Abstract
Diclofenac is a non-steroidal anti-inflammatory drug (NSAID). Here, we use double electron–electron resonance (DEER, also known as PELDOR) to study the interaction of spin-labeled diclofenac (diclofenac-SL) with three types of model membranes consisting of palmitoyl-2-oleoyl-sn-glycero-3-phosphocholine (POPC), an equimolar mixture of 1,2-dipalmitoyl-sn-glycero-3-phosphocholine (DPPC) and 1,2-dioleoyl-sn-glycero-3-phosphocholine (DOPC), and this mixture with the addition of 20 mol% cholesterol. The results suggest that lipid-mediated lateral clustering of diclofenac-SL molecules occurs in all cases. For the POPC bilayer, alternative clustering takes place in two opposite leaflets, with random distribution of the molecules within the clusters. For DOPC/DPPC and DOPC/DPPC/cholesterol bilayers, diclofenac-SL molecules are separated by a distance of at least 1.4 nm. DOPC/DPPC/cholesterol bilayers are known to form nanoscale liquid disordered and liquid ordered lateral structures, the latter called lipid rafts. For this case, diclofenac-SL molecules were found to be captured by lipid rafts, forming a quasi-regular two-dimensional substructure in them with a “superlattice” parameter of ~ 3.0 nm.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Nonsteroidal anti-inflammatory drugs (NSAIDs) are effective antipyretic and analgesic pharmacologic agents [1]. Diclofenac is an NSAID of the phenylacetic acid class which inhibits cyclooxygenase (COX)-2 enzyme [1]. NSAIDs also treat other diseases including cancer [2], arthritis [3, 4], and neurodegenerative diseases [5].
However, side effects of NSAIDs are also known [6]. Many of these effects are related to the membrane activity of these drugs [7,8,9,10]. For diclofenac, this activity may contribute to its gastrointestinal toxicity [6, 11, 12]. Diclofenac can interact with erythrocyte membranes inducing a disordering of the acyl chains of the lipids and changing the erythrocyte morphology [13]. To understand the therapeutic effects of NSAIDs and to develop ways to limit their side effects, understanding the mechanisms at the molecular level of its interaction with the plasma membrane can be extremely useful [7, 14,15,16,17,18,19,20].
Previously, in our laboratory, spin-labeled diclofenac (diclofenac-SL) was synthesized [21]. This new substance allows using electron paramagnetic resonance (EPR) spectroscopy for studying interaction of diclofenac with biological membranes. Here, we study diclofenac-SL in model membranes of three different types: palmitoyl-2-oleoyl-sn-glycero-3-phosphocholine (POPC), equimolar mixture of 1,2-dioleoyl-sn-glycero-3-phosphocholine (DOPC), and 1,2-dipalmitoyl-sn-glycero-3-phosphocholine (DPPC), and this mixture with added 20 mol% of cholesterol. These types of model membranes are often used to mimic real biological membranes. POPC includes in its structure a fully saturated acyl chain and a single-unsaturated acyl chain, DOPC includes two single-unsaturated acyl chains, and DPPC consists of two fully saturated acyl chains. DOPC/DPPC/cholesterol bilayers are known to form nanoscale liquid disordered and liquid ordered lateral structures, the latter called lipid rafts [22,23,24,25,26].
Biological membranes and lipid rafts are nanoscale objects. Therefore, it is reasonable to use experimental methods that allow studying the nanoscale structure. Such a method is double electron–electron resonance (DEER, also known as PELDOR) [27,28,29,30,31].
DEER spectroscopy is based on the electron spin echo (ESE) phenomenon. The DEER signal appears due to modulation by microwave (mw) pulses of magnetic dipole–dipole (d–d) interaction between spins. For some selected pair of spins A and B, the DEER signal from spins A is [27,28,29,30,31]
where
Here, μBohr is the Bohr magneton, gA and gB are the effective g-factors of two interacting spins, and θAB is the angle between the vector rAB connecting two spins and the magnetic field of the spectrometer. The dimensionless parameter pB < 1 is the fraction of electron spins excited by the pumping mw pulse (excitation efficiency).
An important property of ESE spectroscopy in organic and biological solids is that the observable frequency ωAB is the order of 106 rad/s. Therefore, DEER experiment probes the distances rAB of the order of several nanometers.
To take into account all spins in a sample, Eq. (1) is to be multiplied for all pairs of spins.
For mono-spin-labeled molecules randomly distributed in a three-dimensional (3-D) space, theoretical expression for the DEER signal states that [27,28,29,30,31]
where Clocal is the local spin concentration taken in the cm−3 units. It is assumed that gA ≈ gA ≈ g.
For the two-dimensional (2-D) model of the spatial distribution which could be more appropriate for biological membranes, the analogous expression is [32]
where σlocal is the local surface concentration taken in the cm−2 units.
2 Experimental
2.1 Materials and Samples
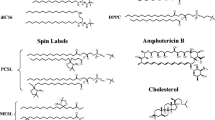
Diclofenac-SL was synthesized as described previously [21]. Cholesterol and lipids POPC, DOPC, and DPPC were from Avanti Polar Lipids (Birmingham, AL, USA). The chemical structures of diclofenac and diclofenac-SL are shown in Fig. 1.
Bilayer were prepared either from POPC lipids (POPC bilayer) or from an equimolar mixture of DOPC and DPPC (DOPC/DPPC bilayer) or from this mixture with the addition of 20 mol% of cholesterol (DOPC/DPPC/cholesterol bilayer). For the latter composition, it is known that at room temperature, the phases Lo + Ld + Lβ and Lo + Ld coexist [33]. Freshly prepared diclofenac-SL was added to these lipids in a desired amount between 0.15 and 2.5 mol%. The substances were first dissolved separately in chloroform, and then the solutions were mixed in required proportions. The solvent was removed in a stream of nitrogen, and the residue was stored then in vacuo for 4 h. Then, an excess of phosphate-buffered saline (pH 7.0) was added in a proportion of 10:1. The sample was stirred and then stored for 2 h at room temperature. Then, the sample was centrifuged at 14,000 g for 4 min, and the final lipid/water ratio was 1:3, w/w. It was shown earlier [21] that diclofenac-SL is completely dissolved in the bilayers after these operations. EPR glass tube of 3 mm o.d. was employed. For low-temperature measurements, the samples were quickly frozen by immersion into liquid nitrogen.
2.2 Measurements
An X-band Bruker ELEXSYS E580 EPR spectrometer was employed. Conventional continuous-wave (CW) EPR spectra were obtained with a Bruker ER 4118X-MD5 dielectric resonator, the modulation amplitude was 0.01 mT, the output microwave power was 200 mW, the microwave attenuation was 25 dB, and the sweep time and the spectrometer time constant were 10.49 s and 20.48 ms, respectively.
In pulsed EPR measurements, the spectrometer was equipped with a split-ring Bruker ER 4118 X-MS-3 resonator. Two-pulse ESE signal was generated with a pulse sequence π/2 – τ − π − τ – echo. The lengths of the first and second pulses applied were 16 and 32 ns, respectively. The DEER measurements utilized a three-pulse measurement scheme without dead time with signal correction by “blank” excitation [34, 35]. The DEER pulse sequence was (π/2)νA − t − πνB − (t − τ) − πνA − τ − echoνA, where the subscripts denote mw frequencies. The lengths of the pulses at the frequency νA were 16 and 32 ns, respectively, the pumping pulse at the frequency νB had duration of 36 ns, and its amplitude was selected to ensure a turning angle π. The time delay τ was set optionally between 200 and 800 ns. The delay t was scanned with a step of 4 ns, starting from a negative value t = − 180 ns before the first detection pulse. Frequency νB corresponded to the excitation of the maximum of the echo-detected EPR spectrum, and frequency νA corresponded to its high-field shoulder (so, the difference νB – νA was 85 MHz).
For low-temperature measurements, the resonator was placed in an Oxford Instruments CF-935 cryostat and cooled with a stream of cold nitrogen. The temperature was stabilized by a Bruker ER4131VT temperature controller.
3 Results
3.1 CW EPR Spectra
The CW EPR spectra of diclofenac-SL taken at room temperature for the three types of bilayer studied are shown in Fig. 2, for different molar diclofenac-SL concentrations χ. One can see that all spectra look very similar. Their lineshape indicates retarded molecular motion typical of spin-labeled molecules entrapped into the membranes [36] and only slightly depend on χ.
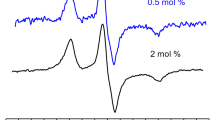
Figure 3 shows CW EPR spectra taken at 200 K for all three types of the bilayers, obtained also for different concentrations χ. It is seen that here the linewidth increases with increasing χ; to visualize this increase, all spectra are normalized to the same value of the second integral over the magnetic field.
At 200 K, molecular motion is frozen, so the increase of the linewidth in EPR spectra occurs only because d–d interaction. It can be seen from data in Fig. 3 that increase of diclofenac-SL concentration χ results in remarkable broadening of EPR lines. For POPC bilayer, broadening is much less than for DOPC/DPPC and DOPC/DPPC/cholesterol bilayers. However, because of presence of different sources contributing to the lineshape, analysis of d–d interaction is not easy in CW EPR.
3.2 DEER Data
DEER spectroscopy is specially designed to extract the pure contribution of the d–d interaction. Previously [37], there was shown that in lipid bilayers, DEER time traces at high concentrations of spin-labeled molecules may depend on the time separation τ between the echo-forming pulses. Therefore, we performed measurements with different τ here as well. Representative data from these measurements are shown in Fig. 4.
Semilogarifmic plot of three-pulse raw representative DEER data for diclofenac-SL in POPC (A), DOPC/DPPC (B), and DOPC/DPPC/cholesterol (C) bilayers taken for various molar concentration χ and for different delays τ. Temperature is 80K. The data are shifted vertically for the convenience of presentation
It can be seen that at high value of concentration χ, the τ-dependence does occur (especially for DOPC/DPPC and DOPC/DPPC/cholesterol bilayers): the DEER signal decays noticeably faster with increasing τ. The reason for such a dependence is interference of additional dipolar pathway arising for the spins under detection (spins A) [38]: for a high concentration of spins, there is an “instantaneous diffusion” effect in the system of spins A. This effect leads to a faster echo decay for closely spaced spins, so that at large τ spins with large spacing contribute mostly to the echo signal. Then, the DEER decay at large τ should indeed be slower than at small τ—this is exactly what can be seen in Fig. 4B, C.
Such dependence on τ is, of course, undesirable, since it leads to the loss of the observed spins. Therefore, the value of τ should be small enough, so that this dependence can be ignored. From the data in Fig. 4, it follows that this occurs at τ about 300 ns for POPC bilayer and at τ near 200 ns for DOPC/DPPC and DOPC/DPPC/cholesterol bilayer.
Another point that can be observed in Fig. 4 is the non-linearity of the data at t > 0. This means that Eq. (3) cannot be applied, which is not surprising, since a 3-D spatial distribution is hardly expected for bilayers. Figure 5 shows the DEER data plotted in coordinates suitable for comparison with Eq. (4) written for the 2-D spatial distribution.
Semilogarithmic plot of three-pulse DEER time traces for diclofenac-SL in POPC (A), DOPC/DPPC (B), and DOPC/DPPC/cholesterol (C) bilayers taken for various molar concentrations χ of ibuprofen-SL [indicated in (A)]. The data are plotted vs. t2/3 for comparison with Eq. (4). Delays τ are 300 ns in (A) and 200 ns in (B) and (C). Dashed lines in (A) are drawn to emphasize the agreement with Eq. (4). Open circles in (B) and (C) present the results of simulation within a model of inaccessibility (see text). Temperature is 80 K. Data are shifted vertically for convenience
The data of Fig. 5A (POPC bilayer) show fairly good linear dependencies for the experimental DEER decays, which implies agreement with theoretical Eq. (4) for 2-D spatial distribution.
For DOPC/DPPC (Fig. 5B) and DOPC/DPPC/cholesterol (Fig. 5C) bilayers, the linearity of the data does not takes place: a noticeable retardation of the decays is seen at the beginning, when t < 0.13/2. The non-linearity of the similar type was observed previously in DOPC/DPPC/cholesterol bilayer for spin-labeled ibuprofen [37] and cholestane [39], and was explained by non-random nanoscale spatial distribution of the molecules in their close contact.
As mentioned above, DEER cannot be applied at large times because of the effect of τ-dependence appearing due to the “instantaneous diffusion” effect for spins under observation (spins A). From the other hand, this effect has the same physical ground as the DEER effect—both are induced by the modulation of d–d interactions by mw pulses. In principle, simple two-pulse echo can also be used to extract information on d–d interactions, like the DEER does. However, this simple approach faces several obstacles—the dead time restriction, intervening electron–nuclear interactions (the so-called ESEEM effect), and fast signal decay due to spin relaxation. The latter two drawbacks can be suppressed by dividing the two time traces obtained at different field positions [40]. Figure 6 shows the result of such division for 2.5 mol% for diclofenac-SL in DOPC/DPPC and DOPC/DPPC/cholesterol bilayers. One can see that temporal behavior in these two cases is substantially different.
4 Discussion
For the POPC bilayer, the experimental DEER data in Fig. 5A can be explained in a rather obvious way, since they obey the theoretical equation [Eq. (4)] for the 2-D spatial distribution. The value of pB = 0.25 ± 0.01 in this equation was found for diclofenac-SL in glassy solution in the same way as it was done in [37] for spin-labeled ibuprofen. The local diclofenac-SL surface concentrations σlocal obtained from using Eq. (4) and the data in Fig. 5A are plotted in Fig. 7.
Local diclofenac-SL surface concentration in the POPC bilayer as a function of its molar concentration χ. The two dashed lines present a dependence σrandom = 0.01χ/AL expected for random spatial distribution and its doubled version, 2σrandom. Insert shows the model of alternative clustering of diclofenac-SL in two opposite leaflets (see text)
Figure 7 presents also the dependence σrandom = 0.01χ/AL where AL is the area per lipid which may be taken for POPC bilayer as 0.60 nm2 [41]. This dependence is expected for random spatial distribution of diclofenac-SL molecules. It can be seen that σlocal exceeds σrandom everywhere up to χ = 2 mol%. This exceeding unambiguously implies clustering of the molecules. From the other side, at small χ < 1 mol%, σlocal is close to 2⋅σrandom. This proximity was previously observed for different types of spin-labeled molecules in the POPC bilayer [37, 42] and was attributed to alternative clustering in two opposite leaflets of the bilayer (a chess-box model). At χ ≥ 1 mol%, this alternation is obviously broken.
The physical nature of these clustering features is currently unclear. We can only state that alternative clustering implies that diclofenac-SL molecules in one leaflet prevent their appearance in the opposite leaflet, and that this restriction is removed at high concentrations.
The situation with DOPC/DPPC and DOPC/DPPC/cholesterol bilayers is more complicated. First, the τ-dependence here is much stronger—see Fig. 4—especially in the presence of cholesterol. Second, DEER decays when plotted against t2/3 show retardation at the beginning of the decay—see Fig. 5B and C. These two features have been observed previously for spin-labeled ibuprofen in DOPC/DPPC/cholesterol bilayers [37]. The τ-dependence was explained by the influence of d–d interactions on the echo decay for the observed A spins at their high concentration [38]; this effect can become important in the case of non-random spatial distribution of spin labels.
And the retardation at the beginning appears also for non-random (regular or quasi-regular) spatial distribution. We suggest that the explanation given in [37] for spin-labeled ibuprofen can be applied here for diclofenac-SL as well. The “inaccessibility model” suggested in [37] assumes that spin labels are randomly distributed in a 2-D space but cannot approach each other closer than some distance Rmin. Besides Rmin, this model has for fitting only one more free parameter, σlocal. The results of this fitting in Fig. 5B and C are given as open circles. Rmin in all these simulations was taken as 1.4 nm. It can be seen a rather good agreement between experiment and simulations. The best-fitted σlocal values are given in Fig. 8.
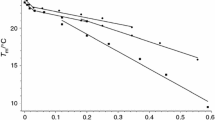
Local diclofenac-SL surface concentration in the DOPC/DPPC and DOPC/DPPC/cholesterol bilayers as a function of molar concentration χ. The solid lines are drawn to guide the eye. Two dashed lines present functions σlocal = 0.01χ/AL expected for random spatial distribution in these two types of bilayer
The dashed straight lines drawn in Fig. 8 present the situation of random 2-D distribution of spin labels in the membrane (a description of how AL were adopted from the data in the literature is given in [37]). The fact that the σlocal values found lie significantly higher clearly indicates the clustering of spin-labeled molecules. Note that the clustering features here is different from the case of the POPC bilayer (cf. Figure 7): σlocal values here are higher and clustering continues up to high molecular concentrations. Another difference is that it is necessary to use an inaccessible distribution model.
Data in Fig. 5B and C in the absence and presence of cholesterol, respectively, look very similar. This similarity, however, may not imply the similar character of the spatial distribution, because the time interval for DEER where the signal can be studied is too small. From the similarity of the data for DOPC/DPPC and DOPC/DPPC/cholesterol bilayers, we may conclude only that the short-range structure of surrounding is nearly the same in these two bilayers. This surrounding is most likely formed by saturated DPPC or unsaturated DOPC molecules enveloping diclofenac-SL.
The two-pulse ESE data may be considered as an “extension” of DEER data for larger time delays (instead of the time delay t in DEER, in the case of two-pulse ESE the time delay corresponds to τ) [40]. These ESE data are presented in Fig. 6; as longer time delays imply probing longer distances, these data show that the long-range order in the DOPC/DPPC and DOPC/DPPC/cholesterol bilayers is quite different. We note that for the latter case, the two-pulse ESE time dependence looks very similar to that observed in [43] for galvinoxyl free radicals in fullerene-containing polymeric matrix where this temporal behavior was attributed to formation of octahedral cluster of the radicals. Probably, such kind of clustering occurs here as well.
However, a detailed analysis of the time dependence of the two-pulse ESE, as well as the stitching of these data with the DEER results, is a task of our future studies. We can only state now that the diclofenac-SL molecules form mutually ordered (nano)clusters in the DOPC/DPPC/cholesterol bilayer. By other words, these ordered clusters form some kind of “superlattice”. Note that the average distance between diclofenac-SL molecules assessed from Fig. 8 as (σlocal)−1/2 at large χ is about 3 nm; this value can be attributed to the parameter of this superlattice.
The DOPC/DPPC/cholesterol bilayer of the studied molar composition (0.4:0.4:0.2) is known to form lipid rafts at physiological temperatures [33]; such membrane heterogeneity is also known to persist upon cooling [44]. Then, the data in Fig. 6 can be interpreted as evidence that diclofenac-SL molecules are captured by lipid rafts. The same conclusion was done in [37] for spin-labeled ibuprofen in the DOPC/DPPC/cholesterol bilayer of the same composition.
5 Conclusions
DEER study of mono-spin-labeled molecules allows getting information on the mutual spatial arrangement of spin-labeled molecules. The results obtained here for mono-spin-labeled diclofenac-SL indicate clustering of its molecules in the studied lipid bilayers. In the POPC bilayer, clustering occurs as alternative assembling in two opposite membrane leaflets, with random 2-D spatial distribution within the clusters. In the DOPC/DPPC and DOPC/DPPC/cholesterol bilayers, the short-range structure is likely formed by DPPC or DOPC molecules enveloping diclofenac-SL molecules; this envelopment prevents the molecules from contacting closer than 1.4 nm.
In the DOPC/DPPC/cholesterol bilayer, the data can be explained by suggesting that diclofenac-SL is taken up by lipid rafts, which are assumed to exist in this system, even at low temperatures. This explanation is based on data from the two-pulse ESE experiment: the time dependence of the echo signal revealed features known for the long-range order of spin-labeled molecules. The superlattice parameter obtained from the found surface density of diclofenac-SL is about 3.0 nm. The presence of such a superlattice indicates that lipid rafts have an internal substructure. The stitching of DEER and two-pulse ESE data is the subject of further studies that may possibly allow to give more information about this internal substructure of lipid rafts.
Data availability
All data necessary to support the findings of this study are included in this article.
References
R. Altman, B. Bosch, K. Brune, P. Patrignani, C. Yang, Drugs 75, 859–877 (2015)
C.M. Ulrich, J. Bigler, J.D. Potter, Nat. Rev. Cancer 6, 130–140 (2006)
L.J. Crofford, Arthritis Res. Ther. 15, S2 (2013)
M.A. González-Gay, C. González-Juanatey, Nat. Rev. Cardiol. 14, 69–70 (2017)
A. Lleo, E. Galea, M. Sastre, Cell. Mol. Life Sci. 64, 1403–1418 (2007)
T.D. Warner, F. Giuliano, I. Vojnovic, A. Bukasa, J.A. Mitchell, J.R. Vane, Proc. Natl. Acad. Sci. 96, 7563–7568 (1999)
M. Manrique-Moreno, L. Heinbockel, M. Suwalsky, P. Garidel, K. Brandenburg, Biochim. Biophys. Acta 1858, 2123–2131 (2016)
C.P. Leite, C. Nunes, S. Reis, Prog. Lipid Res. 52, 571–584 (2013)
Y. Zhou, S.J. Plowman, L.M. Lichtenberger, J.F. Hancock, J. Biol. Chem. 285, 35188–35195 (2010)
A.M. Seddon, D. Casey, R.V. Law, A. Gee, R.H. Templer, O. Ces, Chem. Soc. Rev. 38, 2509–2519 (2009)
E. Fernandes, T. B. Soares, H. Gonçalves, S. Bernstorff, M. E. C. D. Real Oliveira, C. M. Lopes, M. A. Lúcio, Int. J. Mol. Sci. 19, 3411 (2018)
L.M. Lichtenberger, Y. Zhou, E.J. Dial, R.M. Raphael, J. Pharm. Pharmacol. 58, 1421–1428 (2006)
M. Suwalsky, M. Manrique, F. Villena, C.P. Sotomayor, Biophys. Chem. 141, 34–40 (2009)
V.K. Sharma, E. Mamontov, M. Tyagi, Biochim. Biophys. Acta Biomembr. 1862, 183100 (2020)
S.L. Yefimova, T.N. Tkacheva, N.A. Kasian, J. Appl. Spectrosc. 84, 284–290 (2017)
S. Ramadurai, N.K. Sarangi, S. Maher, N. Macconnell, A.M. Bond, D. McDaid, D. Flynn, T.E. Keyes, Langmuir 35, 8095–8109 (2019)
M. Wood, M. Morales, E. Miller, S. Braziel, J. Giancaspro, P. Scollan, J. Rosario, A. Gayapa, M. Krmic, S. Lee, Langmuir 37, 4468–4480 (2021)
S. Sun, A.M. Sendecki, S. Pullanchery, D. Huang, T. Yang, P.S. Cremer, Langmuir 34, 10782–10792 (2018)
J. Kremkow, M. Luck, D. Huster, P. Müller, H.A. Scheidt, Biomolecules 10, 1384 (2020)
E. Aloi, B. Rizzuti, R. Guzzi, R. Bartucci, Arch. Biochem. Biophys. 654, 77–84 (2018)
D.S. Baranov, A.S. Kashnik, A.N. Atnyukova, S.A. Dzuba, Molecules 28, 5991 (2023)
D. Lingwood, J. Ries, P. Schwille, K. Simons, Proc. Natl. Acad. Sci. U.S.A. 105, 10005–100010 (2008)
L.J. Pike, J. Lipid Res. 47, 1597–1598 (2006)
D. Marsh, Biochim. Biophys. Acta – Biomembr. 1788, 2114–2123 (2009)
M.R. Elkins, A. Bandara, G.A. Pantelopulos, J.E. Straub, M. Hong, J. Phys. Chem. B 125, 1825–1837 (2021)
I. Levental, Nat. Rev. Mol. Cell Biol. 21, 420 (2020)
A. Milov, K. Salikhov, M. Shchirov, Soviet Phys. Solid State 23, 565–569 (1981)
A.D. Milov, A.G. Maryasov, Y.D. Tsvetkov, Appl. Magn. Reson. 15, 107–143 (1998)
O. Schiemann, T.F. Prisner, Q. Rev, Biophys. 40, 1–53 (2007)
G. Jeschke, Ann. Rev. Phys. Chem. 63, 419–446 (2012)
O. Schiemann, C.A. Heubach, D. Abdullin, K. Ackermann, M. Azarkh, E.G. Bagryanskaya, M. Drescher, B. Endeward, J.H. Freed, L. Galazzo et al., J. Am. Chem. Soc. 143, 17875–17890 (2021)
A.S. Kashnik, D.S. Baranov, S.A. Dzuba, Membranes 12, 1077 (2022)
J.H. Davis, J.J. Clair, J. Juhasz, Biophys. J. 96, 521–539 (2009)
A.D. Milov, Y.A. Grishin, S.A. Dzuba, Y.D. Tsvetkov, Appl. Magn. Reson. 41, 59–67 (2011)
S.A. Dzuba, J. Magn. Reson. Open 14–15, 100100 (2023)
D. Marsh, Spin-label electron paramagnetic resonance spectroscopy (CRC Press, Boca Raton FL USA, 2020)
A.S. Kashnik, D.S. Baranov, S.A. Dzuba, J. Phys. Chem. B 128, 3652–3661 (2024)
K.M. Salikhov, I.T. Khairuzhdinov, R.B. Zaripov, Appl. Magn. Reson. 45, 573–619 (2014)
V.V. Unguryan, E.A. Golysheva, S.A. Dzuba, J. Phys. Chem. B 125, 9557–9563 (2021)
E.A. Golysheva, A.S. Smorygina, S.A. Dzuba, Appl. Magn. Reson. 53, 685–698 (2022)
N. Kučerka, M.P. Nieh, J. Katsaras, Biochim. Biophys. Acta 1808, 2761–2771 (2011)
A.S. Kashnik, V.N. Syryamina, B. Biondi, C. Peggion, F. Formaggio, S.A. Dzuba, Appl. Magn. Reson. 54, 401–414 (2023)
M.N. Uvarov, L.V. Kulik, S.A. Dzuba, Phys. Chem. Chem. Phys. 25, 26219–26224 (2023)
D.V. Leonov, S.A. Dzuba, N.V. Surovtsev, RSC Adv. 9, 34451–34456 (2019)
Funding
This work was supported by the Russian Federal Ministry of Science and Higher Education (the core funding, # FWGF-2021-0003).
Author information
Authors and Affiliations
Contributions
A.S.K. and A.N.A. made all researches and prepared figures. D.S.B. performed synthesis. S.A.D. initiated this work, interpreted the data and wrote the main manuscript text
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Ethics approval
This research did not contain any studies involving animal or human participants, nor did it take place in any private or protected area.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Kashnik, A.S., Atnyukova, A.N., Baranov, D.S. et al. DEER Study of Spatial Arrangement of Spin-Labeled Diclofenac in Lipid Bilayers of Different Composition. Appl Magn Reson 55, 1145–1157 (2024). https://doi.org/10.1007/s00723-024-01674-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00723-024-01674-8