Abstract
Morphological and molecular characters were analysed to investigate diversity within isolates of the Glomus claroideum/Glomus etunicatum species group in the genus Glomus. The inter- and intra-isolate sequence diversity of the large subunit (LSU) rRNA gene D2 region of eight isolates of G. claroideum and G. etunicatum was studied using PCR-single strand conformational polymorphism (SSCP)-sequencing. In addition, two isolates recently obtained from Southern China were included in the analysis to allow for a wider geographic screening. Single spore DNA isolation confirmed the magnitude of gene diversity found in multispore DNA extractions. An apparent overlap of spore morphological characters was found between G. claroideum and G. etunicatum in some isolates. Analysis of the sequence frequencies in all G. etunicatum and G. claroideum isolates (ten) showed that four LSU D2 sequences, representing 32.1% of the clones analysed for multispore extraction (564) were found to be common to both species, and those sequences were the most abundant in four of the ten isolates analysed. The frequency of these sequences ranged between 23.2% and 87.5% of the clones analysed in each isolate. The implications for the use of phenotypic characters to define species in arbuscular mycorrhizal fungi are discussed. The current position of G. claroideum/G.etunicatum in the taxonomy of the Glomeromycota is also discussed.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
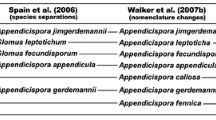
Until recently, the arbuscular mycorrhizal fungi (AMF) were placed within the order Glomales, and comprised five families, seven genera and around 150 formally accepted species (Walker and Trappe 1993; Morton 2000; Morton and Redecker 2001). Recently, however, there has been a published set of corrections and a new taxonomic structure proposed based on small sub-unit (SSU) ribosomal RNA (rRNA) gene sequences (Schüßler et al. 2001b). All AMF and related fungi have been placed within the phylum Glomeromycota, which comprises four orders: Paraglomerales, Archaeosporales, Diversisporales (containing the Acaulosporaceae and Gigasporaceae) and the Glomerales. The grouping of Acaulosporaceae with Gigasporaceae in Diversisporales based on rRNA gene sequences alone is controversial, and debate will continue until evidence from other genes can be found (Morton 2003). The Glomerales comprises two clades of species of Glomus. The genus Glomus includes more than 50% of all described species (Simon et al. 1993a; Stürmer and Morton 1997). In many cases, however, species have been described that are clearly synonymous with other species (Rosendahl et al. 1994; Bentivenga et al. 1997; Walker and Vestberg 1998; Morton 2003). Considerable genetic diversity has been reported within the genus Glomus (Dodd et al. 1996; Lloyd-MacGilp et al. 1996; Simon 1996; Clapp et al. 2001; Schwarzott et al. 2001), but the morphological diversity of spores appears to be lower than that in other genera (Stürmer and Morton 1997).
There is increasing evidence that the genus Glomus is polyphyletic/paraphyletic (Rosendahl et al. 1994; Schüßler et al. 2001a, 2001b; Schwarzott et al. 2001). Several phylogenetic analyses have used data sets comprising partial or complete sequences from the SSU rRNA genes (Helgason et al. 1998; Sawaki et al. 1998; Schüßler 1999; Kramadibrata et al. 2000; Redecker et al. 2000a, 2000b; Schwarzott et al. 2001). These studies have shown the existence of more than one clade in the genus Glomus. These analyses have indicated that Glomus claroideum (Schenck and Smith) Walker and Vestberg and Glomus etunicatum Becker and Gerdemann (named as Glomus—group B by Schüßler et al. 2001b) are distantly related to other Glomus species (named as Glomus—group A by Schüßler et al. 2001b). Studies of spore development of several species of Glomus (Stürmer and Morton 1997) concluded that G. etunicatum and G. clarum share common ontogeny and hyphal structures, but diverge from the group formed by G. intraradices and G. claroideum. In contrast, phylogenetic trees based on complete SSU rRNA gene sequences indicate that G. etunicatum and G. claroideum cluster together in a distinct group that diverges from a main cluster that includes G. intraradices and G. manihotis (Schüßler 1999; Schüßler et al. 2001b). Similarly, analysis of the highly conserved 5.8S rRNA gene grouped G. etunicatum and G. claroideum independently from G. clarum (Redecker et al. 1999). However, when homologous regions of the 5.8S and ITS (internal transcribed spacer) regions were analysed for specific primer design, G. etunicatum, G. claroideum and G. intraradices clustered separately from the “G. mosseae group” (Millner et al. 2001). Furthermore, the resolution of both the SSU and ITS regions has been questioned when closely related species or isolates are compared (Simon et al. 1993b; Helgason et al. 1998; Kjoller and Rosendahl 2000). Therefore, our investigation here concentrates on the large sub-unit (LSU) rRNA gene.
Little is known of the range of diversity within species of AMF because of the difficulty in identifying them in the field and the frequent problems or lack of resolve in obtaining them in pure culture. As a consequence, only a few isolates are normally included in diversity analyses, considerably limiting interpretation of AMF variability. In the case of the main phylogenetic cluster of G. etunicatum/G. claroideum—Glomus Group B (Schüßler et al. 2001b), there has been no investigation on the diversity within the two species and how different isolates cluster within this group. In addition, analysis of genetic diversity in the Glomeromycota has often failed to consider the occurrence of multiple copies of the rRNA genes within single spores of AMF and the analysis of the G. etunicatum and G. claroideum clade is no exception (Millner et al. 2001; Schüßler et al. 2001a; Schwarzott et al. 2001). Recently, the use of sequences from the LSU rRNA gene D2 region, after pre-screening for variation with single strand conformational polymorphism (SSCP), has proved to be a valuable tool in the study of gene diversity and structure of species in the Glomeromycota. This approach has been applied to study four, well defined, and closely related, species in the genus Glomus (Clapp et al. 2001). The analysis of seven Glomus coronatum isolates (representing 80% of the available germplasm for this species) along with Glomus mosseae, Glomus constrictum and Glomus geosporum revealed a possible genetic continuum between these species. Sequences forming a main “core cluster” of G. coronatum were identified. However, several sequences from spores of G. mosseae, G. constrictum and G. geosporum also fell within the G. coronatum cluster. These were only detected because of the large number of clones investigated.
The main purpose of this study was to investigate diversity across the Glomus etunicatum/Glomus claroideum species group in a comparative analysis using morphological, and molecular characters. There is a growing consensus nowadays that intra-specific diversity is of major importance in assessing biodiversity of these fungi as it has been suggested that AMF diversity is larger at intra-specific levels than at other taxonomic levels. In total, ten AMF isolates of G. etunicatum/G. claroideum isolated in pure culture from around the world were subjected to both morphological and molecular analysis. The second objective was to determine whether the two species could be clearly distinguished by analysing different characters. In addition, morphological analysis of 35 isolates belonging to 12 AMF species was included to broaden the perspective on diversity across the genus Glomus. The PCR-SSCP sequencing technique described in Clapp et al. (2001) was used to investigate diversity of the LSU rRNA gene, concentrating on the variable D2 region, and to further analyse the diversity composition and sequence relationships within the group G. etunicatum/G. claroideum.
Materials and methods
Maintenance of the isolates
Isolates of AMF were obtained from the culture collection at the International Institute of Biotechnology (IIB; Sittingbourne, Kent, UK) and the culture collection at Pruhonice, Czech Republic. Isolates obtained from agricultural regions in Southern China were also included in this analysis. The identity of all isolates used was checked. The fungi were maintained either in an attapulgite clay product (Agsorb 8/16 LVM-6A; Oil-Dri, Wisbech, UK), a durite sand (a calcined flint of pH 8.5 in water), or a mixture of the two at IIB, whilst a mixture 3:1 of sand:original-site-soil was employed in the Czech Republic. Isolate G. claroideum V43a was grown in a soil-peat mixture. The cultures were started with either single or multi-spore inocula—but all have been sub-cultured as multi-spores over many subsequent generations—and employed a range of host plants (Zea mays, Thymus sp., Allium porrum or Plantago lanceolata). For further details on culturing history of these isolates, please refer to the BEG, La Banque Européenne des Glomales now known as the International Bank for the Glomeromycota [http://www.kent.ac.uk/bio/beg]. Arbuscular mycorrhizal fungi were maintained in open pot cultures in a glasshouse (min 18°C/max 40°C, relative humidity 60–80%, photoperiod 18 h/6 h light/dark). Supplementary metal halide lighting (400 W) was used, giving a photon flux density of 400–600 µmol m−2 s−1 measured at the leaf surface (between and directly under the lights, respectively). Pot cultures were watered every two days with deionised water and weekly with 1.4 g l−1 Vitafeed 102 (Vitax, Leicester, UK) and NK (18–36) fertiliser with trace elements.
Morphological characterisation
Table 1 lists the isolates used in this study and their origins. All cultures were checked for monospecificity and spores were observed to correspond with original species descriptions. Pot culture material from each isolate was passed through 710 μm and 45 μm sieves and the finer fraction collected. This material was centrifuged in a water-sucrose 60% (w/v) step-gradient for 3 min. Spores were collected from the water-sucrose interface and used either for making permanent herbarium slides, for transmission electron microscopy (TEM), or for DNA analysis.
Diagnostic slides containing 20 broken and 20 unbroken spores were prepared according to a standard protocol (http://www.kent.ac.uk/bio/beg/). Apparently healthy mature spores were selected using spore colour and size as indicators. Spore colour was determined using a stereomicroscope with a split fibre-optic light source (colour temperature of 3,200°K) at magnifications up to 50×, by comparison with the INVAM (Morton 2003) colour chart for spores. Colour values comprised percentage cyan, magenta, yellow and black (C-M-Y-B). Spore diameter, spore wall thickness and structure were assessed under a dissecting Zeiss-Axioskop microscope with bright field optics and differential interference contrast, at magnifications of up to 200×. Presence of an outer mucilaginous wall layer and the spore wall lamination were recorded and included as binomial data for the analysis.
DNA analyses
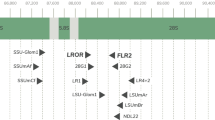
All DNA extractions, amplifications and cloning included in this study were performed as described by Clapp et al. (2001). DNA was extracted from multi-spore preparations (150 spores) of seven isolates of G. etunicatum (BEG136, BEG137, BEG92, BEG146, S329, BEG186 and BEG151), and three isolates of G. claroideum (BEG23, BEG3, and V198). A G. lamellosum ex-holotype, G. intraradices BEG145 and BEG193, G. manihotis BEG112 and G. mosseae BEG185 were also used for multispore DNA extraction and included in the analysis of sequence relationships. In addition, single-spore DNA isolations from G. claroideum BEG3 and V198 were used for PCR amplifications to confirm the sequence variability found within multispore preparations. Single-spore DNA isolations were performed using the methodology described by Schwarzott and Schüßler (2001). PCR amplification of the LSU rRNA gene D2 region used primers ALF01 and NDL22 as described by Clapp et al. (2001). Subsequently, purified amplicons were ligated into the vector pGEM-T Easy (Promega) and transformed into competent Escherichia coli JM109 (Promega).
PCR-SSCP and sequence analyses of the D2 region of the LSU rRNA gene
Between 38 and 64 cloned inserts from each selected isolate of G. claroideum and G. etunicatum were screened for sequence variation using PCR-SSCP of cloned amplicons, and the frequency of each SSCP type recorded. Clones representative of each SSCP type were purified and their inserts sequenced. PCR-SSCP, sequencing and phylogenetic analyses were carried out as described in Clapp et al. (2001). All sequences were checked for the possibility that they were chimeric using Chimera Check [http://rdp.cme.msu.edu/html/]. All sequences obtained were checked against existing sequences using BLAST [http://www.ncbi.nlm.nih.gov/index.html]. A group of inserts obtained from G. intraradices BEG145 and BEG193, G. manihotis BEG112, G. mosseae BEG185 and G. lamellosum ex-holotype were also selected, purified, sequenced and analysed as described above. Previously published sequences from other Glomus species, and from the Gigasporaceae and Acaulosporaceae were also included in the alignment (see Clapp et al. 2001; Rodriguez et al. 2001). In addition, sequences from the ribosomal sequence database that spanned the D2 domain of Basidiomycota, Ascomycota and Zygomycota sequences were included as out-groups. Altogether, 220 sequences (209 in-group taxa) were assessed over 470 characters. Phylogenetic analysis used PAUP (Swofford 1999) and analyses were carried out using both neighbour joining (Saitou and Nei 1987) and maximum parsimony (Swofford 1999) analysis. Maximum parsimony analysis was performed using heuristic search options with 100 bootstrap replications (Felsenstein 1985) branch swapping by tree-bisection-reconnection (TBR), branches able to collapse to yield polytomies, and with parsimony uninformative characters excluded. Single, unambiguously aligned gaps were treated as fifth character states. Trees were displayed using Treeview (Page 1996).
Results
Spore morphology study
The majority of isolates used in this study were observed to fit their original species descriptions. Spore size, wall thickness, hyphal attachment and colour were all within the range originally described (Fig. 1). A range of morphological diversity of spores among isolates of G. claroideum and G. etunicatum was, however, observed (Fig. 2). The analysis of spore dimensions, spore wall structure and colour among isolates of G. claroideum and G. etunicatum showed a clear overlap (data not shown). It was, however, possible to establish differences between isolates of G. claroideum and G. etunicatum on the basis of spore colour (Fig. 1). Glomus lamellosum could also not be separated on the basis of spore morphology from G. claroideum (Fig. 1).
Size and colour of spores from the several Glomus species investigated. * Data in the first and second columns were taken directly from the respective species descriptions or re-descriptions. Spore colour data are presented on the scale used on the INVAM colour chart. The third and fourth columns are combined data from isolates included in the analysis (n number of isolates). In the fourth column, modal values with error bars are presented. Spore colour presented as percentage cyan (C), magenta (M), yellow (Y) and black (B). ** Chinese isolates are not included
Spore morphologies of isolates of arbuscular mycorrhizal fungi (AMF) used in this study. a G. claroideum BEG3 showing the presence of an innermost wall layer “endospore” as described by Walker and Vestberg (1998). b G. claroideum V43a, specimen showing the typical spore wall structure. c G. lamellosum ex-holotype spore wall pigmentation quite similar to G. claroideum specimens. Note the thick spore wall and the wall lamination. d G. etunicatum BEG146 specimen showing typical hyphal attachment and spore wall structure. e G. claroideum A6 specimen showing the mucigel-like outermost wall layer as described by Walker and Vestberg (1998) present in some specimens. f G. etunicatum BEG136 specimen showing spore wall typically comprising two spore wall layers. g G. etunicatum S329 specimen showing the typical wall structure with an additional innermost layer (arrow), which becomes thicker at maturity. h G. claroideum BEG93 crushed spore showing no obvious innermost separable layer visible as noted in other isolates of G. claroideum. i G. claroideum V198 here the innermost wall layer appears as an “endospore” after gentle crushing. j G. manihotis BEG112 outermost wall layer giving “halo” effect and an inner pigmented wall layer. Note extensive wall thickening of hyphal attachment that differentiates this species. k G. etunicatum BEG137 showing pigmented wall layer observed in G. etunicatum isolates and small remnants of evanescent outer hyaline wall layer (arrow). l G. geosporum BEG11 used as an outgroup in the morphological analysis, with typical spore wall structure and hyphal attachment. BEG La Banque Européenne des Glomales, now International Bank of Glomeromycota (http://www.kent.ac.uk/bio/beg). Bars 50 µm
Particularly interesting was the case of G. etunicatum S329, which had, in addition to an outermost hyaline mucilaginous layer (L1) and a second unit layer (L2), an innermost wall layer (L3) not found in other isolates of G. etunicatum (Fig. 2). Spore walls of G. etunicatum S329 and BEG137 viewed with TEM showed that the innermost layer (L3) was unique to the former and appeared as an amorphous electron translucent layer, which became thicker on more mature spores (data not shown).
PCR-SSCP analysis and sequencing
The LSU D2 region of the rRNA gene was successfully amplified for each isolate studied, and a typical product 460 bp in length was obtained from all isolates. Sequences were also obtained from G. intraradices BEG145 and BEG193, G. manihotis BEG112, G. mosseae BEG185 and G. lamellosum ex-holotype. In total, 658 cloned inserts were analysed for sequence polymorphisms as follows: 38, 48, 63, 53, 64, 55 and 62 from G. etunicatum BEG136, BEG137, S329, BEG146, BEG92, BEG186 and BEG151, respectively, and 56, 64, and 61 clones were analysed from G. claroideum BEG3, V198, BEG23, respectively, all from multispore DNA preparations. Also included were 48 and 46 inserts from single-spore DNA extractions of G. claroideum BEG3 and V198, respectively.
No non-glomeromycotan sequences were recovered. Similar PCR-SSCP patterns resulted in identical sequences. In addition to the 64 sequences obtained from multispore (51) and single-spore (13) DNA extractions of isolates of G. claroideum and G. etunicatum (accession numbers: AY541810–AY541851, AY541873–AY541894), 10 sequences were obtained from G. lamellosum ex-holotype (accession numbers: AY541858–AY541867), 6 from G. intraradices BEG145 (accession numbers: AY541852–AY541857), and 5 from G. manihotis BEG112 (accession numbers: AY541868–AY541872). Also included in the analysis were 11 sequences from G. intraradices BEG193 (accession numbers: AY541895–AY541906) and 12 sequences from G. mosseae BEG185 (accession numbers: AY541907–AY541918) isolated from China.
LSU D2 domain rRNA sequence frequencies
A total of 564 cloned inserts obtained from multispore DNA extractions of 10 G. etunicatum and G. claroideum isolates, were used for the sequence diversity analysis described in this section. Of the 564 clones, 452 (80%), comprising 51 PCR-SSCP patterns, were identified as either G. claroideum or G. etunicatum after representative clones were sequenced. The remaining 112 (20%) clones comprised 81 PCR-SSCP patterns that were not sequenced but included in the analysis of total and individual (per isolate) diversity.
The intra-isolate diversity composition, based on the relative frequencies of each PCR-SSCP pattern found in each isolate, is shown in Fig. 3. The number of different sequences (PCR-SSCP patterns) among isolates ranged from four (G. claroideum V198) to 26 (G. etunicatum BEG146). The frequency of those patterns was between 0.17% and 12.2%, as calculated with respect to the total number of clones analysed for all isolates (564).
Charts showing the frequencies of each sequence type within each isolate. Black bars Unique sequences within each isolate, coloured bars sequences found in more than one isolate. BEG La Banque Européenne des Glomales, now International Bank of Glomeromycota (http://www.kent.ac.uk/biol/beg)
Five of the isolates studied, G. etunicatum—BEG136, BEG137 and S329 and G. claroideum BEG23 and V198—had one sequence that comprised a high proportion of the clones analysed in each case (Table 2). The frequencies of these five more abundant sequences ranged from 47.5% to 87.5% of the clones analysed in each isolate. Of these sequences, three were found in more than one isolate, BEG136-02, BEG137-01 and BEG3-11 (Table 2).
Seven of the ten isolates had sequences in common. Both Chinese isolates included in this study—Glomus etunicatum BEG186 and BEG151—had no sequences found in other isolates analysed. Similarly, G. etunicatum S329 had only unique sequences. Seven sequences were found in more than one isolate (AY541823, AY541825, AY541840, AY541841, AY541842, AY541843 and AY541846) but each was not necessarily found in every isolate. Their frequencies are shown in Table 2. The range of frequencies of these sequences was between 0.5% and 12.2% of the total number of clones analysed (564). Sequence G. etunicatum BEG137-01 was the most abundant, with 12.2% of the total clones containing this sequence (Table 2). The seven isolates found to have sequences in common, G. claroideum BEG23, BEG3 and V198 and G. etunicatum BEG92, BEG136, BEG137 and BEG146, can be classified into three different groups. The first group corresponds to isolates that had many different sequences with one or two sequences in common, with those common sequences being found at low frequency (G. etunicatum BEG146 and BEG92). The second group corresponds to isolates that had few sequences in common but those sequences were the most abundant in every case (G. etunicatum BEG136 and BEG137 and G. claroideum V198). In the third group, isolates had several sequences in common with other isolates and those sequences were found in a high number of clones (G. claroideum BEG23 and BEG3) (Fig. 3). Analysis of sequence diversity found that most of the clones analysed in G. claroideum V198 contained either G. etunicatum BEG137-01 (24/64 clones) or BEG136-02 (38/64 clones).
Sequence relationships
The analysis was based on 220 taxa, including sequences from G. etunicatum and G. claroideum isolates, obtained in both multispore and single-spore DNA isolations. In total, 470 characters were analysed, 355 (75.5%) parsimony informative positions out of the alignment of 430 (91.4%) variable bases; 75 uninformative characters were excluded from the analysis. The neighbour-joining and maximum parsimony trees, with minor differences in topology, gave almost identical results. Figure 4 shows the results of the phylogenetic analysis performed. All sequences isolated from G. claroideum and G. etunicatum clustered together in an independent clade within Glomus. Sequences from the G. lamellosum ex-holotype grouped within the G. claroideum/G. etunicatum clade. The G. claroideum/G. etunicatum clade comprised two sub-clusters; one contained the majority of sequences isolated from these species (52 out of 69 sequences), and seven of the ten sequences isolated from the G. lamellosum ex-holotype. In addition, all G. claroideum and G. etunicatum sequences from the GenBank database included in this analysis fell into this sub-cluster. Two G. claroideum BEG14 sequences (AJ271928 and AF235007) already in the databases clustered in distinct groups within this sub-cluster, showing that different sequences from the same isolate had been found before in the G. claroideum group. Two discrete groups of sequences isolated from G. etunicatum BEG92 and S329 were recognised, which included unique sequences obtained from both isolates, representing 58% and 90.5% of the clones analysed in each case, respectively. Additionally, a sequence isolated from G. etunicatum S329 by other authors (Boehm and Hock, personal communication) (AF389015) fell within our G. etunicatum S329 cluster, corroborating our results (Fig. 4). The second sub-cluster found in the G. claroideum/G. etunicatum clade comprised sequences isolated from both species associated with the remaining three sequences obtained from G. lamellosum ex-holotype. All sequences obtained from G. intraradices BEG145 and G. manihotis BEG112 formed a discrete cluster outside the G. etunicatum/G. claroideum clade, which also contained G. intraradices and G. microaggregatum sequences from the database (Fig. 4).
Neighbour-joining tree. Topologies shown with bootstrap values in percentage. The Glomus claroideum/Glomus etunicatum clade includes sequences obtained in this study from both species and sequences isolated from G. lamellosum ex-holotype. Sequences obtained in previous phylogenetic analyses of the LSU D2 rRNA gene were included (cited in the text), and also sequences isolated from G. intraradices BEG145 and BEG193, G. mosseae BEG185 and G. manihotis BEG112 obtained in this study. Homologous sequences from the GenBank were also included. Note that sequences obtained from single-spore DNA extractions are indicated in the tree by an S before complete code, e.g. SBEG3-01. Cluster Glomus-1 includes sequences from isolates of G. coronatum, G. mosseae including BEG185, G. constrictum BEG130, G. fragilistratum B05 AF145747 G. caledonium BEG86 AF145746 and G. caledonium BEG20 AF145745, G. geosporum as reported by Clapp et al. (2001), as well as G. intraradices BEG145-01. Cluster Glomus-2 includes sequences from G. coronatum as reported by Clapp et al. (2001). Cluster Glomus-3 includes five sequences isolated from G. intraradices BEG145, one isolated from G. manihotis BEG112 and the already deposited sequences from G. intraradices AF389024, AF389022 and X99640. Cluster Glomus-4 includes four sequences isolated from G. manihotis BEG112 and 11 sequences from G. intraradices BEG193. Cluster Gigaspora includes sequences from Gigaspora rosea BEG143 and G. rosea Y12076, also sequences from Entrophospora infrequens as reported by Rodriguez et al. (2001). Cluster Scutellospora includes sequences from Scutellospora heterogama BEG40 as reported by Rodriguez et al. (2001). Cluster Acaulospora-1 includes 2 sequences from Acaulospora longula BEG8 (AF389005 and AF389007) as reported by Rodriguez et al. (2001). Cluster Acaulospora-2 includes sequences from Acaulospora spinosa BEG10 and Acaulospora tuberculata BEG41 as reported by Rodriguez et al. (2001). Cluster Entrophospora colombiana includes four sequences from E. colombiana BEG39 (AF389003, AF389016, AF389017 and AF389018). BEG La Banque Européenne des Glomales, now International Bank of Glomeromycota (http://www.kent.ac.uk/biol/beg)
Discussion
Glomeromycotan sequence diversity
The high levels of sequence diversity first found in isolates of G. claroideum and G. etunicatum analysed by multispore extractions was confirmed by single-spore DNA analyses, adding weight to the high levels of polymorphisms present in the rRNA genes of AMF spores, already reported for other AMF species (Antoniolli et al. 2000; Pringle et al. 2000; Clapp et al. 2001; Rodriguez et al. 2001). This result also refutes the arguments used by other studies, which state that contamination of cultures, i.e. several AMF present together in pure pot cultures, is the reason for such high variability in sequences obtained from one fungus (Jansa et al. 2002; Schüßler et al. 2003). Furthermore, characterisation of individual isolates from agricultural soils in South-East China, using the molecular approach described in this paper, has reached similar conclusions (Robinson et al. 2002). The sequence diversity analysis of the LSU rRNA gene D2 domain of G. etunicatum BEG186 and BEG151 isolated from Chinese agricultural soils yielded 11 unique sequences in each case. The observation of multiple variants in the rRNA genes of AMF spores from pure cultures seems to be a global phenomenon, and reinforces our reservations regarding the use of rRNA sequences as molecular markers in these fungi in ecological studies without sufficient analysis of a range of isolates of each species group.
The analysis of the diversity of isolates of G. claroideum and G. etunicatum based on PCR-SSCP profiles showed that these species have several LSU sequences in common, and in several isolates those sequences were the most abundant. As an indication of the amount of variation within these isolates, sequences alignment as described by Clapp et al. 2001 showed that the Chinese isolate G .etunicatum BEG186 has up to 19% bp differences within the sequences analysed. This compares with G. mosseae, a much tighter species group, which only has up to 2% of differences (sequences reported here and in Clapp et al. 2001). It is interesting to note that the most frequently found sequence from the Chinese isolate G. mosseae BEG185 was identical to that of BEG25 G. mosseae (accession no. AF304983), isolated from West Sussex in England. All of the other sequences analysed were equal to or in excess of 98% similar to sequences recorded for G. mosseae BEG25.
The LSU phylogeny presented in this study indicates that G. claroideum and G. etunicatum represent a separate lineage from other species in the genus Glomus, and supports the two Glomus clades proposed by previous work using SSU rRNA gene sequences (Schüßler et al. 2001b). The large gene diversity found among isolates of G. etunicatum and G. claroideum is highlighted, and a close relationship is also reported between these species and G. lamellosum. The latter species is differentiated mainly by the thickness of its spore wall at maturity. Difficulties in delimiting specific boundaries in AMF have been consistently mentioned by several workers in different contexts and with different implications, particularly in the genus Glomus (Bentivenga and Morton 1994; Simon 1996; Stürmer and Morton 1997; Walker and Vestberg 1998). Few studies, however, have attempted to analyse inter- and intra-specific relationships in the Glomeromycota, utilising different approaches, encompassing a large number of isolates and taking into account the multinucleate state of the spores of these fungi until recently (Schüßler et al. 2001b).
Spore morphology comparisons
The analysis of some spore morphological characters from 45 isolates of G. etunicatum and G. claroideum underlines the high levels of diversity of spores of these species in culture. Spore colour at maturity was the chief discriminating morphological character separating isolates of G. claroideum and G. etunicatum amongst the spore morphological traits investigated. Obviously a taxonomist would evaluate more criteria in allocating a species name to a fungus, but here we refer to five important characters normally employed. The ontogeny of the spore is important but features such as extra wall layer presence at spore maturity do not appear to be consistent, taxonomically useful, features (see Walker and Vestberg 1998 for discussion of inner wall of G. claroideum). Dodd et al. (1996) have reported a possible similar morphological continuum between Glomus coronatum and G. mosseae. These authors found that spore colour was also the only character that could differentiate G. coronatum from G. mosseae. In that study a multimodal approach was proposed as being more appropriate for the assessment and identification of species in this genus. A multidisciplinary approach was later used in a larger scale screening by Clapp et al. (2001) to investigate gene diversity of G. coronatum, G. mosseae, G. geosporum and G. constrictum. They demonstrated that the four species analysed could be part of a genetic continuum (Clapp et al. 2001). A similar approach was used in this investigation and the results strongly suggest a stronger continuum between G. etunicatum and G. claroideum. This does not infer that these species cannot be separated, but rather implies that there are anomalous fungi within the phylogenetically distinct cluster that have some morphologically distinct characters (thicker inner wall layers) or fall between the species descriptions.
On the basis of ontogenic analysis, Stürmer and Morton (1997) suggested that G. etunicatum was closely allied with G. clarum, while G. intraradices was more similar to G. claroideum. In addition, these authors proposed that G. claroideum diverged from the other three species studied and represented a separate lineage based on the formation of an additional innermost flexible sublayer (Stürmer and Morton 1997). Walker (1992) also used this innermost layer to propose polyphyly in the genus Glomus. A recent study on the spore wall structure of G. claroideum, however, reinterpreted the innermost wall as an artefact of specimen preparation (Walker and Vestberg 1998). Our comparative analysis showed that the spore morphologies of G. claroideum and G. etunicatum overlap, and that spore colour at maturity was needed to discriminate all isolates of these two species. In addition, we found that Glomus etunicatum S329 had an additional innermost layer (L3) and also that not all G. claroideum isolates analysed had the innermost flexible layer described by Stürmer and Morton (1997). Stürmer and Morton (1997) likewise reported an innermost sublayer in spores of G. etunicatum UT315, which resembles the flexible layer observed in spores of G. claroideum.
The high levels of morphological diversity found among isolates of G. claroideum and G. etunicatum, in terms of spore diameter, spore colour and wall thickness, has been noted by other authors. For example, the re-analysis of the G. claroideum type specimen in comparison with other isolates of this species from different geographical sites resulted in a re-description of G. claroideum (Walker and Vestberg 1998). Thus, Walker and Vestberg (1998) observed the wide morphological variability in the important taxonomic characters (spore colour, spore diameter) among isolates of G. claroideum, unlike previously reported by Stürmer and Morton (1997). More importantly, some isolates (G. claroideum V198, BEG3 and BEG96 and G. etunicatum S329, BEG92 and CZ1) analysed in our study exhibit intermediate characteristics between G. etunicatum and G. claroideum. Walker (1992) indicated the possibility of morphologically intergrading species in the Glomeromycota. Our evidence supports this view, as isolates of G. claroideum and G. etunicatum have many similarities and appear to have a wide range of spore morphologies.
Multiple sequences vs single sequences in AMF analyses
Very divergent multiple sequences have been reported within single spores of AMF, for example Glomus-like sequences isolated from a species in the Gigasporaceae (Clapp et al. 1999; Hosny et al. 1999) and vice versa (Clapp et al. 2001). Moreover, Glomeraceae-like and Gigasporaceae-like sequences have been found existing in single spores of Entrophospora infrequens (Rodriguez et al. 2001). These examples clearly show that the assumption that sequences from one spore or spores from one isolate cluster together (Schüßler et al. 2001a) is potentially imprecise. The heterogeneity of the multiple copies of rRNA genes within single spores of AMF has been shown to be exceptionally large (Antoniolli et al. 2000; Pringle et al. 2000) and unavoidable for the analysis of the relationships among the Glomeromycota at different taxonomic levels. Schüßler et al. (2001a) acknowledge the presence of multiple sequences but dismiss their importance for phylogenetic analyses, stating that the differences between sequences are minor and they cluster together. This is usually true, if only a few sequences are examined. However, if a greater sample size is used, sequences begin to be found that often cluster with different species (Clapp et al. 2001) or different genera (Rodriguez et al. 2001). Realistically, this will impinge more on studies using sequences to track introduced AMF within colonised root systems. Kjoller and Rosendahl (2001) could not assign sequences from pea roots grown in the field, even cross-checked against single LSU sequences from 17 species of AMF (five had been isolated from the same field).
Another example showing differences with data interpretation is the case of G. microaggregatum BEG56; spores basically have similar morphology to those of G. intraradices but are smaller. This species clustered with 13 sequences isolated from two G. intraradices isolates and G. manihotis BEG112 when incorporated into our LSU phylogeny. This is in contrast to the SSU phylogenetic analysis by Schwarzott et al. (2001), where G. microaggregatum DAOM215235 appeared with the G. claroideum/G. etunicatum clade. The difference could be the result of misidentification of the isolate, as has happened before (reported by Clapp et al. 2002); a chance SSU sequence which is atypical, or a mixed culture. Whatever the reason, this needs further cross-examination. The data, however, do support the SSU phylogeny proposed by Schwarzott et al. (2001) linking G. clarum/G. manihotis with G. intraradices type fungi and should prompt a reassessment of the descriptions of G. proliferum and G. vesiculiferum as to whether their spore morphologies are distinct enough to separate them from a general grouping of G. intraradices or G. aggregatum, which appear to be synonyms.
Pure cultures of AMF
Monospecificity in pot cultures of AMF, free from cross-contamination, requires the employment of numerous and laborious strategies to accomplish this preliminary step in the study of the Glomeromycota. The pursuit of a monospecific culture supposes a ‘purification’ process in which, ideally, morphologically homogeneous spores are obtained after several pot culture cycles. This ‘purification’ process generally starts with multiple spores isolated from trap cultures containing field soil. Thereafter, another process commences which involves the selection of single spores or small groups (5–100 spores) of the desired spore morphotype. Subsequently, many scientists may just sub-culture using a mixed inoculum of spores, hyphae and colonised root fragments, or maintain a sub-culturing cycle using only spores. Selection pressures are applied during this process, which can continue for as long as the isolate is maintained in culture.
Our results suggest that the age of the culture and the subculturing process might explain the differences in sequence diversity found among isolates of G. claroideum and G. etunicatum. Indeed, sequence diversity expressed as a proportion of the occurrence of a given PCR-SSCP pattern varied among the isolates. The highly diverse isolates had numerous sequences, in some cases up to 26, i.e. G. etunicatum BEG146, and no predominant sequences were identified, as previously found in isolates of G. coronatum (Clapp et al. 2001). Isolates with lower diversity, only four sequences in some cases, e.g. G. claroideum V198, often had sequences that occurred in most of the clones analysed. This result was further corroborated by analyses using single spore DNA isolations. Therefore, isolates held in long term culture with frequent subculturing involving single-spore selection may lead to a reduction in their genetic diversity. These ‘monospecific’ cultures, said to be desirable for taxonomic studies (Schüßler et al. 2001a), may simply represent a purified group of morphologically homogeneous spores that in reality are part of a genetic continuum.
There was evidence of genetic drift in lines of G. mosseae BEG12, held for more than 12 years at different European laboratories (Wyss and Bonfante 1993). Gene variation was also reported in subcultures of the same isolate of G. coronatum, maintained in different laboratories for around six years (Clapp et al. 2001). Further studies should be attempted to establish the real effects on AMF genetic diversity of the different selection pressures that we apply when pot cultures are established and maintained. This may also be connected to the loss of effectiveness of certain isolates of AMF sometimes observed on certain hosts. Feldmann (1998) observed that long-term cultivation of G. etunicatum isolates on parsley led to a decrease in effectiveness of those strains on that host. The recommendations that employing continual single-spore culturing, or even production in vitro (Declerck et al. 1996), should be used for culture production, may lead inadvertently to a reduction in genetic diversity and even to attenuation of cultures over time.
The combined analysis of spore morphology and LSU rRNA D2 region sequences from isolates of G. claroideum and G. etunicatum in this study, showed that there are some isolates that overlap in certain characteristics and others that are unique but clearly fall within the phylogenetic cluster—Glomus Group B (Schüßler et al. 2001b). Isolates with intermediate morphological characteristics between the original descriptions of G. claroideum and G. etunicatum were identified, which paralleled the high number of sequences in common (35% of total clones analysed) found in isolates of these species. Moreover, those common sequences were also the most abundant for four of the isolates analysed, suggesting that these are intergrading species, which comprise a wide morphological range. Differences in LSU D2 region sequence diversity found in isolates of the G. claroideum/G. etunicatum complex, may reflect differences in culture age and subculturing methods in the process of ‘purification’ of these species. The use of rRNA sequences for the analysis of AMF phylogeny requires adequate sampling, both in terms of number of isolates analysed, and sequences included, due to the high gene diversity within the Glomeromycota and because of the polymorphic nature of their rRNA genes, which implies that “anomalous” sequences are only detected in large sample sizes. This has obvious implications for the use of such data to “detect” species of AMF in field studies following inoculation, based on the use of “specific” sequences. This is supported by recent work that employed Glomus specific sequences from the LSU rRNA gene in a nested PCR to investigate molecular diversity of AMF in field grown peas (Kjoller and Rosendahl 2001). They also used SSCP to screen the nested PCR products to detect the presence of multiple sequence types and found that some of the root derived sequences could not be aligned with those from 17 isolates of Glomus held in culture, even though five of these had been isolated from the same field.
There would appear to be clear evidence that species groups, as defined by clustering of rRNA sequences along with spore morphological characters, could lead to a far more comprehensible taxonomy and phylogeny for the Glomeromycota (Schüßler et al. 2001b). The use of such sequences, however, to “identify” these species groups in roots in agricultural or natural ecosystems may be far more difficult if single sequences alone are used to track the presence or absence of an individual AMF. On the other hand, the use of chosen LSU D2 region sequences, for example the most abundant ones, could be applied for specific purposes, i.e. in situ detection of specific isolates after the range of diversity within species is known.
References
Antoniolli ZI, Schachtman DP, Ophel-Keller K, Smith SE (2000) Variation in rDNA ITS sequences in Glomus mosseae and Gigaspora margarita spores from a permanent pasture. Mycol Res 104:708–715
Bentivenga SP, Morton JB (1994) Stability and heritability of fatty-acid methyl-ester profiles of glomalean endomycorrhizal fungi. Mycol Res 98:1419–1426
Bentivenga SP, Bever JD, Morton JB (1997) Genetic variation of morphological characters within a single isolate of the endomycorrhizal fungus Glomus clarum. Am J Bot 84:1211–1216
Clapp JP, Young JPW, Fitter AH (1999) Ribosomal small subunit sequence variation within spores of an arbuscular mycorrhizal fungus, Scutellospora sp. Mol Ecol 9:915–921
Clapp JP, Rodriguez A, Dodd JC (2001) Inter- and intra-isolate rRNA large subunit variation in Glomus coronatum spores. New Phytol 149:539–554
Clapp JP, Rodriguez A, Dodd JC (2002) Glomales rRNA gene diversity—all that glistens is not necessarily glomalean? Mycorrhiza 12:269–270
Declerck S, Strullu DG, Plenchette C (1996) In vitro mass-production of the arbuscular mycorrhizal fungus, Glomus versiforme, associated with Ri T-DNA transformed carrot roots. Mycol Res 100:1237–1242
Dodd JC, Rosendahl S, Giovannetti M, Broome A, Lanfranco L, Walker C (1996) Inter- and intraspecific variation within the morphologically-similar arbuscular mycorrhizal fungi Glomus mosseae and Glomus coronatum. New Phytol 133:113–122
Feldmann F (1998) The strain inherent variability of arbuscular mycorrhizal effectiveness: II. Effectiveness of single spores. Symbiosis 25:131–143
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 133:783–793
Helgason T, Daniell TJ, Husband R, Fitter AH, Young JPW (1998) Ploughing up the wood-wide web? Nature 394:431
Hosny M, Hijri M, Passerieux E, Dulieu H (1999) rDNA units are highly polymorphic in Scutellospora castanea (Glomales, Zygomycetes). Gene 226:61–71
Jansa J, Mozafar A, Banke S, McDonald B, Frossard E (2002) Intra- and intersporal diversity of ITS rDNA sequences in Glomus intraradices assessed by cloning and sequencing, and by SSCP analysis. Mycol Res 106:670–681
Kjoller R, Rosendahl S (2000) Detection of arbuscular mycorrhizal fungi (Glomales) in roots by nested PCR and SSCP (single stranded conformation polymorphism). Plant Soil 226:189–196
Kjoller R, Rosendahl S (2001) Molecular diversity of glomalean (arbuscular mycorrhizal) fungi determined as distinct Glomus specific DNA sequences from roots of field grown peas. Mycol Res 105:1027–1032
Kramadibrata K, Walker C, Schwarzott D, Schüßler A (2000) A new species of Scutellospora with a coiled germination shield. Ann Bot 86:21–27
Lloyd-MacGilp SA, Chambers SM, Dodd JC, Fitter AH, Walker C, Young JPW (1996) Diversity of the internal transcribed spacers within and among isolates of Glomus mosseae and related arbuscular mycorrhizal fungi. New Phytol 133:103–111
Millner PD, Mulbry WW, Reynolds SL (2001) Taxon-specific oligonucleotide primers for the detection of Glomus etunicatum. Mycorrhiza 10:259–265
Morton JB (2000) Evolution of endophytism in arbuscular mycorrhizal fungi of glomales. In: Bacon CW, White JF Jr (eds) Microbial endophytes. Dekker, New York, pp 121–140
Morton JB (2003) Glomeromycota http://invam.caf.wvu.edu/fungi/taxonomy/glomales.htm
Morton JB, Redecker D (2001) Two new families of Glomales, Archaeosporaceae and Paraglomaceae, with two new genera Archaeospora and Paraglomus, based on concordant molecular and morphological characters. Mycologia 93:181–195
Page RDM (1996) TREEVIEW: an application to display phylogenetic trees on personal computers. Comput Appl Biosci 12:357–358
Pringle A, Moncalvo JM, Vilgalys R (2000) High levels of variation in ribosomal DNA sequences within and among spores of a natural population of the arbuscular mycorrhizal fungus Acaulospora colossica. Mycologia 92:259–268
Redecker D, Hijri M, Dulieu H, Sanders IR (1999) Phylogenetic analysis of a dataset of fungal 5.8S rDNA sequences shows that highly divergent copies of internal transcribed spacers reported from Scutellospora castanea are of ascomycete origin. Fungal Genet Biol 28:238–244
Redecker D, Kodner R, Graham LE (2000a) Glomalean fungi from the Ordovician. Science 289:1920–1921
Redecker D, Morton JB, Bruns TD (2000b) Ancestral lineages of arbuscular mycorrhiza fungi (Glomales). Mol Phylogenet Evol 14:276–284
Robinson L, Rodriguez A, Clapp JP, Knight E, Wong YH, Jeffries P, Dodd JC (2002) Diversity of the D2 domain of the LSU rRNA gene of several Chinese Glomus species. In: Avio L, Sbrana C, Strani P (eds) AM Research in Europe—the dawning of a new millennium. Cost 838 meeting. Area della Ricerca Pisa, 10–12th October 2002, session 1, S.T.A.R. Servizio Tecnografico Area della Ricerca del CNR, Pisa, pp 28
Rodriguez A, Dougall T, Dodd JC, Clapp JP (2001) The large sub-unit ribosomal RNA genes of Entrophospora infrequens comprise sequences related to two different glomalean families. New Phytol 152:159–167
Rosendahl S, Dodd JC, Walker C (1994) Taxonomy and phylogeny of Glomales. In: Gianinazzi S, Schüepp H (eds) Impact of arbuscular mycorrhizas on sustainable agriculture and natural ecosystems. Birkhäuser, Basel, pp 1–12
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Sawaki H, Sugawara K, Saito M (1998) Phylogenetic position of an arbuscular mycorrhizal fungus, Acaulospora gerdemannii, and its synanamorph Glomus leptotichum, based upon 18S rRNA gene sequence. Mycoscience 39:477–480
Schüßler A (1999) Glomales SSU rRNA gene diversity. New Phytol 144:205–207
Schüßler A, Gehrig H, Schwarzott D, Walker C (2001a) Analysis of partial Glomales SSU rRNA gene sequences: implications for primer design and phylogeny. Mycol Res 105:5–15
Schüßler A, Schwarzott D, Walker C (2001b) A new fungal phylum, the Glomeromycota: phylogeny and evolution. Mycol Res 105:1413–1421
Schüßler A, Schwarzott D, Walker C (2003) Glomeromycota rRNA genes—the diversity of myths? Mycorrhiza 13:233–236
Schwarzott D, Schüßler A (2001) A simple and reliable method for SSU rRNA gene DNA extraction, amplification, and cloning from single AM fungal spores. Mycorrhiza 10:203–207
Schwarzott D, Walker C, Schüßler A (2001) Glomus, the largest genus of the arbuscular mycorrhiza fungi (Glomales), is non-monophyletic. Mol Phylogenet Evol 21:190–197
Simon L (1996) Phylogeny of the Glomales: deciphering the past to understand the present. New Phytol 133:95–101
Simon L, Bousquet J, Lévesque RC, Lalonde M (1993a) Origin and diversification of endomycorrhizal fungi and coincidence with vascular land plants. Nature 363:67–69
Simon L, Lévesque RC, Lalonde M (1993b) Identification of endomycorrhizal fungi colonizing roots using fluorescent SSCP-PCR. Appl Environ Microbiol 5:4211–4215
Stürmer SL, Morton JB (1997) Developmental patterns defining morphological characters in spores of four species in Glomus. Mycologia 89:72–81
Swofford DL (1999) PAUP*. Phylogenetic analysis using parsimony (*and other methods). Sinauer, Sunderland, Mass
Walker C (1992) Systematics and taxonomy of the arbuscular endomycorrhizal fungi (Glomales)—a possible way forward. Agronomie 12:887–897
Walker C, Trappe JM (1993) Names and epithets in the Glomales and Endogonales. Mycol Res 97:339–344
Walker C, Vestberg M (1998) Synonymy amongst the arbuscular mycorrhizal fungi: Glomus claroideum, G. maculosum, G. multisubstenum and G. fistulosum. Ann Bot 82:601–624
Wyss P, Bonfante P (1993) Amplification of genomic DNA of arbuscular mycorrhizal (AM) fungi by PCR using short arbitrary primers. Mycol Res 97:1351–1357
Acknowledgements
This work was partly supported by funds contributed from the following sources: The International Institute of Biotechnology (IIB), the BEGNET project EU Framework IV, contract number BIO4-CT97-2225 and NATO grant ENVIR.LG.950923. Alia Rodriguez wishes to express her thanks to COLFUTURO, Carrera 15 No. 37–15, Santafe de Bogota (Colombia), Research School of Biosciences, University of Kent and IIB for the financial support of her Ph.D. studies. The authors would like to thank Drs. David Sylvia, Miroslav Vosatka and Mauritz Vestberg for providing some of the isolates included in this study. Professors Xiaolin Li (China Agricultural University, Beijing), Bin Zhao (Hauzhong Agricultural University, Wuhan) and Yuen Ha Wong (Hong Kong Polytechnic University, Hong Kong, P.R. China) are also acknowledged for the provision of original Chinese soil samples or cultures. The Mychintec (ICA4-CT-2000-30014) project (partner IIB) from the EU INCO-DEV FP5 programme supported Louisa Robinson and the sequencing of Chinese isolates of AMF.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Rodriguez, A., Clapp, J.P., Robinson, L. et al. Studies on the diversity of the distinct phylogenetic lineage encompassing Glomus claroideum and Glomus etunicatum . Mycorrhiza 15, 33–46 (2005). https://doi.org/10.1007/s00572-003-0291-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00572-003-0291-0