Abstract
Proteus mirabilis is responsible for a wide range of infections that affect the urinary tract, the respiratory tract, burns, wounds and the feet of individuals with diabetes. They are highly resistant to antimicrobial agents, and new therapeutic options are therefore needed to combat this pathogen. The use of bacteriophages is one option that may be useful in treating multidrug-resistant (MDR) Proteus mirabilis infections, especially biofilm-based infections. The aim of this study was to control biofilms formed by MDR Proteus mirabilis using bacteriophages. Proteus mirabilis isolates were identified based on biochemical tests, and their resistance profiles were determined by the disk diffusion method. The biofilm-forming capacity of the isolates was assessed by the spectrophotometric method. Bacteriophages attacking Proteus mirabilis were isolated from sewage. The effect of phage on biofilm formation was investigated by the viable count method. A high rate of drug resistance was found (87.2%). Strong biofilm formation was observed in 80.5% of isolates, while moderate production was found in 19.5%. Five bacteriophages were isolated from sewage and were tested for their ability to eliminate biofilms. Significant disruption of pre-formed biofilms was observed that reached up to 99.9% decrease in the number of viable cells. The use of bacteriophages is considered a promising strategy against the biofilm infections caused by MDR Proteus mirabilis isolates.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Proteus mirabilis is a Gram-negative bacillus that belongs to the family Morganellaceae [3]. It is an opportunistic pathogen that can cause many types of infections, with urinary tract infections being the most prevalent [31]. The risk of infection is high in patients who are catheterized for long periods of time or have structural or physiological abnormalities of the urinary tract [63]. These bacteria can also cause diabetic foot ulcers in diabetic patients [58], respiratory tract infections, and infections of burns and wounds [55].
The routine use of antibiotics has led to the development of antibiotic resistance, particularly in Gram-negative bacteria [67]. Multidrug resistance (MDR) also varies depending on the isolation source, as reported by Abbas et al. [2].
Biofilms are communities of cells that adhere to a surface and are enclosed within a matrix of polysaccharides, proteins and nucleic acids [30]. Bacteria within a biofilm exhibit high levels of resistance to biocides and antimicrobial agents. These agents are needed at high concentrations, which can be 1000-fold higher than that required in the case of planktonic cells to produce an antibacterial effect [10]. The most widely investigated P. mirabilis biofilms are those that occur in the urinary tract, especially in catheterized patients. The formation of crystalline biofilms on catheters can lead to catheter encrustation and obstruction [31]. Moreover, a study performed by Abbas and Gad [1] investigated the biofilm-forming capacity of different bacteria isolated from diabetic foot infections, and it was found that the most prevalent isolates were P. mirabilis and that these all showed strong biofilm formation capacity.
Biofilm resistance to antimicrobial agents may be due to several mechanisms, including slow growth rates and low metabolic activities. Moreover, antimicrobial agents can be inactivated by the negatively charged extracellular matrix, which limits the penetration of positively charged molecules such as aminoglycosides. As a result, the penetration of these antibiotics is significantly retarded. In addition, a low oxygen level can prevent the action of some antibiotics [26, 42, 46, 62]. Another reason for resistance is the presence of dormant “persister cells” within the biofilm matrix that are resistant to antibiotic therapy [43].
To treat biofilm-related infections, new approaches are required. One of these is the use of bacteriophages [25], which can infect and lyse bacterial cells present in biofilms [24, 41]. Various studies have investigated the effect of phage on biofilm removal. Lehman and Donlan [41] and Melo et al. [47] studied the effect of pre-treatment of silicone hydrogel catheters with phage and observed a reduction in biofilm formation caused by P. mirabilis in artificial urine medium. Moreover, Nzakizwanayo et al. [51] utilized a phage tablet to effectively remove P. mirabilis from catheters in an in vitro bladder model system. Furthermore, the use of lytic bacteriophages in the prevention and eradication of biofilms of Proteus mirabilis and Escherichia coli was reported by Carson et al. [9].
The use of bacteriophages provides several advantages over traditional antibiotic therapy. They are host specific and do not affect natural microflora, they are not harmful to humans or animals, and there are many possible routes of administration. Moreover, phage preparation production is rapid, simple and inexpensive [14, 52, 60]. In this study, we investigated the ability of phages to eliminate Proteus mirabilis biofilms.
Materials and methods
Media and chemicals
Mueller Hinton broth and agar, tryptone soya broth (TSB) and agar (TSA), and MacConkey agar were purchased from Oxoid (St. Louis, USA). Cystine lactose electrolyte-deficient (CLED) medium, triple sugar iron (TSI) agar, and Simmons’ citrate agar were obtained from LAB-M (UK). Other chemicals were of pharmaceutical grade, including alpha naphthol, sodium chloride, potassium hydroxide, hydrogen peroxide, dipotassium hydrogen phosphate, barium chloride, sulphuric acid, methyl red, glycerol, 95% ethanol, isoamyl alcohol, methanol, glacial acetic acid and hydrochloric acid, which were obtained from the El Nasr Pharmaceutical Chemical Company, Cairo, Egypt. Crystal violet and p-dimethyl amino benzaldehyde (DMAB) were obtained from Avondale Laboratories, Oxon, England. The antibiotic disks that were used in this study were obtained from Oxoid (Hampshire, England). These disks contained ampicillin (AM; 10 µg), piperacillin (PRL; 100 µg), ampicillin-sulbactam (SAM; 20 µg), cefotaxime (CTX; 30 µg), cefoperazone (CEP; 75 µg), ceftazidime (CAZ; 30 µg), cefepime (FEB; 30 µg), imipenem (IPM; 10 µg), meropenem (MEM; 10 µg), aztreonam (ATM; 10 µg), gentamicin (CN; 10 µg), amikacin (AK; 30 µg), tetracycline (TE; 30 µg), ciprofloxacin (CIP; 5 µg), levofloxacin (LEV; 5 µg), sulphamethoxazole-trimethoprim (SXT; 5 µg) or chloramphenicol (C; 30 µg).
Bacterial strains and growth conditions
A total of 41 P. mirabilis isolates were used in this study. They were recovered from 400 specimens (Table 1). Specimens were obtained from patients with urinary tract infections, diabetic foot ulcers, burn infections, wound infections, and respiratory tract infections hospitalized in Zagazig University Hospital and Al Ahrar Hospital in Zagazig Sharkia Governorate, Egypt. The strains were identified based on morphology and biochemical activity according to Koneman et al. [35]. Clinical isolates of Escherichia coli, Klebsiella pneumoniae, Serratia marcescens, Proteus penneri, Proteus vulgaris and Pseudomonas aeruginosa were obtained from the stock culture of the Department of Microbiology and Immunology, Faculty of Pharmacy, Zagazig University. Salmonella Typhimurium ATCC 14028 was kindly provided by Egyptian Pharmaceutical Industries Company (EPICO), (Table 2). All isolates were maintained in Mueller Hinton broth with 10-15% glycerol and kept at -80 °C.
Antibiotic susceptibility testing
Antibiotic susceptibility was tested using the Kirby-Bauer disc diffusion method according to Clinical Laboratory Standards Institute (CLSI) guidelines [15]. The bacterial suspensions were prepared from overnight cultures on Muller-Hinton agar (Oxoid, Hampshire, England). Suspension densities were adjusted to the approximate turbidity of 0.5 McFarland standard (1.5 × 108 colony-forming units [CFU]/ml). The surface of the Muller-Hinton agar plate was evenly inoculated with the suspensions using a sterile swab. The plates were dried before applying antibiotic disks. The disks were placed on the surface no less than 24 mm apart from center to center. The plates were incubated overnight at 37 °C, after which the diameters of the inhibition zones around the disks were measured. The results were interpreted according to Clinical Laboratory Standards Institute guidelines [15].
Quantitative assessment of biofilm formation by spectrophotometry
The ability of Proteus mirabilis isolates to form biofilms was tested according to Stepanovic et al. [61]. P. mirabilis suspensions with turbidity matching that of 0.5 McFarland standard were diluted 1:100 in tryptone soya broth (TSB). Aliquots of 200 µL of the prepared suspensions were added to individual wells of 96-well U-shaped microtiter plates, which were then incubated for 24 h at 37 °C. Negative-control wells containing 200 µL of TSB alone were included in each plate. After incubation, the contents of wells were decanted and the plates were washed three times with water and left to dry in air.
To fix the biofilm, 150 µL of 99% methanol was added to each well and left for 20 min. The methanol was then removed, and the wells were left to dry in air. The fixed biofilms in the wells were stained for 15 min with 150 µL of crystal violet (1%), and the excess dye was removed by washing with water. The plates were air dried, and the bound dye was dissolved by adding 150 µL of 33% glacial acetic acid to each well. Optical density was measured spectrophotometrically at 570 nm using a Bio Tek synergy HT microplate reader (USA). Measurements were performed in triplicate, and the experiment was repeated three times. The cutoff optical density (ODc) was calculated as three standard deviations above the mean OD of the negative control.
The tested strains were classified according to the criteria of Stepanovic et al. [61] into non-biofilm producer (OD ≤ ODc), weak biofilm producer (OD > ODc, but ≤ 2× ODc), moderate biofilm producer (OD > 2× ODc, but ≤ 4× ODc) and strong biofilm producer (OD > 4× ODc).
Isolation, purification and propagation of bacteriophages
Bacteriophages were isolated from raw sewage water samples collected from different areas in Sharkia Governorate, Egypt, and identified by transmission electron microscopy (TEM). The phages were isolated by the enrichment technique [11]. Briefly, 4.5 mL of the sewage sample was mixed with 0.5 mL of Proteus mirabilis culture in early exponential phase (4-h cultures) in TSB and 5 mL of TSB in 15-mL Falcon tubes. The tubes were incubated at 37 °C for 24 h with shaking. After centrifugation at 4000 rpm for 20 min at 4 °C using a Hermle cooling centrifuge (Germany), the supernatants were filtered using 0.22-µm membrane filters (Millipore Corporation, Bedford, MA, USA) and were screened for presence of phages using the double-layer agar (DLA) technique [7]. Aliquots of 100 μL of the filtrates were mixed with 100 μL of an early-log-phase culture Proteus mirabilis and added to 3 mL of prewarmed molten soft TSA and poured into TSA plates. The plates were incubated at 37 °C for 24 h, and the presence of phages was indicated by the appearance of clear zones (plaques) on bacterial lawns.
Phages were purified by multiple rounds of single-plaque selection, resuspended in 1 mL of TSB, serially diluted, and plated until a single plaque morphology was observed [34]. The purified phages were propagated to obtain stocks with a high titer according to Carey-Smith et al. [7]. Purified phages were serially diluted in saline to give a concentration that would provide confluent lysis of the host in a soft agar overlay plate. For each dilution, three to five plates were overlaid, and after incubation, plates with almost confluent lysis were chosen. To recover the phages, 5 mL of saline was added to the plates with confluent lysis, and after 1 h at room temperature, the liquid was poured into centrifuge tubes and centrifuged. The supernatants containing phages were then filtered and stored at 4 °C [7].
Transmission electron microscopy (TEM)
Bacteriophages were visualized using transmission electron microscopy (TEM) as described by Fard et al. [20]. A single drop of purified phage was deposited on a carbon-coated copper grid (200 mesh) for 60 s, and the excess liquid was removed. The suspension was then negatively stained with 1% phosphotungstate, air dried, and observed using a JEM-2100 transmission electron microscope (JEOL Ltd.).
Host-range test
The phage host specificity against different bacterial species was tested according to Jun et al. [32] with some modifications. Aliquots of 100 μL of early-log-phase cultures of the tested bacterial suspensions were mixed with 3 mL of soft TSA (0.7%) and plated on TSA plates (1.5%). After solidification of the upper agar layer, 10 μL of a phage stock of 1010 plaque-forming units per mL (PFU/mL) was applied to the surface. The plates were then dried, incubated at 37 °C for 24 h and checked for the presence of clear zones.
Lytic spectra of isolated bacteriophages
Phage activity was tested against different clinical isolates of Proteus mirabilis according to Jun et al. [32] with some modifications. Ten microliters of diluted phage stock (1010 PFU/mL) was spotted on a bacterial lawn. After incubation, the spots were described as showing strong bacterial lysis, weak bacterial lysis, very weak bacterial lysis, or no bacterial lysis.
Anti-biofilm activity of bacteriophages
Phage isolates showing strong lytic activity were tested for their ability to remove established biofilms as described by Harrison et al. [28] with some modifications. Biofilms were formed on acid-washed 4-mm sterile glass beads [66]. Equal quantities of glass beads were added to 15-mL Falcon tubes containing 5 mL of TSB and sterilized by autoclaving. Then, 50 microliters of P. mirabilis suspensions with turbidity matching that of 0.5 McFarland standard was added to the tubes containing the beads and incubated for 24 h at 37 °C to allow biofilm formation.
The beads were washed three times with 0.9% saline to remove unattached bacteria and transferred to fresh Falcon tubes, and 2 mL of phage (109 PFU/bead) was added. Untreated beads were used as a control. The control and treated beads were incubated for 6 h at 37 °C without shaking, and the beads were washed again. For counting viable bacteria, glass beads with biofilms were transferred to fresh Falcon tubes containing 5 mL saline and disrupted by vortexing for 5 min. A serial dilution was made in saline, and 100 μL of each dilution was spread on CLED agar plates. Visible colonies were counted after 24 h of incubation at 37 °C. The number of total viable cells was calculated from the plate count. The results were expressed as log10 CFU/cm2.
Statistical analysis
Statistical analysis was performed using OneWay ANOVA test followed by Dunnett’s Multiple Comparison Test, GraphPad Prism® Software version 5.0.1 (San Diego, CA, USA). A probability value (P-value) <0.05 was considered significant.
Results
Isolation and identification of Proteus mirabilis
From a total of 400 specimens, 47 isolates of P. mirabilis were recovered with an overall frequency of 11.75% (Table 1). The frequency of P. mirabilis isolation from different sources was as follows: 15.2% from urinary tract infections, 12% from endotracheal aspirates of mechanically ventilated patients, 11.8% from burn infections, 11.4% from diabetic foot infections, and 8% from surgical wound infections. P. mirabilis isolates were identified as Gram-negative rods that did not ferment lactose on MacConkey agar, showed swarming motility on nutrient agar, and had a characteristic foul odor. Complete identification of the isolates was performed based on their biochemical characteristics (Table 3).
Antimicrobial susceptibility
It was found that 41 (87.2%) P. mirabilis isolates were resistant to at least one antibiotic of three or more different classes and were therefore considered multi-drug resistant (MDR). An antibiotic susceptibility test showed a variable degree of resistance to different antibiotics (Table 4).
Quantitative assessment of biofilm formation by spectrophotometry
Biofilm quantification was performed for all 41 MDR P. mirabilis isolates. The cutoff optical density (ODc) was calculated to be 0.08. According to the OD of the bacterial biofilm, isolates were divided into four categories: not biofilm-forming (OD ≤ 0.08), weakly biofilm-forming (OD > 0.08, but ≤ 0.16), moderately biofilm-forming (OD > 0.16, but ≤ 0.32), and strongly biofilm-forming (OD > 0.32). Among the 41 P. mirabilis isolates, strong biofilm formation was observed in 80.5%, while moderate production was found in 19.5%.
Isolation, purification and propagation of Proteus mirabilis phages
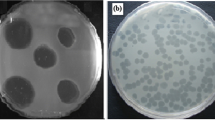
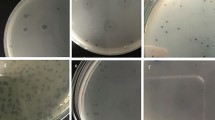
Five different phages specific for Proteus mirabilis were isolated from raw sewage water. They were named PMP1 (Proteus mirabilis phage 1), PMP2, PMP3, PMP4 and PMP5. PMP1, PMP2 and PMP5 produced large, circular, clear plaques surrounded by a halo, PMP3 produced small, circular, clear plaques surrounded by a halo, and PMP4 produced small, circular, clear plaques with no halo (Fig. 1).
Transmission electron microscopy (TEM) imaging of Proteus mirabilis phages
TEM (Fig. 2) showed that PMP1 had an icosahedral head of 45.10 × 44.41 nm and a short tail of 21.38 nm in length and 7.75 nm in width, suggesting that it probably belongs to the family Podoviridae.
Transmission electron micrographs of P. mirabilis phages (PMP1, PMP2, PMP3, PMP4 and PMP5) using a JEM-2100 transmission electron microscope (JEOL). Bars = 200 nm. Phage PMP1 has a head of 45.10 × 44.41 nm and short tail of 21.38 nm length and 7.75 nm width. PMP2 has a head of 45.31 × 43.76 nm, PMP3 has a head of 45.59 × 45.23 nm, PMP4 has a head of 43.10 × 41.60 nm PMP5 has a head of 49.75 × 49.16 nm
No tail was observed with the phages PMP2-5. PMP2 appeared to be pleomorphic, with a head size of 45.31 × 43.76 nm diameter and thus probably belong to Plasmaviridae family. PMP3, 4 and 5 had icosahedral heads with sizes of 45.59 × 45.23 nm, 43.10 × 41.60 nm, and 49.75 × 49.16 nm, respectively. These phages could belong to any of the families Microviridae, Corticoviridae, Tectiviridae, Leviviridae or Cystoviridae.
Host range test
The five isolated phages were tested against six bacterial isolates belonging to different bacterial species in the order Enterobacteriales (Salmonella Typhimurium, Escherichia coli, Klebsiella pneumoniae, Serratia marcescens, Proteus penneri and Proteus vulgaris) and Pseudomonas aeruginosa. No plaque formation was observed with the tested bacteria.
Phage lytic spectra of isolated bacteriophages
The susceptibility of the 41 multidrug-resistant Proteus mirabilis isolates to bacteriophages was tested using a modified double-layer agar technique. Only 10 clinical isolates showed lysis by one or more of the tested phages. PMP1, PMP3 and PMP4 exhibited a narrow spectrum of activity. Each lysed only 10% of the tested isolates, producing a strong lysis zone. PMP5 had a broader spectrum and lysed 20% of the bacterial isolates. A strong lytic zone was seen in 12% of the tested isolates, and a very weak lysis zone was seen in 8% of the tested isolates. PMP2 had the broadest spectrum of lysis activity. It lysed 22% of the bacterial isolates with a strong lysis area in 20% of the tested isolates and a weak lysis area in 2% of the tested isolates (Table 5).
Biofilm disruption by bacteriophages
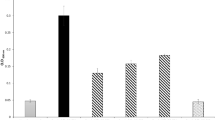
The viability of the cells in the biofilms was measured by counting. The tested bacteriophages had different abilities to reduce the number of viable cells in the biofilms (Table 6). Statistically significant reduction in biofilm in the presence and absence of phages was found at <0.05 (Fig. 3). PMP1, PMP2 and PMP3 were able to cause more than a 90% reduction in the count of bacterial cells embedded in the biofilm matrix. PMP4 caused a 3.8% to 40.2% reduction in viable counts for all of the isolates tested except for PM36, in which a 92% reduction in viability was achieved. PMP5 also had a limited effect on the viable count, with reduction ranging between 2.4% and 35.1%.
Discussion
Proteus mirabilis is the etiological agent of many human infections [13, 16, 19, 53, 57]. P. mirabilis shows high antimicrobial resistance, and its resistance to multiple drugs complicates therapy [5].
Bacteriophage therapy is a potential alternative to control biofilm infections [6, 27]. Some phages produce polysaccharide depolymerases. These enzymes degrade exopolysaccharides (EPS), allowing phages to enter bacteria embedded in the biofilm matrix [56, 59, 64]. Bacteriophages have high specificity for their hosts [36], thus minimizing overall damage to the microbial population [48]. However, this high specificity also means that phage therapy using natural lytic phages requires isolation and identification of appropriate phages, which may delay treatment. This can be partially overcome by using phage cocktails to broaden the phage host range [12, 22]. There is still a lack of properly documented clinical research on phage therapy, and there are no established protocols for the route of administration, dose, frequency or duration of treatment. There is also limited knowledge regarding phage behavior in vivo [21]. Bacteria can also become phage resistant through the lack or loss of a receptor [44] and through structural modifications and/or masking of the receptor [40]. Phages can be used in cocktails or in combination with antibiotics to overcome phage resistance [29, 38].
In this study, five different phages specific for Proteus mirabilis isolates were isolated from raw sewage samples. On culture plates, these phages produced plaques that varied in size and morphology (turbid/clear appearance, presence/absence of a halo). Some produced large plaques, while others produced small ones. All of the phages produced round, clear plaques, most of which were surrounded by a halo. These data were similar to those reported by Morozova et al. [50], who isolated four lytic P. mirabilis phages that produced large, clear plaques surrounded by a halo. Comparable results were reported by Morozova et al. [49], who isolated a phage against a Proteus mirabilis isolate that formed large clear plaques surrounded by a halo on a culture plate. Nzakizwanayo et al. [51] also isolated three phages with halo-producing plaques. Yazdi et al. [68] isolated a phage that produced clear plaques. However, Melo et al. [47] isolated two different phages infecting Proteus mirabilis. One formed clear plaques and the other formed less-clear ones. Numerous phages that induce enzymes capable of degrading the EPS of many Gram-negative bacteria, including bacteria capable of biofilm formation, have been isolated. On culture plates, such phages are characterized by halos of different sizes, surrounding the plaques obtained after the infection of a single bacterial cell. The halos are formed by bacteria from which the EPS has been removed by excess phage enzyme released during the lysis of infected cells [64].
For further identification, TEM imaging was performed on the isolated phages. It was found that the phages PMP1 and PMP2 probably belong to the families Podoviridae and Plasmaviridae, respectively, while the other three remaining phages could belong to any of the families Microviridae, Corticoviridae, Tectiviridae, Leviviridae or Cystoviridae. Further studies using techniques such as DNA sequencing are needed to determine to which families these phages belong. The present results agree with those of Nzakizwanayo et al. [51], Morozova et al. [49], and Morozova et al. [50], who showed that their phages belonged to the family Podoviridae. Moreover, Melo et al. [47] isolated two phages infecting Proteus mirabilis that belonged to the families Podoviridae and Myoviridae. However, TEM studies by Yazdi et al. [68] revealed that their phages belonged to the family Siphoviridae, and three phages reported by Thompson [65] were found to belong to the families Siphoviridae, Myoviridae and Podoviridae.
The tested phages could infect P. mirabilis isolates, but could not infect any of the other bacterial species tested. This specificity might be due to the presence of specific phages receptors present on the P. mirabilis cell surface. Receptors used by bacteriophages include flagella, polysaccharides, lipopolysaccharides, capsules, cell wall proteins, pili, and fimbria [23]. Variations in these receptors among various bacterial species have a strong effect on phage specificity [54].
In many previous reports, only a limited number of phages and hosts were used [47, 49, 50, 65, 68]. The results of this study are similar to those reported by Afriani et al. [4], who used P. mirabilis, enteropathogenic E. coli, Salmonella spp., Bacillus pumilus, and Photobacterium damselae to determine P. mirabilis phage specificity. They showed that the isolated phage exhibited high specificity against P. mirabilis. A phage isolated by Morozova et al. [49] against P. mirabilis was strain specific, lysed only one clinical strain, and did not infect the other 12 P. mirabilis isolates used in the test. However, Melo et al. [47] isolated two different phages specific for P. mirabilis isolates and tested their specificity against 43 stains of the family Enterobacteriaceae and found that both phages exhibited a much broader spectrum than in the current study; one of them lysed 16 out of 26 of Proteus strains tested, including (18 P. mirabilis strains, six P. vulgaris strains, one P. hauseri strain and one P. penneri strain), while the other lysed 26 out of 26 isolates. Both had no activity against any other tested strains in family Enterobacteriaceae.
Biofilm infections pose a therapeutic problem due to their extreme resistance to antimicrobial agents. In order to treat such infections, very high concentrations of antimicrobials are required. These concentrations may be toxic for the patient. Thus, alternative therapeutic approaches are necessary. One such approach is the use of bacteriophages [6], which have several advantages over antibiotics. First, they cause only minimal disruption of the natural microflora. Second, new phages can be discovered quickly, and their production cost is low. Third, phages can be administered by various routes and are also available in many formulations, such as creams and liquids [8, 37,38,39]. Fourth, they can be used at low dosage. Finally, no side effects have been reported with phage application, in contrast to antibiotics [45].
In this study, three bacteriophages were able to eliminate more than 60% of a pre-formed biofilm, while the other two phages had low (1%-30%) and intermediate (31%-60%) activity. The biofilm eradication activity of the bacteriophages used in the current study was higher than that reported by Carson et al. [9], who found that there was nearly complete eradication of pre-formed biofilms caused by Escherichia coli and only 10% reduction in case of Proteus mirabilis biofilm. Furthermore, the results of the current study were similar to those reported by Danis-Wlodarczyk et al. [18], who applied Pseudomonas aeruginosa PAO1 phage treatment for 4 h at 37 °C. Colony counts revealed that both phages used in the study were able to decrease a 24- to 72-h-old Pseudomonas aeruginosa PAO1 biofilm by 70%-90%. This was also reported by Cornelissen et al. [17], who observed maximum biofilm degradation 8 h after addition of high doses (104 and 106 PFU) of T7-related Pseudomonas putida phage φ15 (96% for 106 PFU) and development of resistance after 24 h. This might indicate that there is a negative correlation between biofilm degradation and time of exposure to phage. Some authors have studied the relationship between biofilm eradication and exposure time, such as Karaca et al. [33], who applied P22 phage at different titers (104-108 PFU/mL) to 24-h-old Salmonella Typhimurium DMC4 biofilms for 24h at 37 °C. They found that the highest eradication activity was 57.1% and 69% for a 24-h-old biofilm at phage titers of 107 and 108 PFU/mL, respectively. This was also reported by Danis-Wlodarczyk et al. [18]. Moreover, the resistance of biofilm to phage therapy has been studied as a time-dependent process. Yazdi et al. [68] studied the capability of a P. mirabilis phage to degrade a 24-h-old biofilm formed by P. mirabilis and found that at a multiplicity of infection (MOI) of 100, this phage removed 89% of the biofilm’s biomass after 8 h of phage treatment. However, this proportion decreased to 70% after 24 h. The development of resistance was also reported by Cornelissen et al. [17].
Conclusion
Proteus mirabilis is an opportunistic Gram-negative bacterium that can cause many types of infections in humans, such as infections of the urinary tract, respiratory tract, burns and wounds, in addition to infections of diabetic foot ulcers. High rates of drug resistance were found in this study. P. mirabilis showed high capability of biofilm formation. Biofilm formation is considered a virulence factor and contributes to antibiotic resistance. Bacteriophages have been used as an alternative to treat biofilm-based P. mirabilis infections and have been effective in elimination of biofilms. We recommend the use of bacteriophages in controlling biofilm-based infections caused by MDR P. mirabilis strains.
References
Abbas HA, Gad AI (2014) Eradication of biofilms formed by bacteria isolated from diabetic foot infections by potential antibiofilm agents alone and in combination with ciprofloxacin. Afr J Microbiol Res 8:3882–3892
Abbas HA, El-Saysed MA, Ganiny AM, Fattah AA (2018) Antimicrobial resistance patterns of Proteus mirabilis isolates from urinary tract, burn wound and diabetic foot infections. Res J Pharm Tech 11:249–252
Adeolu M, Alnajar S, Naushad S, Gupta RS (2016) Genome-based phylogeny and taxonomy of the ‘Enterobacteriales’: proposal for Enterobacterales ord. nov. divided into the families Enterobacteriaceae, Erwiniaceae fam. nov., Pectobacteriaceae fam. nov., Yersiniaceae fam. nov., Hafniaceae fam. nov., Morganellaceae fam. nov., and Budviciaceae fam. nov. Int J Syst Evol Microbiol 66:5575–5599
Afriani R, Rusmana I, Budiarti S (2014) Characterization of Proteus mirabilis lytic phage from Situ Letik River Bogor Indonesia. Int J Innovat Res Sci Eng 2:2347–3207
Akova M (2016) Epidemiology of antimicrobial resistance in bloodstream infections. Virulence 7:252–266
Azeredo J, Sutherland IW (2008) The use of phages for the removal of infectious biofilms. Curr Pharm Biotechnol 9:261–266
Carey-Smith GV, Billington C, Cornelius AJ, Hudson JA, Heinemann JA (2006) Isolation and characterization of bacteriophages infecting Salmonella spp. FEMS Microbiol Lett 258:182–186
Carlton RM (1999) Phage therapy: past history and future prospects. Arch Immunol Ther Exp (Warsz) 47:267–274
Carson L, Gorman SP, Gilmore BF (2010) The use of lytic bacteriophages in the prevention and eradication of biofilms of Proteus mirabilis and Escherichia coli. FEMS Immunol Med Microbiol 59:447–455
Ceri H, Olson ME, Stremick C, Read RR, Morck D, Buret A (1999) The Calgary Biofilm Device: new technology for rapid determination of antibiotic susceptibilities of bacterial biofilms. J Clin Microbiol 37:1771–1776
Cerveny KE, DePaola A, Duckworth DH, Gulig PA (2002) Phage therapy of local and systemic disease caused by Vibrio vulnificus in iron-dextran-treated mice. Infect Immun 70:6251–6262
Chan BK, Abedon ST, Loc-Carrillo C (2013) Phage cocktails and the future of phage therapy. Fut Microbiol 8:769–783
Chen C-Y, Chen Y-H, Lu P-L, Lin W-R, Chen T-C, Lin C-Y (2012) Proteus mirabilis urinary tract infection and bacteremia: risk factors, clinical presentation, and outcomes. J Microbiol Immunol Infect 45:228–236
Clark JR, March JB (2006) Bacteriophages and biotechnology: vaccines, gene therapy and antibacterials. Trends Biotechnol 24:212–218
CLSI-Clinical and Laboratory Standards Institute (2016) Performance standards for antimicrobial susceptibility testing, CLSI document M100-S-26. CLSI-Clinical and Laboratory Standards Institute, Wayne
Coker C, Poore CA, Li X, Mobley HL (2000) Pathogenesis of Proteus mirabilis urinary tract infection. Microb Infect 2:1497–1505
Cornelissen A, Ceyssens P-J, T’syen J, Van Praet H, Noben J-P, Shaburova OV, Krylov VN, Volckaert G, Lavigne R (2011) The T7-related Pseudomonas putida phage φ15 displays virionassociated biofilm degradation properties. PLoS One 6:e18597
Danis-Wlodarczyk K, Olszak T, Arabski M, Wasik S, Majkowska-Skrobek G, Augustyniak D, Gula G, Briers Y, Jang HB, Vandenheuvel D, Duda KA, Lavigne R, Drulis-Kawa Z (2015) Characterization of the newly isolated lytic bacteriophages KTN6 and KT28 and their efficacy against Pseudomonas aeruginosa biofilm. PLoS One 10:e0127603
Endimiani A, Luzzaro F, Brigante G, Perilli M, Lombardi G, Amicosante G, Rossolini GM, Toniolo A (2005) Proteus mirabilis bloodstream infections: risk factors and treatment outcome related to the expression of extended-spectrum beta-lactamases. Antimicrob Agents Chemother 49:2598–2605
Fard RMN, Barton MD, Heuzenroeder MW (2010) Novel Bacteriophages in Enterococcus spp. Curr Microbiol 60:400–406
Ghannad MS, Mohammadi A (2012) Bacteriophage: time to re-evaluate the potential of phage therapy as a promising agent to control multidrug-resistant bacteria. Iran J Basic Med Sci 15:693–701
Goodridge LD (2010) Designing phage therapeutics. Curr Pharm Biotechnol 11:15–27
Gurnev PA, Oppenheim AB, Winterhalter M, Bezrukov SM (2006) Docking of a single phage lambda to its membrane receptor maltoporin as a time-resolved event. J Mol Biol 359:1447–1455
Gutiérrez D, Vandenheuvel D, Martínez B, Rodríguez A, Lavigne R, García P (2015) Two phages, phiIPLA-RODI and phiIPLA-C1C, lyse mono-and dual-species Staphylococcal biofilms. Appl Environ Microbiol 81:3336–3348
Hall-Stoodley L, Costerton JW, Stoodley P (2004) Bacterial biofilms: from the natural environment to infectious diseases. Nat Rev Microbiol 2:95–108
Hall-Stoodley L, Stoodley P (2009) Evolving concepts in biofilm infections. Cell Microbiol 11:1034–1043
Harper D, Enright M (2011) Bacteriophages for the treatment of Pseudomonas aeruginosa infections. J Appl Microbiol 111:1–7
Harrison JJ, Turner RJ, Joo DA, Stan MA, Chan CS, Allan ND, Vrionis HA, Olson ME, Ceri H (2008) Copper and quaternary ammonium cations exert synergistic bactericidal and anti-biofilm activity against Pseudomonas aeruginosa. Antimicrob Agents Chemother 52:2870–2881
Ho K (2001) Bacteriophage therapy for bacterial infections. Rekindling a memory from the pre-antibiotics era. Perspect Biol Med 44:1–16
Hoiby N, Bjarnsholt T, Givskov M, Molin S, Ciofu O (2010) Antibiotic resistance of bacterial biofilms. Int J Antimicrob Agents 35:322–332
Jacobsen SM, Shirtliff ME (2011) Proteus mirabilis biofilms and catheter-associated urinary tract infections. Virulence 2:460–465
Jun JW, Kim JH, Shin SP, Han JE, Chai JY, Park SC (2013) Characterization and complete genome sequence of the Shigella bacteriophage pSf-1. Res Microbiol 164:979–986
Karaca B, Akcelik N, Akcelik M (2015) Effects of P22 bacteriophage on Salmonella enterica subsp. enterica serovar Typhimurium DMC4 strain biofilm formation and eradication. Arch Biol Sci 67:1361–1367
Karumidze N, Kusradze I, Rigvava S, Goderdzishvili M, Rajakumar K, Alavidze Z (2013) Isolation and characterisation of lytic bacteriophages of Klebsiella pneumoniae and Klebsiella oxytoca. Curr Microbiol 66:251–258
Koneman E, Winn WC, Allen S, Janda W, Procop G, Schreckenberger P, Woods G (2006) Koneman’s color atlas and textbook of diagnostic microbiology, 6th edn. Lippincott Williams & Wilkins, Pennsylvania
Koskella B, Meaden S (2013) Understanding bacteriophage specificity in natural microbial communities. Viruses 5:806–823
Krylov VN (2001) Phagotherapy in terms of bacteriophage genetics: hopes, perspectives, safety, limitations. Genetika 37:869–887
Kutateladze M, Adamia R (2010) Bacteriophages as potential new therapeutics to replace or supplement antibiotics. Trends Biotechnol 28:591–595
Kutter E, De Vos D, Gvasalia G, Alavidze Z, Gogokhia L, Kuhl S, Abedon ST (2010) Phage therapy in clinical practice: treatment of human infections. Curr Pharm Biotechnol 11:69–86
Labrie SJ, Samson JE, Moineau S (2010) Bacteriophage resistance mechanisms. Nat Rev Microbiol 8:317–327
Lehman SM, Donlan RM (2015) Bacteriophage-mediated control of a two-species biofilm formed by microorganisms causing catheter-associated urinary tract infections in an in vitro urinary catheter model. Antimicrob Agents Chemother 59:1127–1137
Lewis K (2001) Riddle of biofilm resistance. Antimicrob Agents Chemother 45:999–1007
Lewis K (2010) Persister cells. Annu Rev Microbiol 64:357–372
Liu M, Deora R, Doulatov SR, Gingery M, Eiserling FA, Preston A, Maskell DJ, Simons RW, Cotter PA, Parkhill J, Miller JF (2002) Reverse transcriptase-mediated tropism switching in Bordetella bacteriophage. Science 295:2091–2094
Loc-Carrillo C, Abedon ST (2011) Pros and cons of phage therapy. Bacteriophage 1:111–114
Lynch AS, Robertson GT (2008) Bacterial and fungal biofilm infections. Annu Rev Med 59:415–428
Melo LD, Veiga P, Cerca N, Kropinski AM, Almeida C, Azeredo J, Sillankorva S (2016) Development of a phage cocktail to control Proteus mirabilis catheter-associated urinary tract infections. Front Microbiol 7:1024–1035
Modi SR, Collins JJ, Relman DA (2014) Antibiotics and the gut microbiota. J Clin Invest 124:4212–4218
Morozova V, Kozlova Y, Shedko E, Kurilshikov A, Babkin I, Tupikin A, Yunusova A, Chernonosov A, Baykov I, Capital Kabul CI, Kabilov M, Ryabchikova E, Vlassov V, Tikunova N (2016) Lytic bacteriophage PM16 specific for Proteus mirabilis: a novel member of the genus Phikmvvirus. Arch Virol 161:2457–2472
Morozova V, Kozlova Y, Shedko E, Babkin I, Kurilshikov A, Bokovaya O, Bardashova A, Yunusova A, Tikunov A, Tupikin A, Ushakova T, Ryabchikova E, Tikunova N (2018) Isolation and characterization of a group of new Proteus bacteriophages. Arch Virol 163:2189–2197
Nzakizwanayo J, Hanin A, Alves DR, McCutcheon B, Dedi C, Salvage J, Knox K, Stewart B, Metcalfe A, Clark J (2015) Bacteriophage can prevent encrustation and blockage of urinary catheters by Proteus mirabilis. Antimicrob Agents Chemother 60:1530–1536
Parasion S, Kwiatek M, Gryko R, Mizak L, Malm A (2014) Bacteriophages as an alternative strategy for fighting biofilm development. Pol J Microbiol 63:137–145
Perim MC, Borges Jda C, Celeste SR, Orsolin Ede F, Mendes RR, Mendes GO, Ferreira RL, Carreiro SC, Pranchevicius MC (2015) Aerobic bacterial profile and antibiotic resistance in patients with diabetic foot infections. Rev Soc Br Med Trop 48:546–554
Projan S (2004) Phage-inspired antibiotics? Nat Biotechnol 22:167–168
Różalski A, Sidorczyk Z, Kotelko K (1997) Potential virulence factors of Proteus bacilli. Microbiol Mol Biol Rev 61:65–89
Rydman PS, Bamford DH (2002) The lytic enzyme of bacteriophage PRD1 is associated with the viral membrane. J Bacteriol 184:104–110
Sekhar S, Vyas N, Unnikrishnan M, Rodrigues G, Mukhopadhyay C (2014) Antimicrobial susceptibility pattern in diabetic foot ulcer: a pilot study. Ann Med Health Sci Res 4:742–745
Shanmugam P, Jeya M, Susan SL (2013) The bacteriology of diabetic foot ulcers, with a special reference to multidrug resistant strains. J Clin Diagn Res 7:441–445
Shapiro OH, Kushmaro A (2011) Bacteriophage ecology in environmental biotechnology processes. Curr Opin Biotechnol 22:449–455
Skurnik M, Strauch E (2006) Phage therapy: facts and fiction. Int J Med Microbiol 296:5–14
Stepanović S, Vuković D, Hola V, Bonaventura GD, Djukić S, Ćirković I, Ruzicka F (2007) Quantification of biofilm in microtiter plates: overview of testing conditions and practical recommendations for assessment of biofilm production by staphylococci. Acta Pathol Microbiol Immunol Scand B Microbiol 115:891–899
Stewart PS (2002) Mechanisms of antibiotic resistance in bacterial biofilms. Int J Med Microbiol 292:107–113
Stickler DJ (2014) Clinical complications of urinary catheters caused by crystalline biofilms: something needs to be done. J Intern Med 276:120–129
Sutherland IW, Hughes KA, Skillman LC, Tait K (2004) The interaction of phage and biofilms. FEMS Microbiol Lett 232:1–6
Thompson R (2018) The isolation and characterisation of Proteus mirabilis bacteriophages and their effect on the colonisation and blockage of urinary catheters. (PhD thesis), Faculty of Health and Applied Sciences, University of the West of England, Bristol
Trachoo N (2004) Biofilm removal technique using sands as a research tool for accessing microbial attachment on surface. Songklanakarin J Sci Technol 26:109–115
Yah S, Enabulele I, Yusuf E, Eghafona N (2006) Emerging quinolones resistant transfer genes among gram-negative bacteria isolated from faeces of HIV/AIDS patient attending some clinic and hospital in the city of Benin, Edo State, Nigeria. Online J Health Allied Sci 5:61–91
Yazdi M, Bouzari M, Ghaemi EA (2018) Isolation and characterization of a lytic bacteriophage (vB_PmiS-TH) and its application in combination with ampicillin against planktonic and biofilm forms of Proteus mirabilis isolated from urinary tract infection. J Mol Microbiol Biotechnol 28:37–46
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Handling Editor: T. K. Frey.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Gomaa, S., Serry, F., Abdellatif, H. et al. Elimination of multidrug-resistant Proteus mirabilis biofilms using bacteriophages. Arch Virol 164, 2265–2275 (2019). https://doi.org/10.1007/s00705-019-04305-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-019-04305-x