Abstract
The multi-copper oxidase (MCO) MnxG from marine Bacillus bacteria plays an essential role in geochemical cycling of manganese by oxidizing Mn2+(aq) to form manganese oxide minerals at rates that are three to five orders of magnitude faster than abiotic rates. The MCO MnxG protein is isolated as part of a multi-protein complex, denoted as Mnx, which includes one MnxG unit and a hexamer of MnxE3F3 subunit. During the oxidation of Mn2+(aq) catalyzed by the Mnx protein complex, an enzyme-bound Mn(III) species was trapped recently in the presence of pyrophosphate (PP) and analyzed using parallel-mode electron paramagnetic resonance (EPR) spectroscopy. Herein, we provide a full analysis of this enzyme-bound Mn(III) intermediate via temperature dependence studies and spectral simulations. This Mnx-bound Mn(III) species is characterized by a hyperfine-coupling value of A(55Mn) = 4.2 mT (corresponding to 120 MHz) and a negative zero-field splitting (ZFS) value of D = − 2.0 cm−1. These magnetic properties suggest that the Mnx-bound Mn(III) species could be either six-coordinate with a 5B1g ground state or square-pyramidal five-coordinate with a 5B1 ground state. In addition, as a control, Mn(III)PP is also analyzed by parallel-mode EPR spectroscopy. It exhibits distinctly different magnetic properties with a hyperfine-coupling value of A(55Mn) = 4.8 mT (corresponding to 140 MHz) and a negative ZFS value of D = − 2.5 cm−1. The different ZFS values suggest differences in ligand environment of Mnx-bound Mn(III) and aqueous Mn(III)PP species. These studies provide further insights into the mechanism of biological Mn2+(aq) oxidation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Manganese oxide minerals (MnOx) are widely distributed over the Earth’s surface and are among the most powerful natural oxidants in the environment [1]. MnOx minerals serve as the electron sink for microbial metabolism in the absence of O2 and can degrade xenobiotic organic compounds to low-molecular-mass compounds [2, 3]. Therefore, geochemical redox cycling of MnOx minerals is globally important. In the upper photic zone of the ocean, MnOx minerals undergo photo-reductive dissolution to Mn2+(aq) [1, 4], while this dissolution is counterbalanced by diurnal oxidation of Mn2+(aq) back to the minerals [5], where microorganisms are known to play an essential role [2, 3]. Biological Mn2+(aq) oxidation by molecular oxygen in seawater (pH 8.16) occurs at rates that are three to five orders of magnitude faster than abiotic pathways [6]. In marine Bacillus species (PL-12, SG-1 and MB-7), one gene, mnxG, encodes a putative multi-copper oxidase (MCO) that has been identified as an enzymatic catalyst for Mn2+(aq) oxidation [7,8,9,10].
The MCO enzymes are a family of proteins found in bacteria, fungi, plants, and animals, which contain three types of copper cofactors [11, 12]: Type 1 “blue” copper (T1Cu), Type 2 “normal” copper (T2Cu), and the “coupled binuclear” Type 3 copper (T3Cu). All are required for oxidase activity. The mechanism of the enzymatic substrate oxidation coupled with O2 being reduced to H2O in solution is well-established by Solomon and coworkers [11, 12]. Briefly, the T1Cu site accepts electrons from the substrate and shuttles them via intramolecular electron transfers (IET, kIET ≈ 0.11 s−1 at 4 °C) [13] over 13 Å through a T1–Cys–His–T3 pathway to the trinuclear T2/3Cu site (consisting of one T2Cu and one dinuclear T3Cu cluster) [11, 12]. Then, in the trinuclear T2/3Cu site, exogenous O2 binds and is rapidly reduced to water (with a second-order rate constant of keff ≈ 106 M−1 s−1) [13,14,15].
The well-studied MCOs are categorized into two groups based on their substrates. One group, including plant laccase, fungal laccase, ascorbate oxidase and bilirubin oxidase, uses organic compounds as substrates [11, 12]. The other group uses metal ions as substrates, such as Fet3p [16] and human ceruloplasmin [16,17,18] that can oxidize Fe(II), or the MCO enzyme CueO from E. coli that can oxidize Cu(I) during copper homeostasis in bacteria [19]. It is particularly noteworthy that the MnxG protein is a rare version of an MCO enzyme that can oxidize its aqueous metal ion substrate (Mn2+) to form insoluble biogenic metal oxides (MnOx) [7, 10, 20,21,22].
The MCO MnxG protein was recently isolated as part of a multi-protein complex, denoted as Mnx, where the MnxG subunit is combined with several copies of MnxE and MnxF accessory protein subunits. Mass spectrometric analysis shows that the Mnx protein complex has a molecular weight of ca. 211 kDa, with a composition of one MnxG unit (≈ 138 kDa) along with a hexamer of MnxE3F3 subunit (≈ 73 kDa), as illustrated in Fig. 1 [10, 23]. In this Mnx protein complex, two distinct classes of T2CuII sites were identified using continuous-wave (CW) electron paramagnetic resonance (EPR) spectroscopy [24]. One class of T2CuII site (denoted as T2Cu-A) resides in the MCO MnxG unit and presents magnetic parameters of g|| = 2.320 and A||(63Cu) = 510 MHz. The other class of T2CuII site (denoted as T2Cu-B) presents g|| = 2.210 and A||(63Cu) = 615 MHz and is located in the hexametric MnxE3F3 subunit [24]. These different magnetic properties correlate with their different Cu(II/I) reduction potentials; namely, as the g||-value decreases, corresponding to a larger energy gap between the dxy and \(d_{{x^{2} - y^{2} }}\) Cu(II)-based molecular orbitals (MO), the reduction potential (E°) is lowered, which is consistent with our previous protein film voltammetric studies [24,25,26]. The reduction potential of T2Cu-B sites residing in the MnxE3F3 subunit was found to be ca. 350 mV (vs NHE, normal hydrogen electrode, pH 7.8), which is ca. 50 mV lower than that of T2Cu-A sites in the MnxG unit [25, 26].
Cartoon showing the Mnx protein complex and its Cu centers. Mnx protein complex (≈ 211 kDa) consists of one MCO MnxG unit (≈ 138 kDa) and a putative MnxE3F3 hexamer (E3F3, ≈ 73 kDa) [24]. Previous EPR studies show that there are three copper sites (T2Cu-B) per MnxE3F3 hexamer. The MCO MnxG unit contains two T2Cu sites (T2Cu-A) as well as one T1Cu and one dinuclear T3Cu site. The X-band CW EPR spectra of T2Cu-B (orange trace) and T2Cu-A (green trace) are adapted from Ref. [24]
In terms of the T1Cu site bound within MnxG that accepts electrons from the substrate, the reduction potential is determined to be ca. 380 mV (vs NHE, pH 7.8) via the poised potential titration method [27], as the T1Cu site is inaccessible for direct electron transfer from the electrode [25, 26]. The reduction potential of this T1Cu is near the lower end of the known range of T1 sites in MCOs [11]. The question then becomes how does the Mnx protein with this low-potential T1Cu oxidize Mn2+(aq)? Our early EPR-spectroscopic work [20] showed that there is a Mnx-bound mononuclear Mn(II) species (denoted in that work as a class ii Mn(II) species) that is coordinated to one nitrogenous ligand and there is also a weakly exchanged-coupled dimeric Mn(II) (denoted as class iii) species. Kinetic studies [27] further suggested that the Mnx protein complex takes advantage of the polynuclear chemistry of manganese to adjust the potential of Mn(III/II) couple for an efficient electron transfer to the low-potential T1Cu, and to ultimately form MnOx minerals. Briefly, the Mnx protein complex requires an activation step, forming a hydroxide-bridged binuclear complex, Mn(II)(µ-OH)Mn(II), to decrease the reduction potential of Mn(III/II) below that of the T1Cu site [27]. Oxidation leads to a dihydroxide-bridged binuclear Mn(III) intermediate, which disproportionates in the enzyme to Mn(II) and a binuclear Mn(IV) intermediate, the precursor to MnO2. The Mn(II) is then recycled to the substrate site, allowing the turnover to continue.
We were unable to find EPR-spectroscopic signatures of higher oxidation state manganese intermediates. This could be due to the instability of the mononuclear Mn(III) ion which disproporationates rapidly unless strongly ligated to a compound such as pyrophosphate (PP), or the manganese intermediates are binuclear and spin-coupled and are, therefore, EPR silent [28]. However, in our recent work, we did find the EPR spectrum of a mononuclear Mn(III) species when PP was present to trap Mn(III) that dissociates during turnover [29].
In the present work, we provide a full analysis of this Mnx-bound Mn(III) intermediate using parallel-mode EPR spectroscopy. Electronic-structure parameters including 55Mn hyperfine-coupling A and zero-field splitting (ZFS) D of this Mn(III) species coordinated within Mnx protein are derived via temperature dependence studies and spectral simulations. These magnetic properties are distinct from those of the aqueous Mn(III)PP complex employed as a control in this work.
Experimental procedures
Mnx protein expression and purification
The Mnx protein complex was expressed and purified by optimizing previous methods described in Ref. [10] using the plasmid containing a mnxEFG gene construct inserted into the pASK/IBA3plus vector. This plasmid was transformed into E. coli BL21(DE3) and grown by shaking (≈ 200 rpm) at 37 °C to an O.D.600 ≈ 0.5–0.6 a.u. in a Luria–Bertani (LB) broth containing 0.2 mM CuSO4, and 100 mg/L ampicillin. The cells were then cooled down to 17 °C on ice (≈ 20 min) and induced by adding 100 μL of 2 mg/mL anhydrotetracycline. Protein expression was continued for 18 h by shaking (≈ 180 rpm) at 17 °C. Then, CuSO4 was added into the culture to a final concentration of 2 mM and the shaking function was turned off for another 22–24 h at 17 °C, to enable the micro-aerobic uptake of copper ions into the E. coli cytoplasm as described by Durao et al. [30].
The cells were harvested by centrifugation (6000×g at 4 °C for 30 min) and re-suspended in Strep-tactin equilibration buffer (20 mM Tris–HCl, 150 mM NaCl, pH 8.0, and 50 µM CuSO4) supplemented with 10 mM CaCl2, 1 mM CuSO4, and an EDTA-Free SIGMAFAST™ Protease Inhibitor Cocktail Tablet. The cells were lysed by two rounds of French press at 1000 psi and the crude extract was clarified by heat denaturation at 70 °C for 15 min. The cell debris was removed by centrifugation (13,000×g at 4 °C for 30 min) and the supernatant was filtered through a 0.4-µm pore polyvinylidene fluoride (PVDF) filter. The clarified supernatant was added to a 10-mL column volume (CV) of Strep-tactin Superflow Plus resin (QIAGEN) and slowly rotated for 1 h at room temperature. The unbound protein fraction was removed by gravity flowing through the resin and the resin was washed with 100 mL Strep-tactin equilibration buffer. The Mnx protein fraction was eluted by adding 50 mL of 2.5 mM d-desthiobiotin in Strep-tactin equilibration buffer. The resin was regenerated with 200 mL of 1 mM 2-(4-hydroxyphenylazo)benzoic acid and washed with 500 mL Strep-tactin equilibration buffer. The eluted Mnx protein fraction was concentrated to < 1.0 mL in the filtration with 100 kDa molecular weight cutoff (Millipore) and loaded into HiLoad™ 16/600 Superdex™ 200 pg (GE Healthcare) gel-filtration column equilibrated with 20 mM HEPES buffer (pH 7.8) with 50 mM NaCl and 5% d-glucose (weight/volume) at 4 °C.
Fractions corresponding to a single broad peak (≈ 211 kDa protein complex) were collected, concentrated, and dialyzed three times (at least 3 h each) with a volume of 1 L HEPES-buffered solution (20 mM HEPES, 50 mM NaCl, pH 7.8) for every 1 mL protein sample at 4 °C. The protein was further dialyzed with 500 mL Tris-buffered solution (20 mM Tris–HCl, 50 mM NaCl, pH 8.0) for 1–2 h to remove exogenous Cu(II) to give a clean EPR spectrum (≈ 6 Cu(II)/Mnx) [24]. The protein was quantified by Thermo Scientific Pierce bicinchoninic acid (BCA) protein assay or by the extinction coefficient of T1Cu (ε590 nm = 5600 M−1cm−1) [24] in the Mnx protein complex, giving a yield of ≈ 2 mg/L culture. The final protein solution was flash frozen in liquid nitrogen and stored at −80 °C.
Sample preparation
Mn(III) pyrophosphate (Mn(III)PP) was prepared by aerobically adding Mn(III) acetate to a Na4P2O7 buffer solution (20 mM HEPES, 20 mM NaCl, pH 7.8). Then the precipitate was filtered using a 0.4-µm PVDF filter. The concentration of the resulting transparent pink Mn(III)PP solution was determined via the extinction coefficient of ε258 nm = 6200 M−1 cm−1 [31, 32]. Mn(III)PP reaction sample with Mnx protein was prepared by first transferring 100 μL as-isolated Mnx protein (200 μM) into the X-band EPR tube, followed by aerobically adding 100 μL Mn(III)PP solution (400 μM). After being allowed to react for 5 min, the sample was frozen in liquid nitrogen and analyzed by CW EPR spectroscopy.
The Mnx-bound Mn(III) intermediate was generated as follows: 100 μL of 200 μM as-isolated Mnx protein, pre-incubated with 1 mM Na4P2O7, was aerobically mixed with 100 μL of 800 μM MnSO4 buffer solution (20 mM HEPES, 20 mM NaCl, pH 7.8). After being allowed to react for 3 min, the sample was frozen in liquid nitrogen and analyzed by CW EPR spectroscopy.
A reaction mixture of the Mnx protein with Mn2+(aq) in the presence of the inhibitor sodium azide (NaN3) was prepared as follows: 100 μL of 100 μM as-isolated Mnx protein and 1 mM NaN3 were aerobically mixed with 100 μL of 400 μM MnSO4 buffer solution (20 mM HEPES, 20 mM NaCl, pH 7.8). After being allowed to react for 2 min, samples were frozen in liquid nitrogen and analyzed by CW EPR spectroscopy.
EPR spectroscopy
X-band CW EPR spectra were recorded using a Bruker (Billerica, MA) EleXsys E500 spectrometer. Cryogenic temperatures were achieved and controlled using an ESR900 liquid helium cryostat in conjunction with a temperature controller (Oxford Instruments ITC503) and gas flow controller. For all the parallel-mode EPR experiments (B0 || B1), a dual-mode resonator (ER4116DM) was employed, while for perpendicular-mode EPR (B0 ⊥ B1) a super-high Q resonator (ER4122SHQE) was used. All CW EPR data were collected under slow-passage, non-saturating conditions. Spectrometer settings were as follows: conversion time = 40 ms, modulation amplitude = 0.8 mT, and modulation frequency = 100 kHz; other settings are given in corresponding figure captions. Simulations of the spectra were performed using the Easyspin 5.1.10 toolbox [33, 34] within the Matlab 2014a software suite (The Mathworks Inc., Natick, MA).
Results and discussion
Five- or six-coordinate Mn(III) ions (3d4) are typically high-spin with a total spin of S = 2. For such integer spin system, due to generally large ZFS values (D ≈ 2 cm−1), the conventional perpendicular polarization (the oscillating magnetic field of the incident microwave radiation B1 is applied perpendicular to the direction of the static magnetic field, B0 ⊥ B1) EPR techniques reveal no spin transitions (∆Ms = ± 1) at X-band microwave frequencies (~ 9.38 GHz, corresponding to ~ 0.3 cm−1). However, using parallel polarization (B0 ‖ B1), transitions between the Ms= | ± 2 > spin manifold of S = 2 integer spin system (e.g., Mn(III) [35, 36] and Fe(II) [37, 38]) can be detected. Besides the parallel-mode EPR, high-frequency and high-field EPR (HFEPR) [39, 40] can also be applied to the integer spin system (e.g., Mn(III) [40,41,42] and Cr(II) [43]), see Ref. [40] for details.
The high-spin 3d4 Mn(III) ion has a 5D ground term. As illustrated in Fig. 2, when placed in a ligand field, a five-coordinate Mn(III) could either have a trigonal-bipyramidal (TBP) geometry with a 5A1 ground state or a square-pyramidal (SP) geometry with a 5B1 ground state. As for a six-coordinate center, Mn(III) adopts an octahedral (Oh) geometry. In an octahedral field, the 5D ground term will first split into a 5T2g excited state and a 5Eg ground state. Further, due to the spin–orbit coupling and Jahn–Teller distortions, the degeneracy of the 5Eg ground state is lifted, giving rise to either a 5A1g or 5B1g ground state [36, 44]. For the 5A1 ground state in a TBP geometry [45] or the 5A1g ground state in an Oh geometry, the lowest unoccupied metal-centered MO is 3 \(d_{{z^{2} }}\)-based, giving a positive ZFS value D due to the second-order effects of the spin-orbital coupling [35, 46, 47]. For the 5B1 ground state in a SP geometry or the 5B1g ground state in an Oh geometry, the lowest unoccupied metal-centered MO is 3 \(d_{{x^{2} - y^{2} }}\)-based, giving a negative ZFS value D (with an exception of a tetragonally elongated Mn(III) compound [Mn(cyclam)I2]I (cyclam = 1,4,8,11-tetraazacyclotetradecane), which has a positive D [48]) [45, 49].
EPR characterization of Mn(III)PP
A representative X-band (9.38 GHz) parallel-mode CW EPR spectrum of the aqueous Mn(III)PP standard is presented in Fig. 3 (red trace). A sextet is centered at the magnetic field position (81.0 mT) corresponding to geff = 8.20, and shows 55Mn (I = 5/2) hyperfine splittings of ~ 4.8 mT (corresponding to 140 MHz). This small 55Mn A-value is comparable to that reported for six-coordinate Mn(III) species, such as Mn(III) salen [49], Mn(III) in Bacillus subtilis oxalate decarboxylase [50] and the Mn(III)-hexaaqua ion [51, 52] (see Table 1 for details). Mn(III)PP EPR signals (Fig. 4a) exhibit Curie law behavior—the signal intensity is inversely proportional to the temperature—indicating that the Ms= | ± 2 > spin manifold is populated at all temperatures from 5 to 20 K. This is true only when the Ms= | ± 2 > spin manifold lies lowest in energy, corresponding to a negative sign of the ZFS. The temperature dependence is well-simulated (Fig. 4b) using the ZFS parameters of D = − 2.5 cm−1 and E = 0.25 cm−1, as shown in Fig. 4c. This temperature-dependent behavior is in contrast to that of Mn(III) species with positive ZFS, such as certain forms of manganese superoxide dismutase (EcMnSOD, D = + 2.1 cm−1, E = 0.24 cm−1 [35], CaMnSOD, D = + 1.90 cm−1, E = 0.20 cm−1) [47], in which the Ms= | ± 2 > spin manifold lies highest in energy. Thus, the EPR transitions within this manifold become apparent only by increasing the temperature to thermally populate the donor spin level.
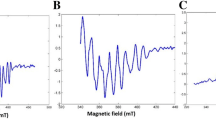
X-band (9.38 GHz) parallel-mode EPR spectra of Mn(III)PP (red trace) and two equivalents of Mn(III)PP mixed with the as-isolated Mnx protein for 5 min aerobically (blue trace). Experimental parameters: temperature = 8 K; microwave frequency = 9.385 GHz; microwave power = 10 mW; conversion time = 40 ms; modulation amplitude = 0.8 mT; modulation frequency = 100 kHz
Temperature dependence of the parallel-mode EPR spectra of Mn(III)PP (a) and the corresponding simulated spectra obtained using the parameters of 55Mn A = 140 MHz, D = − 2.5 cm−1 and E = 0.25 cm−1 (b). Experimental parameters: microwave frequency = 9.385 GHz; microwave power = 10 mW (no saturation); conversion time = 40 ms; modulation amplitude = 0.8 mT; modulation frequency = 100 kHz. c Temperature dependence of the peak-to-peak amplitudes of the fourth EPR feature of Mn(III) sextet (peak at 84.38 mT). The peak-to-peak amplitudes are plotted as a function of the inversion of temperature, with the red circles corresponding to the experimental data shown in a and the blue stars corresponding the simulation data given in b
The small hyperfine-coupling value of A = 4.8 mT, as well as the negative ZFS value of D = − 2.5 cm−1, suggests that Mn(III)PP could either have a hexa-coordinated Oh geometry with a 5B1g ground state or a five-coordinated square-pyramidal geometry with a 5B1 ground state, in which the lowest unoccupied metal-centered MO is 3 \(d_{{x^{2} - y^{2} }}\) –based.
EPR characterization of the trapped Mn(III) species in Mnx protein
To gain insights into the Mn(III)-binding sites in Mnx, we first aerobically mixed the as-isolated Mnx protein complex with two equivalents of Mn(III)PP in a HEPES-buffered solution. After aging for 5 min, the sample was frozen in liquid nitrogen and analyzed by X-band (9.38 GHz) parallel-mode CW EPR spectroscopy, shown in Fig. 3 (blue trace). No distinct new signals corresponding to Mn(III) species were detected, only the sextet identical to the Mn(III)PP signals was seen. Also, the perpendicular-mode EPR spectrum only shows EPR signals from copper sites in the Mnx protein complex (data not shown), indicating that there is no Mn(II) species being generated from the disproportionation of Mn(III)PP during this modest incubation time. Therefore, the Mn(III) ion must have much higher binding affinity to the chelator PP (kcomplexation > 1020) [53] than to the Mnx protein complex. This result is consistent with previous kinetic studies [29], which suggest that the reaction between Mn(III)PP and Mnx is second order by forming a hydroxide-bridged dinuclear Mn(III) enzyme-bound intermediate. This dinuclear Mn(III) species is expected to be spin-coupled, and possibly not EPR detectable [28].
As a control, we looked for the Mn(III) intermediate during Mnx-catalyzed Mn2+(aq) oxidation by employing sodium azide (NaN3), which inhibits MCO enzymes by binding to the trinuclear copper center [54, 55]. The as-isolated Mnx protein complex, pre-incubated with ten equivalents of NaN3, was mixed with four equivalents of MnSO4 in a HEPES-buffered solution. After being allowed to react for 2 min, the sample was frozen in liquid nitrogen and analyzed by X-band (9.38 GHz) CW EPR spectroscopy. The perpendicular-mode EPR spectrum (Fig. 5, blue trace) shows signals from copper sites in the Mnx protein complex [24] and signals corresponding to mononuclear (class ii) and dinuclear (class iii) Mn(II) species bound to Mnx [20]. No Mn(III) species were detected by parallel-mode EPR spectroscopy, consistent with our previous observation that Mn(II) is not oxidized by the Mnx T1Cu in the absence of molecular oxygen [27].
X-band (9.38 GHz) perpendicular-mode EPR spectra of Mnx protein aerobically mixed with four equivalents of Mn(II) in the presence of ten equivalents of NaN3 for 2 min (blue trace) or in the presence of five equivalents of PP for 3 min (green trace, see Experimental Procedure for details). Experimental parameters: temperature = 15 K; microwave frequency = 9.38 GHz; microwave power = 0.2 mW; conversion time = 40 ms; modulation amplitude = 0.8 mT; modulation frequency = 100 kHz
Our previous EPR observation of a Mnx-produced Mn(III) species was achieved by oxidizing Mn(II) with Mnx and O2 in the presence of PP [29]. Mn(III) is released from the enzyme and is trapped by the PP, based on the appearance of a ligand-to-metal charge-transfer electronic absorption band at ~ 258 nm of Mn(III)PP (see Fig. 6) [7, 29]. Under the conditions used for kinetic studies, only Mn(III) complexes in solution can be detected by absorption spectroscopy since they are present in much higher concentration than the enzyme. However, if concentrated enzyme was used, as in the present study, the spectral contribution of the integer spin enzyme-bound Mn(III) could be distinguished from the aqueous Mn(III)PP signal via parallel-mode EPR spectroscopy.
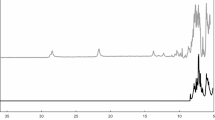
UV–Vis absorption spectra recorded at the indicated times during the oxidation of 100 μM MnSO4 by 50 nM Mnx, in the presence of 0.5 mM NaPP in 10 mM sodium phosphate buffer, pH 7.8. The band at 258 nm is due to the Mn(III) intermediate in solution trapped by PP; the ~ 370 nm band is due to nanoparticulate MnO2 enzymatic product. The inset shows the time profiles for the Mn(III)PP (black line), MnO2 (red line), and MnSO4 (green line) components of the Mnx assay, as described in Ref. [29]
We thus prepared a reaction sample by incubating the as-isolated Mnx protein with five equivalents of PP, and then adding four equivalents of MnSO4 to initiate the reaction. After being allowed to react for 3 min, this sample was frozen in liquid nitrogen and analyzed by CW EPR spectroscopy. The perpendicular-mode EPR spectrum (Fig. 5, green trace) is similar to the spectrum of the reaction mixture of Mnx, Mn2+(aq), and O2 in the presence of NaN3 (Fig. 5, blue trace), with signals from copper sites in Mnx and bound Mn(II) species (vide supra). However, the parallel-mode EPR spectrum (Fig. 7, green trace, as well as Fig. 8), possesses a new sextet appearing at a slightly lower geff = 8.13 (centered at the magnetic field of 82.0 mT) and with a smaller 55Mn hyperfine-coupling value of A = 4.2 mT (corresponding to 120 MHz), indicating that the generated Mn(III) species is distinct from aqueous Mn(III)PP. This new signal could arise from a Mn(III) species coordinated in Mnx protein, denoted as “Mn(III)–Mnx”, which is possibly stabilized by being partially ligated to PP. We reported this spectrum in our recent paper [29] as the first spectroscopic evidence for Mnx-bound Mn(III) intermediate during the Mn(II) oxidation. As mentioned above, previous kinetic studies implicated a hydroxide-bridged dinuclear Mn(III) intermediate, which is not expected to be EPR detectable [28]. Indeed, no Mn(III) EPR signal is seen in the absence of PP. However, the kinetic studies also revealed a pronounced slowing of the enzyme reaction in the presence of PP, which could enable the freezing of an EPR-detectable mononuclear Mnx-bound Mn(III) species at high enzyme concentration, before a second electron transfer leads to formation of the putative binuclear Mn(III) intermediate. It is also possible that PP can stabilize the mononuclear Mnx-bound Mn(III). Indeed, a ferroxidase ceruloplasmin, which is the closest structural homologue of MCO MnxG, oxidizes the substrate Fe(II) and then translocates its oxidation product Fe(III) over several angstroms towards a holding site near the surface [18]. It seems likely that in MnxG (see Fig. 9), after the first round of electron transfer, a Mn(III) species is translocated to near the protein surface, and, in the presence of PP, the resulting Mn(III) is stabilized against disproportionation by being partially ligated to PP.
X-band (9.38 GHz) parallel-mode EPR spectra of Mn(III)PP (red trace, which is also shown in Fig. 3) and the as-isolated Mnx protein aerobically mixed with four equivalents of Mn(II) in the presence of five equivalents of PP for 3 min (green trace, with the corresponding perpendicular-mode spectrum shown in Fig. 5). Experimental parameters: temperature = 8 K; microwave frequency = 9.385 GHz; microwave power = 10 mW; conversion time = 40 ms; modulation amplitude = 0.8 mT; modulation frequency = 100 kHz. The black traces are the simulated spectra using the parameters of A = 140 MHz, D = − 2.5 cm−1, E = 0.25 cm−1 for standard Mn(III)PP and using the parameters of A = 120 MHz, D = − 2.0 cm−1, E = 0.20 cm−1 for Mnx-bound Mn(III) species. The two experimental spectra are adapted from Ref. [29]
Temperature dependence of the parallel-mode EPR spectra of Mnx-bound Mn(III) species (a) and the corresponding simulated spectra using the parameters of A = 120 MHz, D = − 2.0 cm−1 and E = 0.20 cm−1 (b). Experimental parameters: microwave frequency = 9.385 GHz; microwave power = 10 mW (no saturation); conversion time = 40 ms; modulation amplitude = 0.8 mT; modulation frequency = 100 kHz. c Temperature dependence of the peak-to-peak amplitudes of the fourth EPR feature of Mn(III) sextet (peak around 85.0 mT). The reason we chose the fourth 55Mn hyperfine peak is due to its distinguishable signal/noise intensity. The peak-to-peak amplitudes are plotted as a function of the inversion of temperature, with the red triangles corresponding to the experimental data shown in a and the green stars corresponding the simulation data given in b. The black starred peak shown in a is a signal from background
Structural model of the catalytic sites in MCO MnxG, based on the generated I-TASSER MnxG homology model [23]. The MnxG structural model was aligned to the X-ray structure of human ceruloplasmin (PDB:1KCW) [18], using the conserved T1Cu ligands. This structural model shows a trinuclear T2/T3Cu centers (brown spheres) and a mononuclear T1Cu center (blue sphere) from ceruloplasmin structure, together with the MnxG conserved ligands around the copper centers (blue sticks). The red sphere is the substrate Fe(II) from ceruloplasmin structure, and the pink sphere is the expected location of the oxidation product Fe(III) at the holding site with the ceruloplasmin residues showing as yellow sticks. The residue E935 was proposed to translocate Fe(III) from the substrate site to the holding site [18]. In ceruloplasmin, the oxidation product Fe(III) is guided to the protein exterior by residues E932, E753, and D921, as predicted in Ref. [58]. The possible holding site in MnxG (residues shown as blue sticks) is located close to the solvent interface, permitting access of pyrophosphate
In what follows, we further probe this Mn(III) species using temperature dependence measurements and EPR spectral simulations. The temperature dependence of the Mnx-bound Mn(III) EPR signals shown in Fig. 8a again exhibits Curie law behavior, suggesting that the ZFS D is negative. A 55Mn hyperfine-coupling value of A = 120 MHz and ZFS parameters of D = − 2.0 cm−1 and E = 0.20 cm−1 were employed in the simulation (Figs. 7, 8). However, the temperature dependence of the simulated spectra of Mnx-bound Mn(III) does not fit the experimental data (Fig. 8c) very well, which could be due to the low signal intensity of this intermediate compared with that of Mn(III)PP. The negative ZFS D suggests that Mnx-bound Mn(III) either has a hexa-coordinated Oh geometry with a 5B1g ground state or a five-coordinated SP geometry with a 5B1 ground state, in which the lowest unoccupied metal-centered MO is 3 \(d_{{x^{2} - y^{2} }}\)-based. To be noted, although we cannot determine the exact geometry of Mnx-bound Mn(III) species currently, the same hyperfine-coupling value of A = 4.2 mT is also reported for Mn(III) salen complex with additive of 4-phenylpyridine-N-oxide (4-PPNO), which has an axially elongated octahedral geometry (see Table 1 for details) [49].
Origin of the variation in ZFS values D
For Mn(III) species with a 5B1g/5B1 ground state, i.e., aqueous Mn(III)PP and the Mnx-bound Mn(III) in this work, the variation in the negative D values could be related to the energy of the excited 3E spin triplet [56, 57]. The lower energy of 3E corresponding to small tetragonal distortions leads to larger magnitude of D. As the ZFS value D for aqueous Mn(III)PP and the Mnx-bound Mn(III) is determined to be − 2.5 and − 2.0 cm−1, respectively, Mnx-bound Mn(III) species has larger tetragonal distortion in comparison with Mn(III)PP. Therefore, it is mostly likely that in the presence of PP, the Mn(III) species, generated in Mnx via oxidation of the bound Mn(II), is stabilized by being partially ligated to PP, with a lower-symmetry ligand environment in comparison with Mn(III)PP.
Conclusions
In this work, a Mn(III) species bound to the Mnx protein complex is exposed during the oxidation of Mn2+(aq) by trapping the Mn(III) with PP. A parallel-mode EPR signal is observed at geff = 8.13 with a 55Mn hyperfine-coupling value of A = 4.2 mT (corresponding to 120 MHz). Temperature dependence studies and spectral simulations indicate a negative ZFS value of D = − 2.0 cm−1. These magnetic properties suggest that the Mnx-bound Mn(III) species could be either six-coordinate in the Mnx protein complex with a 5B1g ground state or square-pyramidal five-coordinate with a 5B1 ground state. In addition, aqueous Mn(III)PP is also analyzed by parallel-mode EPR spectroscopy. It exhibits a distinctly different sextet of hyperfine signals, with geff = 8.20 and a larger hyperfine-coupling value of A = 4.8 mT. Temperature dependence studies and spectral simulations indicate a negative ZFS value of D = − 2.5 cm−1. The variation in the negative D values of Mn(III) species with a 5B1g/5B1 ground state could be related to the energy of excited 3E spin triplet. The lower energy of 3E corresponding to small tetragonal distortions leads to larger magnitude of D. Therefore, it is reasonable that Mnx-bound Mn(III) species has larger tetragonal distortion in comparison with aqueous Mn(III)PP, resulting in small magnitude of D. Collectively, these results provide direct evidence that a Mn(III) species is formed in MCO-containing Mnx protein during the biological Mn(II) oxidation.
References
Post JE (1999) Manganese oxide minerals: crystal structures and economic and environmental significance. Proc Natl Acad Sci USA 96(7):3447–3454
Tebo BM, Bargar JR, Clement BG, Dick GJ, Murray KJ, Parker D, Verity R, Webb SM (2004) Biogenic manganese oxides: properties and mechanisms of formation. Annu Rev Earth Planet Sci 32(1):287–328
Spiro TG, Bargar JR, Sposito G, Tebo BM (2010) Bacteriogenic manganese oxides. Acc Chem Res 43(1):2–9
Waite TD, Szymczak R (1994) Photoredox transformation of iron and manganese in marine systems: review of recent field investigations. Lewis Publishers, London
Sunda WG, Huntsman SA (1988) Effect of sunlight on redox cycles of manganese in the southwestern Sargasso Sea. Deep Sea Res 35(8):1297–1317
Morgan JJ (2005) Kinetics of reaction between O2 and Mn(II) species in aqueous solutions. Geochim Cosmochim Acta 69(1):35–48
Butterfield CN, Soldatova AV, Lee SW, Spiro TG, Tebo BM (2013) Mn(II, III) oxidation and MnO2 mineralization by an expressed bacterial multicopper oxidase. Proc Natl Acad Sci USA 110(29):11731–11735
Waasbergen LG, Hildebrand M, Tebo BM (1996) Identification and characterization of a gene cluster involved in manganese oxidation by spores of the marine Bacillus sp. strain SG-1. J Bacteriol 178(12):3517–3530
Dick GJ, Torpey JW, Beveridge TJ, Tebo BM (2008) Direct identification of a bacterial manganese(II) oxidase, the multicopper oxidase MnxG, from spores of several different marine Bacillus species. Appl Environ Microbiol 74(5):1527–1534
Butterfield CN, Tao L, Chacon KN, Spiro TG, Blackburn NJ, Casey WH, Britt RD, Tebo BM (2015) Multicopper manganese oxidase accessory proteins bind Cu and Heme. Biochim Biophys Acta 1854(12):1853–1859
Solomon EI, Szilagyi RK, George SD, Basumallick L (2004) Electronic structures of metal sites in proteins and models: contributions to function in blue copper proteins. Chem Rev 104(2):419–458
Solomon EI, Sundaram UM, Machonkin T (1996) Multicopper oxidases and oxygenases. Chem Rev 96(7):2563–2605
Lee SK, George SD, Antholine WE, Hedman B, Hodgson KO, Solomon EI (2002) Nature of the intermediate formed in the reduction of O2 to H2O at the Trinuclear Copper Cluster active site in native Laccase. J Am Chem Soc 124(21):6180–6193
Solomon EI, Augustine AJ, Yoon J (2008) O2 reduction to H2O by the multicopper oxidases. Dalton Trans 30:3921–3932
Heppner DE, Kjaergaard CH, Solomon EI (2013) Molecular origin of rapid versus slow intramolecular electron transfer in the catalytic cycle of the multicopper oxidases. J Am Chem Soc 135(33):12212–12215
Quintanar L, Gebhard M, Wang TP, Kosman DJ, Solomon EI (2004) Ferrous binding to the multicopper oxidases Saccharomyces cerevisiae Fet3p and human ceruloplasmin: contributions to ferroxidase activity. J Am Chem Soc 126(21):6579–6589
Machonkin TE, Solomon EI (2000) The thermodynamics, kinetics, and molecular mechanism of intramolecular electron transfer in human ceruloplasmin. J Am Chem Soc 122(50):12547–12560
Lindley PF, Card G, Zaitseva I, Zaitsev V, Reinhammar B, Selin-Lindgren E, Yoshida K (1997) An X-ray structural study of human ceruloplasmin in relation to ferroxidase activity. J Biol Inorg Chem 2(4):454–463
Singh SK, Roberts SA, McDevitt SF, Weichsel A, Wildner GF, Grass GB, Rensing C, Montfort WR (2011) Crystal structures of multicopper oxidase CueO bound to Copper(I) and Silver(I): functional role of a methionine-rich sequence. J Biol Chem 286(43):37849–37857
Tao L, Stich TA, Butterfield CN, Romano C, Tebo BM, Casey WH, Britt RD (2015) Mn(II) binding and subsequent oxidation by the multicopper oxidase Mnx investigated by electron paramagnetic resonance spectroscopy. J Am Chem Soc 137(33):10563–10575
Su J, Deng L, Huang L, Guo S, Liu F, He J (2014) Catalytic oxidation of manganese(II) by multicopper oxidase CueO and characterization of the biogenic Mn oxide. Water Res 56:304–313
Su J, Bao P, Bai T, Deng L, Wu H, Liu F, He J (2013) CotA, a multicopper oxidase from Bacillus pumilus WH4, exhibits manganese-oxidase activity. PLoS One 8:e60573
Romano CA, Zhou M, Song Y, Wysocki VH, Dohnalkova AC, Kovarik L, Paša-Tolić L, Tebo BM (2017) Biogenic manganese oxide nanoparticle formation by a multimeric multicopper oxidase Mnx. Nat Commun 8(1):746
Tao L, Stich TA, Liou S-H, Soldatova AV, Delgadillo DA, Romano CA, Spiro TG, Goodin DB, Tebo BM, Casey WH, Britt RD (2017) Copper binding sites in the manganese-oxidizing Mnx protein complex investigated by electron paramagnetic resonance spectroscopy. J Am Chem Soc 139(26):8868–8877
Tao L, Simonov AN, Romano CA, Butterfield CN, Fekete M, Tebo BM, Bond AM, Spiccia L, Martin LL, Casey WH (2017) Biogenic manganese-oxide mineralisation is enhanced by an oxidative priming mechanism for the multi-copper oxidase, MnxEFG. Chem Eur J 23(6):1346–1352
Tao L, Simonov AN, Romano CA, Butterfield CN, Tebo BM, Bond AM, Spiccia L, Martin LL, Casey WH (2018) Probing electron transfer in the manganese-oxide-forming MnxEFG protein complex using Fourier transformed AC voltammetry: understanding the oxidative priming effect. ChemElectroChem 5(6):872–876
Soldatova AV, Tao L, Romano CA, Stich TA, Casey WH, Britt RD, Tebo BM, Spiro TG (2017) Mn(II) oxidation by the multicopper oxidase complex Mnx: a binuclear activation mechanism. J Am Chem Soc 139(33):11369–11380
Mathur P, Crowder M, Dismukes GC (1987) Dimanganese(II) complexes of a septadentate ligand. Functional analogs of the manganese pseudocatalase. J Am Chem Soc 109(17):5227–5233
Soldatova AV, Romano CA, Tao L, Stich TA, Casey WH, Britt RD, Tebo BM, Spiro TG (2017) Mn(II) oxidation by the multicopper oxidase complex Mnx: a coordinated two-stage Mn(II)/(III) and Mn(III)/(IV) mechanism. J Am Chem Soc 139(33):11381–11391
Durão P, Chen Z, Fernandes AT, Hildebrandt P, Murgida DH, Todorovic S, Pereira MM, Melo EP, Martins LO (2008) Copper incorporation into recombinant CotA laccase from Bacillus subtilis: characterization of fully copper loaded enzymes. J Biol Inorg Chem 13(2):183–193
Webb SM, Dick GJ, Bargar JR, Tebo BM (2005) Evidence for the presence of Mn(III) intermediates in the bacterial oxidation of Mn(II). Proc Natl Acad Sci USA 102(15):5558–5563
Archibald FS, Fridovich I (1982) The scavenging of superoxide radical by manganous complexes: in vitro. Arch Biochem Biophys 214(2):452–463
Stoll S, Britt RD (2009) General and efficient simulation of pulse EPR spectra. Phys Chem Chem Phys 11(31):6614–6625
Stoll S, Schweiger A (2006) EasySpin, a comprehensive software package for spectral simulation and analysis in EPR. J Magn Reson 178(1):42–55
Campbell KA, Yikilmaz E, Grant CV, Gregory W, Miller A, Britt RD (1999) Parallel polarization EPR characterization of the Mn(III) center of oxidized manganese superoxide dismutase. J Am Chem Soc 121(19):4714–4715
Campbell KA (1999) University of California Davis, Ph.D. dissertation
Hendrich MP, Debrunner PG (1989) Integer-spin electron paramagnetic resonance of iron proteins. Biophys J 56(3):489–506
Münck E, Ksurerus K, Hendrich MP (1993) Combining Mössbauer spectroscopy with integer spin electron paramagnetic resonance. Methods Enzymol Acad Press 227:463–479
Sharma A, Gaidamakova EK, Grichenko O, Matrosova VY, Hoeke V, Klimenkova P, Conze IH, Volpe RP, Tkavc R, Gostinčar C, Gunde-Cimerman N, DiRuggiero J, Shuryak I, Ozarowski A, Hoffman BM, Daly MJ (2017) Across the tree of life, radiation resistance is governed by antioxidant Mn2+, gauged by paramagnetic resonance. Proc Natl Acad Sci 114(44):E9253–E9260
Krzystek J, Telser J, Pardi LA, Goldberg DP, Hoffman BM, Brunel L-C (1999) High-frequency and -field electron paramagnetic resonance of high-spin manganese(III) in porphyrinic complexes. Inorg Chem 38(26):6121–6129
Goldberg DP, Telser J, Krzystek J, Montalban AG, Brunel L-C, Barrett AGM, Hoffman BM (1997) EPR spectra from “EPR-Silent” species: high-field EPR spectroscopy of manganese(III) porphyrins. J Am Chem Soc 119(37):8722–8723
Anne-Laure B, Dante G, Roberta S, Luca AG, Andrea C, Fabretti AC, Uytterhoeven MG (1997) Electronic structure of Manganese(III) compounds from high-frequency EPR spectra. Angew Chem Int Ed Engl 36(21):2329–2331
Telser J, Pardi LA, Krzystek J, Brunel L-C (1998) EPR spectra from “EPR-Silent” species: high-field EPR spectroscopy of aqueous chromium(II). Inorg Chem 37(22):5769–5775
Campbell KA, Force DA, Nixon PJ, Dole F, Diner BA, Britt RD (2000) Dual-mode EPR detects the initial intermediate in photoassembly of the photosystem II Mn cluster: the influence of amino acid residue 170 of the D1 polypeptide on Mn coordination. J Am Chem Soc 122(15):3754–3761
Whittaker JW, Whittaker MM (1991) Active site spectral studies on manganese superoxide dismuase. J Am Chem Soc 113(17):5528–5540
Griffith JS (1971) The theory of transition-metal ions. Cambridge University Press, London
Sheng Y, Stich TA, Barnese K, Gralla EB, Cascio D, Britt RD, Cabelli DE, Valentine JS (2011) Comparison of two yeast MnSODs: mitochondrial Saccharomyces cerevisiae versus cytosolic Candida albicans. J Am Chem Soc 133(51):20878–20889
Mossin S, Weihe H, Barra A-L (2002) Is the axial zero-field splitting parameter of tetragonally elongated high-spin Manganese(III) complexes always negative? J Am Chem Soc 124(30):8764–8765
Campbell KA, Lashley MR, Wyatt JK, Nantz MH, Britt RD (2001) Dual-mode EPR study of Mn(III) salen and the Mn(III) salen-catalyzed epoxidation of cis-β-methylstyrene. J Am Chem Soc 123(24):5710–5719
Zhu W, Wilcoxen J, Britt RD, Richards NGJ (2016) Formation of hexacoordinate Mn(III) in Bacillus subtilis oxalate decarboxylase requires catalytic turnover. Biochemistry 55(3):429–434
Krivokapic I, Noble C, Klitgaard S, Tregenna-Piggott P, Weihe H, Barra AL (2005) Anisotropic hyperfine interaction in the manganese(III) hexaaqua ion. Angew Chem Int Ed 44(23):3613–3616
Tregenna-Piggott PLW, Weihe H, Barra A-L (2003) High-field, multifrequency EPR study of the [Mn(OH2)6]3+ cation: influence of π-bonding on the ground state zero-field-splitting parameters. Inorg Chem 42(25):8504–8508
Klewicki JK, Morgan JJ (1998) Kinetic behavior of Mn(III) complexes of pyrophosphate, EDTA, and citrate. Environ Sci Technol 32(19):2916–2922
Gromov I, Marchesini A, Farver O, Pecht I, Goldfar D (1999) Azide binding to the trinuclear copper center in laccase and ascorbate oxidase. Eur J Biochem 266(3):820–830
Soldatova AV, Butterfield C, Oyerinde OF, Tebo BM, Spiro TG (2012) Multicopper oxidase involvement in both Mn(II) and Mn(III) oxidation during bacterial formation of MnO2. J Biol Inorg Chem 17(8):1151–1158
Dugad LB, Behere DV, Marathe VR, Mitra S (1984) Magnetic properties and electronic structure of manganese(III) porphyrins. Chem Phys Lett 104(4):353–356
Kennedy BJ, Murray KS (1985) Magnetic properties and zero-field splitting in high-spin manganese(III) complexes. 2. Axially ligated manganese(III) porphyrin complexes. Inorg Chem 24(10):1557–1560
White KN, Conesa C, Sánchez L, Amini M, Farnaud S, Lorvoralak C, Evans RW (2012) The transfer of iron between ceruloplasmin and transferrins. Biochim Biophys Acta Gen Subjects 1820(3):411–416
Acknowledgements
The work was supported by the National Science Foundation Award Numbers CHE-1213699, CHE-1665455 to RDB, EAR-1231322 to WHC, CHE-1410688 to BMT, and CHE-1410353 to TGS. The EPR spectrometers at the CalEPR facility used in this study were funded by the National Institutes of Health (S10-RR021075) and the NSF (CHE-1048671).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Tao, L., Stich, T.A., Soldatova, A.V. et al. Mn(III) species formed by the multi-copper oxidase MnxG investigated by electron paramagnetic resonance spectroscopy. J Biol Inorg Chem 23, 1093–1104 (2018). https://doi.org/10.1007/s00775-018-1587-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00775-018-1587-z