastDNA Kit (Qbiogene, Irvine, CA, USA), and the genomic DNA was then isolated according to the manufacturer’s instructions.
Genomic DNA library construction and activity screening
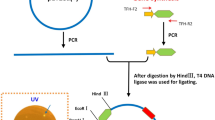
The genomic DNA isolated from the waste lagoon samples was partially digested with ApoI restriction enzyme. The digest was separated on an agarose gel by electrophoresis, and the fragments sized 4–10 kB were excised and purified using the QiaExII gel purification kit (Qiagen, Valencia, CA, USA). These fragments were then ligated to the EcoRI-digested Lambda ZAP II vector and packaged into lambda phage (Lambda ZAP II Vector and Gigapack III Packaging Extract, Stratagene, La Jolla, CA, USA).
The phage genomic DNA library was screened for xylanase activity using RBB-xylan (4-O-methyl-D-glucurono-D-xylan-remazol brilliant blue R), a dye-labeled substrate (Sigma, St. Louis, MO, USA). XL1-Blue MRF’ cells (Stratagene) were infected with phage and spread onto NZY agar plates. A 0.2% RBB-xylan overlay was poured over the infected cells and incubated overnight at 37°C. Phage plaques that resulted in clearings in the RBB-xylan overlay were isolated and rescreened for activity until purified plaques were obtained.
Subcloning xyn8 gene
The purified phage clones with the genomic DNA fragments encoding the xylan-degrading activity were subjected to in vivo excision according to the manufacturer’s protocol (Stratagene) resulting in a plasmid containing the genomic DNA fragment in the pBluescript vector backbone. The genomic DNA was sequenced in its entirety, and predicted open reading frames were determined with the Vector NTI bioinformatics package (Invitrogen, Carlsbad, CA, USA). BLAST analysis identified potential xylanase gene sequences to be studied (Altschul et al. 1990).
The xyn8 gene was amplified by subjecting the corresponding genomic DNA fragment to PCR using the following primers:
lagNA-X8-5: CGCCATATGCATGACAGAGGTGCTTTTTATACAGG
lagNA-X8-3: GCGCTCGAGTTCCGGTTTCCATATTTTAAAATCACCG
The 5′ and 3′ PCR primers were designed with NdeI and XhoI restriction enzymes sites, respectively, adjacent to the gene coding sequences. The PCR product and pET-22b plasmid (Novagen, Madison, WI, USA) were digested with NdeI and XhoI restriction enzymes and ligated. The resulting expression vector, pET22-X8, was designed to express the Xyn8 enzyme fused to a 6X-histidine tag at the C-terminus.
Expression and purification of Xyn8
The pET22-X8 plasmid was transformed into BL21::DE3(pLysE) (Novagen). Liquid Luria-Bertani (LB) cultures were grown at 37°C, and protein expression was induced by adding isopropyl-β-D-thiogalactopyranoside (IPTG) to 1 mM when the culture reached an optical density of 0.5 at 600 nm. Three hours after induction, the bacterial cells were harvested by centrifugation. The cell pellet was resuspended in 1/20 original culture volume in 25 mM Tris (pH 8), 300 mM sodium chloride, and 20 mM imidazole. The suspension was frozen in liquid nitrogen, thawed, sonicated, and centrifuged. The soluble protein was collected, applied to a HisTrap HP column (GE Healthcare, Piscataway, NJ, USA), and eluted with an imidazole gradient in buffer containing 50 mM sodium phosphate (pH 8) and 300 mM sodium chloride. The buffer of the elution fractions containing purified Xyn8 enzyme was exchanged by applying the fractions to a protein-desalting column (Pierce, Rockford, IL, USA) that had been equilibrated with 25 mM sodium acetate (pH 5.5) and 25 mM sodium chloride.
Xylanase activity assays
Enzyme activity on solid media was assayed by spotting BL21::DE3(pLysE) clones transformed with expression constructs on LB agar plates supplemented with 0.1% RBB-xylan and 1 mM IPTG. Plates were incubated overnight at 37°C.
Liquid enzyme activity was determined for various carbohydrate substrates: beech wood xylan, birch wood xylan, wheat arabinoxylan, rye arabinoxylan, carboxymethyl cellulose, Avicel, laminarin, and lichenin. The wheat and rye arabinoxylan were obtained from Megazyme (Bray, Ireland), and all others were obtained from Sigma. Reactions were conducted at various temperatures (4–60°C) and various pHs (Universal buffer: 8.3 mM each of citric acid, monobasic potassium phosphate, and boric acid). Temperature stabilities were determined by incubating the enzyme at various temperatures for 45 min at pH 7, cooling the reaction on ice, and then adding the enzyme to beech wood xylan and measuring the activity at 20°C. The pH stabilities were determined by incubating the enzyme at various pHs for 17 h at 20οC, and then adding the enzyme to substrate and measuring the activity at pH 7. All determinations of temperature and pH optimums and stabilities were conducted with 5 mg/ml beech wood xylan. All activity assays were conducted three times.
Extent of enzymatic digest was quantified by measuring the release of reducing sugar. Briefly, 100 μl of the reaction was added to 150 μl of DNSA reagent (1% dinitrosalicyclic acid and 30% potassium sodium tartrate in 0.5 M sodium hydroxide) and heated to 100°C for 5 min. The absorption of the resulting solution was measured on a microplate reader at 562 nm. Xylose standards were used to plot a calibration curve.
Analysis of xylanase hydrolysis products
1.9 μg of xylanase was incubated 16 h at 30°C with substrate in a 25 μl reaction (Universal buffer (pH 7)). Beech wood xylan was used at 10 mg/ml, and xylose, xylobiose, xylotriose, xylotetraose, and xylopentaose were used at 1 nmol/μl. Xylose was obtained from Sigma, and xylooligomers were obtained from Megazyme. Reactions were stopped by adding 80 μl of 100% ethanol to each tube. The tubes were incubated on ice for 10 min, centrifuged, and the supernatant was collected and lyophilized. The reducing sugar ends were labeled with a fluorophore using the Carbohydrate Labeling and Analysis Kit (Beckman Coulter, Fullerton, CA, USA). Samples were then analyzed on an eCAP N-CHO-coated capillary installed on a P/ACE MDQ Capillary Electrophoresis System with a 488 nm laser detection module (Beckman Coulter).
Results and discussion
Cloning and sequence analysis of xyn8 gene
An environmental sample was collected from a dairy farm waste lagoon. The total number of aerobic and anaerobic bacteria was 1×106 colony-forming units/ml based on viable plate counts (Jeffery McGarvey, personal communication), although there likely were many more organisms that were not accounted for because they could not grow under laboratory conditions.
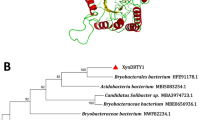
Genomic DNA was isolated directly from an uncultured sample of the waste lagoon water and used to create a phage library. 5×106 plaque-forming units were screened for activity on xylan substrate. One of the clones that had activity against xylan contained a 1,194 base pair-open reading frame (xyn8) (Genbank DQ241404) that was predicted to encode an endoxylanase (Table 1). The putative catalytic domain of the enzyme classified it as a member of the GH family 8. There are only four other xylanases in the public domain that have been categorized in GH family 8. Alignment of the Xyn8 protein sequence to these other enzymes indicates the presence of the conserved catalytic amino acids of this family (Fig. 1). Glu73 is the predicted catalytic acid, and Asp281 is most likely the catalytic base (Guerin et al. 2002). However, Asp133 has also been suggested to be either the catalytic base or play some other critical role in overall catalysis (Ozaki et al. 1994; Alzari et al. 1996; Van Petegem et al. 2003). The Xyn8 was most homologous to X6806 (Fig. 1) with 42% amino acid identity over the length of the enzyme (data not shown). The Xyn8 enzyme does not encode a predicted signal peptide sequence (Nielsen et al. 1997) or any known carbohydrate binding modules. The Xyn8 enzyme is likely of bacterial origin given the high homology to other bacterial xylanases and the lack of introns in the gene.
Expression and characterization of Xyn8 enzyme
The xyn8 gene was subcloned into a plasmid vector to express enzyme in bacteria. Clones transformed with the expression vector were spotted onto solid media supplemented with xylan substrate to demonstrate that active enzyme was produced (Fig. 2a). Xyn8 enzyme was collected and affinity purified from large-scale liquid culture. Polyacrylamide electrophoresis established that the size of the purified enzyme was approximately the predicted molecular weight of 45.9 kD (Fig. 2b).
The Xyn8 enzyme had activity against a variety of natural xylan substrates and beech wood xylan was determined to be the most susceptible to digestion (Fig. 3). There was negligible activity against cellulose substrates (carboxymethyl cellulose and Avicel) or other glucose-based substrates (laminarin and lichenin). The products of the enzymatic digest were analyzed by capillary electrophoresis (Table 1). When beech wood xylan was used as substrate, the primary products were xylose, xylobiose, and xylotriose. When xylooligomers were used as substrates, only xylotetraose and xylopentaose were hydrolyzed by the enzyme. Xylotetraose was hydrolyzed to xylotriose and xylose; xylopentaose was hydrolyzed to xylotriose and xylobiose.
Susceptibility of carbohydrate substrates to xylanase digestion. All reactions were conducted at 20°C and pH 7 with 10 mg/ml substrate. Be beech wood xylan; Bi birch wood xylan; W-AX wheat arabinoxylan; R-AX rye arabinoxylan; CMC carboxymethyl cellulose; Avi Avicel; Lam laminarin (algal β-glucan); and Lic lichenin (lichen glucan)
Using beech wood xylan as substrate, it was determined that the enzyme had an optimal activity at pH 6–7 (Fig. 4a.) The enzyme was also stable over a broad pH range (Fig. 4a). More than 60% of the enzyme activity was present even after incubation at pH 4 or pH 11. The Xyn8 enzyme had an optimal temperature of 20°C (Fig. 4b). At 60°C, there was no detectable activity; however, the enzyme demonstrated significant activity (29% of optimal) at the coldest temperature tested (5°C). The enzyme was stable when preincubated up to 50°C, but lost all activity at 60°C.
pH and temperature optimums and stabilities of Xyn8 enzyme on beech wood xylan. a Open squares are relative activities at various pH. Closed triangles are relative activities of the enzyme after incubation at various pHs followed by measuring the activity at pH 7. All reactions were conducted at 20°C. b Open squares are relative activities at various temperatures. Closed triangles are relative activities after incubation at various temperatures followed by measuring the activity at 20°C. All reactions were conducted at pH 7. The error bars are approximately the size of the symbols
Kinetic parameters were determined for Xyn8 using beech wood xylan at 20°C and 4°C (Table 1). Kinetic studies have been reported for only four other cold-active xylanase enzymes. Xylanase A from the Antarctic krill has an optimal activity at 40°C, but retains 30% activity at 4°C against oat xylan (Turkiewicz et al. 2000). The K m value of the krill xylanase at its optimal temperature is comparable to the Xyn8 enzyme (K m=4.1 and 5.3 mg/ml, respectively). However, the V max of Xyn8 enzyme is a 100-fold higher than that of the krill xylanase (V max=768 and 7.4 μmol/min/mg, respectively). When tested against beech wood xylan at 20°C, the cold-active Xyn10 enzyme isolated from a Flavobacterium sp. had a lower K m than the Xyn8 enzyme (K m=1.8 and 5.3 mg/ml, respectively), but Xyn10 enzyme had a much lower V max than Xyn8 enzyme (V max=142 and 768 μmol/min/mg, respectively) (Lee et al. 2006). Another cold-active xylanase is the enzyme isolated from Pseudoalteromonas haloplanktis that also happens to be a GH8 family member and was assayed against birch wood xylan (Collins et al. 2002). At 25°C, the P. haloplanktis enzyme has a higher catalytic constant than Xyn8 at its optimal temperature (20°C) (k cat=1,247 and 588/s, respectively). However, the Xyn8 has a much lower K m than the P. haloplanktis enzyme (K m=5.3 and 28 mg/ml, respectively). The only previously reported cold-active xylanase that has been characterized in detail at lower temperatures is from Cryptococcus adeliea using oat xylan as substrate (Petrescu et al. 2000). At 4°C, the C. adeliea enzyme has a k cat of 14.8/s which is eightfold less than that of Xyn8 (123/s). Thus, compared to previously reported cold-active xylanases, the Xyn8 enzyme had favorable biochemical characteristics.
References
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Alzari PM, Souchon H, Dominguez R (1996) The crystal structure of endoglucanase CelA, a family 8 glycosyl hydrolase from Clostridium thermocellum. Structure 4:265–275
Amann RI, Ludwig W, Schleifer KH (1995) Phylogenetic identification and in situ detection of individual microbial cells without cultivation. Microbiol Rev 59:143–169
Andrews SR, Taylor EJ, Pell G, Vincent F, Ducros VM, Davies GJ, Lakey JH, Gilbert HJ (2004) The use of forced protein evolution to investigate and improve stability of family 10 xylanases. The production of Ca2+-independent stable xylanases. J Biol Chem 279:54369–54379
Bajpai P (1997) Microbial xylanolytic enzyme system: properties and applications. Adv Appl Microbiol 43:141–194
Beg QK, Kapoor M, Mahajan L, Hoondal GS (2001) Microbial xylanases and their industrial applications: a review. Appl Microbiol Biotechnol 56:326–338
Biely P, MacKenzie CR, Puls J, Schneider H (1986) Cooperativity of esterases and xylanases in the enzymatic degradation of acetyl xylan. Biotechnology 4:731–733
Brennan Y, Callen WN, Christoffersen L, Dupree P, Goubet F, Healey S, Hernandez M, Keller M, Li K, Palackal N, Sittenfeld A, Tamayo G, Wells S, Hazlewood GP, Mathur EJ, Short JM, Robertson DE, Steer BA (2004) Unusual microbial xylanases from insect guts. Appl Environ Microbiol 70:3609–3617
Collins T, Gerday C, Feller G (2005) Xylanases, xylanase families and extremophilic xylanases. FEMS Microbiol Rev 29:3–23
Collins T, Meuwis MA, Stals I, Claeyssens M, Feller G, Gerday C (2002) A novel family 8 xylanase, functional and physicochemical characterization. J Biol Chem 277:35133–35139
Gerday C, Aittaleb M, Bentahir M, Chessa JP, Claverie P, Collins T, D’Amico S, Dumont J, Garsoux G, Georlette D, Hoyoux A, Lonhienne T, Meuwis MA, Feller G (2000) Cold-adapted enzymes: from fundamentals to biotechnology. Trends Biotechnol 18:103–107
Guerin DM, Lascombe MB, Costabel M, Souchon H, Lamzin V, Beguin P, Alzari PM (2002) Atomic (0.94 A) resolution structure of an inverting glycosidase in complex with substrate. J Mol Biol 316:1061–1069
Henrissat B, Bairoch A (1996) Updating the sequence-based classification of glycosyl hydrolases. Biochem J 316:695–696
Lee CC, Smith M, Kibblewhite-Accinelli RE, Williams TG, Wagschal K, Robertson GH, Wong DWS (2006) Isolation and characterization of a cold-active xylanase enzyme from Flavobacterium sp. Curr Microbiol 52:112–116
Lee YE, Lim PO (2004) Purification and characterization of two thermostable xylanases from Paenibacillus sp. DG-22. J Microbiol Biotechnol 14:1014–1021
Lorenz P, Liebeton K, Niehaus F, Eck J (2002) Screening for novel enzymes for biocatalytic processes: accessing the metagenome as a resource of novel functional sequence space. Curr Opin Biotechnol 13:572–577
Nielsen H, Engelbrecht J, Brunak S, von Heijne G (1997) Identification of prokaryotic and eukaryotic signal peptides and prediction of their cleavage sites. Protein Eng 10:1–6
Ozaki K, Sumitomo N, Hayashi Y, Kawai S, Ito S (1994) Site-directed mutagenesis of the putative active site of endoglucanase K from Bacillus sp. KSM-330. Biochim Biophys Acta 1207:159–164
Petrescu I, Lamotte-Brasseur J, Chessa JP, Ntarima P, Claeyssens M, Devreese B, Marino G, Gerday C (2000) Xylanase from the psychrophilic yeast Cryptococcus adeliae. Extremophiles 4:137–144
Poutanen K, Tenkanen M, Korte H, Puls J (1991) Accessory enzymes involved in the hydrolysis of xylans. In: Leatham GF, Himmel ME (eds) Enzymes in biomass conversion, vol 460. American Chemical Society, Washington DC, pp 426–436
Saha BC, Bothast RJ (1999) Enzymology of xylan degradation. In: Imam SH, Greene RV, Zaidi BR (eds) Biopolymers. American Chemical Society, Washington DC, pp 167–194
Streit WR, Daniel R, Jaeger KE (2004) Prospecting for biocatalysts and drugs in the genomes of non-cultured microorganisms. Curr Opin Biotechnol 15:285–290
Subramaniyan S, Prema P (2002) Biotechnology of microbial xylanases: enzymology, molecular biology, and application. Crit Rev Biotechnol 22:33–64
Sun JY, Liu MQ, Xu YL, Xu ZR, Pan L, Gao H (2005) Improvement of the thermostability and catalytic activity of a mesophilic family 11 xylanase by N-terminus replacement. Protein Expr Purif 42:122–130
Turkiewicz M, Kalinowska H, Zielinska M, Bielecki S (2000) Purification and characterization of two endo-1,4-beta-xylanases from Antarctic krill, Euphausia superba Dana. Comp Biochem Physiol B Biochem Mol Biol 127:325–335
van den Broek LA, Lloyd RM, Beldman G, Verdoes JC, McCleary BV, Voragen AG (2005) Cloning and characterization of arabinoxylan arabinofuranohydrolase-D3 (AXHd3) from Bifidobacterium adolescentis DSM20083. Appl Microbiol Biotechnol 67:641–647
Van Petegem F, Collins T, Meuwis MA, Gerday C, Feller G, Van Beeumen J (2003) The structure of a cold-adapted family 8 xylanase at 1.3 A resolution. Structural adaptations to cold and investgation of the active site. J Biol Chem 278:7531–7539
Ward OP, Moo-Young M (1989) Enzymatic degradation of cell wall and related plant polysaccharides. Crit Rev Biotechnol 8:237–274
Yoon KH, Yun HN, Jung KH (1998) Molecular cloning of a Bacillus sp. KK-1 xylanase gene and characterization of the gene product. Biochem Mol Biol Int 45:337–347
Acknowledgement
We are grateful to Jeffery McGarvey for assistance with collecting wastewater samples and supplying viable plate count data.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by K. Horikoshi
Reference to a company and/or products is only for the purposes of information and does not imply approval or recommendation of the product to the exclusion of others which may also be suitable. All programs and services of the U.S. Department of Agriculture are offered on a nondiscriminatory basis without regard to race, color, national origin, religion, sex, age, marital status, or handicap.
Rights and permissions
About this article
Cite this article
Lee, C.C., Kibblewhite-Accinelli, R.E., Wagschal, K. et al. Cloning and characterization of a cold-active xylanase enzyme from an environmental DNA library. Extremophiles 10, 295–300 (2006). https://doi.org/10.1007/s00792-005-0499-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00792-005-0499-3