Abstract
Superoxide dismutases (SODs) protect cells against oxidative stress by disproportionating O2 − to H2O2 and O2. The recent finding of a nickel-containing SOD (Ni-SOD) has widened the diversity of SODs in terms of metal contents and SOD catalytic mechanisms. The coordination and geometrical structure of the metal site and the related electronic structure are the keys to understanding the dismutase mechanism of the enzyme. We performed Q-band 14N,1/2H continuous wave (CW) and pulsed electron–nuclear double resonance (ENDOR) and X-band 14N electron spin echo envelope modulation (ESEEM) on the resting-state Ni-SOD extracted from Streptomyces seoulensis. In-depth analysis of the data obtained from the multifrequency advanced electron paramagnetic resonance techniques detailed the electronic structure of the active site of Ni-SOD. The analysis of the field-dependent Q-band 14N CW ENDOR yielded the nuclear hyperfine and quadrupole coupling tensors of the axial Nδ of the His-1 imidazole ligand. The tensors are coaxial with the g-tensor frame, implying the g-tensor direction is modulated by the imidazole plane. X-band 14N ESEEM characterized the hyperfine coupling of Nε of His-1 imidazole. The nuclear quadrupole coupling constant of the nitrogen suggests that the hydrogen-bonding between Nε–H and OGlu-17 present for the reduced-state Ni-SOD is weakened or broken upon oxidizing the enzyme. Q-band 1H CW ENDOR and pulsed 2H Mims ENDOR showed a strong hyperfine coupling to the protons(s) of the equatorially coordinated His-1 amine and a weak hyperfine coupling to either the proton(s) of a water in the pocket at the side opposite the axial Nδ or the proton of a water hydrogen-bonded to the equatorial thiolate ligand.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Superoxide dismutases (SODs) catalyze disproportionation of superoxide (O2 −) to hydrogen peroxide (H2O2) and molecular oxygen (O2) to protect cells against oxidative stress [1–4]. SODs are found in all living organisms and all have redox-active transition metals in their active sites. The most well known one, found in both eukaryotes and prokaryotes, contains copper and zinc (CuZn-SOD). Other SODs contain manganese or iron. Crystallographic studies of the SODs revealed histidine-rich metal centers. In CuZn-SOD, copper and zinc are bridged by the imidazolate of a histidine residue, whereas manganese-containing SOD and iron-containing SOD have mononuclear transition metal centers. In the resting state of CuZn-SOD, the Cu(II) site is pentacoordinated with four histidine residues, including the bridging histidine, and one solvent-accessible ligand site. The Zn(II) site has three histidine ligands, including the bridging histidine, and one oxygen ligand of the carboxylate group of an aspartate residue [5–8]. The metal centers of both the oxidized manganese-containing SOD and the oxidized iron-containing share identical trigonal bipyramidal geometries. In those, two nitrogen atoms of histidine side chains and one oxygen atom of aspartate occupy the three equatorial ligand positions of the bipyramidal coordination, and a solvent molecule and one histidine imidazole occupy the remaining two axial positions [9–14].
The diversity of SODs in terms of metal contents has become even wider. Recent isolation of SODs from Streptomyces species revealed a nickel-containing SOD (Ni-SOD) [15, 16]. The amino acid sequence analysis of the enzyme exhibited no sequence homology to any known SODs [17]. The finding of Ni-SOD is also of interest because copper, iron, and manganese ions can catalyze disproportionation of superoxide anions in aqueous solution but nickel ion does not, indicating that Ni-SOD is a new class among the SODs. The 3D coordination environments and the electronic structure of the metal site of Ni-SOD should play a key role both in fine-tuning the redox potential of the metal site and in the enzymatic mechanism.
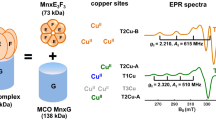
Ni-SOD isolated from Streptomyces seoulensis was originally reported as a tetramer but now is known as a hexamer in its catalytically active form. Each subunit consists of 117 amino acids and one nickel ion with molecular mass of 13.4 kDa [18, 19]. Electron paramagnetic resonance (EPR) studies of the oxidized (resting) Ni-SOD showed that the active site contains Ni(III) ion with g = [2.30 2.24 2.01] (S = 1/2) and that an axial position is occupied by a nitrogen ligand [16, 18–20]. X-ray crystallographic analyses of both the oxidized and the reduced Ni-SODs have shown that the nickel ion resides in the N-terminal loop. The active site of the reduced form [Ni(II)] has a square-planar coordination with the amino group of His-1, the amide group of Cys-2, and two thiolate groups of Cys-2 and Cys-6 [18, 19]. The oxidized [Ni(III)] Ni-SOD has the axial ligand of the imidazole group of His-1 in addition to the equatorial ligands, forming a pentacoordinated square pyramid. The mechanistic conversion between the reduced and oxidized forms and the Ni(II)/Ni(III) redox potential are finely tuned by rotating the imidazole plane about the Cβ–Cγ bond of His-1 (Fig. 1). The mechanism of the dismutation by Ni-SOD is still controversial. X-ray structural analysis has suggested outer-sphere electron transfer [18], whereas EPR and X-ray crystallographic studies of Ni-SOD with inhibitor/inactivator have proposed inner-sphere electron transfer in which the superoxide ion is directly coordinated to the nickel ion in both oxidized and reduced forms at the side opposite His-1 imidazole [19].
Among the known redox-active nickel-containing enzymes, both carbon monoxide dehydrogenase and acetyl-coenzyme A synthase contain an active nickel center linked to an Fe–S cluster [21]. Methyl-coenzyme M reductase has a mononuclear nickel center, but there are no endogenous sulfur-donor ligands [22, 23]. [NiFe] hydrogenase also has a redox-active nickel center and is viewed as a Ni–Fe cluster bridged by two μ-thiolate sulfur atoms of cysteine residues [24]. Therefore, the active-site structure of Ni-SOD is a completely new class in redox-active nickel-containing enzymes, and the coordination and geometrical structure of the site and the related electronic structure are the keys to specifically controlling the dismutase activity of the enzyme. In the present study, we performed Q-band (35 GHz) 14N,1/2H continuous wave (CW) and pulsed electron–nuclear double resonance (ENDOR) and 14N X-band (9 GHz) electron spin echo envelope modulation (ESEEM) on the resting-state Ni(III)-SOD extracted from S. seoulensis. In-depth analyses of the data obtained from the multifrequency advanced-EPR techniques detail the electronic structure of the active site of Ni-SOD.
Materials and methods
Protein isolation
The Ni-SOD sample was prepared from S. seoulensis as described previously [15, 16], with some modifications. S. seoulensis was cultured in yeast extract/malt extract medium supplemented with 100 μM NiCl2 to increase the expression of Ni-SOD. Mycelial cells of S. seoulensis harvested by aspiration on filter paper were resuspended in buffer A (20 mM phosphate buffer, pH 7.4). After cell disruption with an Omni-mixer (Omni International), cell debris was removed by centrifugation at 12,000g for 20 min. The supernatant was brought to a final 35% ammonium sulfate saturation and the precipitate was removed by centrifugation at 12,000g for 30 min. The resulting supernatant was loaded onto a phenyl-Sepharose CL-4B FastFlow column (5 cm × 30 cm) equilibrated with buffer A containing ammonium sulfate at 35% saturation, and was eluted with a linear gradient of ammonium sulfate between 35 and 0% saturation. The fractions containing SOD activity were collected and desalted with a Sephadex G-25 column (8 cm × 60 cm) equilibrated with buffer A. The desalted fractions were applied to a Protein-Pak DEAE 5PW column (2.15 cm × 15 cm) using a Waters Delta Prep 4000 chromatography system and were separated using a linear gradient of 0.1–0.3 M NaCl. Active fractions were pooled and desalted using a YM10 membrane (Amicon), and were separated by preparative electrophoresis with a Bio-Rad model 491 prep cell apparatus using 7.5% nondenaturating polyacrylamide gel. After the active fractions had been concentrated using an Amicon YM10 membrane, the concentrated sample was loaded onto a Superdex 200 column (HiLoad 16/60) equilibrated with buffer A and was eluted with the same buffer by a fast protein liquid chromatography system (Pharmacia). The active fractions were further purified on a Mono Q column (HR 5/5) with a linear gradient of 0–0.3 M NaCl using the same fast protein liquid chromatography system. Finally, the purified Ni-SOD sample was mixed with an equal volume of ethylene glycol and loaded into the X-band or Q-band EPR quartz tube to be frozen at liquid-nitrogen temperature.
EPR/ENDOR/ESEEM measurements
X-band EPR spectra were collected at liquid-nitrogen temperature with a Bruker EMX spectrometer. Q-band CW EPR and 1H,14N ENDOR spectra were collected using a modified Q-band Varian E-110 spectrometer equipped with a liquid-helium immersion Dewar at 2 K a under rapid passage condition [25]. Q-band 2H Mims ENDOR data were obtained with a locally built Q-band pulsed-EPR spectrometer [26]. Three-pulse echo (π/2−τ−π/2−T−π/2−τ−echo) intensities were recorded as a function of radio frequency to generate 2H Mims ENDOR data [27].
The first-order ENDOR spectrum of a nucleus N with I = 1/2 in a paramagnetic center is a doublet with frequencies given by [28]
Here, ν N is the nuclear Larmor frequency and A N is the angle-dependent hyperfine coupling constant. For a nucleus with I ≥ 1, each ν ± transition is further split into 2I lines by the nuclear quadrupole interaction,
where P N is the angle-dependent quadrupole coupling constant and –I + 1 ≤ m I ≤ I. For a frozen-solution sample with a rhombic EPR signal (g 1, g 2, g 3), the full hyperfine and quadrupole tensors, including both the principal values and the orientations in the g-tensor axis frame, of the hyperfine-coupled nucleus can be obtained from the analysis of the 2D datasets comprising numerous ENDOR spectra collected across the EPR envelope, as described elsewhere [29–32].
X-band 14N ESEEM experiments were performed on a locally built X-band pulsed-EPR spectrometer at 4 K [33]. Three-pulse echo (π/2−τ−π/2−T−π/2−τ−echo) intensities were recorded as a function of the time interval T to construct the time-domain ESEEM data [34]. The π/2 microwave pulse duration was 16 ns with power of approximately 1 W. The frequency-domain spectra were obtained through Fourier transform of the time-domain data by modifying the “dead-time reconstruction” procedure originally developed by Mims [35].
Results
Q-band 14N CW ENDOR
The resting-state Ni-SOD shows a well-resolved rhombic S = 1/2 EPR signal with g = [2.30 2.24 2.01] originating from a Ni(III) center. The g 3 (2.01) EPR signal is further split into three lines by 25 G (70 MHz) of the hyperfine coupling to an axial nitrogen (I = 1) ligand [16, 18–20]. Figure 2a shows the 20–45-MHz region of the Q-band CW ENDOR spectra obtained across the EPR envelope of the resting-state Ni-SOD. The “single-crystal-like” ENDOR spectrum obtained at the high-field edge (g 3) of the EPR spectrum shows two doublets centered at 35.2 MHz (circles in Fig. 2a) and each doublet splits by 3.0 MHz (“goal-post” marks in Fig. 2a). The centers of the two doublets are separated by 7.7 MHz, which is twice the 14N Larmor frequency at the experimental magnetic field, indicating that the signal originates from a hyperfine-coupled 14N nucleus; the observed ENDOR signal is hyperfine-centered [35.2 MHz = A(g 3)/2] and the separation within each doublet arises from the nuclear quadrupole coupling [3.0 MHz = 3P(g 3)] (Eq. 2). Such a four-line pattern is also well observed at g 1, the other “single-crystal-like” field position. The 14N ENDOR signals are well followed across the EPR absorption envelope, although ENDOR line distortion is observed around g 2 because of slow nuclear relaxation, which is often the case in Q-band “rapid passage” ENDOR experiments. The observed field-dependent (orientation-selective) 14N ENDOR data are well simulated, as shown in Fig. 2b, with the hyperfine coupling tensor A = [55 55 70.4] MHz and the nuclear quadrupole coupling tensor P = [0.4 0.6 −1.0] MHz, both of whose principal axes are coaxial to the g-tensor frame [29–32] (Table 1).
a Q-band (35 GHz) 14N continuous wave (CW) electron–nuclear double resonance (ENDOR) spectra of the resting-state (oxidized-state) Ni-SOD and b their corresponding numerical simulations. Dashed lines guide the change of the ENDOR pattern across the electron paramagnetic resonance (EPR) envelope. The experimental conditions were as follows: microwave frequency 35.252 GHz, modulation amplitude 1.3 G, radio-frequency (RF) power 20 W, RF sweep speed 1 MHz/s, and temperature 2 K. The simulation parameters were as follows: EPR linewidth 300 MHz and ENDOR linewidth 1 MHz. Other parameters were as in Table 1
X-band 14N ESEEM
Figure 3a displays frequency-domain 14N ESEEM spectra, obtained from Fourier transform of the time-domain ESEEM data, across the EPR of Ni-SOD. The numerical simulation of the data was obtained with the method developed previously [36]. Figure 3b depicts the results in a contour plot. The 14N ESEEM collected at g 3 shows three well-resolved lines at about 0.55, about 1.05, and about 1.6 MHz, whose frequencies are almost invariant across the EPR envelope. Also one relatively broad band is observed at about 5.0 MHz in the 14N ESEEM spectrum of g 3. The position of this band shifts to lower frequencies as the g-value increases (or the magnetic field decreases).
a X-band (9 GHz) 14N three-pulse electron spin echo envelope modulation (ESEEM) frequency-domain (Fourier transform) spectra of the resting-state (oxidized-state) Ni-SOD and b their corresponding numerical simulations in a 2D contour plot. The experimental conditions were as follows: microwave frequency 9.714 GHz, τ ~ 140 ns, shot repetition time 24 ms, and temperature 4 K. For the simulation, the 14N nuclear hyperfine and quadrupole tensors were as in Table 1
This four-line pattern (three field-independent lines and one field-dependent band) of 14N ESEEM is often observed when the nuclear Zeeman and half the nuclear hyperfine interactions are comparable. For this coupling condition, the nuclear Zeeman and hyperfine interactions are opposed and largely cancel in one electron-spin manifold, resulting in the nuclear states becoming nuclear quadrupole states [30]. All three nuclear transition lines from this manifold are primarily determined by the nuclear quadrupole coupling constant, which is independent of the external magnetic field. Therefore, the lines are usually narrow and the frequencies are mostly independent of the field strength. In the other electron spin manifold, the nuclear Zeeman and hyperfine interactions are additive. Single-quantum transitions (Δm I = ±1) in this manifold depend on the orientation of the magnetic field and give rise to a broad powder-ESEEM spectrum, seldom observed in time-domain experiment. However, the double-quantum transition (Δm I = ±2) is weakly dependent on the orientation of the magnetic field and thus is readily detected. The double-quantum transition frequency varies according to the strength of the magnetic field because it is dependent on the nuclear Zeeman interaction as well as the nuclear hyperfine and quadrupole tensors [36].
Numerical simulation (Fig. 3b), based on the frequency-domain orientation-selective 14N ESEEM analysis method [36], shows that the observed 14N nucleus has a nuclear hyperfine coupling tensor of A = [1.9 2.0 2.7] MHz and a nuclear quadrupole coupling constant of e 2 qQ = 1.65 MHz with an asymmetry parameter of η = 0.57. The simulation found that the lineshape and the field-dependent frequency position of the double-quantum transition band were sensitive to the relative orientation of the hyperfine tensor with respect to the g-tensor frame, whereas those of the three narrow transition lines of the nuclear quadrupole states were sensitive to the relative orientation of the quadrupole tensor, yielding Euler angles of [α β γ] = [90° 10° 90°] for the hyperfine tensor and [α β γ] = [0° 65° 0°] for the quadrupole tensor with respect to the g-tensor frame. An unresolved 14N ESEEM feature found in the 2–4-MHz region (Fig. 3a) may come from weakly coupled 14N or the Δm I = ±1 transition.
Q-band 1/2H ENDOR
The left panel in Fig. 4a shows the Q-band 1H CW ENDOR spectra obtained across the EPR envelope of the resting-state Ni(III)-SOD. The ENDOR spectrum collected at the low-field edge (g 1) exhibits at least three doublets, centered at the 1H Larmor frequency, arising from the protons hyperfine-coupled to the Ni(III) center. The field-dependent patterns of the signals are not completely resolved owing to overlaps of the powder-ENDOR lineshapes. However, the outermost ν+ pattern of the strongly coupled (10.2 MHz at g 1) 1H signals are easily followed (dark lines in Fig. 4a). These strongly coupled signals were absent in the 1H ENDOR spectra of Ni-SOD incubated in D2O buffer, indicating the signals arise from a solvent-exchangeable proton. As seen in the right panel in Fig. 4a, the ENDOR pattern of the exchangeable proton is well simulated by a through-space point-dipole approximation with an effective distance of r eff(Ni(III)–1H) = 2.55 Å and an isotropic coupling of A iso(1H) = 1.0 MHz. The Ni(III) → 1H vector is estimated as lying at θ = 70° and ϕ = 30° with respect to the g-tensor frame. The 2H nucleus corresponding to the exchangeable proton can be detected by 2H Mims ENDOR of Ni-SOD incubated in D2O buffer. Figure 4b shows such data (solid lines) along with the simulation (dotted lines) using the corresponding 2H hyperfine coupling constant (|A(1H)/A(2H)| = 6.51) and the orientation vector, confirming that the strongly coupled proton observed by 1H CW ENDOR is solvent-exchangeable. 2H Mims ENDOR also reveals other solvent-exchangeable protons with |A(2H)| ≤ 0.75 MHz, corresponding to |A(1H)| ≤ 5.0 MHz (r eff(Ni–H) ≥ 3.2 Å).
a Q-band (35 GHz) 1H CW ENDOR spectra of the resting-state (oxidized-state) Ni-SOD (left) and their corresponding numerical simulations for the solvent-exchangeable strongly coupled proton (right). The spectra are centered at the proton Larmor frequencies. Dark lines denote the outermost pattern of the ν+ line of the solvent-exchangeable strongly coupled 1H signals. The experimental conditions were as follows: microwave frequency 35.255 GHz, modulation amplitude 1.3 G, RF power 20 W, RF sweep speed 1 MHz/s, and temperature 2 K. The simulation parameters were as follows: EPR linewidth 200 MHz and ENDOR linewidth 0.5 MHz. The other parameters were as in Table 1. b Q-band (35 GHz) 2H Mims ENDOR spectra of the resting-state (oxidized-state) Ni-SOD in D2O buffer (solid lines) and their corresponding numerical simulations for the strongly coupled 2H (dashed lines). The spectra are centered at the deuterium Larmor frequencies. Corresponding proton frequencies centered at the proton Larmor frequencies are marked at the top. Dark lines denote the outermost patterns of the ν+ and ν− lines of the solvent-exchangeable strongly coupled 2H signals. The experimental conditions were as follows: microwave frequency 34.858 GHz, τ = 332 ns, microwave pulse width 52 ns, RF pulse width 30 μs, shot repetition time 10 ms, and temperature 2 K. The simulation parameters were as follows: EPR linewidth 200 MHz, ENDOR linewidth 0.05 MHz, the 2H hyperfine coupling tensor values were 1/6.51th of the 1H hyperfine coupling tensor values and the orientation was the same, and the nuclear quadrupole tensor P = [0.07 0.07 −0.14] MHz with Euler angles of (90° 35° 0°)
Discussion
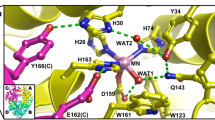
The EPR spectrum of the resting-state Ni-SOD shows a well-resolved rhombic S = 1/2 EPR signal with g = [2.30 2.24 2.01]. The g-tensor feature is well characterized as a Ni(III) center with a \( d_{{z}^{2}} \) ground state in a tetragonally distorted octahedral or square-pyramidal geometry. A slight contribution of \( d_{{{{x}}^{2} - {{y}}^{2} }} \) to the ground state causes g 3 (2.01) to be slightly larger than 2.00. The rhombicity of the g tensor indicates that the d xz and d yz orbitals are not degenerate. The g 3 (2.01) EPR signal is further split into three lines by 25 G (70 MHz) of the hyperfine coupling to an axial nitrogen (I = 1) ligand. The axial nitrogen has been identified as Nδ of His-1 imidazole of Ni-SOD by X-ray crystallographic studies of the oxidized enzyme [18, 19]. Further, the structural study revealed that the equatorial plane of the Ni(III) site is occupied by the amino group of His-1, the amide group of Cys-2, and two thiolate groups of Cys-2 and Cys-6, to form a square-pyramidal coordination geometry with the axial nitrogen.
Analysis of Q-band 14N CW ENDOR data (Fig. 2) yields the hyperfine coupling tensor, A = [55 55 70.4] MHz, and the nuclear quadrupole coupling tensor, P = [−0.4 −0.6 1.0] MHz, of the axial nitrogen (Table 1). The large isotropic hyperfine coupling (A iso = 60 MHz) is comparable to those of axial nitrogen ligands of Ni(III) complexes with square-pyramidal geometry [37–41] and results from spin delocalization through the σ-bond between the metal \( d_{{z}^{2}} \) and the axial nitrogen (Nδ), as is common for Ni(III)–imidazole complexes.
For metal–nitrogen complexes, the 14N nuclear quadrupole coupling is sensitive to details of the metal-nitrogen bond. The largest 14N nuclear quadrupole coupling can be either along the metal–nitrogen bond (nitrogen lone pair direction) or normal to the metal–nitrogen bond. The former shows a negative coupling constant, whereas the latter, occurring with increasing nitrogen-to-metal donation as found in copper(II) tetraphenylporphyrin and aquometmyoglobin, shows a positive coupling constant [42]. In Ni-SOD, the 14N nuclear quadrupole and nuclear hyperfine coupling tensors of the axial nitrogen are collinear with the g tensor (Table 1) and the largest 14N nuclear quadrupole coupling (P 3) is along the Ni(III)–Nδ bond. For the tetragonally distorted (or square-pyramidal) S = 1/2 Ni(III), the orientations of the d yz and d xz orbitals affect the directions of the g 1 (equivalent to g x ) and g 2 (equivalent to g y ) components of the g tensor, respectively. The collinearity of the g and 14N nuclear quadrupole tensors can be explained by metal-to-nitrogen π back-donation. A favorable orientation of either the filled d yz or the filled d xz orbital for back-donation to the empty nitrogen p π orbital [43], normal to the imidazole plane, strengthens the metal–nitrogen bond and causes the coincidence of the g and P tensors. In other words, for the tetragonally distorted (or square-pyramidal) S = 1/2 Ni(III) complexes with axial imidazole, the g-tensor direction is modulated by the imidazole plane.
X-band 14N ESEEM measurements (Fig. 3) identified one well-defined signal from a 14N atom with the hyperfine tensor A = [1.9 2.0 2.7] MHz, plus the possibility of other weakly coupled nitrogen(s). There are four nitrogen atoms present within the magnetic-coupling range of the Ni(III) center of Ni-SOD; the imidazole Nδ and Nε of His-1, the terminal amine nitrogen of His-1, and the amide nitrogen of Cys-2. The strongly coupled imidazole Nδ of His-1 was characterized by Q-band 14N CW ENDOR (Fig. 1). For spin-bearing metal-bound imidazole, the hyperfine coupling of the nitrogen directly bound to the metal is in general 20–30 times bigger than that of the remote nitrogen. This enables us to assign the well-defined nitrogen signal seen by ESEEM to the imidazole Nε of His-1 (remote nitrogen). In-plane coordinated 14N atoms have minimal coupling to an electron in the \( d_{{z}^{2}} \) orbital.
The crystallographic studies of the reduced Ni-SOD have shown that the imidazole of His-1 donates two hydrogen bonds, Nδ–H···OVal-8 and Nε–H···OGlu-17. Upon oxidation of the enzyme, the hydrogen bond of Nε–H···OGlu-17 remains, whereas that of Nδ–H···OVal-8 is broken and Nδ is bound to Ni(III) [18, 19]. The 14N nuclear quadrupole coupling parameters of the remote nitrogen of a metal-bound imidazole have been widely utilized to study the chemical environment, which tunes the electron density of the remote nitrogen. The 14N nuclear quadrupole coupling constant e 2 qQ = 1.67 MHz and the asymmetry parameter η = 0.57 of the remote nitrogen (Nε) of His-1 of Ni-SOD are close to the values expected when there is a weak or no hydrogen bonding to the remote N–H [44, 45]. This suggests that there is a subtle change in the hydrogen-bonding network around the active sites in crystal and solution phases in the Ni(III) state.
Q-band 1H CW ENDOR and pulsed 2H Mims ENDOR show that the hyperfine couplings (10.2 MHz at g 1) to the strongly coupled solvent-exchangeable proton(s) are mostly dipolar (Fig. 4). From the point-dipole approximation, assuming 100% spin on Ni(III), the effective distance (r eff) between Ni(III) and the proton is estimated to be approximately 2.55 Å (Table 1). The Ni(III) → H vector with θ = 70° and ϕ = 30° in the g-tensor frame indicates that the coupled proton(s) is near the g 1–g 2 plane (equatorial ligand plane). The crystal structure shows the distance from Ni(III) to the amine nitrogen of His-1 is 2.182 Å. Assuming the bond length of the amine N–H is about 1 Å and the Ni(III)–N–H angle adopts near tetrahedral geometry, we estimate the Ni(III)–Hamine distance to be approximately 2.3 Å. This enables us to assign the proton giving rise to the hyperfine coupling to the proton(s) of the terminal amine nitrogen of His-1. The difference between ENDOR and crystallographic distances reflects the delocalization of the spin density.
Q-band 1/2H ENDOR also found weakly coupled solvent-exchangeable proton(s), for which the effective distance is no less than 3.2 Å (|A(1H)| ≤ 5.0 MHz). In the framework of the protein backbone, solvent-exchangeable protons are not present within the detected hyperfine-coupling range. The crystal structure shows a vacant pocket, enclosed by the backbone and side chains of Cys-2 through Cys-6, on the side opposite the axial Nδ. This pocket embeds a water molecule at a distance of 3.427 Å from Ni(III) in the crystal. The weakly coupled solvent-exchangeable proton thus seems to be the proton(s) of this water. But, as a kind reviewer pointed, the possibility of the proton of a water molecule being hydrogen-bonded to the equatorial thiolate ligand cannot be ruled out.
In summary, we performed ENDOR and ESEEM experiments to obtain details on the electronic structure and magnetic couplings of the Ni(III) site of Ni-SOD. The analysis of the field-dependent Q-band 14N CW ENDOR determined the nuclear hyperfine and quadrupole coupling tensors of the axial 14N ligand of Ni(III), Nδ of His-1 imidazole. The tensors are coaxial with the g-tensor frame, implying the g-tensor direction is modulated by the imidazole plane. X-band 14N ESEEM characterizes the weakly coupled Nε of His-1 imidazole. The nuclear quadrupole coupling constant of this nitrogen suggests that the hydrogen-bonding between Nε–H and OGlu-17 present for the reduced-state Ni-SOD is weakened or broken in the Ni(III) enzyme in aqueous phase. Q-band 1H CW ENDOR and pulsed 2H Mims ENDOR distinguished two kinds of solvent-exchangeable protons: one is the equatorial amine proton(s) of His-1 and the other is either the proton(s) of a water in the pocket at the side opposite the axial Nδ or the proton of a water hydrogen-bonded to the equatorial thiolate ligand.
Abbreviations
- CuZn-SOD:
-
Copper- and zinc-containing superoxide dismutase
- CW:
-
Continuous wave
- ENDOR:
-
Electron–nuclear double resonance
- EPR:
-
Electron paramagnetic resonance
- ESEEM:
-
Electron spin echo envelope modulation
- Ni-SOD:
-
Nickel-containing superoxide dismutase
- SOD:
-
Superoxide dismutase
References
Fridovich I (1997) J Biol Chem 272:18515–18517
Cabelli DE, Riley D, Rodriguez JA, Valentine JS, Zhu H (2000). In: Meunier B (ed) Biomimetic oxidations catalyzed by transition metal complexes. Imperial College Press, London, chap 10
Miller A-F (2003) In: Que L Jr, Tolman W (eds) Coordination chemistry in the biosphere and geosphere. Pergamon, Amsterdam, pp 479–506
Miller A-F (2004) Curr Opin Chem Biol 8:162–168
Tainer JA, Getzoff ED, Richardson JS, Richardson DC (1983) Nature 306:284–287
Bordo D, Djinovic-Carugo K, Bolognesi M (1994) J Mol Biol 238:366–368
Djinovic-Carugo K, Battistoni A, Carri M, Polticelli F, Desideri A, Rotilio G, Coda A, Wilson K, Bolognesi M (1996) Acta Crystallogr D 52:176–188
Bordo D, Matak D, Djinovic-Carugo K, Rosano C, Pesce A, Bolognesi M, Stoppolo ME, Falconi M, Battistoni A, Desideri A (1999) J Mol Biol 285:283–296
Parker MW, Blake CC (1988) J Mol Biol 199:649–661
Ludwig ML, Metzger AL, Pattridge KA, Stallings WC (1991) J Mol Biol 219:335–358
Borgstahl GE, Parge HE, Hickey MJ, Beyer WF Jr, Hallewell RA, Tainer JA (1992) Cell 71:107–118
Edwards RA, Whittaker MM, Whittaker JW, Jameson GB, Baker EN (1998) J Am Chem Soc 120:9684–9685
Lah MS, Dixon MM, Pattridge KA, Stallings WC, Fee JA, Ludwig ML (1995) Biochemistry 34:1646–1660
Guan Y, Hickey MJ, Borgstahl GEO, Hallewell RA, Lepock JR, O’Connor D, Hsieh Y, Nick HS, Silverman DN, Tainer JA (1998) Biochemistry 37:4722–4730
Youn H-D, Kim E-J, Roe J-H, Hah YC, Kang S-O (1996) Biochem J 318:889–896
Youn H-D, Youn H, Lee J-W, Yim Y-I, Lee J-K, Hah YC, Kang S-O (1996) Arch Biochem Biophys 334:341–348
Dupont CL, Neupane K, Shearer J, Palenik B (2008) Environ Microbiol 10:1831–1843
Wuerges J, Lee J-W, Yim Y-I, Yim H-S, Kang S-O, Djinovic-Carugo K (2004) Proc Natl Acad Sci USA 101:8569–8574
Barondeau DP, Kassmann CJ, Bruns CK, Tainer JA, Getzoff ED (2004) Biochemistry 43:8038–8047
Choudhury SB, Lee J-W, Davidson G, Yim Y-I, Bose K, Sharma ML, Kang S-O, Cabelli DE, Maroney MJ (1999) Biochemistry 38:3744–3752
Evans DJ (2005) Coord Chem Rev 249:1582–1595
Kraulis PJ (1991) J Appl Crystallogr 24:946–950
Merritt EA, Murphy MEP (1994) Acta Crystallogr D 50:869–873
Fontecilla-Camps JC, Volbeda A, Cavazza C, Nicolet Y (2007) Chem Rev 107:4273–4303
Werst MM, Davoust CE, Hoffman BM (1991) J Am Chem Soc 113:1533–1538
Davoust CE, Doan PE, Hoffman BM (1996) J Magn Reson 119:38–44
Schweiger A, Jeschke G (2001) Principles of pulse electron paramagnetic resonance. Oxford University Press, Oxford
Abragam A, Bleaney B (1986) Electron paramagnetic resonance of transition ions. Dover, New York
Hoffman BM, Martinsen J, Venters RA (1984) J Magn Reson 59:110–123
Hoffman BM, Venters RA, Martinsen J (1985) J Magn Reson 62:537–542
Hoffman BM, DeRose VJ, Doan PE, Gurbiel RJ, Houseman ALP, Telser J (1993) Biol Magn Reson 13:151–218
Doan PE (2003) In: Telser J (ed) Paramagnetic resonance of metallobiomolecules. American Chemical Society, Washington, pp 55–81
Fan CL, Doan PE, Davoust CE, Hoffman BM (1992) J Magn Reson 98:62–72
Dikanov SA, Tsvetkov YD (1992) Electron spin echo envelope modulation (ESEEM) spectroscopy. CRC, Boca Raton
Mims WB (1984) J Magn Reson 59:291–306
Lee H-I, Doan PE, Hoffman BM (1999) J Magn Reson 140:91–107
Lappin AG, Murray CK, Margerum DW (1978) Inorg Chem 17:1630–1634
Lovecchio FV, Gore ES, Busch DH (1974) J Am Chem Soc 96:3109–3118
Pinho D, Gomes P, Freire C, de Castro B (2001) Eur J Inorg Chem 2001(6):1483–1493
Wang YL, Beach MW, Pappenhagen TL, Margerum DW (1988) Inorg Chem 27:4464–4472
Seth J, Palaniappan V, Bocian DF (1995) Inorg Chem 34:2201–2206
Brown TG, Hoffman BM (1980) Mol Phys 39:1073–1109
Telser J, Fann Y-C, Renner MW, Fajer J, Wang S, Zhang H, Scott RA, Hoffman BM (1997) J Am Chem Soc 119:733–743
Jiang F, McCracken J, Peisach J (1990) J Am Chem Soc 112:9035–9044
Colaneri MJ, Peisach J (1992) J Am Chem Soc 114:5335–5341
Mathews J, Walker RL (1965) Mathematical methods of physics. Benjamin, Elmsford
True E, Nelson MJ, Venters RA, Orme-Johnson WH, Hoffman BM (1988) J Am Chem Soc 110:1935–1943
Acknowledgments
This work was supported by the Korea Research Foundation (KRF-2006-312-C00219, H.I.L.), the Research Fellowship of BK21 project (S.O.K.), and the NSF (MCB0723330, B.M.H.).
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Lee, HI., Lee, JW., Yang, TC. et al. ENDOR and ESEEM investigation of the Ni-containing superoxide dismutase. J Biol Inorg Chem 15, 175–182 (2010). https://doi.org/10.1007/s00775-009-0581-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00775-009-0581-x