Abstract
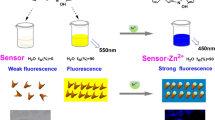
Fluorescent zinc complexes have recently attracted a lot of interest owing to their vast applications in cellular imaging. We report the synthesis as well as physical, chemical and biological studies of a novel zinc glyoxalbis(4-methyl-4-phenyl-3-thiosemicarbazone), [Zn(GTSC)]3, complex. As compared with the well-studied zinc biacetylbis(4-methyl-3-thiosemicarbazone), Zn(ATSM), complex, which was used as a reference, [Zn(GTSC)]3 had 2.5-fold higher fluorescence. When cellular fluorescence was measured using flow cytometry, we observed that [Zn(GTSC)]3 had 3.4-fold to 12-fold higher fluorescence than Zn(ATSM) in various cell lines (n = 9) of different tissue origin. Confocal fluorescence microscopy results showed that [Zn(GTSC)]3 appeared to have a nuclear localization within 30 min of addition to MCF7 cells. Moreover, [Zn(GTSC)]3 showed minimal cytotoxicity compared with Zn(ATSM), suggesting that [Zn(GTSC)]3 may be less deleterious to cells when used as an imaging agent. Our data suggest that the novel [Zn(GTSC)]3 complex can potentially serve as a biocompatible fluorescent imaging agent for live cells.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
There is widespread interest in developing novel imaging agents that can be effectively and reliably used in many disease conditions, including cancer, therapy and treatment monitoring [1–4]. Several reports have demonstrated the use of small organic fluorophores for imaging, but these dyes have been found to have serious side effects. For example, commercial dyes such as rhodamine 123 and fluorescein isothiocyanate can cause cytotoxicity and even lung fibrosis in rodents. They also appear to produce symptoms such as nausea and vomiting [5–8]. The use of metal complexes for live cell imaging, however, remains a less explored area. Although several radioactive copper (64Cu) bis(thiosemicarbazone) complexes have been tested for imaging and treatment, the corresponding zinc-based complexes are excellent alternatives for fluorescent imaging [9–25]. Not only are zinc complexes readily available, they are also generally less cytotoxic to cells and are conveniently monitored using simple fluorescence techniques.
Recently, Cowley et al. [26] reported an intrinsically fluorescent zinc biacetylbis(4-methyl-3-thiosemicarbazone), Zn(ATSM), complex whose cellular uptake and distribution in live cells could be monitored by confocal fluorescence microscopy. Zn(ATSM) was shown to accumulate in significant concentrations at the nucleus of PC-3 (prostate cancer) cells, whereas in the IGROV (ovarian cancer) cell line, the uptake is greater in acidic subcellular organelle lysosomes [22]. Selective cellular localization can prove valuable for understanding various mechanisms of action during therapeutic studies. Analogous zinc complexes such as zinc acenaphthenequinonebis(4-allyl-3-thiosemicarbazone) have also shown cellular fluorescence. However, their use as imaging agents is likely to be limited by their cytotoxicity. The latter complex has been shown to be as cytotoxic (LC50 12.5 μM) as cisplatin in the MCF-7 cell line [27, 28]. Thus, there is a need to develop new imaging agents which exhibit high fluorescence in cells, low background fluorescence and low cellular toxicity. We report here a novel trimeric zinc complex formed by a glyoxalbis(4-methyl-4-phenyl-3-thiosemicarbazone) (GTSCH2) ligand. The complex shows good fluorescence response and low cytotoxicity, making it a promising agent for live cell imaging.
Materials and methods
All the reagents and solvents were obtained from commercial sources and used without further purification. (Amines and CS2 used in the study were purified by standard methods prior to experiments.) NMR spectra were recorded with a Bruker AMX 400 spectrometer with tetramethylsilane as the internal standard. IR spectra were recorded in the solid state using KBr pellets with a PerkinElmer Spectrum One spectrometer. All UV–vis experiments were carried out with a PerkinElmer Lambda 35 instrument and the fluorescence measurements were carried out with an Horiba Jobin Yvon Fluoromax-4 spectrofluorimeter. A standard solution of zinc was purchased from Fluka Analytical and the amount of zinc in cells was estimated using a PerkinElmer Optima 2000 DV inductively coupled plasma optical emission spectroscopy (ICP-OES) instrument. All reagents used for cell culture study and DNA binding experiments were obtained from Sigma.
Synthesis and characterization of bis(thiosemicarbazone) ligands and their zinc complexes
Biacetylbis(4-methyl-3-thiosemicarbazone)
The ligand biacetylbis(4-methyl-3-thiosemicarbazone) (ATSMH2) was prepared following the literature procedure and was characterized by spectroscopic techniques [20].
Glyoxalbis(4-methyl-4-phenyl-3-thiosemicarbazone)
An ethanolic solution (30 mL) of 4-methyl-4-phenylthiosemicarbazide (0.8 g, 4.4 mmol) was heated to 60 °C and stirred until the 4-methyl-4-phenylthiosemicarbazide dissolved completely. The clear solution was cooled to room temperature and glyoxal (0.317 g, 2.2 mmol) was added as a 40% aqueous solution. The color changed from colorless to yellow. Addition of one drop of concentrated H2SO4 brought about the immediate precipitation of the product, which was collected by filtration. The residue was washed repeatedly with ethanol and dried in air for 12 h to give GTSCH2 as a light-yellow solid. Yield: 54%. 1H NMR (400 MHz, CDCl3): δ = 3.72 (s, 3H, NCH 3), 7.39 (s, 1H, CH), 7.21–7.24 (dd, 2H, Ph), 7.45 (m, 1H, Ph) 7.50–7.53 (m, 2H, Ph), 8.60 (s, 1H, NH). Electrospray ionization (ESI) mass spectroemetry (MS) calcd for [M+Na+]+: m/z: 407.11; found: m/z (%): 407.60 (100) [M+Na+]+, ESI(−): calcd for [M–H+]−: m/z: 383.11; found: m/z (%): 383.67 [M–H+]− (100); elemental analysis calcd (%) for C18H20S2N6: C 55.76, H 5.33, N 20.80, S 16.22; found: C 56.10, H 5.47, N 21.22, S 16.19; IR (KBr): νmax (cm−1) 3,230 (N–H), 1,522 (C=N), 1,320 (thioamide); UV–vis (dimethyl sulfoxide, DMSO): λmax(ε) = 354 nm (37,300 M−1 cm−1).
Benzilbis(4-pyrrolidinyl-3-thiosemicarbazone)
To an ethanolic solution of benzil (0.5 g, 2.4 mmol) and 4-pyrrolidinyl-3-thiosemicarbazide (0.7 g, 4.8 mmol) were added a few drops of concentrated H2SO4. The resulting reaction mixture was refluxed for 4 h. The solid product was isolated and washed with ethanol. This crude product was further recrystallized from a 1:1 mixture of chloroform and ethanol to give benzilbis(4-pyrrolidinyl-3-thiosemicarbazone) (BTSCH2) as a yellow crystals. Yield: 61%. 1H NMR (400 MHz, CDCl3): δ = 1.62 (quintet, 2H, NCH2CH 2), 1.73 (quintet, 2H, NCH2CH 2), 2.01 (quintet, 2H, NCH2CH 2), 2.07 (quintet, 2H, NCH2CH 2), 2.96 (t, 2H, NCH 2), 3.70 (t, 2H, NCH 2), 3.90 (t, 2H, NCH 2), 4.08 (bs, 2H, NCH 2), 7.29-7.58 (m, 10H, Ph), 8.59, 8.77 (s, 2H, NH, exchanges with D2O). 13C NMR (100.6 MHz, d 6-DMSO): δ = 24.17, 24.90 (NCH2 CH2), 49.8, 50.3 (NCH2) 127.45, 127.69, 128.26, 128.71, 130.45, 131.71, 134.18 (Ph), 144.55 (C=N), 176.10 (C=S); ESI–MS calcd for [M+Na+]+: m/z: 487.17; found: m/z (%): 487.87 (100) [M+Na+]+; elemental analysis: calcd for C24H28S2N6: C 62.04, H 6.07, N, 17.64; found: C 61.77, H 6.25, N 17.91; IR (KBr): νmax (cm−1) 3,175 (N–H), 1,580 (C=N), 1,448 (thioamide); UV–vis (DMSO): λmax(ε) = 302 nm (29,500 M−1 cm−1).
Zinc biacetylbis(4-methyl-3-thiosemicarbazone)
The Zn(ATSM) complex was prepared from zinc acetate (0.09 g, 0.410 mmol) and an ethanolic solution (10 mL) of ATSMH2 (0.1 g, 0.384 mmol) following the literature procedure [20]. After the addition of zinc acetate, the color changed from light yellow to intense yellow and the solution was refluxed for 4 h. The yellow precipitate formed was collected by filtration, washed with ethanol followed by diethyl ether and dried in vacuo. Yield: 56%. 1H NMR (400 MHz, d 6-DMSO): δ = 2.19 (s, 3H, CH 3–C=N), 2.81 (s, 3H, CH 3–N), 7.21 (bs, 1H, NH); 13C NMR (100.6 MHz, d 6-DMSO): δ = 14.79 (CH3), 30.15 (NCH3), 146.15 (C=N), 176.62 (C = S); IR (KBr): νmax (cm−1) 3,335 (NH), 1,525 (C=N), 1,372 (thioamide); UV–vis (DMSO): λmax(ε) = 313, 435 nm (11,700, 11,900 M−1 cm−1).
Zinc glyoxalbis(4-methyl-4-phenyl-3-thiosemicarbazone)
Zinc glyoxalbis(4-methyl-4-phenyl-3-thiosemicarbazone), [Zn(GTSC)]3, was prepared using GTSCH2 (0.14 g, 0.36 mmol) and zinc nitrate (0.1 g, 0.36 mmol) in ethanol (10 mL) using the procedure used to prepare the Zn(ATSM) complex. About 50 mg of the complex was isolated as an orange solid. The single crystal used for the X-ray diffraction study was grown in chloroform. Yield: 31%. 1H NMR (400 MHz, CDCl3): δ = 3.59 (s, 3H, NCH 3), 7.29–7.40 (m, 3H, Ph), 7.50–7.53 (m, 2H, Ph), 8.30 (s, 1H, CH). 13C NMR (100.6 MHz, CDCl3): δ = 41.21 (NCH3), 127.29, 127.86, 129.12, 146.18 (Ph), 146.88, 144.55 (C=N), 176.10 (C=S); ESI–MS calcd for [M+H+]+: m/z: 447.30: found: m/z (%): 447.33 [M+H+]+ (100); elemental analysis: calcd for C18H18S2N6Zn: C 48.27, H 4.05, N 18.76, S 14.32; found: C 48.08, H 4.29, N 18.19, S 13.71; IR (KBr): νmax (cm−1) 1,437 (C=N), 1,367 (thioamide); UV–vis (DMSO): λmax(ε) = 339, 475 nm (21,900, 10,900 M−1 cm−1).
Zinc benzilbis(4-pyrrolidinyl-3-thiosemicarbazone)
Zinc benzilbis(4-pyrrolidinyl-3-thiosemicarbazone), [Zn(BTSC)(DMSO)], was prepared following the procedure used for preparing Zn(ATSM) except that zinc nitrate (0.068 g, 0.228 mmol) in ethanol (5 mL) was used with BTSCH2 (0.099 g, 0.213 mmol) dissolved in chloroform (5 mL). The product, [Zn(BTSC)], was isolated as a bright-orange solid. Recrystallization of [Zn(BTSC)] from DMSO gave single crystals of [Zn(BTSC)DMSO] suitable for X-ray diffraction studies. Yield: 36%. 1H NMR (400 MHz, d 6-DMSO): δ = 1.78 (s, 4H, NCH2CH 2), 3.60 (s, 4H, NCH 2), 7.13–7.15 (3H, m, Ph), 7.28 (2H, d, J H, H = 7.2, Ph). 13C NMR (100.6 MHz, d 6-DMSO): δ = 25.35 (NCH2 CH2), 49.5, 50.20 (NCH2) 127.91, 128.74, 131.15, 134.57 (Ph), 145.01 (C=N), 176.48 (C=S); ESI–MS calcd for [M+H+]+: m/z: 527.10: found: m/z (%): 527.0 [M+H+]+ (100); elemental analysis: calcd for C24H26S2N6Zn: C 54.59, H 4.96, N 15.91; found: C 53.80, H 4.85, N 15.43; IR (KBr): νmax (cm−1) 1,437 (C=N), 1,367 (thioamide); UV–vis (DMSO): λmax(ε) = 333, 490 nm (10,500, 11,100 M−1 cm−1).
Crystal structure determination
X-ray crystallographic studies were carried out using a Bruker CCD diffractometer with graphite-monochromatized MoKα radiation (λ = 0.71073) controlled by a Pentium-based PC running the SMART (version 5.05, 1998; Bruker AXS single crystal diffractometer; Bruker, Madison, WI, USA) software package. Single crystals were mounted on glass fibers at room temperature, and data were collected at room temperature (20 °C). The structures were solved by direct methods (SIR-92) followed by the full-matrix least-squares procedure of F 2 for all reflections (SHELXL-97). All non-hydrogen atoms were refined by anisotropic displacement parameters and hydrogen atoms were fixed at idealized positions [29, 30].
Cell culture
All cell lines were grown in Dulbecco’s modified Eagle medium (DMEM) supplemented with 10% fetal calf serum (FCS), penicillin and streptomycin. Cell cultures were maintained at 37 °C and 5% CO2 in an incubator.
Estimation of zinc in cells by ICP-OES
MCF-7 cells were treated with [Zn(GTSC)]3 for 1 h at 37 °C and harvested using a cell scraper. The cell suspension was centrifuged at 2,000 rpm for 5 min at 4 °C. The cell pellet so obtained was washed with phosphate-buffered saline (PBS) and an aliquot was taken for protein analysis. The suspension was again centrifuged at 2,000 rpm for 5 min at 4 °C and the cell pellet was collected. Dry cell pellets were then lysed using one freeze–thaw cycle and 1% HNO3 solution. The final volume was made up to 10 mL and used for analysis with the ICP-OES instrument. The instrument was calibrated for zinc using solutions containing 10, 50, 100, 500 and 1,000 ppb zinc. The concentration of zinc obtained was normalized to cellular protein concentration (measured using Bradford protein assay) [31, 32].
Confocal fluorescence imaging
Fluorescence imaging studies were done using a Leica confocal fluorescence microscope (TCS SP5) in HCX APO L 20.0 × 1.00 water-objective mode. Cells were grown in appropriate media in six-well dishes containing circular cover slips. On the day of the experiment, the medium was replaced with phenol red free DMEM (pH 7.4) containing 10% FCS, penicillin and streptomycin. The cover slip was placed on the imaging chamber, zinc complex/ligand (1.5 mL in 1% DMSO in phenol red free DMEM) was added and fluorescence was recorded using 488-nm excitation and emission was collected in the 500–600-nm region. Quantification of microscopy images was done using ImageJ (NIH) [33].
Flow cytometry
Cell pellets were collected following trypsinization from exponentially growing cultures and washed with PBS (pH 7.4). The pellet was then resuspended in 1 mL PBS buffer (pH 7.4), zinc complexes were added and fluorescence was recorded in the FL1 channel of a Beckman Coulter CyAn flow cytometer. The mean fluorescence intensity was represented as an average of 10,000 cells gated according to forward and side scatter.
3-(4,5-Dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide assay
3-(4,5-Dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) assay was carried out to measure cell viability. An appropriate number of cells were seeded onto the wells of a 96-well plate. After 24 h, various concentrations of the zinc complexes were added and incubation was carried out for 48 h in a 37 °C incubator. After 45 h incubation, MTT was added and the absorbance at 550 nm was read 3 h later on a microplate reader (Molecular Devices). A graph of the concentration versus percentage cell viability was plotted and the concentration at which 50% cell death occurred was used as the IC50 value.
DNA binding constant estimation by ethidium bromide displacement assay
The DNA binding experiments were done in aqueous solutions of sodium perchlorate (400 mM) and the zinc complexes dissolved using minimum amounts of DMSO. The purity of calf thymus DNA was evaluated from the ratio of the absorbance at 260 and 280 nm, which was found to be more than 1.8, and the concentration was calculated from the absorbance at 260 nm. DNA was added to ethidium bromide (1.3 μM) solution until the fluorescence intensity at 602 nm saturated (λex = 560 nm). Subsequently, quenching was observed in the presence of increasing amounts of zinc complexes. The apparent binding constant (K app) was calculated from the equation [EtBr] × K ErBr = [complex]50% × K app, where [EtBr] refers the concentration of ethidium bromide and [complex]50% is the concentration required for 50% quenching of the DNA–EtBr adduct. The reported values are the average of two independent experiments.
Results and discussion
Synthesis and characterization
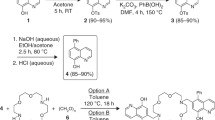
The bis(thiosemicarbazone) ligands and their zinc complexes were synthesized (Fig. 1) by following known protocols with minor modifications (Scheme S1). The ligands and zinc complexes were well characterized by 1H NMR spectroscopy, 13C NMR spectroscopy, UV–vis spectroscopy, IR spectroscopy, CHN analysis, ESI–MS and fluorescence spectroscopic techniques. The crystal structures of BTSCH2 ligand and [Zn(GTSC)]3 and [Zn(BTSC)(DMSO)] complexes were characterized using single-crystal X-ray crystallography (Figs. S2, S3, S4, Table S5).Footnote 1
The versatile coordinating behaviour of bis(thiosemicarbazone) ligands and the structural flexibility of zinc leads to the formation of a variety of zinc bis(thiosemicarbazone) complexes with different coordination numbers and geometries. The formation of monomers, dimers and polymeric zinc bis(thiosemicarbazone) complexes in both square pyramidal and tetrahedral geometry around zinc have been reported [14, 34–38]. However, our crystallographic analysis of the zinc complex of GTSCH2 revealed the formation of an unusual trimeric [Zn(GTSC)]3 complex. This trimeric structure resembles a trigonal prism with each face capped by a ligand. Each zinc is coordinated to two ligands through sulphur and nitrogen ligation, thus forming a ZnS2N2 framework where zinc is in a distorted tetrahedral geometry (Fig. 2). [Zn(GTSC)]3 is structurally similar to the previously reported zinc (1,2-ethanediyliden)-bis(S-methylhydrazonecarbodithionate) complex [39]. In the case of [Zn(BTSC)(DMSO)], the complex is a monomer and DMSO is coordinated to zinc as a fifth ligand, resulting in a distorted square pyramidal structure.
Crystal structure of the trimeric [Zn(GTSC)]3 complex. Hydrogen atoms are omitted for clarity. Selected bond lengths (Å): Zn(2)–S(3), 2.264(16); Zn(2)–S(2), 2.255(16); Zn(2)–N(4), 2.085(4); Zn(2)–N(6), 2.071(4); S(3)–C(26), 1.724(6); S(2)–C(11), 1.745(5); N(5)–C(11), 1.325(6); N(8)–C(26), 1.326(5); N(4)–N(5), 1.366(5); N(6)–N(8), 1.362(5)
The absence of bulky groups on the diketone and the presence of methyl and phenyl groups on the nitrogen of thiosemicarbazide enforce an s-trans conformation for the GTSC ligand in [Zn(GTSC)]3. In most complexes, the s-cis conformation is preferred as shown by the [Zn(BTSC)(DMSO)] complex, resulting in a distorted square pyramidal geometry. Our examination of bond lengths in the ligands and in zinc complexes revealed elongation of the C=S bond and shortening of the adjacent C–N bond length relative to the free ligand, indicating extensive delocalization and formation of the thiolate form in zinc complexes.
UV–vis and fluorescence studies
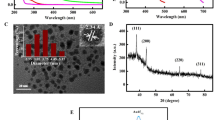
UV–vis absorption spectra were recorded for the complexes in DMSO. [Zn(GTSC)]3 and [Zn(BTSC)(DMSO)] complexes showed a characteristic absorption band centered at 475 and 490 nm, respectively, whereas Zn(ATSM) showed a band centered at 435 nm. These lower-energy bands in [Zn(GTSC)]3 and [Zn(BTSC)(DMSO)] can be attributed to charge transfer transitions from sulphur to zinc (Fig. 3A).
Fluorescence spectra of complexes dissolved in DMSO (10 μM in zinc) were recorded and the results are shown in Fig. 3B and C. The studies were also repeated using 5% DMSO in PBS (Fig. 3D) and 1% DMSO in water (Fig. S6) to ascertain the fluorescence response in aqueous media. It was observed that in aqueous media the fluorescence intensity was significantly quenched. The quantum yields for [Zn(GTSC)]3, [Zn(BTSC)(DMSO)] and Zn(ATSM) complexes were estimated in DMSO (0.02 mM) using [Ru(bpy)3](PF6)2 (bpy is bipyridyl) as a standard [40–42]. The observed values for [Zn(GTSC)]3 (ϕ = 3.0 × 10−3) and [Zn(BTSC)(DMSO)] (ϕ = 2.1 × 10−3) were 2.5 and 1.7 times higher, respectively, than the value for Zn(ATSM) (ϕ = 1.2 × 10−3) under identical conditions. Photophysical properties of all the synthesized zinc complexes are listed in Table 1. The stock solutions of [Zn(GTSC)]3 and [Zn(BTSC)(DMSO)] were found to be stable for at least 10 days in DMSO. Emission spectra recorded with excitation at 488 nm showed that [Zn(GTSC)]3 and [Zn(BTSC)(DMSO)] exhibit fluorescence with maxima at 565 and 575 nm, respectively, and have better a fluorescence response than Zn(ATSM).
Interestingly, the commonly available excitation wavelength 488 nm is close to the absorption maxima of both complexes, making them more convenient cellular imaging agents. The [Zn(BTSC)(DMSO)] complex showed a much lower fluorescence quantum yield than [Zn(GTSC)]3 and was not investigated further. The unique trimeric nature of [Zn(GTSC)]3 is presumably responsible for the greater fluorescence of this complex.
In vitro cellular uptake studies by flow cytometry
We then investigated the ability of [Zn(GTSC)]3 to enter cells and fluoresce in an intracellular environment. To optimize the concentration and time required for the maximum cellular uptake of [Zn(GTSC)]3, MCF-7 breast cancer cells were treated with various concentrations of [Zn(GTSC)]3, were incubated for either 30 or 60 min and fluorescence was measured by flow cytometry. A dose-dependent cellular uptake of [Zn(GTSC)]3 was observed (Fig. 4A). There was no significant difference in uptake at 30 or 60 min of incubation, suggesting that maximum uptake of the complex occurs within 30 min of the treatment (Fig. 4B). Furthermore, the dose-dependent increase in intracellular zinc concentration following treatment with [Zn(GTSC)]3 (Fig. 4C), as measured by ICP-OES, provided evidence for entry of [Zn(GTSC)]3 into cells. On the basis of these experimental results, 10 μM [Zn(GTSC)]3 and 30 min incubation were taken as the ideal concentration and time for internalization and were used for all further experiments.
A MCF-7 cells were treated with different concentrations of [Zn(GTSC)]3 for 30 min and fluorescence was recorded using a flow cytometer. B Flow cytometry quantification for MCF-7 cells treated with different concentrations of [Zn(GTSC)]3 for two different time points, 30 and 60 min. C Elemental analysis was carried out using inductively coupled plasma optical emission spectroscopy and intracellular zinc concentration was estimated in MCF-7 cells treated with either 25 or 50 μM (in zinc) [Zn(GTSC)]3. MFI mean fluorescence intensity
To compare the uptake of [Zn(GTSC)]3 with that of the previously known Zn(ATSM) complex, MCF-7 and normal keratinocyte HaCat cells were treated with either 10 μM [Zn(GTSC)]3 or 10 μM Zn(ATSM) for 30 min and fluorescence was measured using flow cytometry. In MCF-7 cells, [Zn(GTSC)]3 showed a 12-fold higher fluorescence response (45.5 ± 1.1) than Zn(ATSM) (3.8 ± 0.2) (Fig. 5A). When HaCat cells were treated with [Zn(GTSC)]3, 6.5-fold higher fluorescence was observed (18.6 ± 1.1 for [Zn(GTSC)]3 vs. 2.8 ± 0.1 for Zn(ATSM)) (Fig. 5B). In another breast cancer cell line, MDA-MB-231, [Zn(GTSC)]3 showed 3.4-fold higher fluorescence (17.2 ± 1.3 for [Zn(GTSC)]3 vs. 5.1 ± 0.4 for Zn(ATSM)). Under our experimental conditions, no increase above the background level was seen when either MCF-7 or HaCat cells were treated with Zn(ATSM) (Fig. 5). These data show that [Zn(GTSC)]3 has higher fluorescence than Zn(ATSM) inside the cells as well.
Cellular localization by confocal fluorescence imaging
To visualize the cellular localization and cellular retention time of [Zn(GTSC)]3, MCF-7 cells were treated with either 10 μM [Zn(GTSC)]3 or 10 µM Zn(ATSM) and uptake was monitored immediately and followed up to 30 min by confocal fluorescence microscopy. As shown in Fig. 6A and B, [Zn(GTSC)]3 localizes on the cell membrane/cytoplasm of MCF-7 cells initially and then becomes internalized into the nucleus. In agreement with flow cytometry data, [Zn(GTSC)]3 showed a higher fluorescence response than Zn(ATSM) (Fig. 6C, D) in MCF-7 cells. Figure 6E shows the quantification of cellular fluorescence from images taken from 2 to 30 min of [Zn(GTSC)]3 incubation. It appears that cytoplasmic fluorescence decreases and nuclear fluorescence increases within the first 30 min. However, the overall cellular fluorescence remains constant for more than 60 min (Fig. 6F), suggesting that the complex does not efflux from the cell for at least 1 h. To ensure that the cellular viability is not affected by [Zn(GTSC)]3 treatment, we carried out MTT-based cell viability assay following treatment with [Zn(GTSC)]3 for 15, 30, 45 or 60 min. We also examined the cell morphology for any signs of stress following [Zn(GTSC)]3 treatment. We observed that neither the cell morphology (Fig. S7a–e) nor the cell viability (Fig. S7f) was affected by [Zn(GTSC)]3 treatment, suggesting that the complex is non-toxic. To investigate whether our complex enters the cell, we carried out co-localization measurement of the fluorescence and bright field images following treatment of fixed MCF-7 cells with [Zn(GTSC)]3. When the cells were treated with [Zn(GTSC)]3, intense intracellular fluorescence was observed at the cytoplasmic region after 15 min, and cellular morphology was also retained. This fact is evident from the overlaid fluorescence and bright field images, as shown in Fig. 7B. No fluorescence was observed from the cells with an addition of an equal amount of DMSO under identical conditions (Fig. 7A). This suggests that [Zn(GTSC)]3 enters the cell and is able to exhibit stable fluorescence in the intracellular environment. To show further that the zinc complex is essential for the intracellular fluorescence, a confocal fluorescence imaging experiment was performed in live MCF-7 cells treated with [Zn(GTSC)]3 or GTSCH2 ligand (10 μM) and the fluorescence was monitored for 30 min. We observed significant fluorescence from the cells treated with [Zn(GTSC)]3 (Fig. 7D). The observed intracellular fluorescence was quenched by the addition of an excess (100 μM) of N,N,N′,N′-tetrakis(2-pyridylmethyl)ethylenediamine, a cell-permeable, non-fluorescent and high-affinity zinc chelator (Fig. 7F). It is apparent that the intracellular fluorescence was quenched by the process of trans chelation. Very faint fluorescence was observed for the cells treated with GTSCH2 ligand alone under similar conditions (Fig. 7E). A control experiment with untreated MCF-7 cells showed little fluorescence (Fig. 7C). All these results confirm that internalization of the trimeric zinc complex [Zn(GTSC)]3 is essential for the observed intracellular fluorescence.
Representative images of MCF-7 cells following treatment with [Zn(GTSC)]3 (A, B) or Zn(ATSM) (10 μM in zinc) (C, D) recorded using a confocal fluorescence microscope. E MCF-7 cells were treated with 10 μM [Zn(GTSC)]3 for 60 min. Images were recorded using a confocal microscope after every 2 min, generating a total of 30 images. For quantification, ten cells were selected per image and fluorescence was measured using ImageJ (NIH). The average of the fluorescence of ten cells is shown in the graph. F MCF-7 cells were treated with [Zn(GTSC)]3 (10 μM in zinc) and images were recorded over a period of 30 min using a confocal fluorescence microscope. The fluorescence intensity of ten cells and that of corresponding nuclei were quantified using ImageJ. The latter was subtracted from the former to get the fluorescence intensity of the cytoplasm. The graph represents the change in fluorescence of the cytoplasm and the nucleus over 30 min and error bars represent the standard deviation (SD) from ten different cells
A Autofluorescence of fixed MCF-7 cells. B MCF-7 cells were treated with [Zn(GTSC)]3 (10 μM in zinc) in an eight-chamber slide. After 15 min, spent medium was removed and cells were fixed using 4% paraformaldehyde and fluorescence images were recorded using a Zeiss confocal fluorescence microscope (×40 magnification) with 488-nm excitation. Simultaneously, bright field images were recorded and the two images were overlaid to visualize co-localization. For both A and B, a the fluorescence image, b the bright field image and c the co-localization of images a and b. C Autofluorescence of live MCF-7 cells was recorded without any fixation. D Live MCF-7 cells were treated with [Zn(GTSC)]3 (10 μM) in Dulbecco’s modified Eagle medium and fluorescence was recorded after 30 min incubation with a Leica confocal fluorescence microscope (×20 magnification). E Live MCF-7 cells were treated with GTSCH2 ligand (10 μM) alone and fluorescence was recorded. F Live MCF-7 cells were treated as in D and subsequently quencher N,N,N′,N′-tetrakis(2-pyridylmethyl)ethylenediamine (TPEN) (100 μM) was added; fluorescence was recorded after 5 min incubation with TPEN
In HaCat cells as well, [Zn(GTSC)]3 showed higher fluorescence than Zn(ATSM) (Fig. 8). As observed in the case of MCF-7 cells, [Zn(GTSC)]3 appears to localize in the nucleus of HaCat cells as well. In another breast cancer cell line, MDA-MB-231, nuclear localization was visible only at lower concentrations of [Zn(GTSC)]3 (Fig. S8). This suggests that [Zn(GTSC)]3 is cell-permeable and can fluoresce in an intracellular environment of cancer or normal cells. Since the fluorescence of the complex is partially quenched in aqueous media, and the intracellular environment is mostly aqueous, we hypothesize that the observed intracellular fluorescence is due to the binding of [Zn(GTSC)]3 with hydrophobic membranes of intracellular organelles.
Cell viability studies
To ascertain the effect of these complexes on cellular viability after prolonged treatment, we performed MTT assay following 48 h treatment with both [Zn(GTSC)]3 and Zn(ATSM). The IC50 value obtained for [Zn(GTSC)]3 was nearly 1.7-fold to 3.6-fold higher in cells than that for Zn(ATSM) even after 48 h, showing that the complex is relatively non-toxic (Table 2; Fig. S9). Moreover, the concentration at which maximum biological fluorescence was observed (10 μM) is below the IC50 value for [Zn(GTSC)]3 and cells had 60–80% viability.
We also found [Zn(GTSC)]3 to be less toxic (1.7 and 3.3 times) to normal cells (immortalized human keratinocyte cells—HaCat) than to MCF-7 and MDA-MB-231 breast cancer cells (Table 2). In addition, IC50 was twofold higher for [Zn(GTSC)]3 than for Zn(ATSM) in HaCat cells (Table 2). To investigate the possible reason behind the lower toxicity of [Zn(GTSC)]3 compared with Zn(ATSM), we measured the binding ability of [Zn(GTSC)]3 and Zn(ATSM) for DNA, a possible target for these complexes, by ethidium bromide displacement assay. We observed that the apparent DNA binding constant of [Zn(GTSC)]3 (4.6 ± 1.4 × 104 M−1) was nearly twofold lower than that of Zn(ATSM) (8.3 ± 0.1 × 104 M−1), suggesting that it may cause less DNA damage and hence low cytotoxicity. The possibility of the ligand leaching out from the metal complex, inside the cell, is excluded as these ligands possess lower cytotoxicity than their zinc complexes (data not shown). Thus, [Zn(GTSC)]3 appears to be safer and biocompatible, which bodes well for its use as an imaging agent for cancer cells and normal cells.
Quantification of complex uptake in different cell lines
One of the desired qualities of an imaging agent is that it should be able to fluoresce in various types of tissues. To determine the utility of [Zn(GTSC)]3 in cancers of different tissue origin, we tested its fluorescence by flow cytometry in nine cancer cell lines of six different tissue types. In all cell lines, we observed that [Zn(GTSC)]3 had higher fluorescence than Zn(ATSM) (Table 3; Fig. 9).
A Fluorescence of [Zn(GTSC)]3 (10 μM in zinc) (blue curve) in comparison with autofluorescence (black curve) and fluorescence of Zn(ATSM) (10 μM) (red curve) in nine different cancer cell lines of different tissue origin. B Quantification of A. Error bars represents the SD from three independent samples
Conclusion
In conclusion, we have shown that the novel trimeric zinc complex [Zn(GTSC)]3 is a good imaging agent, is easy to synthesize and exhibits enhanced fluorescence owing to its unique trimeric structure. It is significantly better than the other zinc complexes owing to its low cytotoxicity. Since the fluorescence of [Zn(GTSC)]3 is quenched in the aqueous phase, background fluorescence is minimal. [Zn(GTSC)]3 appears to be stable in an intracellular environment and fluoresces for a long time (more than 1 h) without being quenched in the hydrophobic regions of the cell. It shows high fluorescence in several cell lines, and in some it shows nuclear localization. Additionally, it is relatively non-toxic to two breast cancer cell lines and a normal cell line tested in this study. These features make it an excellent agent for live cell imaging.
Notes
The Cambridge Crystallographic Data Centre (CCDC) numbers for [Zn(GTSC)]3, [Zn(BTSC)(DMSO)] and BTSCH2 are CCDC 776332, CCDC 776330 and CCDC 776331.
References
Cai W, Rao J, Gambhir SS, Chen X (2006) Mol Cancer Ther 5:2624–2633
Caroll L, Bejot R, Hueting R, King R, Bonnitcha P, Bayly S, Christlieb M, Dilworth JR, Gee AD, Declerck J, Gouverneour V (2010) Chem Commun 46:4052–4054
Ferreira MF, Martins AF, Martins JA, Ferreira PM, Toth E, Geraldes CFGC (2009) Chem Commun 42:6475–6477
Craig P, Benjamin MS, Elizabeth NJ, Pal R, Parker D (2009) Acc Chem Res 42:925–937
Castro DJ, Saxton RE, Fetterman HR, Castro DJ, Ward PH (1992) J Clin Laser Med Surg 10:83–90
Lampidis TJ, Bernal SD, Summerhayes IC, Chen LB (1983) Cancer Res 43:716–720
Roberts SN, Howie SE, Wallace WA, Brown DM, Lamb D, Ramage EA, Donaldson K (1995) J Pathol 176:309–318
Rumbaut RE, Sial AJ (1999) Microcirculation 6:205–213
Lewis JS, Laforest R, Buettner TL, Song SK, Fujibayashi Y, Connett JM, Welch MJ (2001) Proc Natl Acad Sci USA 98:1206–1211
Kraker A, Krezoski S, Schneider J, Minkel D, Petering DH (1985) J Biol Chem 260:13710–13718
Fujibayashi Y, Taniuchi H, Yonekura Y, Ohtani H, Konishi J, Yokoyama A (1997) J Nucl Med 38:1155–1160
Vavere AL, Lewis JS (2007) Dalton Trans 4893–4902
Xiao Z, Donnelly PS, Zimmermann M, Wedd AG (2008) Inorg Chem 47:4338–4347
Holland JP, Aigbirhio FI, Betts HM, Bonnitcha PD, Burke P, Christlieb M, Churchill GC, Cowley AR, Dilworth JR, Donnelly PS, Green JC, Peach JM, Vasudevan SR, Warren JE (2007) Inorg Chem 46:465–485
Katherine AP, Aphrodite C, Paterson BM, Filiz G, Volitakis I, Masters CL, Barnham KJ, Donnelly PS, Crouch PJ, White AR (2009) J Med Chem 52:6606–6620
Holland JP, Barnard PJ, Collison D, Dilworth JR, Edge R, Green JC, Heslop JM, McInnes EJL, Salzmann CG, Thompson AL (2008) Eur J Inorg Chem 22:3549–3560
Beraldo H, Boyd LP, West D (1998) Transition Met Chem 23:67–71
Cowley AR, Dilworth JR, Donnelly PS, Labisbal E, Sousa A (2002) J Am Chem Soc 124:5270–5271
Holland JP, Fisher V, Hickin JA, Peach JM (2010) Eur J Inorg Chem 48–58
Holland JP, Barnard PJ, Collison D, Dilworth JR, Edge R, Green JC, McInnes JL (2008) Chem Eur J 14:5890–5907
Dearling JLJ, Blower PJ (1998) Chem Commun 2531–2532
Lim SC, Price KA, Chong SF, Paterson BM, Caragounis A, Barnham KJ, Crouch PJ, Peach JM, Dilworth JR, White AR, Donnelly PS (2010) J Biol Inorg Chem 15:225–235
Pascu SI, Waghorn PA, Kennedy BWC, Arrowsmith RL, Bayly SR, Dilworth JR, Christlieb M, Tyrrell RM, Churchill GC, Aigbirhio FI (2010) Chem Asian J 5:506–519
You Y, Tomat E, Hwang K, Atanasijevic T, Nam W, Jasanoff AP, Lippard SJ (2010) Chem Commun 46:4139–4141
Gao Y, Wu J, Li Y, Sun P, Zhou H, Yang J, Zhang S, Jin B, Tian Y (2009) J Am Chem Soc 131:5208–5213
Cowley AR, Davis J, Dilworth JR, Donnelly PS, Dobson R, Nightingale A, Peach JM, Shore B, Kerr D, Seymour L (2005) Chem Commun 845–847
Pascu SI, Waghorn PA, Conry TD, Betts HM, Dilworth JR, Churchill GC, Pokrovska T, Christlieb M, Aigbirhio FI, Warren JE (2007) Dalton Trans 43:4988–4997
Pascu SI, Waghorn PA, Conry TD, Lin B, Betts HM, Dilworth JR, Sim RB, Churchill GC, Aigbirhio FI, Warren JE (2008) Dalton Trans 16:2107–2110
Walker N, Stuart D (1983) Acta Crystallogr Sect A 39:158–166
Altomare A, Cascarano G, Giacovazzo C, Gualardi A (1993) J Appl Crystallogr 26:343–350
Donnelly PS, Caragounis A, Du T, Laughton KM, Volitakis I, Cherny RA, Sharples RA, Hill AF, Li QX, Masters CL, Barnham KJ, White AR (2008) J Biol Chem 283:4568–4577
Deng Z, Dailey LA, Soukup J, Stonehuerner J, Richards JD, Callaghan KD, Yang F, Ghio AJ (2009) Biometals 22:803–815
Rasband WS (1997–2004) ImageJ. National Institutes of Health, Bethesda. http://rsb.info.nih.gov/ij/
Christlieb M, Struthers HSR, Bonnitcha PD, Cowley AR, Dilworth JR (2007) Dalton Trans 5043–5054
Al-Karawi AJM, Clegg W, Harrington RW, Henderson RA (2009) Dalton Trans 564–570
Zhong X, Yi J, Sun J, Wei HL, Liu WS, Yu KB (2006) Eur J Med Chem 41:1090–1092
Buncic G, Donnelly PS, Paterson BM, White JM, Zimmermann M, Xiao Z, Wedd AG (2010) Inorg Chem 49:3071–3073
Torres EL, Mendiola MA, Pastor CJ, Perez BS (2004) Inorg Chem 43:5222–5230
Hennig C, Hallmeier KH, Zahn G, Tschwatschal F, Hennig H (1999) Inorg Chem 38:38–43
Caspar JV, Meyer TJ (1983) J Am Chem Soc 105:5583–5590
Williams ATR, Winfield SA, Miller JN (1983) Analyst 108:1067–1071
Forgues SF, Lavabre D (1999) J Chem Educ 76:1260–1264
Acknowledgments
D.D. thanks the Indian Instutite of Science (IISc) for an IISc Centenary Postdoctoral Fellowship. D.P. acknowledges the Council of Scientific & Industrial Research for a Senior Research Fellowship. K.S. and A.G.S. thank the Department of Science and Technology (New Delhi) for a research grant. We thank IRIS, IISc and Pushpa Janardhan for assistance with the confocal fluorescence microscopy experiments.
Author information
Authors and Affiliations
Corresponding authors
Additional information
D. Dayal and D. Palanimuthu contributed equally.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Dayal, D., Palanimuthu, D., Shinde, S.V. et al. A novel zinc bis(thiosemicarbazone) complex for live cell imaging. J Biol Inorg Chem 16, 621–632 (2011). https://doi.org/10.1007/s00775-011-0764-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00775-011-0764-0